Abstract
AP-1 is a dimeric complex that is composed of JUN, FOS, ATF and MAF protein families. FOS-related antigen 1 (FRA1) which encoded by FOSL1 gene, belongs to the FOS protein family, and mainly forms an AP-1 complex with the protein of the JUN family to exert an effect. Regulation of FRA1 occurs at levels of transcription and post-translational modification, and phosphorylation is the major post-translational modification. FRA1 is mainly regulated by the mitogen-activated protein kinases signaling pathway and is degraded by ubiquitin-independent proteasomes. FRA1 can affect biological functions, such as tumor proliferation, differentiation, invasion and apoptosis. Studies have demonstrated that FRA1 is abnormally expressed in many tumors and plays a relevant role, but the specific condition varies from the target organs. FRA1 is overexpressed in breast cancer, lung cancer, colorectal cancer, prostate cancer, nasopharyngeal cancer, thyroid cancer and other tumors. However, the expression of FRA1 is decreased in cervical cancer, and the expression of FRA1 in ovarian cancer and oral squamous cell carcinoma is still controversial. In this review, we present a detailed description of the regulatory factors and functions of FRA1, also, the expression of FRA1 in various tumors and its function in relative tumor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 1988, Cohen DR and Curran T isolated a new cDNA by screening rat c-DNA libraries with FOS DNA probe. This gene is very similar to FOS and is named FOSL1. FOSL1 encodes a protein (FOS-related antigen 1, FRA1) of 275 amino acids. With the ability to induced rapidly by serum in the presence of protein synthesis inhibitors, FOSL1 is considered as cellular immediate-early gene [1]. Subsequently, they went further and found that the FRA1 protein is localized in the nucleus and cytoplasm, and is mainly modified by post-translational phosphorylation. Like c-FOS, FRA1 can bind to JUN to recognize the AP-1 site [2]. By the same method, Matsui et al. confirmed the existence of FOSL1 in human cells in 1990, which is 90% similar with rat FOSL1 gene [3]. Since then, a large number of researches on FRA1 have been published.
In this review, we summarize the regulatory factors and functions of FRA1, especially its expression and corresponding functions in tumors.
AP-1 and FRA1
The FOSL1 gene is located at the 11q13 locus, which encodes the FRA1 protein consisting of 271 amino acids. FRA1 belongs to the FOS protein family, and other members of the family include c-FOS, FOSB, and FRA2. FOS is an important member of the transcription factor AP-1. AP-1 is a dimeric complex that is composed of the JUN (c-JUN, JUNB, JUND), FOS, ATF (ATF1–4, ATF-6, b-ATF, ATFx) and MAF (c-MAF, MAFA, MAFB, MAFG/F/K, and NRL) protein families [4, 5]. In mammals, AP-1 is mainly composed of JUN and FOS. Among them, FOS can only form JUN-FOS heterodimer with JUN, but JUN itself can also form JUN–JUN homodimer. The formation of AP-1 dimer is dependent on the basic leucine zipper (bZIP) domain on JUN and FOS, which also binds to DNA, while AP-1 can regulate target genes through this bZIP domain by binding to the TRE (TGAC/GTCA), a specific DNA sequence on the promoter or enhancer of target genes, thereby affecting the proliferation, differentiation, invasion and apoptosis of tumor cells [6, 7]. In addition, the bZIP domain of FRA1 is located at 115–168 region.
The FOS family has the highly homologous bZIP domain, but compared to c-FOS and FOSB, FRA1 and FRA2 lack a C-terminal transactivation domain and are therefore considered to be poor activators of transcription [8]. Even, since FRA1 can inhibit the expression of c-FOS from the promoter level and inhibit c-FOS-mediated transcriptional activation, it is considered to be a repressor of AP-1 transcriptional activity [9, 10]. In turn, other members of AP-1 can upregulate FOSL1 transcriptional activity by binding to its promoter [11].
Regulation of FRA1
Regulation of FRA1 by post-translational modification
The regulation of FRA1 expression is multifaceted. In addition to the transcriptional regulation of other members of AP-1 and some upstream genes (i.e. PI3K, WNT3a, STAT3, SIRT1) [12,13,14,15], FRA1 is also regulated by itself. Early studies have used 12-O-tetradecanoylphorbol-13-acetate (TPA) to stimulate bronchial epithelial cells and found that the transcriptional activation of FOSL1 is associated with multiple cis-elements of itself (EBS, a GC box, and TRE) [11], While the transcript elongation of FOSL1 is the result of a series of reactions triggered by the phosphorylation of the serine 10 at histone H3 on FOSL1 enhancer [16].
In all regulation of FRA1, post-translational phosphorylation is crucial.
The common phosphorylation sites of FRA1 are at serine and threonine residues, of which S265 and S252 are the most important. Phosphorylation of FRA1 in the C-terminal tail at S265 and S252 can neutralize its degradation while favouring its stabilization [12, 17]. And also, PKCθ-induced phosphorylation of FRA1 at T217 and T227 can enhance its transcriptional activity [18]. In addition, T223, T230 and T240 are also phosphorylation sites of FRA1, but their influence on FRA1 is not as critical as S265 and S252 [17].
FRA1 can be phosphorylated by some kinases, such as mitogen-activated protein kinase (MAPK), protein kinase C (PKC), cAMP-dependent kinase (PKA), and cyclin-dependent kinase 1-cdc2 (CDC2) [19]. FRA1 is directly phosphorylated by MAPKs, and occasionally MAPKs also indirectly regulate FRA1 phosphorylation by RSK1/2 [17]. ERK1/2 and Ste20-related proline-alanine-rich kinase (SPAK) are two mechanisms of PKCθ-induced phosphorylation of FRA1 [20]. PKCα can significantly up-regulate the phosphorylation level of FRA1 without affecting the total FRA1 level [21].
In addition to phosphorylation, FRA1 has other post-translational modifications. With the help of the HDAC6 deacetylase, IL6/STAT3 axis can deacetylate the lys-116 residue of FRA1, and ending with the acquisition of stemness in colon cancer cell [22].
Degradation of FRA1
As a transcription factor protein, FRA1 itself is an unstable protein with a short half-life, and its instability is caused by a single destabilizer located within 30–40 amino acid residues at the C-terminus [17, 23]. The degradation of FRA1 mainly relies on ubiquitin-independent proteasome action. First, the ubiquitin-independent proteasome recognition is initiated by the 19S proteasome subunit-TBP1, and the subsequent proteolytic process is performed by the C-terminal degron [24].
In addition to reducing the degradation of FRA1 by down-regulating the expression level of TBP1, its stability is enhanced mainly by its phosphorylation, as we mentioned above.
Regulation of FRA1 by MARKs pathway
Among the upstream regulatory pathways of FRA1, MAPKs pathway topped the list. As mentioned above, phosphorylation and transcriptional activity of FRA1 are mainly dependent on MAPKs. In mammals, there are at least four different MAPK signaling pathways: extracellular signal-related kinases (ERK)-1/2, Jun amino-terminal kinases (JNK1/2/3), p38 MAPK and ERK5 [25].
The MAPK pathway is a tertiary cascade reaction consisting of MAPKKK–MAPKK–MAPK. When the external stimulus such as growth factors stimulate cells, it can trigger the activation of the proto-oncogene RAS, thereby affecting the downstream RAF gene and activating the MEK1/2-ERK1/2 pathway. Low level of ERK-MAPK activity mainly regulates the transcription of the FOSL1 gene, while a higher level of ERK activity increase FRA1 accumulation by phosphorylating it and preventing its proteasome-dependent degradation [26]. Moreover, recent studies have further revealed that the production rate of FRA1 protein has a linear relationship with the total activity of ERK, and there is also a linear relationship between the total expression levels of FRA1 with the duration of ERK activity [27]. To remove the effects of exogenous stimuli, mutations in the proto-oncogenes RAS and RAF are themselves very common in the development of tumors. However, this is not the only way, RAS can also activate FRA1 by triggering the PI3K/AKT pathway [12]. The regulation of FRA1 by PKC is divided into RAS/ERK-dependent pathway and non-RAS/ERK-dependent pathway [20]. However, when external stimuli are stress signals, inflammatory factors, etc., the activated MAPK pathway is not RAS/ERK, but MLKs, MEKKs, TAK1 and ASK1, some of them activate p38 through MKK3/6, and the other activates JNK1/2/3 through MKK4/7, the expression of FRA1 is then regulated [28] (Fig. 1).
Regulation of FRA1 by microRNAs
MicroRNAs (miRNAs) are a class of endogenous small RNAs with approximately 22 nucleotides that have a variety of important regulatory roles in cells. Recently, many miRNAs have been shown to inhibit tumorigenesis by downregulating FRA1 expression. These include but not limited to the miR-19a/b in breast cancer and cervical cancer [29, 30], miR-138 in squamous cell carcinoma [31], miR-497 in colorectal cancer [32], miR-130a in breast cancer [33], miR-195 in prostate cancer [34], miR-34 in breast cancer and colon cancer [35, 36] (Fig. 2). Based on the role of miR-34 in breast cancer, researchers have found a nanohybrids targeted miR-34a could inhibit tumor growth in vivo [37]. These miRNAs modulate FRA1 expression through directly bind the seed sequence of itself to its partially complementary seed match sequence in the 3′ untranslated region (UTR) of FOSL1. However, miR-138 was found to bind not only to the 3′UTR of FOSL1 but also to its 5′UTR and coding sequences (CDS). Canonical and non-canonical targeting sites work together to inhibit FRA1 expression [38]. In contrast, certain miRNAs are also FRA1-targeting genes. FRA1 can indirectly affect the expression of ZEB by regulating the transcription of miR-221/222 [39]. In ovarian cancer, up-regulation of FRA1 can increase the expression level of miR-134 and enhance the chemotherapy resistance of ovarian cancer [40].
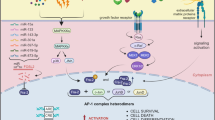
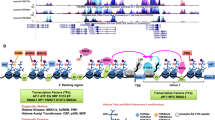
Regulation and functions of FRA1 in tumor. Fra-1 is primarily regulated by transcriptional levels and post-translational phosphorylation and plays a crucial role in tumor progression. The RAS/RAF pathway, PKC, PI3K, WNT/APC, IL-6/STAT3, SIRT1 and miRNAs can all regulate the transcription and post-translational phosphorylation of FRA1. p-FRA1 and p-JUN form heterodimers, which then affect tumor proliferation by regulating cycle-associated proteins, affect tumor invasion and metastasis through MMPs and EMT-TFs, and affect tumor apoptosis through p53 pathway
The expression of FRA1 in tumors
At first, the researchers thought that FRA1 is involved in embryonic development and bone formation. Later, more and more studies have confirmed that FRA1 is abnormally expressed in many tumors, which plays an important role in tumorigenesis and progression. So far, studies on FRA1 have covered tumors of almost all parts of the body, such as breast cancer, colorectal cancer, lung cancer, cervical cancer, ovarian cancer, skin cancer, melanoma, and esophageal cancer. Interestingly, the abnormal expression of FRA1 in various tumors and its effect on tumors vary depending on the type of tumor [14, 34, 40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71] (Table 1).
Breast cancer
Breast cancer is the most common malignant tumor among Chinese women and the leading cause of death among women under 45 years of age in all tumors [72]. Among the cancer in female worldwide, the diagnosis rate and mortality rate of breast cancer have exceeded lung cancer and ranked No. 1 [73], which seriously threatens women’s health and life. Among all the tumors that have been studied for FRA1, breast cancer was the most studied.
Very early studies have found that FRA1 is highly expressed in breast cancer and plays an important role in the malignant progression of breast cancer. However, due to the differences in hormone receptor levels, there are still some differences in the expression levels and functions of FRA1 in different subtypes of breast cancer. In vitro, compared to less aggressive, estrogen receptor-positive (ER+) breast cancer cell line MCF7, highly invasive breast cancer cell lines MDA-MB231, BT549 and HS578T that are estrogen receptor-negative (ER−) have a higher expression level of FRA1 [74]. In vivo, the researchers confirmed that the expression of FRA1 was enhanced with the severity of the lesions through a large number of clinical specimens [42], and was negatively correlated with the patient’s distant metastasis survival and overall survival [75]. At the meantime, compared with the normal tissues adjacent to the cancer, the expression of FRA1 in cancer tissues was found to be higher than that in the adjacent tissues.
According to the further analysis of the subtypes, it was found that FRA1 expression is only slightly increased in the HER-2 type, while the expression in the basal like type is increased significantly compared with the luminal type [43, 76]. Similar results were also obtained in Oliveira-Ferrer. L’s study: ER- patients had higher FRA1 expression level than ER+ patients, and their survival was generally shorter than ER+ patients. Moreover, it was reported that the prognosis was significantly correlated with the level of FRA1 expression between ER+ patients, but there is no correlation between ER− patients [77]. However, unlike above studies, a recent study found that FRA1 can be used as a clinical outcome indicator for ER− breast cancer patients: the ER− patients with a higher expression level of FRA1 showed a shorter overall survival, but which was not found in ER+ breast cancer. In addition, the study also found that triple-negative breast cancer (NTBC) cells have a higher expression level of FRA1 than ER+ cells due to the presence of more enhancers on FOSL1 sequence [78]. By copy number analysis of the DNA extracted from tissues, it was found that the copy number alterations of FOSL1 in NTBC was significantly higher than that in luminal and HER2+ [79]. Moreover, as an oncogenic coactivator of FRA1, DDT5 can enhance the transcriptional activity of FOSL1 and the proliferative effect of it. Like FRA1, DD5 is expressed more highly in TNBC than other subtypes [80]. Similarly, the architectural chromatin protein HMGA1 is also highly expressed in TNBC, and FRA1 can recruit RNA Polymerase II to its promoter by binding to its enhancer, promoting its transcription and malignant effects on TNBC [81]. To some extent, this explains why FRA1 has different effects in different breast cancer cells, but how this variation which depends on hormone receptor level produced has not been further studied.
Lung cancer
As the cancer with highest morbidity and mortality in the world, lung cancer accounts for 11.6% of all cancer patients, and accounts for 18.4% of total cancer deaths. It is the leading cause of cancer death in men and ranks third among women [73].
KRAS is one of the most common mutations in non-small cell lung cancer (NSCLC). KRAS can regulate FRA1 through ERK1/2, ERK5 and JNK [61]. KRAS can also activate the PI3K-mTOR pathway. In general, the combination of inhibitors for two pathways is used as a treatment for this mutation [82]. In vivo, FRA1 participates in KRAS-induced lung cancer, by regulating the anti-oxidation and anti-apoptosis-related genes. The FRA1 deficiency mice reduce the mutant KRAS-induced lung tumorigenesis and show a longer survival time [83].
It is well realized that many environmental factors such as smoking, silica, asbestos, and DEP are high risk factors for causing lung cancer. The effect of these risk factors on lung cancer is closely related to FRA1 [84, 85]. For example, smoking can promote the expression of FRA1 by stimulating the MMPs-EGFR-ERK/JNK/p38 pathway of lung epithelial cells, thereby promoting the development of lung cancer [86]. EGFR-associated FRA1 expression which stimulated by smoking can also be regulated by the PI3K-PAK1-MEK1/2-ERK1 pathway, without the involvement of AKT [87]. Smoking can induce c-JUN/FRA1 to bind to the promoter of SPRR3 and promote its upregulation, while SPRR3 is an indicator of pathogenic keratinization [88]. Besides, smoking can also synergize with Nanoceria on FRA1 to enhance the growth and migration of lung cancer cells [89]. Asbestos can change the redox state of cells by regulating the glutathione, promoting the phosphorylation of ERGF, and then induce the expression of FRA1 [90].
Colorectal cancer
So far, the mortality of colorectal cancer ranks second in the world. It has been estimated that more than 860,000 patients worldwide will die from colorectal cancer in 2018 [73].
Although the expression level of FRA1 in tumor tissues is higher than that in normal tissues, it is not essential for growth and proliferation of proto-oncological lesions. The regulation of FRA1 on colorectal cancer is mainly manifested in the invasion and metastasis ability of cancer [91].
The effect of FRA1 on the invasiveness of colorectal cancer can be reflected by the results of IHC: FRA1 is not stained in normal epithelial cells and strongly stained in the nucleus of cancer cells, and its staining in marginal cancer cells with invasiveness and inflammation is stronger than the center of lesion. Moreover, it has been found that FRA1 staining in liver metastases is stronger than the primary lesion [14, 92].
Cervical cancer
Cervical cancer has the highest incidence and death in female genital malignant tumors [73]. Unlike the tumors mentioned above, FRA1 has been shown to be lowly expressed in cervical cancer, inhibiting the malignant phenotype of cervical cancer cells.
Cervical cancer is one of the few cancers with a clear cause to date, and human papillomavirus (HPV) (especially HPV16/18) infection is the leading cause of cervical cancer. The oncoproteins E6 and E7 play a key role in the malignant progression of HPV-induced cervical cancer. In the process of HPV-induced cervical cancer formation, reorganization of AP-1 dimer (FRA1 down-regulation and c-FOS up-regulation) plays an important role [93]. Similarly, in HPV infection-associated tongue cancer, by FRA2 knockdown, the researchers found that FRA1 and p53 were up-regulated, while MMP9, cyclinD and HPV E6/E7 were down-regulated, suggesting that FRA1 has anti-tumor effects [53]. In HPV-positive (higher AP-1 activity) esophageal cancer, FRA1 expression is also very low, while FRA1 is highly expressed in HPV-negative esophageal cancer. Unlike in cervical cancer, FRA1 promotes the progression of esophageal cancer [68], which maybe the interaction of HPV and FRA1 in esophageal cancer is not dominant. Therefore, the effect of FRA1 on HPV-associated tumors is associated with host cell types. How HPV infection affects the biological behavior of related tumor cells through FRA1 still needs further exploration.
Ovarian cancer
Although the incidence of ovarian cancer is not high compared to other gynecological malignancies, the degree of malignancy is the first in gynecological malignancies. Late diagnosis and treatment resistance are the two major causes of high mortality in ovarian cancer. The common chemotherapy regimen for ovarian cancer is the addition of paclitaxel to platinum, but patients eventually die due to chemotherapy resistance and cancer recurrence [94].
At present, there are few studies on FRA1 and ovarian cancer, and there is no unified recognition of the expression and specific effects of AP-1 and FRA1 in ovarian cancer. Oleg I Tchernitsa et al. found that the expression level of FRA1 in ovarian epithelial cells of rat with KRAS mutation was significantly increased. It has been demonstrated that silencing of FRA1 reverses some of the growth-promoting effects caused by KRAS mutations [95]. However, S Mahner et al. found that c-FOS is an independent factor for ovarian cancer, but not FRA1. Decrease of c-FOS expression can promote ovarian cancer progression and reduce progression-free survival and overall survival in patients [96], which was consistent with subsequent findings that c-FOS is beneficial for the prognosis of ovarian cancer, and overexpression of c-FOS can promote tumor cell apoptosis [97]. However, a recent study indicated that TGF-β induced the development of ovarian cancer by stimulating the c-FOS/c-JUN via the MAPKs pathway [98]. S Mahner et al. compared the expression levels of AP-1 in benign ovarian tumors, borderline ovarian cancer and malignant ovarian cancer. It was found that the JUN family was generally more highly expressed in malignant tumors than benign tumors. Whereas there was no significant difference in the expression level of FRA1 in ovarian specimens with different malignant degrees (including benign tumors). Although the number of specimens used in the study was small, the results of in vitro experiments also showed that there was no significant relationship between the expression level of FRA1 and the proliferation, invasion and metastasis of ovarian cancer cells [49].
Other tumors
In liver cancer, the expression of FRA1 is positively correlated with the level of alpha fetoprotein (AFP) and the degree of vascular invasion [62]. The high expression of FRA1 corresponds to the poor prognosis of liver cancer, suggesting that FRA1 can be used as a prognostic biomarker for liver cancer [63]. As the fifth most common cancer worldwide, the incidence of gastric cancer has a clear regional difference. China is a region with high incidence of gastric cancer. Studies have shown that this is related to a functional single nucleotide polymorphisms (SNPs) rs1892901 in FOSL1, which can enhance the expression of FRA1 and promote the development of gastric cancer [99]. Recent study has found that the expression of FRA1 is regulated by Helicobacter pylori and plays an important role in the development of Helicobacter pylori-mediated gastric cancer [100]. The oncoprotein TAX encoded by HTLV-1 is critical for the development of adult T cell leukemia (ATL), and previous studies have found that TAX can affect the transcription of FOSL1 [101]. Recent studies have revealed that this is associated with TAX-induced activation of the PI3K/AKT-AP-1 pathway [5]. Cigarette smoke can activate the expression of FRA1 via CHRNA7 signaling, allowing it to bind to the promoter of the RNA polymerase II-associated factor, thereby inducing stem cell features in pancreatic cancer cells [102].
Similar to ovarian cancer, the expression of FRA1 in oral squamous cell carcinoma is also controversial. The positive rate of FRA1 in oral squamous cell carcinoma specimens was initially found to be lower than that in normal tissues [52], which was consistent with the results obtained by Gupta et al. in tongue cancer specimens [53]. However, through in vitro experiments, it was found that Yes-associated protein can promote the proliferation and invasion of oral squamous cell carcinoma cells by activating FRA1. And the results of IHC also showed that the expression of FRA1 in the edge of the invasive tumor was higher than that within the tumor [103]. Similarly, recent studies have also indicated that the expression of c-FOS is higher in normal tissues than in cancer, and the overexpression rate of FRA1 is elevated with the increased degree of malignancy and variation of clinical classification, along with the loss of c-FOS. The high expression of FRA1 is negatively correlated with the 5-year survival of patients [104, 105].
Role of FRA1 in tumor
Proliferation
FRA1 can directly regulate the expression of cell cycle-related protein, including cyclin-dependent kinases (CDKs) and cyclins to promote mitotic progression, thereby promoting cell proliferation (Fig. 2). Early studies analyzed the relationship between the seven members of the AP-1 family and cell cycle-associated proteins, and found that the predominance expression of FRA1, similar to FRA2, c-FOS and JUND, can lead to G1-S transition. Moreover, FRA1 expression level is closely related to the expression of cyclinE and p16 [106]. i.e. Silica can increase the expression of FRA1 and the accumulation of lung epithelial cells in S phase [107]. Similarly, in gastric cancer, FRA1 overexpression can increase DNA synthesis and promote cell proliferation by accumulate cancer cells in S phase [64]. In osteosarcoma, the knockdown of FRA1 can significantly down-regulate the expression of cyclinD1 and cyclinD3, and decrease the G1-S phase conversion rate [71]. In thyroid cancer, FRA1 can bind to the promoter of ACCNA2 (encodes cyclinA) and convert cell from G2 to M phase. When the expression of FRA1 is decreased, most of the cells are retained in G2 phase, while parts of them continuation of mitosis, but ended in failure [108]. In oral squamous cell carcinoma and esophageal squamous cell carcinoma, proliferation is associated with overexpression of FRA1 and its downstream cyclinD [103, 109]. In malignant mesothelioma, HGF-induced cell proliferation is mainly due to increased expression of proliferating cell nuclear antigen (PCNA) caused by FRA1 [110]. The growth-promoting effect of UBE2N, a K63-specific ubiquitin conjugase, on melanoma is also closely related to the activation of MEK/FRA1 pathway [111]. As for breast cancer, in addition to directly affecting the expression of CDKs, cyclinD, and cyclinE, by using ChIP-qPCR, the researchers also found that FRA1/c-JUN can bind to the third intron of CLCA2, a gene negatively-associated with proliferation, and decrease its expression, promote cell proliferation [112]. The anti-proliferation effect of psoralen on breast cancer cells is due to the up-regulation of Axin2 and the down-regulation of FRA1 expression, resulting in G0/G1 phase and G2/M phase arrest in MCF-7 cells and MDA-MB-231 cells, respectively [113]. In neuroblastoma and nasopharyngeal carcinoma, cell proliferation is also performed by the c-JUN/FRA1 complex [114, 115]. However, in gliomas and cervical cancers, overexpressed FRA1 inhibits the proliferation of related cells [48, 116]. In prostate cancer, dihydrotestosterone can increase the proliferation of androgen receptor (AR)-positive LNCaP cell line by up-regulating the expression of FRA1, but has no effect on AR-negative PC-3 cell line, thus inferring the hormonal sensitivity of target organs maybe join in the relationship between FRA1 and cell proliferation [117].
Invasiveness and metastasis
The metastasis of cancer cells is an important feature of malignant tumors, and is also the main cause of inability to perform surgery, as well as death in cancer patients. The occurrence of epithelial-mesenchymal transition (EMT) is a key step in cancer invasiveness and metastasis [118]. The effect of FRA1 on tumor invasion and metastasis is mainly achieved by regulating the expression of EMT- inducing transcription factors (EMT-TFs) and matrix metalloproteinases (MMPs) (Fig. 2).
For breast cancer, its strong invasiveness is closely related to FRA1. FRA1 can directly bind to the promoters of PLAU (encoding uPA) [74], MMP1 [119] and MMP9 [120], promoting the aggressiveness and non-adherent growth of cancer cells. Through the MLK3-KO human cell lines tested by murine xenografts, the researchers also found that the MLK3-ERK/JNK-FRA1-MMP1/9 signal cascade existed not only in the primary tumor but also in the circulating tumor cells, which was of great significance for the distant metastasis of cancer cells [121]. Radix Glycyrrhiza is used for the treatment of breast cancer because its main component Glycyrrhetinic acid (GA) can inhibit the FRA1/MMPs signal axis, but the main pathway of action is the p38MAPK/FRA1 pathway [122]. Studies have confirmed that FRA1 is involved in the EMT process of ER- breast cancer by direct binding to the promoters of ZEB1 and ZEB2 [75, 112]. FRA1 can also affect the expression of SLUG by targeting TGFβ [75], and inhibit E-cadherin, promoting EMT process. MiR-130a inhibits the EMT process of MDA-MB-231 cells by up-regulating the expression of ZO-1 through inhibition of FOSL1 transcription [33]. In addition, recent studies have found that FRA1 is involved in the process of CD137-induced monocyte/macrophages migration to the tumor microenvironment and differentiation into osteoclasts, of which favors bone metastasis [123].
The effect of FRA1 on invasiveness of lung cancer is correlated to the enhanced phosphorylation of EGFR mediated by stimulation of MMP2 and MMP9 by FRA1, and MMPs can also form a positive feedback loop with ERK to further promote the expression of FRA1 [124]. In KRAS-mutant lung cancer, even newly discovered molecular targets, such as differentiation-1, promote lung cancer progression and liver metastasis by regulating the level of FRA1 [125].
By using bioinformatics analysis, it has been found that six easily mutated genes (APC, KRAS, BRAF, PIK3CA, SMAD4 and p53) in colorectal cancer are associated with invasiveness and metastasis [126]. FRA1 is involved in the regulation of colorectal cancer cell invasion by these genes. Moreover, FRA1 can form a complex with c-JUN and galectin3, and further bind to the promoter of MUC2 to regulate its transcription, thereby enhancing the invasiveness of colorectal cancer [127]. FRA1 is involved in the effects of IL-6/STAT3 and SIRT1 on EMT processes in colorectal cancer cells [14, 15]. Studies have found that FRA1 can bind to the promoter of vimentin, directly promoting the expression of vimentin with interstitial cell characteristics [128].
In prostate cancer, FRA1 enhance cell metastasis by upregulating N-cadherin and SNAIL and downregulating E-cadherin [50]. In pancreatic cancer, FRA1 promotes EMT and metastasis of tumor upon the stimulation of MUC1 [60]. Moreover, the carcinogen benzidine can promote EMT of bladder cancer cells via ERK5-AP-1 (c-FOS, c-JUN, FRA1) [129]. It has been shown that the formation of complex between αB-Crystallin and 14-3-3ζ protein can promote EMT of liver cancer cells via KRAS-RAF-MEK1/2-ERK1/2-FRA1-SLUG pathway, as well as the resistance production of cancer cells to Sorafenib [130]. CTHRC1 is one of the most highly expressed genes in esophageal squamous cell carcinoma. It can activate FRA1 through RAF-MEK1/2-ERK1/2, and then up-regulate SNAIL, MMP14 and HMGA1 to promote invasion of cancer cells [109, 131]. The EB virus-encoded latent membrane protein (LMP) is an important cause of nasopharyngeal carcinoma. LMP2A and LMP1 can promote the phosphorylation of FRA1/c-JUN through ERK, thereby activating MMP9 and promoting the invasion of nasopharyngeal carcinoma cells [132]. In addition, LMP1 can also activate the upstream pathway of FRA1, PI3K pathway [115].
The most common mutated gene in melanoma is BRAF. If the tumor suppressor gene PTEN is also silenced, HMGA1 expression can be induced by BRAF-ERK1/2-FRA1 and PI3K-AKT-mTOR-FRA1 pathways, resulting in down-regulation of MITF/AXL ratio, which finally lead to the melanocyte reprogramming and transformation [66, 133]. NRAS/BRAF pathway can induce the conversion of melanocytes into malignant melanoma cells by reorganization of EMT-TFs, that is the conversion of SNAIL2, ZEB2 to TWIST1 and ZEB1. The dedifferentiation and malignant switch are FRA1-dependent [134].
Apoptosis
Although the relationship between FRA1 and tumor cell apoptosis varies greatly from tissue to tissue, it is closely related to the p53 pathway (Fig. 2). In lung cancer cells, elevated FRA1 can inhibit p53 and up-regulate the level of its negative regulator MDM2, ultimately inhibiting the apoptosis of lung cancer cells by enhancing apoptosis-related mitochondrial membrane potential (ΔΨm) and down-regulating intracellular ROS and aggregation of Ca2+ [45]. On the contrary, the expression of p53 in cervical cancer is consistent with FRA1, and overexpression of FRA1 can promote apoptosis of cancer cells through p53 [48]. So far, what caused this difference between organizations has not yet been elucidated.
Treatment resistance
According to different treatment methods, tumor resistance can be divided into radiotherapy resistance and chemotherapy resistance.
As mentioned above, BRAF is one of the common mutation genes in colorectal cancer, and patients with BRAFV600E mutations often resist to the MEK1 inhibitor, Selumetinib. The reason is that a key protein CEMIP links the WNT pathway to the MEK1-ERK1/2 pathway, whereas CEMIP expression is induced by β-catenin- and FRA1-dependent pathways [135]. In addition, high expression of FRA1 is also associated with the formation of radiotherapy resistance in colorectal cancer [136].
The radiotherapy resistance of prostate cancer is related to the activation of EGFR and PI3K. Both pathways can increase the expression of AP-1 and enhance prostate cancer cells resistance to radiation [137].
In the treatment of cervical cancer, curcumin is considered a good anticancer agent. The use of curcumin before radiotherapy can increase the activity of ERK1/2 and ROS production, and enhance the radiosensitivity of cervical cancer [138]. Recent studies have confirmed that the radiosensitizer behavior of turmeric is related to its inhibition for AP-1 DNA binding activity and stimulation of AP-1 reorganization, i.e. down-regulation of c-FOS and up-regulation of FRA1 [139]. In other words, FRA1 has a radiosensitizing effect on cervical cancer.
In liver cancer cells, the formation of complex between αB-Crystallin and 14-3-3ζ protein can promote EMT of cells via KRAS-RAF-MEK1/2-ERK1/2-FRA1-SLUG pathway, as well as the resistance production of cancer cells to Sorafenib [130]. In melanoma cells, PI3K/FRA1 is involved in the regulation of FGF1 secretion associated with BRAF inhibitor resistance, providing a basis for the combined application of FGF1 inhibitor and BRAF inhibitor [140].
For ovarian cancer, a study about miR-134 showed that the mRNA level of FOSL1 was higher in cancer than in normal ovaries and positively correlated with the expression of miR-134. In the case of HRAS mutation, the JNK/ERK-FRA1-miR-134-SDS22-JNK/ERK-FRA1 positive feedback loop promotes the proliferation of tumor cells, enhances the chemotherapy resistance of ovarian cancer cells to doxorubicin, and reduces the median survival time of patients [40]. Similarly, cellular studies have shown that occurrence of JQ1 resistance during the treatment of BRD4-associated ovarian cancer cell is due to the activation of RTK-PI3K-AKT and RTK-PI3K-ERK-c-Myc/FRA1 pathways, suggesting that c-MYC/FRA1 can promote the growth of ovarian cancer cells in vitro [141]. Later, the team further studied that the FRA1-related RTK-RAF and RTK-PI3K pathways were also involved in the chemoresistance of MEKi [142]. Furthermore, as the most commonly used chemotherapy drug for ovarian cancer, studies have confirmed that cisplatin resistance is associated with abnormal expression of FRA1. FRA1 participates in ROS-IL-11-JAK2-STAT5-mediated cisplatin resistance by directly binding to the promoter of IL-11 [143].
Similar to ovarian cancer, c-MYC/FRA1 is associated with chemotherapy resistance of mesothelioma. A combination of JQ1 and cisplatin can promote tumor cell apoptosis by affecting FRA1/c-MYC in vitro [144], while trametinib can down-regulate FRA1 and CD44 by inhibiting ERK phosphorylation and elicit anti-tumor effect [145].
In summary, FRA1 has an effect on tumor radioresistance and chemoresistance, which may provide some ideas for future combination therapy. However, we also noticed that although FRA1 serves as a hub for many classical pathways and plays an important role in tumors, there are currently no inhibitors directly targeting FRA1. While, recently, Wei Yang et al. developed a multi-kinase inhibitor, LY-1816, which not only inhibits the phosphorylation level of SRC kinase, but also directly inhibits the expression level of FRA1. The inhibitor has confirmed its tumor suppressing effect in a variety of vitro cells and pancreatic ductal adenocarcinoma xenografts. Although the anti-tumor effect of the inhibitor was found to be stronger than that of gemcitabine and dasatinib, many difficulties still need to be overcome before its clinical trial [146]. In general, the development of small molecule targeted drugs for FRA1 still has a long way to go.
Discussion and perspectives
In this review, we discussed the regulatory factors of FRA1. FRA1 can be regulated by transcription and post-translational modification, mainly by post-translational phosphorylation. The classic regulatory pathway is RAS/ERK signaling pathway. FRA is degraded by proteasome. FRA1 is also regulated by miRNAs. We also discussed the expression and function of FRA1 in various tumors. Whether FOSL1 plays as a proto-oncogene or a tumor suppressor gene is closely related to the type of target organs. In general, FRA1 is highly expressed in most of tumors and promotes the malignant progression of tumors, except cervical cancer and some controversial tumors.
Heterogeneity is an important feature of malignant tumors, including clonal evolution and cell plasticity, and FRA1 plays a crucial role in this process. Readers can refer to the excellent review by AS Dhillon and E Tulchinsky [147]. The cell plasticity is manifested by the cell reprogramming. In melanoma, FRA1 can affect the chromatin remodeling factor HMGA and MITF, which is closely related to melanoma phenotype [66]. In hepatocellular carcinoma, hepatocyte growth factor (HGF), which derived from cancer-associated fibroblasts could regulate tumor-initiating cell plasticity through c-Met/FRA1/HEY1 Signaling [148]. However, in other tumors, the effect of FRA1 on cell reprogramming is mostly focused on altering cell polarity by EMT-TFs. There is no profound study on how FRA1 changes chromatin status, organelle structure, and cytoskeletal rearrangement. In addition, abnormal energy metabolism is one of the top ten characteristics of malignant tumors. As an important factor affecting the development of tumors, the role of FRA1 in energy metabolism is also worth exploring. At present, researchers have found that FRA1 and c-FOS can increase the rate of phospholipid synthesis and promote breast cancer cell proliferation by associating and activating the rate limiting enzyme CDP-DAG synthase, which is a phospholipid synthesis factor. The N-terminal domain plays a key role in this process [149].
The occurrence of tumors is a very complicated process, and the tumor microenvironment (TME) plays a very important role in the occurrence of tumors. Earlier studies have used co-culture techniques to find that breast cancer cells can promote the overexpression of FRA1 in tumor-associated macrophages (TAMs), activate the IL-6/STAT3 pathway, and induce the differentiation of tumor-associated macrophages from M1 to M2, which facilitate the immune evasion of tumor cells, and in turn promotes the aggressiveness of breast cancer cells [150, 151]. miR-19a-3p inhibits the polarization of TAM and the aggressiveness of breast cancer by inhibiting the level of FRA1 in TAMs [29]. Similar phenomenon exists in lung cancer. M2 macrophages can promote the invasion and metastasis of lung cancer by regulating CRYAB expression and activating the ERK1/2-FRA1-SLUG signaling pathway [152]. Based on the correlation between FRA1 and the TME, it is possible that target of FRA1 will be a breakthrough in cancer immunotherapy. But the current progress in this area is still relatively limited (only initial trials in breast cancer) and requires further efforts.
Targeted therapy as an important treatment for malignant tumors, many small molecule targeted drugs targeting specific targets have been developed. However, studies have shown that sustained targeted therapy can induce secretome, leading to drug resistance, and accelerate the spread of tumors, severely limiting the effectiveness of targeted therapy. The BRAF inhibitor vemurafenib reactive secretome is closely related to the down regulation of FRA1 [153]. In addition to simple drug resistance, some tumor cells even have an addictive reaction to drugs, and a sudden withdrawal of drugs will lead to the death of a large number of addicted cells. Using CRISPR technology, Professor Peeper’s team found that the death of melanoma cells induced by drug withdrawal is closely related to the activation of the ERK2-JUNB/FRA 1 pathway [154]. Professor Roger’s group also advocates that DNA damage and cell death caused by drug withdrawal are associated with robust p-ERK-induced up-regulation of the p38-JUNB/FRA1-CDKN1A pathway [155]. Rational use of the drug resistance and addiction of tumor cells can improve the lethality of cancer drugs on addictive cells, which provide new ideas for the treatment of drug-insensitive tumors.
References
Cohen DR, Curran T (1988) Fra-1—a serum-inducible, cellular immediate-early gene that encodes a Fos-Related antigen. Mol Cell Biol 8(5):2063–2069. https://doi.org/10.1128/Mcb.8.5.2063
Cohen DR, Ferreira PCP, Gentz R, Franza BR, Curran T (1989) The product of a Fos-Related Gene, Fra-1, binds cooperatively to the Ap-1 Site with Jun—transcription factor Ap-1 is comprised of multiple protein complexes. Gene Dev 3(2):173–184. https://doi.org/10.1101/gad.3.2.173
Matsui M, Tokuhara M, Konuma Y, Nomura N, Ishizaki R (1990) Isolation of human Fos-Related genes and their expression during monocyte-macrophage differentiation. Oncogene 5(3):249–255
Eferl R, Wagner EF (2003) AP-1: a double-edged sword in tumorigenesis. Nat Rev Cancer 3(11):859–868. https://doi.org/10.1038/nrc1209
Gazon H, Barbeau B, Mesnard JM, Peloponese JM Jr (2017) Hijacking of the AP-1 signaling pathway during development of ATL. Front Microbiol 8:2686. https://doi.org/10.3389/fmicb.2017.02686
Curran T, Franza BR Jr (1988) Fos and Jun: the AP-1 connection. Cell 55(3):395–397
Angel P, Imagawa M, Chiu R, Stein B, Imbra RJ, Rahmsdorf HJ, Jonat C, Herrlich P, Karin M (1987) Phorbol ester-inducible genes contain a common cis element recognized by a TPA-modulated trans-acting factor. Cell 49(6):729–739
Wisdom R, Verma IM (1993) Transformation by Fos proteins requires a C-terminal transactivation domain. Mol Cell Biol 13(12):7429–7438. https://doi.org/10.1128/Mcb.13.12.7429
Ito E, Sweterlitsch LA, Tran PBV, Rauscher FJ, Narayanan R (1990) Inhibition of Pc-12 cell-differentiation by the immediate early gene Fra-1. Oncogene 5(12):1755–1760
Yoshioka K, Deng TL, Cavigelli M, Karin M (1995) Antitumor promotion by phenolic antioxidants—inhibition of Ap-1 activity through induction of Fra expression. Proc Natl Acad Sci USA 92(11):4972–4976. https://doi.org/10.1073/pnas.92.11.4972
Adiseshaiah P, Papaiahgari SR, Vuong H, Kalvakolanu DV, Reddy SP (2003) Multiple cis-elements mediate the transcriptional activation of human fra-1 by 12-O-tetradecanoylphorbol-13-acetate in bronchial epithelial cells. J Biol Chem 278(48):47423–47433. https://doi.org/10.1074/jbc.M303505200
Casalino L, De Cesare D, Verde P (2003) Accumulation of fra-1 in ras-transformed cells depends on both transcriptional autoregulation and MEK-dependent posttranslational stabilization. Mol Cell Biol 23(12):4401–4415. https://doi.org/10.1128/Mcb.23.12.4401-4415.2003
Zhang L, Liu H, Mu X, Cui J, Peng Z (2017) Dysregulation of Fra1 expression by Wnt/beta-catenin signalling promotes glioma aggressiveness through epithelial-mesenchymal transition. Biosci Rep. https://doi.org/10.1042/BSR20160643
Liu H, Ren GP, Wang TY, Chen YX, Gong CJ, Bai YF, Wang B, Qi HY, Shen J, Zhu LJ, Qian C, Lai MD, Shao JM (2015) Aberrantly expressed Fra-1 by IL-6/STAT3 transactivation promotes colorectal cancer aggressiveness through epithelial-mesenchymal transition. Carcinogenesis 36(4):459–468. https://doi.org/10.1093/carcin/bgv017
Cheng F, Su L, Yao C, Liu L, Shen J, Liu C, Chen X, Luo Y, Jiang L, Shan J, Chen J, Zhu W, Shao J, Qian C (2016) SIRT1 promotes epithelial-mesenchymal transition and metastasis in colorectal cancer by regulating Fra-1 expression. Cancer Lett 375(2):274–283. https://doi.org/10.1016/j.canlet.2016.03.010
Zippo A, Serafini R, Rocchigiani M, Pennacchini S, Krepelova A, Oliviero S (2009) Histone crosstalk between H3S10ph and H4K16ac generates a histone code that mediates transcription elongation. Cell 138(6):1122–1136. https://doi.org/10.1016/j.cell.2009.07.031
Basbous J, Chalbos D, Hipskind R, Jariel-Encontre I, Piechaczyk M (2007) Ubiquitin-independent proteasomal degradation of Fra-1 is antagonized by Erk1/2 pathway-mediated phosphorylation of a unique C-terminal destabilizer. Mol Cell Biol 27(11):3936–3950. https://doi.org/10.1128/MCB.01776-06
Belguise K, Cherradi S, Sarr A, Boissiere F, Boulle N, Simony-Lafontaine J, Choesmel-Cadamuro V, Wang X, Chalbos D (2017) PKCtheta-induced phosphorylations control the ability of Fra-1 to stimulate gene expression and cancer cell migration. Cancer Lett 385:97–107. https://doi.org/10.1016/j.canlet.2016.10.038
Gruda MC, Kovary K, Metz R, Bravo R (1994) Regulation of Fra-1 and Fra-2 phosphorylation differs during the cell cycle of fibroblasts and phosphorylation in vitro by MAP kinase affects DNA binding activity. Oncogene 9(9):2537–2547
Belguise K, Milord S, Galtier F, Moquet-Torcy G, Piechaczyk M, Chalbos D (2012) The PKCtheta pathway participates in the aberrant accumulation of Fra-1 protein in invasive ER-negative breast cancer cells. Oncogene 31(47):4889–4897. https://doi.org/10.1038/onc.2011.659
Tam WL, Lu H, Buikhuisen J, Soh BS, Lim E, Reinhardt F, Wu ZJ, Krall JA, Bierie B, Guo W, Chen X, Liu XS, Brown M, Lim B, Weinberg RA (2013) Protein kinase C alpha is a central signaling node and therapeutic target for breast cancer stem cells. Cancer Cell 24(3):347–364. https://doi.org/10.1016/j.ccr.2013.08.005
Wang T, Song P, Zhong T, Wang X, Xiang X, Liu Q, Chen H, Xia T, Liu H, Niu Y, Hu Y, Xu L, Shao Y, Zhu L, Qi H, Shen J, Hou T, Fodde R, Shao J (2019) The inflammatory cytokine IL-6 induces FRA1 deacetylation promoting colorectal cancer stem-like properties. Oncogene 38(25):4932–4947. https://doi.org/10.1038/s41388-019-0763-0
Basbous J, Jariel-Encontre I, Gomard T, Bossis G, Piechaczyk M (2008) Ubiquitin-independent-versus ubiquitin-dependent proteasomal degradation of the c-Fos and Fra-1 transcription factors: is there a unique answer? Biochimie 90(2):296–305. https://doi.org/10.1016/j.biochi.2007.07.016
Pakay JL, Diesch J, Gilan O, Yip YY, Sayan E, Kolch W, Mariadason JM, Hannan RD, Tulchinsky E, Dhillon AS (2012) A 19S proteasomal subunit cooperates with an ERK MAPK-regulated degron to regulate accumulation of Fra-1 in tumour cells. Oncogene 31(14):1817–1824. https://doi.org/10.1038/onc.2011.375
Chang L, Karin M (2001) Mammalian MAP kinase signalling cascades. Nature 410(6824):37–40. https://doi.org/10.1038/35065000
Vial E, Marshall CJ (2003) Elevated ERK-MAP kinase activity protects the FOS family member FRA-1 against proteasomal degradation in colon carcinoma cells. J Cell Sci 116(24):4957–4963. https://doi.org/10.1242/jcs.00812
Gillies TE, Pargett M, Minguet M, Davies AE, Albeck JG (2017) Linear integration of ERK activity predominates over persistence detection in Fra-1 regulation. Cell Syst 5(6):549–563. https://doi.org/10.1016/j.cels.2017.10.019
Gallo KA, Johnson GL (2002) Mixed-lineage kinase control of JNK and p38 MAPK pathways. Nat Rev Mol Cell Bio 3(9):663–672. https://doi.org/10.1038/nrm906
Yang J, Zhang Z, Chen C, Liu Y, Si Q, Chuang TH, Li N, Gomez-Cabrero A, Reisfeld RA, Xiang R, Luo Y (2014) MicroRNA-19a-3p inhibits breast cancer progression and metastasis by inducing macrophage polarization through downregulated expression of Fra-1 proto-oncogene. Oncogene 33(23):3014–3023. https://doi.org/10.1038/onc.2013.258
Xiao SS, Zhou YH, Jiang JF, Yuan L, Xue M (2014) CD44 affects the expression level of FOS-like antigen 1 in cervical cancer tissues. Mol Med Rep 9(5):1667–1674. https://doi.org/10.3892/mmr.2014.2010
Jin Y, Wang C, Liu XQ, Mu WB, Chen ZJ, Yu DS, Wang AX, Dai Y, Zhou XF (2011) Molecular characterization of the MicroRNA-138-Fos-like antigen 1 (FOSL1) regulatory module in squamous cell carcinoma. J Biol Chem 286(46):40104–40109. https://doi.org/10.1074/jbc.C111.296707
Zhang N, Shen Q, Zhang P (2016) miR-497 suppresses epithelial-mesenchymal transition and metastasis in colorectal cancer cells by targeting fos-related antigen-1. Onco Targ Ther 9:6597–6604. https://doi.org/10.2147/OTT.S114609
Chen X, Zhao M, Huang J, Li Y, Wang S, Harrington CA, Qian DZ, Sun XX, Dai MS (2018) microRNA-130a suppresses breast cancer cell migration and invasion by targeting FOSL1 and upregulating ZO-1. J Cell Biochem 119(6):4945–4956. https://doi.org/10.1002/jcb.26739
Wu J, Ji A, Wang X, Zhu Y, Yu Y, Lin Y, Liu Y, Li S, Liang Z, Xu X, Zheng X, Xie L (2015) MicroRNA-195-5p, a new regulator of Fra-1, suppresses the migration and invasion of prostate cancer cells. J Transl Med 13:289. https://doi.org/10.1186/s12967-015-0650-6
Yang S, Li Y, Gao J, Zhang T, Li S, Luo A, Chen H, Ding F, Wang X, Liu Z (2013) MicroRNA-34 suppresses breast cancer invasion and metastasis by directly targeting Fra-1. Oncogene 32(36):4294–4303. https://doi.org/10.1038/onc.2012.432
Wu J, Wu G, Lv L, Ren YF, Zhang XJ, Xue YF, Li G, Lu X, Sun Z, Tang KF (2012) MicroRNA-34a inhibits migration and invasion of colon cancer cells via targeting to Fra-1. Carcinogenesis 33(3):519–528. https://doi.org/10.1093/carcin/bgr304
Xia Y, Deng X, Cao M, Liu S, Zhang X, Xiao X, Shen S, Hu Q, Sheng W (2018) Nanodiamond-based layer-by-layer nanohybrids mediate targeted delivery of miR-34a for triple negative breast cancer therapy. RSC Adv 8(25):13789–13797. https://doi.org/10.1039/c8ra00907d
Jin Y, Wang C, Liu X, Mu W, Chen Z, Yu D, Wang A, Dai Y, Zhou X (2011) Molecular characterization of the microRNA-138-Fos-like antigen 1 (FOSL1) regulatory module in squamous cell carcinoma. J Biol Chem 286(246):40104–40109. https://doi.org/10.1074/jbc.C111.296707
Stinson S, Lackner MR, Adai AT, Yu N, Kim HJ, O’Brien C, Spoerke J, Jhunjhunwala S, Boyd Z, Januario T, Newman RJ, Yue P, Bourgon R, Modrusan Z, Stern HM, Warming S, de Sauvage FJ, Amler L, Yeh RF, Dornan D (2011) miR-221/222 targeting of trichorhinophalangeal 1 (TRPS1) promotes epithelial-to-mesenchymal transition in breast cancer. Sci Signal 4(186):pt5. https://doi.org/10.1126/scisignal.2002258
Wu J, Sun Y, Zhang PY, Qian M, Zhang H, Chen X, Ma D, Xu Y, Chen X, Tang KF (2016) The Fra-1-miR-134-SDS22 feedback loop amplifies ERK/JNK signaling and reduces chemosensitivity in ovarian cancer cells. Cell Death Dis 7(9):e2384. https://doi.org/10.1038/cddis.2016.289
Chiappetta G, Ferraro A, Botti G, Monaco M, Pasquinelli R, Vuttariello E, Arnaldi L, Di Bonito M, D’Aiuto G, Pierantoni GM, Fusco A (2007) FRA-1 protein overexpression is a feature of hyperplastic and neoplastic breast disorders. BMC Cancer 7:17. https://doi.org/10.1186/1471-2407-7-17
Logullo AF, Stiepcich MM, Osorio CA, Nonogaki S, Pasini FS, Rocha RM, Soares FA, Brentani MM (2011) Role of Fos-related antigen 1 in the progression and prognosis of ductal breast carcinoma. Histopathology 58(4):617–625. https://doi.org/10.1111/j.1365-2559.2011.03785.x
Kharman-Biz A, Gao H, Ghiasvand R, Zhao C, Zendehdel K, Dahlman-Wright K (2013) Expression of activator protein-1 (AP-1) family members in breast cancer. BMC Cancer 13:441. https://doi.org/10.1186/1471-2407-13-441
Ma K, Chang D, Gong M, Ding F, Luo A, Tian F, Liu Z, Wang T (2009) Expression and significance of FRA-1 in non-small-cell lung cancer. Cancer Invest 27(3):353–359. https://doi.org/10.1080/07357900802254008
Zhong G, Chen X, Fang X, Wang D, Xie M, Chen Q (2016) Fra-1 is upregulated in lung cancer tissues and inhibits the apoptosis of lung cancer cells by the P53 signaling pathway. Oncol Rep 35(1):447–453. https://doi.org/10.3892/or.2015.4395
Wang HL, Wang J, Xiao SY, Haydon R, Stoiber D, He TC, Bissonnette M, Hart J (2002) Elevated protein expression of cyclin D1 and Fra-1 but decreased expression of c-Myc in human colorectal adenocarcinomas overexpressing beta-catenin. Int J Cancer 101(4):301–310. https://doi.org/10.1002/ijc.10630
Prusty BK, Das BC (2005) Constitutive activation of transcription factor AP-1 in cervical cancer and suppression of human papillomavirus (HPV) transcription and AP-1 activity in HeLa cells by curcumin. Int J Cancer 113(6):951–960. https://doi.org/10.1002/ijc.20668
Xiao S, Zhou Y, Yi W, Luo G, Jiang B, Tian Q, Li Y, Xue M (2015) Fra-1 is downregulated in cervical cancer tissues and promotes cervical cancer cell apoptosis by p53 signaling pathway in vitro. Int J Oncol 46(4):1677–1684. https://doi.org/10.3892/ijo.2015.2873
Hein S, Mahner S, Kanowski C, Loning T, Janicke F, Milde-Langosch K (2009) Expression of Jun and Fos proteins in ovarian tumors of different malignant potential and in ovarian cancer cell lines. Oncol Rep 22(1):177–183
Luo YZ, He P, Qiu MX (2018) FOSL1 enhances growth and metastasis of human prostate cancer cells through epithelial mesenchymal transition pathway. Eur Rev Med Pharmacol 22(24):8609–8615
Mangone FR, Brentani MM, Nonogaki S, Begnami MD, Campos AH, Walder F, Carvalho MB, Soares FA, Torloni H, Kowalski LP, Federico MH (2005) Overexpression of Fos-Related antigen-1 in head and neck squamous cell carcinoma. Int J Exp Pathol 86(4):205–212. https://doi.org/10.1111/j.0959-9673.2005.00423.x
Mishra A, Bharti AC, Saluja D, Das BC (2010) Transactivation and expression patterns of Jun and Fos/AP-1 super-family proteins in human oral cancer. Int J Cancer 126(4):819–829. https://doi.org/10.1002/ijc.24807
Gupta S, Kumar P, Kaur H, Sharma N, Saluja D, Bharti AC, Das BC (2015) Selective participation of c-Jun with Fra-2/c-Fos promotes aggressive tumor phenotypes and poor prognosis in tongue cancer. Sci Rep 5:16811. https://doi.org/10.1038/srep16811
Battista S, de Nigris F, Fedele M, Chiappetta G, Scala S, Vallone D, Pierantoni GM, Mega T, Santoro M, Viglietto G, Verde P, Fusco A (1998) Increase in AP-1 activity is a general event in thyroid cell transformation in vitro and in vivo. Oncogene 17(3):377–385. https://doi.org/10.1038/sj.onc.1201953
Chiappetta G, Tallini G, De Biasio MC, Pentimalli F, de Nigris F, Losito S, Fedele M, Battista S, Verde P, Santoro M, Fusco A (2000) FRA-1 expression in hyperplastic and neoplastic thyroid diseases. Clin Cancer Res 6(11):4300–4306
Kim YH, Oh JH, Kim NH, Choi KM, Kim SJ, Baik SH, Choi DS, Lee ES (2001) Fra-1 expression in malignant and benign thyroid tumor. Korean J Intern Med 16(2):93–97
Serewko MM, Popa C, Dahler AL, Smith L, Strutton GM, Coman W, Dicker AJ, Saunders NA (2002) Alterations in gene expression and activity during squamous cell carcinoma development. Cancer Res 62(13):3759–3765
Sayan AE, Stanford R, Vickery R, Grigorenko E, Diesch J, Kulbicki K, Edwards R, Pal R, Greaves P, Jariel-Encontre I, Piechaczyk M, Kriajevska M, Mellon JK, Dhillon AS, Tulchinsky E (2012) Fra-1 controls motility of bladder cancer cells via transcriptional upregulation of the receptor tyrosine kinase AXL. Oncogene 31(12):1493–1503. https://doi.org/10.1038/onc.2011.336
Bamberger AM, Milde-Langosch K, Rossing E, Goemann C, Loning T (2001) Expression pattern of the AP-1 family in endometrial cancer: correlations with cell cycle regulators. J Cancer Res Clin Oncol 127(9):545–550
Hanson RL, Brown RB, Steele MM, Grandgenett PM, Grunkemeyer JA, Hollingsworth MA (2016) Identification of FRA-1 as a novel player in pancreatic cancer in cooperation with a MUC1: ERK signaling axis. Oncotarget 7(26):39996–40011. https://doi.org/10.18632/oncotarget.9557
Vallejo A, Perurena N, Guruceaga E, Mazur PK, Martinez-Canarias S, Zandueta C, Valencia K, Arricibita A, Gwinn D, Sayles LC, Chuang CH, Guembe L, Bailey P, Chang DK, Biankin A, Ponz-Sarvise M, Andersen JB, Khatri P, Bozec A, Sweet-Cordero EA, Sage J, Lecanda F, Vicent S (2017) An integrative approach unveils FOSL1 as an oncogene vulnerability in KRAS-driven lung and pancreatic cancer. Nat Commun 8:14294. https://doi.org/10.1038/ncomms14294
Gao XQ, Ge YS, Shu QH, Ma HX (2017) Expression of Fra-1 in human hepatocellular carcinoma and its prognostic significance. Tumour Biol 39(6):1010428317709635. https://doi.org/10.1177/1010428317709635
Li L, Zhang W, Zhao S, Sun M (2019) FOS-like antigen 1 is a prognostic biomarker in hepatocellular carcinoma. Saudi J Gastroenterol. https://doi.org/10.4103/sjg.SJG_595_18
He J, Zhu G, Gao L, Chen P, Long Y, Liao S, Yi H, Yi W, Pei Z, Wu M, Li X, Xiang J, Peng S, Ma J, Zhou M, Xiong W, Zeng Z, Xiang B, Tang K, Cao L, Li G, Zhou Y (2015) Fra-1 is upregulated in gastric cancer tissues and affects the PI3K/Akt and p53 signaling pathway in gastric cancer. Int J Oncol 47(5):1725–1734. https://doi.org/10.3892/ijo.2015.3146
Zhu X, Liu H, Xu Z, Zhang Y (2019) Expression and clinical significance of FOS-like antigen 1 in gastric adenocarcinoma. Pathol Res Pract 215(6):152394. https://doi.org/10.1016/j.prp.2019.03.022
Maurus K, Hufnagel A, Geiger F, Graf S, Berking C, Heinemann A, Paschen A, Kneitz S, Stigloher C, Geissinger E, Otto C, Bosserhoff A, Schartl M, Meierjohann S (2017) The AP-1 transcription factor FOSL1 causes melanocyte reprogramming and transformation. Oncogene 36(36):5110–5121. https://doi.org/10.1038/onc.2017.135
Hu YC, Lam KY, Law S, Wong J, Srivastava G (2001) Identification of differentially expressed genes in esophageal squamous cell carcinoma (ESCC) by cDNA expression array: overexpression of Fra-1, Neogenin, Id-1, and CDC25B genes in ESCC. Clin Cancer Res 7(8):2213–2221
Hussain S, Bharti AC, Salam I, Bhat MA, Mir MM, Hedau S, Siddiqi MA, Basir SF, Das BC (2009) Transcription factor AP-1 in esophageal squamous cell carcinoma: alterations in activity and expression during human Papillomavirus infection. BMC Cancer 9:329. https://doi.org/10.1186/1471-2407-9-329
Ramos-Nino ME, Blumen SR, Pass H, Mossman BT (2007) Fra-1 governs cell migration via modulation of CD44 expression in human mesotheliomas. Mol Cancer 6:81. https://doi.org/10.1186/1476-4598-6-81
Wykosky J, Gibo DM, Stanton C, Debinski W (2008) Interleukin-13 receptor alpha 2, EphA2, and Fos-related antigen 1 as molecular denominators of high-grade astrocytomas and specific targets for combinatorial therapy. Clin Cancer Res 14(1):199–208. https://doi.org/10.1158/1078-0432.CCR-07-1990
Han Y, Zhao XY, Sun YF, Sui YT, Liu JG (2019) Effects of FOSL1 silencing on osteosarcoma cell proliferation, invasion and migration through the ERK/AP-1 signaling pathway. J Cell Physiol 234(4):3598–3612. https://doi.org/10.1002/jcp.27048
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66(2):115–132. https://doi.org/10.3322/caac.21338
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. https://doi.org/10.3322/caac.21492
Moquet-Torcy G, Tolza C, Piechaczyk M, Jariel-Encontre I (2014) Transcriptional complexity and roles of Fra-1/AP-1 at the uPA/Plau locus in aggressive breast cancer. Nucleic Acids Res 42(17):11011–11024. https://doi.org/10.1093/nar/gku814
Bakiri L, Macho-Maschler S, Custic I, Niemiec J, Guio-Carrion A, Hasenfuss SC, Eger A, Muller M, Beug H, Wagner EF (2015) Fra-1/AP-1 induces EMT in mammary epithelial cells by modulating Zeb1/2 and TGFbeta expression. Cell Death Differ 22(2):336–350. https://doi.org/10.1038/cdd.2014.157
Annis MG, Ouellet V, Rennhack JP, L’Esperance S, Rancourt C, Mes-Masson AM, Andrechek ER, Siegel PM (2018) Integrin-uPAR signaling leads to FRA-1 phosphorylation and enhanced breast cancer invasion. Breast Cancer Res 20(1):9. https://doi.org/10.1186/s13058-018-0936-8
Oliveira-Ferrer L, Kurschner M, Labitzky V, Wicklein D, Muller V, Luers G, Schumacher U, Milde-Langosch K, Schroder C (2015) Prognostic impact of transcription factor Fra-1 in ER-positive breast cancer: contribution to a metastatic phenotype through modulation of tumor cell adhesive properties. J Cancer Res Clin Oncol 141(10):1715–1726. https://doi.org/10.1007/s00432-015-1925-2
Franco HL, Nagari A, Malladi VS, Li W, Xi Y, Richardson D, Allton KL, Tanaka K, Li J, Murakami S, Keyomarsi K, Bedford MT, Shi X, Li W, Barton MC, Dent SYR, Kraus WL (2018) Enhancer transcription reveals subtype-specific gene expression programs controlling breast cancer pathogenesis. Genome Res 28(2):159–170. https://doi.org/10.1101/gr.226019.117
Serino LTR, Jucoski TS, Morais SB, Fernandes CCC, Lima RS, Urban CA, Cavalli LR, Cavalli IJ, Ribeiro E (2019) Association of FOSL1 copy number alteration and triple negative breast tumors. Genet Mol Biol 42(1):26–31. https://doi.org/10.1590/1678-4685-gmb-2017-0267
He H, Song DD, Sinha I, Hessling B, Li XD, Haldosen LA, Zhao CY (2019) Endogenous interaction profiling identifies DDX5 as an oncogenic coactivator of transcription factor Fra-1. Oncogene 38(28):5725–5738. https://doi.org/10.1038/s41388-019-0824-4
Tolza C, Bejjani F, Evanno E, Mahfoud S, Moquet-Torcy G, Gostan T, Maqbool MA, Kirsh O, Piechaczyk M, Jariel-Encontre I (2019) AP-1 signaling by Fra-1 directly regulates HMGA1 oncogene transcription in triple-negative breast cancers. Mol Cancer Res. https://doi.org/10.1158/1541-7786.MCR-19-0036
Wood SL, Pernemalm M, Crosbie PA, Whetton AD (2015) Molecular histology of lung cancer: from targets to treatments. Cancer Treat Rev 41(4):361–375. https://doi.org/10.1016/j.ctrv.2015.02.008
Elangovan IM, Vaz M, Tamatam CR, Potteti HR, Reddy NM, Reddy SP (2018) FOSL1 promotes Kras-induced lung cancer through amphiregulin and cell survival gene regulation. Am J Respir Cell Mol Biol 58(5):625–635. https://doi.org/10.1165/rcmb.2017-0164OC
Rojewski AM, Zuromski KL, Toll BA (2017) Strategies for smoking cessation among high risk populations to prevent lung cancer. Expert Rev Respir Med 11(2):85–87. https://doi.org/10.1080/17476348.2017.1267571
Vaz M, Rajasekaran S, Potteti HR, Reddy SP (2015) Myeloid-specific Fos-related antigen-1 regulates cigarette smoke-induced lung inflammation, not emphysema, in mice. Am J Respir Cell Mol Biol 53(1):125–134. https://doi.org/10.1165/rcmb.2014-0118OC
Zhang Q, Adiseshaiah P, Reddy SP (2005) Matrix metalloproteinase/epidermal growth factor receptor/mitogen-activated protein kinase signaling regulate fra-1 induction by cigarette smoke in lung epithelial cells. Am J Respir Cell Mol Biol 32(1):72–81. https://doi.org/10.1165/rcmb.2004-0198OC
Zhang Q, Adiseshaiah P, Kalvakolanu DV, Reddy SP (2006) A Phosphatidylinositol 3-kinase-regulated Akt-independent signaling promotes cigarette smoke-induced FRA-1 expression. J Biol Chem 281(15):10174–10181. https://doi.org/10.1074/jbc.M513008200
Hu X, Peng N, Qi F, Li J, Shi L, Chen R (2018) Cigarette smoke upregulates SPRR3 by favoring c-Jun/Fra1 heterodimerization in human bronchial epithelial cells. Future Oncol 14(25):2599–2613. https://doi.org/10.2217/fon-2018-0043
Rubio L, Bach J, Marcos R, Hernandez A (2017) Synergistic role of nanoceria on the ability of tobacco smoke to induce carcinogenic hallmarks in lung epithelial cells. Nanomedicine (Lond) 12(23):2623–2635. https://doi.org/10.2217/nnm-2017-0205
Shukla A, Flanders T, Lounsbury KM, Mossman BT (2004) The gamma-glutamylcysteine synthetase and glutathione regulate asbestos-induced expression of activator protein-1 family members and activity. Cancer Res 64(21):7780–7786. https://doi.org/10.1158/0008-5472.CAN-04-1365
Iskit S, Schlicker A, Wessels L, Peeper DS (2015) Fra-1 is a key driver of colon cancer metastasis and a Fra-1 classifier predicts disease-free survival. Oncotarget 6(41):43146–43161. https://doi.org/10.18632/oncotarget.6454
Diesch J, Sanij E, Gilan O, Love C, Tran H, Fleming NI, Ellul J, Amalia M, Haviv I, Pearson RB, Tulchinsky E, Mariadason JM, Sieber OM, Hannan RD, Dhillon AS (2014) Widespread FRA1-dependent control of mesenchymal transdifferentiation programs in colorectal cancer cells. PLoS ONE 9(3):e88950. https://doi.org/10.1371/journal.pone.0088950
de Wilde J, De-Castro Arce J, Snijders PJ, Meijer CJ, Rosl F, Steenbergen RD (2008) Alterations in AP-1 and AP-1 regulatory genes during HPV-induced carcinogenesis. Cell Oncol 30(1):77–87
Holmes D (2015) Ovarian cancer: beyond resistance. Nature 527(7579):S217. https://doi.org/10.1038/527S217a
Tchernitsa OI, Sers C, Zuber J, Hinzmann B, Grips M, Schramme A, Lund P, Schwendel A, Rosenthal A, Schafer R (2004) Transcriptional basis of KRAS oncogene-mediated cellular transformation in ovarian epithelial cells. Oncogene 23(26):4536–4555. https://doi.org/10.1038/sj.onc.1207585
Mahner S, Baasch C, Schwarz J, Hein S, Wolber L, Janicke F, Milde-Langosch K (2008) C-Fos expression is a molecular predictor of progression and survival in epithelial ovarian carcinoma. Br J Cancer 99(8):1269–1275. https://doi.org/10.1038/sj.bjc.6604650
Oliveira-Ferrer L, Rossler K, Haustein V, Schroder C, Wicklein D, Maltseva D, Khaustova N, Samatov T, Tonevitsky A, Mahner S, Janicke F, Schumacher U, Milde-Langosch K (2014) c-FOS suppresses ovarian cancer progression by changing adhesion. Br J Cancer 110(3):753–763. https://doi.org/10.1038/bjc.2013.774
Hao Y, Zhu L, Yan L, Liu J, Liu D, Gao N, Tan M, Gao S, Lin B (2017) c-Fos mediates alpha1, 2-fucosyltransferase 1 and Lewis y expression in response to TGF-beta1 in ovarian cancer. Oncol Rep 38(6):3355–3366. https://doi.org/10.3892/or.2017.6052
Liu W, Tian T, Liu L, Du J, Gu Y, Qin N, Yan C, Wang Z, Dai J, Fan Z (2017) A functional SNP rs1892901 in FOSL1 is associated with gastric cancer in Chinese population. Sci Rep 7:41737. https://doi.org/10.1038/srep41737
Yang Y, Dong K, Shao S (2019) The effect of Helicobacter pylori on the expression of FRA-1 in gastric epithelial cells and its mechanism. Microb Pathog 129:257–265. https://doi.org/10.1016/j.micpath.2019.02.022
Tsuchiya H, Fujii M, Niki T, Tokuhara M, Matsui M, Seiki M (1993) Human T-cell leukemia virus type 1 Tax activates transcription of the human fra-1 gene through multiple cis elements responsive to transmembrane signals. J Virol 67(12):7001–7007
Nimmakayala RK, Seshacharyulu P, Lakshmanan I, Rachagani S, Chugh S, Karmakar S, Rauth S, Vengoji R, Atri P, Talmon GA, Lele SM, Smith LM, Thapa I, Bastola D, Ouellette MM, Batra SK, Ponnusamy MP (2018) Cigarette smoke induces stem cell features of pancreatic cancer cells via PAF1. Gastroenterology 155(3):892. https://doi.org/10.1053/j.gastro.2018.05.041
Zhang L, Ye DX, Pan HY, Wei KJ, Wang LZ, Wang XD, Shen GF, Zhang ZY (2011) Yes-associated protein promotes cell proliferation by activating Fos Related Activator-1 in oral squamous cell carcinoma. Oral Oncol 47(8):693–697. https://doi.org/10.1016/j.oraloncology.2011.06.003
Xu H, Jin X, Yuan Y, Deng P, Jiang L, Zeng X, Li XS, Wang ZY, Chen QM (2017) Prognostic value from integrative analysis of transcription factors c-Jun and Fra-1 in oral squamous cell carcinoma: a multicenter cohort study. Sci Rep 7(1):7522. https://doi.org/10.1038/s41598-017-05106-5
Krishna A, Bhatt MLB, Singh V, Singh S, Gangwar PK, Singh US, Kumar V, Mehrotra D (2018) Differential expression of c-fos proto-oncogene in normal oral mucosa versus squamous cell carcinoma. Asian Pac J Cancer Prev 19(3):867–874. https://doi.org/10.22034/APJCP.2018.19.3.867
Milde-Langosch K, Bamberger AM, Methner C, Rieck G, Loning T (2000) Expression of cell cycle-regulatory proteins Rb, p16/MTS1, p27/KIP1, p21/WAF1, cyclin D1 and cyclin E in breast cancer: correlations with expression of activating protein-1 family members. Int J Cancer 87(4):468–472
Shukla A, Timblin CR, Hubbard AK, Bravman J, Mossman BT (2001) Silica-induced activation of c-Jun-NH2-terminal amino kinases, protracted expression of the activator protein-1 proto-oncogene, fra-1, and S-phase alterations are mediated via oxidative stress. Can Res 61(5):1791–1795
Casalino L, Bakiri L, Talotta F, Weitzman JB, Fusco A, Yaniv M, Verde P (2007) Fra-1 promotes growth and survival in RAS-transformed thyroid cells by controlling cyclin A transcription. EMBO J 26(7):1878–1890. https://doi.org/10.1038/sj.emboj.7601617
Wang C, Li Z, Shao F, Yang X, Feng X, Shi S, Gao Y, He J (2017) High expression of collagen triple helix repeat containing 1 (CTHRC1) facilitates progression of oesophageal squamous cell carcinoma through MAPK/MEK/ERK/FRA-1 activation. J Exp Clin Cancer Res 36(1):84. https://doi.org/10.1186/s13046-017-0555-8
Ramos-Nino ME, Blumen SR, Sabo-Attwood T, Pass H, Carbone M, Testa JR, Altomare BA, Mossman BT (2008) HGF mediates cell proliferation of human mesothelioma cells through a PI3K/MEK5/Fra-1 pathway. Am J Resp Cell Mol 38(2):209–217. https://doi.org/10.1165/rcmb.2007-0206OC
Dikshit A, Jin YAJ, Degan S, Hwang J, Foster MW, Li CY, Zhang JY (2018) UBE2N promotes melanoma growth via MEK/FRA1/SOX10 signaling. Can Res 78(22):6462–6472. https://doi.org/10.1158/0008-5472.Can-18-1040
Zhao C, Qiao Y, Jonsson P, Wang J, Xu L, Rouhi P, Sinha I, Cao Y, Williams C, Dahlman-Wright K (2014) Genome-wide profiling of AP-1-regulated transcription provides insights into the invasiveness of triple-negative breast cancer. Cancer Res 74(14):3983–3994. https://doi.org/10.1158/0008-5472.CAN-13-3396
Wang X, Xu C, Hua Y, Cheng K, Zhang Y, Liu J, Han Y, Liu S, Zhang G, Xu S, Yang Z (2018) Psoralen induced cell cycle arrest by modulating Wnt/beta-catenin pathway in breast cancer cells. Sci Rep 8(1):14001. https://doi.org/10.1038/s41598-018-32438-7
He WW, Wu YN, Tang XM, Xia Y, He GZ, Min ZQ, Li C, Xiong SQ, Shi Z, Lu YJ, Yuan ZM (2016) HDAC inhibitors suppress c-Jun/Fra-1-mediated proliferation through transcriptionally downregulating MKK7 and Raf1 in neuroblastoma cells. Oncotarget 7(6):6727–6747. https://doi.org/10.18632/oncotarget.6797
Li BB, Wan Z, Huang GL, Huang ZN, Zhang XN, Liao D, Luo SQ, He ZW (2015) Mitogen- and stress-activated Kinase 1 mediates Epstein-Barr virus latent membrane protein 1-promoted cell transformation in nasopharyngeal carcinoma through its induction of Fra-1 and c-Jun genes. BMC Cancer. https://doi.org/10.1186/s12885-015-1398-3
Shirsat NV, Shaikh SA (2003) Overexpression of the immediate early gene fra-1 inhibits proliferation, induces apoptosis, and reduces tumourigenicity of C6 glioma cells. Exp Cell Res 291(1):91–100. https://doi.org/10.1016/S0014-4827(03)00346-X
Kavya K, Kumar MN, Patil RH, Hegde SM, Kiran Kumar KM, Nagesh R, Babu RL, Ramesh GT, Chidananda Sharma S (2017) Differential expression of AP-1 transcription factors in human prostate LNCaP and PC-3 cells: role of Fra-1 in transition to CRPC status. Mol Cell Biochem 433(1–2):13–26. https://doi.org/10.1007/s11010-017-3012-2
Nieto MA, Huang RY, Jackson RA, Thiery JP (2016) Emt: 2016. Cell 166(1):21–45. https://doi.org/10.1016/j.cell.2016.06.028
Henckels E, Prywes R (2013) Fra-1 regulation of Matrix Metallopeptidase-1 (MMP-1) in metastatic variants of MDA-MB-231 breast cancer cells. F1000Res 2:229. https://doi.org/10.12688/f1000research.2-229.v1
Song Y, Qian L, Song S, Chen L, Zhang Y, Yuan G, Zhang H, Xia Q, Hu M, Yu M, Shi M, Jiang Z, Guo N (2008) Fra-1 and Stat3 synergistically regulate activation of human MMP-9 gene. Mol Immunol 45(1):137–143. https://doi.org/10.1016/j.molimm.2007.04.031
Rattanasinchai C, Llewellyn BJ, Conrad SE, Gallo KA (2017) MLK3 regulates FRA-1 and MMPs to drive invasion and transendothelial migration in triple-negative breast cancer cells. Oncogenesis 6(6):e345. https://doi.org/10.1038/oncsis.2017.44
Wang XF, Zhou QM, Lu YY, Zhang H, Huang S, Su SB (2015) Glycyrrhetinic acid potently suppresses breast cancer invasion and metastasis by impairing the p38 MAPK-AP1 signaling axis. Expert Opin Ther Targ 19(5):577–587. https://doi.org/10.1517/14728222.2015.1012156
Jiang P, Gao W, Ma T, Wang R, Piao Y, Dong X, Wang P, Zhang X, Liu Y, Su W, Xiang R, Zhang J, Li N (2019) CD137 promotes bone metastasis of breast cancer by enhancing the migration and osteoclast differentiation of monocytes/macrophages. Theranostics 9(10):2950–2966. https://doi.org/10.7150/thno.29617
Adiseshaiah P, Vaz M, Machireddy N, Kalvakolanu DV, Reddy SP (2008) A Fra-1-dependent, matrix metalloproteinase driven EGFR activation promotes human lung epithelial cell motility and invasion. J Cell Physiol 216(2):405–412. https://doi.org/10.1002/jcp.21410
Roman M, Lopez I, Guruceaga E, Baraibar I, Ecay M, Collantes M, Nadal E, Vallejo A, Cadenas S, Miguel ME, Jang JH, Martin-Uriz PS, Castro-Labrador L, Vilas-Zornoza A, Lara-Astiaso D, Ponz-Sarvise M, Rolfo C, Santos ES, Raez LE, Taverna S, Behrens C, Weder W, Wistuba II, Vicent S, Gil-Bazo I (2019) Inhibitor of differentiation-1 sustains mutant KRAS-driven progression, maintenance, and metastasis of lung adenocarcinoma via regulation of a FOSL1 network. Cancer Res 79(3):625–638. https://doi.org/10.1158/0008-5472.CAN-18-1479
Huang D, Sun W, Zhou Y, Li P, Chen F, Chen H, Xia D, Xu E, Lai M, Wu Y, Zhang H (2018) Mutations of key driver genes in colorectal cancer progression and metastasis. Cancer Metastasis Rev 37(1):173–187. https://doi.org/10.1007/s10555-017-9726-5
Song S, Byrd JC, Mazurek N, Liu K, Koo JS, Bresalier RS (2005) Galectin-3 modulates MUC2 mucin expression in human colon cancer cells at the level of transcription via AP-1 activation. Gastroenterology 129(5):1581–1591. https://doi.org/10.1053/j.gastro.2005.09.002
Andreolas C, Kalogeropoulou M, Voulgari A, Pintzas A (2008) Fra-1 regulates vimentin during Ha-RAS-induced epithelial mesenchymal transition in human colon carcinoma cells. Int J Cancer 122(8):1745–1756. https://doi.org/10.1002/ijc.23309
Sun X, Zhang T, Deng Q, Zhou Q, Sun X, Li E, Yu D, Zhong C (2018) Benzidine induces epithelial-mesenchymal transition of human bladder cancer cells through activation of ERK5 pathway. Mol Cells 41(3):188–197. https://doi.org/10.14348/molcells.2018.2113
Huang XY, Ke AW, Shi GM, Zhang X, Zhang C, Shi YH, Wang XY, Ding ZB, Xiao YS, Yan J, Qiu SJ, Fan J, Zhou J (2013) alphaB-crystallin complexes with 14-3-3zeta to induce epithelial-mesenchymal transition and resistance to sorafenib in hepatocellular carcinoma. Hepatology 57(6):2235–2247. https://doi.org/10.1002/hep.26255
Toyozumi T, Hoshino I, Takahashi M, Usui A, Akutsu Y, Hanari N, Murakami K, Kano M, Akanuma N, Suitoh H, Matsumoto Y, Sekino N, Komatsu A, Matsubara H (2017) Fra-1 regulates the expression of HMGA1, which is associated with a poor prognosis in human esophageal squamous cell carcinoma. Ann Surg Oncol 24(11):3446–3455. https://doi.org/10.1245/s10434-016-5666-5
Lan YY, Hsiao JR, Chang KC, Chang JS, Chen CW, Lai HC, Wu SY, Yeh TH, Chang FH, Lin WH, Su IJ, Chang Y (2012) Epstein-Barr virus latent membrane protein 2A promotes invasion of nasopharyngeal carcinoma cells through ERK/Fra-1-mediated induction of matrix metalloproteinase 9. J Virol 86(12):6656–6667. https://doi.org/10.1128/JVI.00174-12
Dankort D, Curley DP, Cartlidge RA, Nelson B, Karnezis AN, Damsky WE Jr, You MJ, DePinho RA, McMahon M, Bosenberg M (2009) Braf(V600E) cooperates with Pten loss to induce metastatic melanoma. Nat Genet 41(5):544–552. https://doi.org/10.1038/ng.356
Caramel J, Papadogeorgakis E, Hill L, Browne GJ, Richard G, Wierinckx A, Saldanha G, Osborne J, Hutchinson P, Tse G, Lachuer J, Puisieux A, Pringle JH, Ansieau S, Tulchinsky E (2013) A switch in the expression of embryonic EMT-inducers drives the development of malignant melanoma. Cancer Cell 24(4):466–480. https://doi.org/10.1016/j.ccr.2013.08.018
Duong HQ, Nemazanyy I, Rambow F, Tang SC, Delaunay S, Tharun L, Florin A, Buttner R, Vandaele D, Close P, Marine JC, Shostak K, Chariot A (2018) The endosomal protein CEMIP links WNT signaling to MEK1-ERK1/2 activation in Selumetinib-resistant intestinal organoids. Cancer Res 78(16):4533–4548. https://doi.org/10.1158/0008-5472.CAN-17-3149
Endo S, Fujita M, Yamada S, Imadome K, Nakayama F, Isozaki T, Yasuda T, Imai T, Matsubara H (2018) Fra1 enhances the radioresistance of colon cancer cells to Xray or Cion radiation. Oncol Rep 39(3):1112–1118. https://doi.org/10.3892/or.2018.6223
Kajanne R, Miettinen P, Tenhunen M, Leppa S (2009) Transcription factor AP-1 promotes growth and radioresistance in prostate cancer cells. Int J Oncol 35(5):1175–1182. https://doi.org/10.3892/ijo_00000434
Javvadi P, Segan AT, Tuttle SW, Koumenis C (2008) The chemopreventive agent curcumin is a potent radiosensitizer of human cervical tumor cells via increased reactive oxygen species production and overactivation of the mitogen-activated protein kinase pathway. Mol Pharmacol 73(5):1491–1501. https://doi.org/10.1124/mol.107.043554
Tyagi A, Vishnoi K, Kaur H, Srivastava Y, Roy BG, Das BC, Bharti AC (2017) Cervical cancer stem cells manifest radioresistance: association with upregulated AP-1 activity. Sci Rep 7(1):4781. https://doi.org/10.1038/s41598-017-05162-x
Grimm J, Hufnagel A, Wobser M, Borst A, Haferkamp S, Houben R, Meierjohann S (2018) BRAF inhibition causes resilience of melanoma cell lines by inducing the secretion of FGF1. Oncogenesis 7(9):71. https://doi.org/10.1038/s41389-018-0082-2
Kurimchak AM, Shelton C, Duncan KE, Johnson KJ, Brown J, O’Brien S, Gabbasov R, Fink LS, Li Y, Lounsbury N, Abou-Gharbia M, Childers WE, Connolly DC, Chernoff J, Peterson JR, Duncan JS (2016) Resistance to BET bromodomain inhibitors is mediated by kinome reprogramming in ovarian cancer. Cell Rep 16(5):1273–1286. https://doi.org/10.1016/j.celrep.2016.06.091
Kurimchak AM, Shelton C, Herrera-Montavez C, Duncan KE, Chernoff J, Duncan JS (2019) Intrinsic resistance to MEK inhibition through BET protein-mediated kinome reprogramming in NF1-deficient ovarian cancer. Mol Cancer Res 17(8):1721–1734. https://doi.org/10.1158/1541-7786.MCR-18-1332
Zhou W, Sun W, Yung MMH, Dai S, Cai Y, Chen CW, Meng Y, Lee JB, Braisted JC, Xu Y, Southall NT, Shinn P, Huang X, Song Z, Chen X, Kai Y, Cai X, Li Z, Hao Q, Cheung ANY, Ngan HYS, Liu SS, Barak S, Hao J, Dai Z, Tzatsos A, Peng W, Pei H, Han Z, Chan DW, Zheng W, Zhu W (2018) Autocrine activation of JAK2 by IL-11 promotes platinum drug resistance. Oncogene 37(29):3981–3997. https://doi.org/10.1038/s41388-018-0238-8
Zanellato I, Colangelo D, Osella D (2018) JQ1, a BET inhibitor, synergizes with cisplatin and induces apoptosis in highly chemoresistant malignant pleural mesothelioma cells. Curr Cancer Drug Targ 18(8):816–828. https://doi.org/10.2174/1568009617666170623101722
Cho H, Matsumoto S, Fujita Y, Kuroda A, Menju T, Sonobe M, Kondo N, Torii I, Nakano T, Lara PN, Gandara DR, Date H, Hasegawa S (2017) Trametinib plus 4-methylumbelliferone exhibits antitumor effects by ERK blockade and CD44 downregulation and affects PD-1 and PD-L1 in malignant pleural mesothelioma. J Thorac Oncol 12(3):477–490. https://doi.org/10.1016/j.jtho.2016.10.023
Yang W, Meng L, Chen K, Tian C, Peng B, Zhong L, Zhang C, Yang X, Zou J, Yang S, Li L (2019) Preclinical pharmacodynamic evaluation of a new Src/FOSL1 inhibitor, LY-1816, in pancreatic ductal adenocarcinoma. Cancer Sci 110(4):1408–1419. https://doi.org/10.1111/cas.13929
Dhillon AS, Tulchinsky E (2015) FRA-1 as a driver of tumour heterogeneity: a nexus between oncogenes and embryonic signalling pathways in cancer. Oncogene 34(34):4421–4428. https://doi.org/10.1038/onc.2014.374
Lau EY, Lo J, Cheng BY, Ma MK, Lee JM, Ng JK, Chai S, Lin CH, Tsang SY, Ma S, Ng IO, Lee TK (2016) Cancer-associated fibroblasts regulate tumor-initiating cell plasticity in hepatocellular carcinoma through c-Met/FRA1/HEY1 signaling. Cell Rep 15(6):1175–1189. https://doi.org/10.1016/j.celrep.2016.04.019
Racca AC, Prucca CG, Caputto BL (2019) Fra-1 and c-Fos N-terminal deletion mutants impair breast tumor cell proliferation by blocking lipid synthesis activation. Front Oncol 9:544. https://doi.org/10.3389/fonc.2019.00544
Wang Q, Ni H, Lan L, Wei X, Xiang R, Wang Y (2010) Fra-1 protooncogene regulates IL-6 expression in macrophages and promotes the generation of M2d macrophages. Cell Res 20(6):701–712. https://doi.org/10.1038/cr.2010.52
Luo YP, Zhou H, Krueger J, Kaplan C, Liao D, Markowitz D, Liu C, Chen T, Chuang TH, Xiang R, Reisfeld RA (2010) The role of proto-oncogene Fra-1 in remodeling the tumor microenvironment in support of breast tumor cell invasion and progression. Oncogene 29(5):662–673. https://doi.org/10.1038/onc.2009.308
Guo Z, Song J, Hao J, Zhao H, Du X, Li E, Kuang Y, Yang F, Wang W, Deng J, Wang Q (2019) M2 macrophages promote NSCLC metastasis by upregulating CRYAB. Cell Death Dis 10(6):377. https://doi.org/10.1038/s41419-019-1618-x
Obenauf AC, Zou Y, Ji AL, Vanharanta S, Shu W, Shi H, Kong X, Bosenberg MC, Wiesner T, Rosen N, Lo RS, Massague J (2015) Therapy-induced tumour secretomes promote resistance and tumour progression. Nature 520(7547):368–372. https://doi.org/10.1038/nature14336
Kong X, Kuilman T, Shahrabi A, Boshuizen J, Kemper K, Song JY, Niessen HWM, Rozeman EA, Geukes Foppen MH, Blank CU, Peeper DS (2017) Cancer drug addiction is relayed by an ERK2-dependent phenotype switch. Nature 550(7675):270–274. https://doi.org/10.1038/nature24037
Hong A, Moriceau G, Sun L, Lomeli S, Piva M, Damoiseaux R, Holmen SL, Sharpless NE, Hugo W, Lo RS (2018) Exploiting drug addiction mechanisms to select against MAPKi-resistant melanoma. Cancer Discov 8(1):74–93. https://doi.org/10.1158/2159-8290.CD-17-0682
Funding
The work was supported by the Project of Hunan Provincial Natural Science Foundation (No. 2018JJ3782); and Changsha Science and Technology Bureau Foundation (No. kq1701085); and the Fundamental Research Funds for the Central Universities of Central South University (Nos. 2018zzts937 and 2018zzts952); and Graduate research project of central south university (No. 2018-75)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.