Abstract
Paeonia ostii seeds have recently been identified as a new source of α-linolenic acid in China. Studying the gene expression patterns of unsaturated fatty acid-related genes would be helpful for understanding the mechanism of α-linolenic acid accumulation. Quantitative real-time polymerase chain reaction (qRT-PCR) is a useful method for reliably evaluating gene expression, and it is necessary to select reliable reference genes for data normalization in qRT-PCR analysis. In this study, we evaluated the expression stability of 12 candidate reference genes using four mathematical algorithms (∆Ct, BestKeeper, NormFinder, and geNorm). The web-based tool RefFinder was used to integrate the results and to provide a comprehensive ranking order. The expression stability ranking orders of reference genes were different caculated by these four algorithms, and the ranking order analyzed by the RefFinder was UBQ > Tip41 > UCE > EF-1α > α-TUB > PP2A > ACT > GAPDH > SAM > CYP > β-TUB > 18S at the different seed development stages, and UBQ > Tip41 > EF-1α > α-TUB > PP2A > UCE > GAPDH > SAM > ACT > CYP > 18S > β-TUB in P. ostii tissues. UBQ and Tip41 are the two most stable whereas 18S and β-TUB are the two least stable reference genes for gene expression in various tissues and seeds at different developmental stages in P. ostii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tree peony (Paeonia section Moutan DC.), which is a perennial deciduous shrub native to China, is a well-known horticultural and medicinal plant [1]. In 2011, seeds of Paeonia ostii (P. ostii), which belong to a member of tree peony, were identified as novel sources of edible plant oil in China. The seed oil of this species is characterized by abundant unsaturated fatty acids (UFAs) and a high proportion of α-linolenic acid (ALA), which is a type of n-3 fatty acids with many health benefits [2, 3]. The UFA content in P. ostii seed oil exceeds 90% and contains approximately 42.78% ALA, 26.50% linoleic acid (LA) and 22.97% oleic acid (OA) [3, 4]. Consequently, P. ostii seed oil is recognized as an excellent health-promoting edible oil that has great potential economic value. To understand the molecular mechanism of UFA biosynthesis (especially ALA biosynthesis) in P. ostii seeds, transcriptomic analysis was performed during seed development, and numerous key candidate genes associated with UFA biosynthesis were identified [2]. Investigating the expression profiles of these candidate genes will greatly facilitate the understanding of gene function in seed oil biosynthesis.
Quantitative real-time polymerase chain reaction (qRT-PCR) has become the preferred method to quantify transcript abundance because of its high sensitivity, accuracy, specificity and reliability [5, 6]. However, its accuracy is strongly dependent on the constant transcriptional performance of reference genes (RGs), which are often referred to as housekeeping genes and are required for stable expression regardless of experimental conditions [7, 8]. A number of housekeeping genes, which typically encode proteins responsible for the maintenance of basic cellular structures and metabolism, such as glyceraldehyde-3-phosphate dehydrogenase (GAPDH), actin (ACT), 18S ribosomal RNA (18S), elongation factor 1 alpha (EF-1α), ubiquitin-conjugating enzyme (UCE) and ubiquitin (UBQ), are common RGs used for gene expression analysis in many plant species [2, 8,9,10]. However, recent studies have found that there are expression variabilities of these RGs in different tissues, development stages and environmental conditions [7, 8, 11,12,13,14]. There are no universal RGs suitable for all experimental conditions. For example, ACT6, ACT8, and ACT7 were the best RGs for kumquat under salt stress, drought stress and heavy metal stress conditions, respectively [11]. Protein phosphatase 2A subunit A3 (PP2AA3) and 18S were the most stably expressed genes in tea plant (Camellia sinensis) exposed to metal stress [13]. ACT and UCE were the most suitable RGs for normalization in Sacha inchi during the entire growth cycle [12]. Thus, it is important to select reliable RGs as internal controls under individual experimental condition to ensure reproducible and precise gene expression quantification.
In the last two decades, several prevalent mathematical algorithms have been used to identify appropriate RGs for qRT-PCR under different experimental conditions, such as the delta cycle threshold (ΔCt) method [15], BestKeeper [16], geNorm [17], NormFinder [18], and RefFinder [19]. These methods have been used to evaluate candidate RGs in different plant species across different developmental stages, tissues and organs, biotic and abiotic stresses, including Arabidopsis, rice, potato, papaya, tea plant, and Sacha inchi, among others [8, 12,13,14, 20,21,22]. A previous study found that GAPDH and UCE were the two most stable RGs during flower development in ‘Fengdan’, which was the cultivated type originated from P. ostii [23]; however, suitable RGs during seed development and in various vegetative and reproductive tissues have not been identified for normalization of the expression levels of the genes specially involved in seed oil biosynthesis. In this study, we selected 12 candidate RGs (UBQ, 18S, ACT, α-tubulin (α-TUB), β-tubulin (β-TUB), UCE, GAPDH, Protein phosphatase 2A (PP2A), Tip41-like protein (Tip41), S-adenosylmethionine synthetase (SAM), EF-1α, Cyclophilin (CYP) and evaluated the stability of these 12 genes to identify the most suitable RGs for qRT-PCR data normalization in different tissues and seeds at different developmental stages. The reliability of the identified RGs was further validated by analyzing the expression pattern of fatty acid desaturase 2 (FAD2) gene during seed development and in different tissues in P. ostii using the stable and unstable genes for normalization. The results may be useful for selecting reliable RGs suitable for the accurate quantification of gene expression and gene function research in P. ostii.
Materials and methods
Plant materials
Five-year-old P. ostii plants were grown in a field at Zhoukou Normal University, Zhoukou City, Henan Province, China. The roots, stems, leaves, flowers, fruits (5 days after pollination) and seeds of three biological replicates of P. ostii plants were collected for experiments. The seeds at different developmental stages were collected at intervals of 10 days after pollination (DAP) until 120 DAP (containing mature seeds). All tissues prepared for qRT-PCR were immediately frozen in liquid N2 and stored at − 80 °C until use.
RNA extraction and cDNA synthesis
Total RNA was extracted from different P. ostii tissues using a TIANGEN RNA Prep Pure Plant kit (Tiangen Biotech Co. Ltd, Beijing, China) following the manufacturer’s instructions. The RNA integrity, concentration and purity were examined using 1.2% agarose gel electrophoresis and a NanoDrop-2000C spectrophotometer (Thermo Fisher Scientific, USA). Isolated RNA samples, of which the A260/A280 ratio was between 1.9 and 2.1, indicating high purity and no protein contamination, were used for further analysis. First-strand cDNA was synthesized with 0.7 μg total RNA in a 20 μL reaction volume using the PrimeScript® RT Reagent Kit with gDNA Eraser (TAKARA, Dalian, China) according to the manufacturer’s instructions. All cDNA samples were diluted (1:5) with RNase-free water prior to use in qRT-PCR.
Selection of candidate reference genes and qRT-PCR primer design
A total of 12 P. ostii housekeeping genes were selected as candidate RGs for analysis in this study (Table 1). The cDNA sequences of these RGs were obtained from the National Center for Biotechnology Information (NCBI) nucleotide database (http://www.ncbi.nlm.nih.gov/nucleotide) from Paeonia suffruticosa (P. suffruticosa) and a transcriptome data (SRA accession number: SRP051810) from P. ostii submitted by Li et al. [2], and the gene information is shown in Table 1. The primer pairs used for qRT-PCR were designed based on selected sequences of the 12 candidate RGs using Primer Premier 5.0 and are listed in Table 1. To verify the sequences of the RGs, all the amplicons of the 12 candidate genes were analyzed on 1.5% agarose gels and sequenced.
qRT-PCR assay and data analysis
qRT-PCR was conducted using SYBR Premix Ex Taq (TaKaRa, Japan) on a CFX96 Real-Time System (Bio-Rad, USA). The reaction mixture (20 μL) contained 2.0 μL diluted cDNA, 10 μL SYBR Premix Ex Taq, 0.5 μL of each of the forward and reverse primers (10 μM), and 7 μL PCR-grade water. The PCR program was as follows: 95 °C for 2 min; 40 cycles of 95 °C for 10 s, 58 °C for 15 s, and 72 °C for 15 s. The final melting curve was obtained between 65 and 95 °C to verify primer specificity. Each assay was carried out with three biological replicates and three technical replicates. To determine the amplification efficiency of each pair of primers, standard curves were generated from fivefold serial dilutions of cDNA samples from the seeds at 10 DAP. The values of PCR amplification efficiencies, slopes and correlation coefficients were obtained from the standard curves and were calculated using CFX Manager™ software. The quantification cycle (Cq) values were also generated by this software and were used for the gene expression levels.
Four statistical algorithm programs, i.e. ΔCt [15], BestKeeper [16], geNorm [17], and NormFinder [18] were used to evaluate the stability of the candidate genes during seed development. A comparative web-based method, RefFinder (http://150.216.56.64/referencegene), was used to compare and integrate the comprehensive ranking order of the tested candidate RGs.
To verify the expression stability of the recommended candidate RGs, a fatty acid desaturase 2 gene in P. ostii (PoFAD2), which is considered to be one of the key genes involved in UFA biosynthesis, was used as a target gene in the seeds at different stages and diverse tissue samples [2, 24]. The nucleotide sequence of PoFAD2 was obtained from the NCBI database and the primers used in qRT-PCR were designed with the software Primer Premier 5.0 and are listed in Table 1. The qRT-PCR amplification conditions were the same as described above. The two most and least stable RGs, as determined by RefFinder, either alone or in combination (calculated by geometric mean) were used to normalize the expression level of PoFAD2, and the relative expression level of PoFAD2 was calculated using the 2−ΔΔCt method [25].
Results
PCR amplification efficiency and specificity
Twelve P. ostii housekeeping genes, which were obtained from a transcriptome data from P. ostii and the NCBI nucleotide database from P. suffruticosa, were selected as candidate RGs. Alignment results showed that all the RGs shared the identical sequences between P. ostii and P. suffruticosa. All the amplicons of the 12 candidate RGs were also sequenced, and we received the same results between sequences obtained from the test, the sequences from transcriptome data and the sequences in P. suffruticosa. These results indicated that the selected RGs among Paeonia species might very conservative. The gene names, abbreviations, accession numbers, primer sequences and amplicon length, amplification efficiency, correlation coefficient and melting temperature (Tm) values of the 12 candidate RGs are listed in Table 1. The amplicon size ranged from 110 to 210 bp, and the amplification efficiency of each primer pair varied between 98.6% for the 18S gene and 103.9% for UCE. In addition, all correlation coefficient values exceeded 0.99, and the Tm values ranged from 80 °C for CYP to 85 °C for β-TUB. Moreover, the melting curves for the amplified products of all 12 candidate RGs showed a single peak without peaks originated from the formation of primer dimers and amplification of nonspecific PCR products, corresponding to a specific melting temperature (Fig. 1).
Candidate reference gene expression profiles
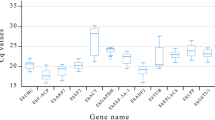
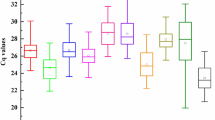
The expression levels of candidate RGs during seed development and in various tissues in P. ostii were determined according to the Cq values obtained by qRT-PCR, and the mean Cq values of these RGs ranged from 10.02 to 33.98 (Table S1, Fig. 2). A low Cq value indicated a high gene expression level, and a high Cq value meaned a low gene expression level. There were wide differences in the transcript levels among these RGs. 18S and ACT were the most and least abundant transcripts in the tested P. ostii tissues, with Cq values ranging from 10.02 to 15.76, and 26.12 to 33.98, respectively (Table S1). The coefficients of variation (CV) of the 12 RGs in all samples were 2.68% (UBQ), 10.13% (18S), 5.73% (ACT), 5.86% (α-TUB), 7.12% (β-TUB), 5.33% (UCE), 7.10% (GAPDH), 5.57% (PP2A), 3.06% (Tip41), 6.64% (SAM), 4.31% (EF-1α) and 6.20% (CYP).
The quantification cycle (Cq) values of the 12 candidate reference genes in seed samples (a) and all tissue samples (b). Lines across the box-plot graph of the Cq values represent the median value. The boxes indicate the 25th and 75th percentiles, and the whisker caps represent the maximum and minimum values
Stability analysis and ranking of reference genes
Four statistical algorithms were used to select the most suitable RGs during seed development and in different tissues of P. ostii, and the stability values of 12 RGs in seeds at different developmental stages and across various tissues were shown in Supplementary Tables S2 and S3, respectively. The two most stable RGs recommended by the four programs were consistent, whereas differences were observed for the least stable RGs (Table 2, Supplementary Tables S2 and S3). The ΔCt method evaluates the expression stability of candidate RGs based on the mean of standard deviation (SD), and the RG with the lowest SD is assumed to be the most stable gene. The BestKeeper program evaluates gene expression stability based on the CV and SD of the Cq values, and the most stable genes have the lowest CV and SD. NormFinder generates the stability values of RGs based on the inter- and intra-group variation in expression level, and lower values indicate higher stability. GeNorm software is a visual basic application tool to select an ideal pair of RGs and creates a stability ranking via the stepwise exclusion of the least stable genes. The stability value (M) of an RG is calculated based on the average pairwise variation between all RGs tested, and is inversely correlated with their exssion stability. Based on ΔCt, NormFinder, GeNorm analysis, UBQ and Tip41 were the two most stable RGs during P. ostii seed development and in all tissues, while 18S and β-TUB were identified as the least most stable RGs (Table 2, Supplementary Tables S2 and S3). Based on BestKeeper analysis, as shown in Table 2, Supplementary Tables S2 and S3, UBQ and Tip 41 were ranked as the two most stable RGs in P. ostii seed and in all tissues, which was consistent with the results from the three other software analyses. However, CYP and β-TUB exhibited low stability and were ranked as the least stable RGs, which was different from the results analyzed by the three other algorithms. GeNorm program was also used to analyze pairwise variation (Vn/Vn + 1) for the assessment of the optimal number of RGs required for reliable normalization. A threshold Vn/Vn + 1 value of 0.15 was adopted to determine whether it is necessary to introduce n + 1 RGs as the internal control. As shown in Supplementary Fig. S1, values (V2/3) lower than the recommended threshold of 0.15 indicated that two RGs were sufficient for the normalization of gene expression during seed development and across diverse tissues in P. ostii.
Finally, the RefFinder tool was used to integrate the above mentioned statistical algorithms and calculate the recommended comprehensive ranking order. The comprehensive ranking order is shown in Table 2, and the ranking order of expression stability was UBQ > Tip41 > UCE > EF-1α > α-TUB > PP2A > ACT > GAPDH > SAM > CYP > β-TUB > 18S at the different seed development stages, and UBQ > Tip41 > EF-1α > α-TUB > PP2A > UCE > GAPDH > SAM > ACT > CYP > 18S > β-TUB in Paeonia ostii tissues. The result analyzed by RefFinder was similar to those of the ΔCt method, geNorm and NormFinder algorithm, but there were some differences in the orders of the middle ranking RGs.
Validation of selected reference genes
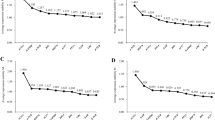
To assess the reliability of the selected RGs, the relative expression pattern of the target gene, PoFAD2, which encodes the enzyme that is essential for UFA biosynthesis in P. ostii, was calculated in seeds at different developmental stages and various tissue samples by qRT-PCR. The primer specificity of PoFAD2 was verified as described for RGs (Table 1). The two most stable RGs (UBQ and Tip41) and the two least stable RGs (18S and β-TUB) identified by RefFinder were used either independently or in combination to normalize the expression levels of PoFAD2 (Fig. 3). Irrespective of normalization by UBQ or Tip41, alone or in combination, approximately the same expression patterns of PoFAD2 were obtained (Fig. 3a). PoFAD2 was predominantly expressed in seeds at the 10–100 DAP stages with the highest expression level at the 70 DAP stage. For other different tissues samples except seed samples, PoFAD2 was found to be mainly expressed in flowers. However, the expression patterns of PoFAD2 were different when using 18S or β-TUB alone or in combination as RGs for normalization (Fig. 3b). During seed development in P. ostti, the highest expression level of PoFAD2 appeared at the 80 DAP stage using β-TUB as the RG for normalization, and at the 90 DAP stage using either 18S alone or in combination with β-TUB for normalization, respectively (Fig. 3b). In addition, the expression pattern of PoFAD2 in various tissues was different when using these two least stable RGs alone or in combination for normalization (Fig. 3b). In addition, when stable RGs were used for normalization, significant differences in the expression levels of PoFAD2 were detected in seeds at 40–90 DAP in comparison with the expression levels in roots, which was used as a calibrator (Fig. 3a). On the other hand, significant differences (P < 0.001, P < 0.01) and no significant difference (P > 0.05) in PoFAD2 gene expression were found between roots and seeds at 40 DAP when 18S, 18S + β-TUB and β-TUB used for normalization, respectively. These results revealed that significant differences in the PoFAD2 expression profiles depending on the RGs used for normalization of the qPCR data, whether stable or unstable. Taken together, these results demonstrate that choosing reliable RGs is important for accurate normalization in qRT-PCR experiments.
Relative expression of the fatty acid desaturase 2 gene in diverse tissue samples using selected reference genes, including the most (a) and least (b) stable reference genes, for normalization in Paeonia ostii. The seed samples were collected at different developmental stages (10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 110 and 120 days after pollination). Each data point represents the mean ± SD of three replicates. Student’s t test was used to determine significant differences from root (calibrator). Significance level: **P < 0.01, ***P < 0.001
Discussion
The accuracy and reliability of gene expression patterns using qRT-PCR technology strongly depend on the use of stable RGs for normalization, which can remove non-biological variations [11, 26]. Previous studies revealed that there were no universal RGs among different plant species, and common RGs, such as ACT, GAPDH and 18S, exhibited expression variation in different plants under different developmental or experimental conditions [12, 27]. Therefore, the selection of reliable RGs for specific developmental or experimental conditions is important in qRT-PCR analysis.
The seed of P. ostii is considered as a new source of ALA in oil production, and studying the mechanism of lipid biosynthesis and the accumulation of ALA during P. ostii seed development has recently received considerable attention. However, suitable RGs in P. ostii seed development have not been identified. In this study, we selected 12 candidate RGs to evaluate their expression stability during seed development and also calculate its stability in different tissues in P. ostii. Four statistical methods (ΔCt, BestKeeper, NormFinder, and geNorm) were used to calculate the stability of gene expression levels. In addition, a web-based tool (RefFinder) was used to integrate the results of these four algorithms and to provide a comprehensive ranking order of all 12 RGs. All four algorithms revealed that UBQ and Tip41 were the two most stable genes among these 12 candidate RGs, but the least stable genes analyzed by these four methods differed (Table 2). The most stable RG ranked by RefFinder was also UBQ (Table 2), which is involved in protein degradation and was also identified as the most stable RG in Platycladus orientalis under NaCl and ABA treatment [28], across different tissues of Rhododendron micranthum Turcz [29], and across all organs and treatments of white clover [26]. However, UBQ is also considered to be inappropriate RG in some plant species. For example, UBQ was unsuitable for normalization in Suaeda aralocaspica under different salt treatments [27]. The 18S gene, which forms part of the eukaryotic ribosome and has been exploited as a reference gene for a long time, was calculated to be the least stable RG by ΔCt, NormFinder, geNorm and RefFinder (Table 2). Previous studies also found that the 18S gene was the worst RG for normalizing gene expression levels in many plant organs and under different conditions, such as Lilium davidii [30], Sacha inchi [12], Chinese tallow [31], Oxytropis ochrocephala Bunge [32] and rubber trees [33]. In contrast, 18S has been demonstrated to be a stable RG in strawberry [34] and rice [21]. In previous study, Tip41, one of the most stable RGs in our study, also showed high stability in the different cultivars of tree peony during flower development, and β-TUB was identified as one of the least stable RG [23], which was similar to the results in our study. It indicated the stability of Tip41 and β-TUB might display consistent trends among Paeonia section Moutan DC cultivars and tissues. These results prove that the most suitable RGs differ among different plant cultivars, plant tissues and under different conditions and it is very important to select reliable RGs according to the specific experiment.
To test the reliability of the selected RGs, the expression pattern of PoFAD2, which encodes a key enzyme in UFA synthesis, was evaluated in this study. The PoFAD2 expression patterns were similar when the gene expression data were normalized using the most stably expressed RGs (UBQ and Tip41) either alone or in combination, with the highest expression level of PoFAD2 appearing at the 70 DAP stage (Fig. 3a). However, the expression patterns of PoFAD2 differed using the least stable RGs (Fig. 3b). For example, the highest expression level of PoFAD2 was observed at the 80 DAP stage using β-TUB as the RG for normalization, and at the 90 DAP stage using either 18S alone or in combination with β-TUB for normalization, respectively. Interestingly, our results showed significant differences in the PoFAD2 expression profiles depending on the RGs used for normalization, whether stable or unstable. The main divergences were observed in the sample of seeds at 40 DAP in comparison with the expression levels in roots (Fig. 3). Therefore, the preliminary requirement for obtaining accurate relative gene expression levels is the use of suitable RGs.
In conclusion, this study evaluated the expression stability and determined the comprehensive rankings of 12 candidate RGs during P. ostii seed development and is diverse tissue samples using four statistical algorithms (ΔCt, BestKeeper, NormFinder, and geNorm) and one web-based method (RefFinder). UBQ and Tip41 were identified as the two most reliable RGs for qRT-PCR, whereas the least stable RGs were 18S and β-TUB. The results obtained in this study provide useful information for accurate qRT-PCR data normalization in P. ostii gene expression studies.
References
Chen FY, Li JJ (1998) Exportation of Chinese tree peonies (Mudan) and their development in other countries I: cultivated. J Northwest Norm Univ 34:109–116
Li SS, Wang LS, Shu QY, Wu J, Chen LG, Shao S, Yin DD (2015) Fatty acid composition of developing tree peony (Paeonia section Moutan DC.) seeds and transcriptome analysis during seed development. BMC Genomics 16:208
Li XQ, Han JG, Liu Z, Liu QH, Hu YH (2014) Economic characteristics investigation and seed oil fatty acid composition analysis of Paeonia ostii plants in different areas. Cereals Oils 27:43–46
Qi JC, Zhou HM, Ma JQ, Li P (2005) Analysis of the chemical constituents in peony seed oil by GC-MS. J Cereals Oils 11:22–23
Bustin SA, Benes V, Nolan T, Pfaffl MW (2005) Quantitative real-time RT-PCR—a perspective. J Mol Endocrinol 34:597–601
Gachon C, Mingam A, Charrier B (2004) Real-time PCR: what relevance to plant studies? J Expl Bot 55:1445–1454
Kanakachari M, Solanke AU, Prabhakaran N, Ahmad I, Dhandapani G, Jayabalan N, Kumar PA (2016) Evaluation of suitable reference genes for normalization of qPCR gene expression studies in Brinjal (Solanum melongena L.) during fruit developmental stages. Appl Biochem Biotechnol 178:433–450
Nicot N, Hausman JF, Hoffmann L, Evers D (2005) Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot 56:2907–2914
Ute A, Kimd B (2008) Gene expression in Citrus sinensis (L.) Osbeck following infection with the bacterial pathogen Candidatus Liberibacter asiaticus causing Huanglongbing in Florida. Plant Sci 175:291–306
Wang X, Liu A (2014) Expression of genes controlling unsaturated fatty acids biosynthesis and oil deposition in developing seeds of Sacha inchi (Plukenetia volubilis L.). Lipids 49:1019–1031
Hu Y, Chen H, Luo C, Dong L, Zhang SW, He XH, Huang GX (2014) Selection of reference genes for real-time quantitative PCR studies of kumquat in various tissues and under abiotic stress. Sci Hortic 174:207–216
Niu LJ, Tao YB, Chen MS, Fu QT, Li CQ, Dong YL, Wang XL, He HY, Xu ZF (2015) Selection of reliable reference genes for gene expression studies of a promising oilseed crop, Plukenetia volubilis, by real-time quantitative PCR. Int J Mol Sci 16:12513–12530
Wang ML, Li QH, Xin HH, Chen X, Zhu XJ, Li XH (2017) Reliable reference genes for normalization of gene expression data in tea plants (Camellia sinensis) exposed to metal stresses. PLoS ONE 12:e0175863
Zhu XY, Li XP, Chen WX, Chen JY, Lu WJ, Lei C, Fu DW (2012) Evaluation of new reference genes in papaya for accurate transcript normalization under different experimental conditions. PLoS ONE 7:e44405
Silver N, Best S, Jiang J, Thein SL (2006) Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol Biol 7:33
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol Lett 26:509–515
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, Paepe AD, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. https://doi.org/10.1186/gb-2002-3-7-research0034
Andersen CL, Jensen JL, Orntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250
Xie F, Xiao P, Chen D, Xu L, Zhang B (2012) miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol Biol 80:75–84
Hofmann J, Grundler FMW (2007) Identification of reference genes for qRT-PCR studies of gene expression in giant cells and syncytia induced in Arabidopsis thaliana by Meloidogyne incognita and Heterodera schachtii. Nematology 9:317–323
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Bioph Res Commun 345:646–651
Pabuayon IM, Yamamoto N, Trinidad JL, Longkumer T, Raorane ML, Kohli A (2016) Reference genes for accurate gene expression analyses across different tissues, developmental stages and genotypes in rice for drought tolerance. Rice 9:32
Li J, Han JG, Hu YH, Yang J (2016) Selection of reference genes for quantitative real-time PCR during flower development in tree peony (Paeonia suffruticosa Andr.). Front Plant Sci 7:516
Okuley J, Lightner J, Feldmann K, Yadav N, Lark E, Browse J (1994) Arabidopsis FAD2 gene encodes the enzyme that is essential for polyunsaturated lipid synthesis. Plant Cell 6:147–158
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Narancio R, John U, Mason J, Spangenberg G (2018) Selection of optimal reference genes for quantitative RT-PCR transcript abundance analysis in white clover (Trifolium repens L.). Funct Plant Biol 45:737–744
Cao J, Wang L, Lan H (2016) Validation of reference genes for quantitative RT-PCR normalization in Suaeda aralocaspica, an annual halophyte with heteromorphism and C4 pathway without Kranz anatomy. PeerJ 4:e1697
Chang E, Shi SQ, Liu JF, Cheng TL, Xue L, Yang XY, Yang WJ, Lan Q, Jiang ZP (2012) Selection of reference genes for quantitative gene expression studies in Platycladus orientalis (Cupressaceae) using real-time PCR. PLoS ONE 7:e33278
Yi SJ, Qian YQ, Han L, Sun ZY, Fan CM, Liu JX, Ju GS (2012) Selection of reliable reference genes for gene expression studies in Rhododendron micranthum Turcz. Sci Hortic 138:128–133
Li XY, Cheng JY, Zhang J, da Silva JAT, Wang CX, Sun HM (2015) Validation of reference genes for accurate normalization of gene expression in Lilium davidii var. unicolor for real time quantitative PCR. PLoS ONE 10:e0141323
Chen X, Mao YJ, Huang SW, Ni J, Lu WL, Hou JY, Wang YT, Zhao WW, Li MH, Wang QJ (2017) Selection of suitable reference genes for quantitative real-time PCR in Sapium sebiferum. Front Plant Sci 8:637
Zhuang H, Fu Y, He W, Wang L, Wei Y (2015) Selection of appropriate reference genes for quantitative real-time PCR in Oxytropis ochrocephala Bunge using transcriptome datasets under abiotic stress treatments. Front Plant Sci 6:475
Long X, He B, Gao X, Qin Y, Yang J, Fang Y, Qi J, Tang C (2015) Validation of reference genes for quantitative real-time PCR during latex regeneration in rubber tree. Gene 563:190–195
Galli V, Borowski JM, Perin EC, Messias RD, Labonde J, Pereira ID, Silva SD, Rombaldi CV (2015) Validation of reference genes for accurate normalization of gene expression for real time-quantitative PCR in strawberry fruits using different cultivars and osmotic stresses. Gene 554:205–214
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31500534), the Startup Fund for Advanced Talents of Zhoukou Normal University (ZKNU2014108), the School-Based Program of Zhoukou Normal University (ZKUNB115203), the Natural Science Foundation of Henan province (162300410346), and the Key Scientific Research Project in Colleges and Universities of Henan Province (15A180024).
Author information
Authors and Affiliations
Contributions
CL and LH designed research; CL, LH, XW, HL, HT and JW performed experiments. CL and LH wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Electronic supplementary material 1 (TIFF 1728 kb)
Fig. S1 Pairwise variation analysis by geNorm to determine the optimal number of reference genes needed for accurate normalization in seeds at different developmental stages and across various tissues in Paeonia ostii. Cutoff value = 0.15
Rights and permissions
About this article
Cite this article
Li, C., Hu, L., Wang, X. et al. Selection of reliable reference genes for gene expression analysis in seeds at different developmental stages and across various tissues in Paeonia ostii. Mol Biol Rep 46, 6003–6011 (2019). https://doi.org/10.1007/s11033-019-05036-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-05036-7