Abstract
Scutellaria baicalensis Georgi is a famous medicinal plant with its dried roots having been used as a traditional Chinese medicinal for more than 2000 years. Although its genome sequence has previously been published and molecular biology methods have been used to study this species, no suitable internal reference genes have been investigated for standardization of gene expression via quantitative real-time polymerase chain reaction (qRT-PCR). Here, the stabilities of 10 candidate reference genes, ACT11, ACT7, α-TUB, β-TUB, GAPDH, UBC, RPL, SAM, HSP70, and PP2A, were analyzed by four different procedures of GeNorm, NormFinder, BestKeeper, and RefFinder. Their expression stabilities were evaluated under various conditions, including different tissue types (root, stem, leaf, and flower), hormone stimuli treatments (methyl jasmonate, salicylic acid, and abscisic acid), and abiotic stresses (heavy metal, salt, drought, cold, and wounding). The results indicated that β-TUB was the most stable gene for all tested samples, while ACT11 was the most unstable. The most stable reference gene was not consistent under different conditions. β-TUB exhibited the highest stability for different tissue types and abiotic stresses, while for hormone stimuli treatments, ACT7 showed the highest stability. To confirm the applicability of suitable reference genes, we selected to SbF6H and SbF8H as target genes to analyze their expression levels in different tissues. This study helps to the accurate quantification of the relative expression levels of interest genes in S. baicalensis via qRT-PCR analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Scutellaria baicalensis Georgi, belonging to the family Lamiaceae, is a well-known medicinal plant that is widely cultivated in China and other East Asian countries for its significant medicinal value [1]. Its dried root, Scutellariae Radix, is a traditional Chinese medicine and has been applied to treat inflammation, dysentery, and liver and lung problems for thousands of years, which was first recorded in “Sheng Nong’s herbal classic” (written between 200 and 300 AD) [2]. The authoritative book “Compendium of Materia Medica”, first published during the Ming Dynasty, reported that Scutellariae Radix could be used in the treatment of a wide range of disorders [3]. Recently, numerous studies have shown that the main bioactive components in Scutellariae Radix are flavonoids, which have a wide range of pharmacological activities, including antitumor, hepatoprotective, neuroprotective, antibacterial, and antiviral activities [4,5,6,7,8]. At the end of 2019, the emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) disease (COVID-19) has investigated large-scale epidemics worldwide and is currently a main public hygiene issue. The discovery of drugs against SARS-CoV-2 is the most promising treatment method for COVID-19. At present, there is no cure. Recently, baicalin and baicalein, two types of natural flavonoids derived from Scutellariae Radix, were appraised as the first non-covalent, non-peptidomimetic inhibitors of SARS-CoV-2 3CLpro (3C-like protease) and revealed powerful anti-virus activity in vitro cell systems [9, 10]. With the increasing understanding of the pharmacological activity, molecular biology methods have been used to study S. baicalensis. The genome sequence of this species is available, and the biosynthetic pathways of some flavonoids and related key enzyme genes have been revealed [11,12,13].
Gene expression levels in plant different tissues along with the abiotic stress tolerance are exceedingly meaningful for understanding the potential functions of the gene [14]. The techniques for detecting gene expression levels were include Northern blotting, microarray, RNase protection analysis, and qRT-PCR [15]. Amongst these, qRT-PCR is a simple and accurate method with strong specificity, repeatability and sensitivity, and has been widely used for evaluating the transcription level of interest gene [16, 17]. Although qRT-PCR is a powerful technology, many factors, such as RNA purity, RNA content, PCR amplification efficiency (E) and reverse transcription efficiency, can affect the results [18]. To eliminate the effects of these factors, one or several reliable reference genes should be used for correction. In different tissues and under different treatments, the transcription levels of ideal reference genes should be unaffected or show the minimal amplitude changes [19]. Therefore, for accurate quantification, selecting suitable reference genes are of great significance and necessity for qRT-PCR analysis.
Glyceraldehyde-3-phosphate dehydrogenase (GAPDH), β-actin (ACT), β-tubulin (TUB), and ubiquitin-conjugating enzyme E2 (UBC) are traditional housekeeping genes and have been often applied as reference genes, which are commonly expressed in all species [20]. Nevertheless, the transcription levels of some housekeeping genes may be unstable under various experimental conditions [21]. When inappropriate reference genes are applied for qRT-PCR normalization, a serious deviation or opposite conclusion in gene expression analysis may occur [22, 23]. Selection and verification of suitable reference genes for qRT-PCR have been studied in quite a few plant species. The results indicate that the most stable reference genes may differ for different species, and the relatively stable genes differ under different experimental conditions. For example, ACT, TUB, and GAPDH are the optimum reference genes ascertained so far in Apium graveolens [24], Petroselinum crispum [25], and Codonopsis pilosula [26], respectively. In C. pilosula, F-box, CACS, APT1, and GAPDH are the most appropriate reference genes for different tissues, different developmental phases, NaCl stress, and methyl jasmonate (MeJA) stimuli, respectively [26]. Consequently, reference genes appropriate for qRT-PCR standardization should be investigated under various experimental conditions for specific species. However, a systematic evaluation of optimum reference genes for qRT-PCR standardization in S. baicalensis has not yet been undertaken.
In this study, 10 candidate reference genes were chosen from the transcriptome sequence of S. baicalensis, and their expression profiles were investigated in different tissues and under diverse treatments. Four different software tools, GeNorm [27], NormFinder [28], BestKeeper [29] and RefFinder [30], were applied to appraise the expression stabilities of them under diverse experimental conditions. Additionally, two target genes, SbF6H (flavone 6-hydroxylase) and SbF8H (flavone 8-hydroxylase), were applied to verify the applicability of the recommended reference gene. The results of this study will guide the option of appropriate internal reference genes under specific experiment condition for acurate quantification of the relative transcription levels of interest genes in S. baicalensis.
Materials and methods
Plant materials
S. baicalensis seeds were collected from Yangcheng County, Shanxi Province, China. They were drenched in running water for 12 h to soften the seed coat, and then planted in pots (3–5 plants per pot) filled with soil mixture and grown in a greenhouse (23 ± 2 °C, Natural light, 60–75% humidity). Three-month-old seedlings received abiotic stress and hormone treatments.
For hormone stimuli treatments, the seedlings in uniform growth state were spritzed with 100 μM MeJA, 100 μM abscisic acid (ABA), or 200 μM salicylic acid (SA), while the control seedlings received only water. For heavy metal and salt stress, the seedlings were watered with 1 mM CrCL (Cr), 200 μM CuSO4 (Cu), or 100 mM NaCl (Na). Wounding treatment involved cutting approximately 50% of the stem and leaf surfaces with a surgical knife. Seedling samples were collected 12 h after hormone, heavy metal, salt, and wounding treatments. For drought stress, the seedlings were imposed by withholding water for 10 days. For cold shock treatment, the seedlings were cultivated in pots at 4 °C for 24 h. Samples of different tissues were taken from one-year-old plants at the flowering stage in the experimental field. All samples were frozen in liquid nitrogen at once and preserved at −80 °C before RNA extraction.
Extraction of total RNA and cDNA synthesis
Total RNA was isolated by the plant rapid RNA isolation kit RP3501 (BioTeke Corporation, Wuxi, China) and quantified by a NanoDrop 2000c spectrophotometer (Thermo Scientific, USA) to measure absorbance ratios. Only RNA samples with A260/A280 of 1.8–2.1 and 260/A230 > 1.8 were used for cDNA synthesis. RNA integrity was observed on 1% agarose gel. The synthesis of cDNA was carried out by taking 1.0 μg of total RNA with a PrimeScript RT kit (Takara, Kusatsu, China). All the cDNA samples were diluted with enzyme-free ultrapure water (1:20) and subjected to qRT-PCR.
Candidate reference gene and primer design
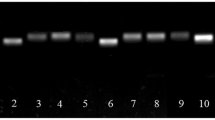
Based on the transcriptome sequences of S. baicalensis root, stem, leaf, and flower, 10 internal reference genes that are commonly used in other plant species and extensively expressed in different tissues were chosen as candidate internal reference genes (Table 1). The candidate reference genes were ACT11, ACT7, α-TUB, β-TUB, GAPDH, UBC, protein phosphatase 2A (PP2A), ribosomal protein L (RPL), S-adenosyl methionine (SAM), and heat shock protein 70 (HSP70). To confirm the dependability of the proposed reference genes, a BLAST search was conducted for the coding sequences of the 10 candidate genes against the TAIR database (http://www.arabidopsis.org/). The sequence with the highest homology with Arabidopsis was identified and presented in Table 1. Based on the unique sequence (Supplementary Table 1), Primer Premier software was used to design specific primers for all genes, with annealing temperatures (Tm) of the primers ranging from 57.9 °C to 61.0 °C, an amplicon length of 112–299 bp, and a primer length of 18–24 bp. Table 1 lists the depiction of candidate reference genes, primer pairs, and qRT-PCR amplification effectiveness. In addition, the specificity of primers was verified by agarose gel electrophoresis and melt-curve analysis.
Conditions and amplification efficiency of qRT-PCR
qRT-PCR was executed on a Roche Light Cycler 96 system (Roche Diagnostics GmbH). Twenty microliters of the reaction mixture, consisting of 10 μL Master Mix (SYBR® Green PCR Master Mix), 2 μL cDNA diluted to 2.5 ng/μL, 0.5 μL 10 mM forward, 0.5 μL 10 mM reverse primer, and 7 μL DNase/RNase-free deionized water, was pipetted into a 96-well plate. The amplification was performed as the following conditions: initial 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s, final melt-curve analysis. From five-fold continuous diluted cDNA (50, 5−1, 5−2, 5−3, 5−4, and 5−5) for each gene, we obtained the standard curve, correlation coefficient (R2), and slope value (S). S was used to calculate the amplification efficiency (E) of PCR in accordance with the following formula: E = (10–1/S − 1) × 100% [31].
Assessment of expression stability
GeNorm, NormFinder, BestKeeper and RefFinder, four conventional software tools for evaluating the stability of the candidate reference genes, were applied under various experimental conditions. The raw Ct values were transformed to relative quantities (Q) applying the following method: Q = 2−△Ct, where △Ct = each corresponding Ct − minimum Ct [32]. Then, the Q-values were input into GeNorm and NormFinder to calculate stability values (SV). GeNorm uses the average stability value (M value) to evaluate the expression stability, which involves the average pairwise variation between each reference gene and others. Genes with M < 1.5 are generally regarded as stable. Moreover, the GeNorm pairwise variation (V) values (Vn/Vn + 1) were applied to define the most appropriate reference gene number. The recommended value is ≤0.15 when opting for an appropriate number of genes [33]. The NormFinder software evaluates expression stability by calculating the SV (lower SV indicates higher stability) when applying reference genes for standardization [34]. In BestKeeper, the most stable reference gene was evaluated by the coefficient of variance (CV) and standard deviation (SD), with the lower CV and SD values mean higher stability. The RefFinder tool (https://www.heartcure.com.au/reffinder/), integrating the outcomes of BestKeeper, Delta Ct,GeNorm and NormFinder analysis, was applied to perform the most stable reference genes.
Validation of reference gene stability
SbF6H and SbF8H, two key genes associated with the biosynthesis of baicalein and scutellarein in S. baicalensis, were chosen as target genes to verify the applicability of the selected reference gene. Their relative transcription levels in different tissues were normalized by the top two most stable reference genes and their combinations, as well as the least stable reference genes. The 2−△△Ct method was applied to calculate the relative transcription levels [35].
Statistical analysis
All experiments were conducted with at least three biological repeats and each qRT-PCR reaction was performed with three technical repeats. Statistical analysis was performed by SPSS 26.0 software (SPSS Inc., Chicago, IL, USA). The relative expression levels of target genes are shown as the mean ± standard deviation (SD). One-way ANOVA was used to analyze the significant difference between the mean values by showing different letters at p < 0.05.
Results
Analysis of primer specificity and expression profilings of 10 candidate reference genes
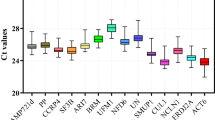
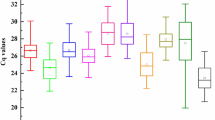
Based on the transcriptome sequences of S. baicalensis, the expression patterns of candidate reference genes were analyzed in diverse experimental conditions, comprising six housekeeping genes (ACT11, ACT7, α-TUB, β-TUB, GAPDH, and UBC) and four novel candidate reference genes (RPL, SAM, HSP70, and PP2A). The results of agarose gel electrophoresis confirmed that all the 10 candidates were specifically amplifed because only one single product of the expected size appeared (Supplementary Fig. 1). Furthermore, the specificity of primers was also verified by the single peak of melting curve (Supplementary Fig. 2). The melting temperatures of the amplicons ranged from 81.6 °C for UBC to 89.2 °C for HSP70 (Supplementary Fig. 2). The E of the primers varied from 92.52% for β-TUB to 118.18% for ACT7, while the R2 ranged from 0.9889 for PP2A to 0.9992 for GAPDH and HSP70. The expression levels of the 10 candidate reference genes were determined by the values of the cycle threshold (Ct). A smaller Ct value indicated a higher expression level. The Ct value of all candidate reference genes under different experimental conditions varied from 21.13 to 32.17 (Supplementary Fig. 3). Among them, ACT11 represented the lowermost transcript abundance with the uppermost average Ct value, while GAPDH had the uppermost transcript abundance with the lowermost average Ct value. In addition, among all samples, the SD of β-TUB was the smallest (SD = 0.79), whereas ACT11 showed the maximum variability (SD = 1.44) (Supplementary Table 2).
Stabilization of candidate internal reference genes
The samples were further divided into three groups according to various tissues, hormone treatments (MeJA, SA, and ABA), and abiotic stresses (heavy metal, salt, drought, cold, and wounding). Based on the M value, GeNorm identified β-TUB as the most stable reference gene for all samples, while ACT11 was identified as the most unstable (Fig. 1a). When considering different tissues and abiotic stress treatments, β-TUB was confirmed as the most appropriate gene, while α-TUB (Fig. 1b) and ACT11 (Fig. 1c) were confirmed as the least stable for different tissues and abiotic stress treatments, respectively. For hormone treatments, ACT7 was regarded as the most appropriate reference gene (Fig. 1d).
Additionally, we applied the average pairwise variation (Vn/Vn + 1) to determine the number of reference gene for normalization of qRT-PCR in S. baicalensis. Among our three experimental sets, the V2/3 values were all below the cut-off threshold value of 0.15 (Fig. 2), which showed that the top two highest-ranked genes were adequate for standardization of qRT-PCR in each experimental set. For the total samples, V2/3 was 0.158. We ascertained that a combination of three genes (β-TUB, UBC, and SAM) was necessary for normalization because V3/4 was lower than 0.15 (Fig. 2).
The results of NormFinder analysis were showed in Supplementary Table 3, which indicated that β-TUB was the most stable gene for different tissues and abiotic stress treatments, and that ACT7 was the most stable for hormone treatments. The most stable genes evaluated by NormFinder were consistent with those of GeNorm.
BestKeeper analysis (Supplementary Table 4) also indicated that β-TUB was the most stable for different tissues and abiotic stress treatments, while UBC was the most stable for hormone treatments.
To verify the results of GeNorm, NormFinder and BestKeeper software, we also used Reffinder tool to generate a comprehensive ranking. As shown in Fig. 3, β-TUB was the most stable reference gene for all samples, while ACT11 was identified as the most unstable (Fig. 3a). When considering different tissues and abiotic stress treatments, β-TUB was confirmed as the most appropriate gene (Fig. 3b, c) were confirmed as the least stable, respectively. For hormone treatments, ACT7 was regarded as the most appropriate reference gene (Fig. 3d).
Confirmation of reference gene stability
To confirm the dependability of the selected reference genes for standardization, we investigated the expression patterns of SbF6H and SbF8H in different tissues. When standardized with the top two most appropriate reference genes (β-TUB and UBC) and their combination, the expression levels of SbF6H and SbF8H were highest in the root (Fig. 4). These findings are coincident with the expression profiles of SbF6H and SbF8H previously reported in S. baicalensis [11]. However, when normalized with the unstable gene ACT11, the expression patterns of SbF6H and SbF8H were quite different from previous report [11], and these two genes showed the highest expression level in the stem and leaf, respectively (Fig. 4). Therefore, the selection of an inappropriate reference gene may cause biased results. Here, analysis of the expression profiles of SbF6H and SbF8H indicted that the chosen reference genes were reliable for qRT-PCR standardization.
Discussion
S. baicalensis, as an important medicinal plant, has attracted the attention of many researchers. With the development of genome sequencing, whole genome information for S. baicalensis is now available [13], providing the basis for molecular biology research of the species. An appropriate reference gene is the prerequisite for credible results of qRT-PCR, which is usually the preferred method for gene expression analysis. Nevertheless, the selection of suitable reference genes in diverse experimental conditions had not been reported previously.
Here, the expression stabilities of 10 candidate reference genes were assessed in different tissues (root, stem, leaf, and flower), hormone treatments (SA, MeJA, and ABA), and abiotic stress treatments (Cr, Cu, salt, drought, cold, and wounding) for qRT-PCR normalization. Additionally, SbF6H and SbF8H, two genes belonging to the CYP450 family that are associated with hydroxyl transferase and are involved in the biosynthesis of baicalein and scutellarin [11], were used as targets for verification of the selected reference genes.
In the current study, we employed GeNorm, NormFinder, BestKeeper, and RefFinder to rank the stability of candidate reference genes among diverse experimental groups and in total samples. Even though the stability rankings of GeNorm and NormFinder were not completely identical, two assessment tools consistently identified ACT7 as the most appropriate reference gene for hormone treatments and β-TUB as the optimum reference gene in different tissue types, abiotic stresses and all tested samples (Fig. 1 and Supplementary Table 3). The rankings generated by BestKeeper differ significantly from those provided by geNorm and NormFinder. In particular, the results of BestKeeper showed that UBC was the most stable reference gene for hormone treatments, while ACT7, the most stable reference gene recommended by geNorm and NormFinder, was only the fourth most stable reference gene by BestKeeper (Supplementary Table 4). We previously found that the results of GeNorm and NormFinder were mostly consistent, while the results of BestKeeper were quite different to those of GeNorm and NormFinder in C. pilosula [26]. Recently, many studies have stated a preference for GeNorm and NormFinder for the evaluation of reference genes rather than BestKeeper to evaluate appropriate reference genes [14, 18, 36]. To evaluate the stability of these genes more reasonably, we utilized RefFinder to obtain the comprehensive ranking of these genes. As shown in Fig. 3, the ranking of RefFinder is highly consistent with those from GeNorm and NormFinder. All three recommender softwares recommend that β-TUB as the best reference gene for different tissue types, abiotic stresses and all tested samples, while ACT7 is recommended as the most stable gene for hormone treatments.
Although β-TUB is an extremely stable reference gene in some species [37,38,39], it is unstable in species such as C. pilosula [26], soybean [40], Vitis vinifera [41], and Raphanus sativus [42]. Here, we found that β-TUB was the most stable among the 10 candidate reference genes in S. baicalensis, followed by UBC and SAM. UBC, which encodes a protein-modifying enzyme that arises in a multi-step reaction, is used as a reference gene in Citrus sinensis [43], Arabidopsis thaliana [44], and Cocos nucifera [45]. SAM, which encodes a key enzyme for the plant synthesis of polyamines, is used as a reference gene in Sinobambusa tootsik [46]. Homologous genes, which show similar structures and functions, are often used as reference genes for gene transcription analysis. Take Fortunella crassifolia, for example, in which the homologous actin genes (ACT6, ACT8, and ACT7) have all been selected as suitable internal control genes [29]. The expression of homologous EF1As (EF1A2a, EF1A2b, and EF1A1a1) are the most stable in Glycine max [47]. Nevertheless, our results indicated that the homologous tubulin genes (α-TUB and β-TUB) exhibit completely different expression levels and stabilities. For hormone treatments, ACT7 was regarded as the most stable reference gene, while ACT11 was the least stable (Figs. 1 and 3 and Supplementary Table 3). Therefore, homologous genes with similar sequences, structures, and functions may have quite different expression levels and stabilities.
To confirm the dependability of the selected reference genes, we used the top two most stable reference genes (β-TUB and UBC), a combination of these, and the most unstable reference gene (ACT11) to standardize the relative expression levels of SbF6H and SbF8H in different tissues. The results indicated that the expression profiles of SbF6H and SbF8H in different tissues were consistent with those reported previously [11] when β-TUB, UBC, or their combination were used for normalization. In contrast, their expression profiles were obviously different when normalized with ACT11 (Fig. 4). The expression analysis of SbF6H and SbF8H indicated that a appropriate reference gene is crucial for obtaining accurate qRT-PCR results.
Conclusion
In summary, four statistical methods, including GeNorm, NormFinder, BestKeeper, and the comprehensive tool RefFinder, were applied to assess the expression stabilities of 10 candidate reference genes in S. baicalensis in diverse experimental conditions, including different tissue types (root, stem, leaf, and flower), hormone stimuli treatments (MeJA, SA, and ABA), and abiotic stresses (heavy metal, salt, drought, cold, and wounding). β-TUB is the optimum reference gene for different tissues, abiotic stress treatments, and all samples, while ACT7 is most stable for hormone treatments. In total samples, β-TUB, UBC, and PP2A were the top three most stable reference genes, while ACT11 was the most unstable gene, followed by GAPDH. The selection and confirmation of appropriate reference genes provides a basis for the accurate quantification of relative transcription levels of genes of interest in S. baicalensis via qRT-PCR analysis.
References
Shang X, He X, He X, Li M, Zhang R, Fan P, Zhang Q, Jia Z (2010) The genus Scutellaria an ethnopharmacological and phytochemical review. J Ethnopharmacol 128(2):279–313. https://doi.org/10.1016/j.jep
Ma JX (2013) Explanatory Notes to ShennongBencao Jing, 3 (Beijing:People’s Medical Publishing House). p 140
Li SZ (1593 and republished in 2012) In: Compendium of materia medica (BencaoGangmu). Huaxia Press, pp 543‑546 (In Chinese)
Tu B, Li RR, Liu ZJ, Chen ZF, Ouyang Y, Hu YJ (2016) Structure-activity relationship study between baicalein and wogonin by spectrometry, molecular docking and microcalorimetry. Food Chem 208:192–198. https://doi.org/10.1016/j.foodchem
Qiao X, Li R, Song W, Miao WJ, Liu J, Chen HB, Guo DA, Ye M (2016) A targeted strategy to analyze untargeted mass spectral data: rapid chemical profiling of Scutellaria baicalensis using ultra-high performance liquid chromatography coupled with hybrid quadrupole orbitrap mass spectrometry and key ion filtering. J Chromatogr A 1441:83–95. https://doi.org/10.1016/j.chroma
Makino T, Hishida A, Goda Y, Mizukami H (2008) Comparison of the major flavonoid content of S. baicalensis, S. lateriflora, and their commercial products. J Nat Med 62(3):294–299. https://doi.org/10.1007/s11418-008-0230-7
Gaire BP, Moon SK, Kim H (2014) Scutellaria baicalensis in stroke management: nature’s blessing in traditional Eastern medicine. Chin J Integr Med 20(9):712–720. https://doi.org/10.1007/s11655-014-1347-9
Yang MD, Chiang YM, Higashiyama R, Asahina K, Mann DA, Mann J, Wang CC, Tsukamoto H (2012) Rosmarinic acid and baicalin epigenetically derepress peroxisomal proliferator-activated receptor γ in hepatic stellate cells for their antifibrotic effect. Hepatology (Baltimore, Md.) 55(4):1271–1281. https://doi.org/10.1002/hep.24792
Su HX, Yao S, Zhao WF, Li MJ, Liu J, Shang WJ, Xie H, Ke CQ, Gao MN, Yu KQ, Liu H, Shen JS, Tang W, Zhang LK, Zuo JP, Jiang H, Bai F, Wu Y, Ye Y, Xu YC (2020) Discovery of baicalin and baicalein as novel, natural product inhibitors of SARS-CoV-2 3CL protease in vitro. bioRxiv. https://doi.org/10.1101/2020.04.13.038687
Liu HB, Ye F, Sun Q, Liang H, Li CM, Lu RJ, Huang BY, Tan WJ, Lai LH (2020) Scutellaria baicalensis extract and baicalein inhibit replication of SARS-CoV-2 and its 3C-like protease in vitro. BioRxiv. https://doi.org/10.1101/2020.04.10.035824
Zhao Q, Cui MY, Levsh O, Yang D, Liu J, Li J, Hill L, Yang L, Hu Y, Weng JK, Chen XY, Martin C (2018) Two CYP82D enzymes function as flavone hydroxylases in the biosynthesis of root-specific 4′-deoxyflavones in Scutellaria baicalensis. Mol Plant 11(1):135–148. https://doi.org/10.1016/j.molp
Zhao Q, Zhang Y, Wang G, Hill L, Weng JK, Chen XY, Xue H, Martin C (2016) A specialized flavone biosynthetic pathway has evolved in the medicinal plant, Scutellaria baicalensis. Sci Adv 2(4):e1501780. https://doi.org/10.1126/sciadv.1501780
Zhao Q, Yang J, Cui MY, Liu J, Fang Y, Yan M, Qiu W, Shang H, Xu Z, Yidiresi R, Weng JK, Pluskal T, Vigouroux M, Steuernagel B, Wei Y, Yang L, Hu Y, Chen XY, Martin C (2019) The reference genome sequence of Scutellaria baicalensis provides insights into the evolution of wogonin biosynthesis. Mol Plant 12(7):935–950. https://doi.org/10.1016/j.molp
Zeng AS, Xu YY, Song LX, Li JQ, Yan JY. (2020). Validation of suitable reference genes for qRT-PCR in cabbage (Brassica oleracea L.) under different abiotic stress experimental conditions. J Plant Biochem Biotechnol 1–12. https://doi.org/10.1007/s13562-020-00556-x
VanGuilder HD, Vrana KE, Freeman W (2008) Twenty-five years of quantitative PCR for gene expression analysis. BioTechniques 44(5):619–626. https://doi.org/10.2144/000112776
Tsai HF, Sammons LR, Zhang X, Suffis SD, Su Q, Myers TG, Marr KA, Bennett JE (2010) Microarray and molecular analyses of the azole resistance mechanism in Candida glabrata oropharyngeal isolates. Antimicrob Agents Chemother 54(8):3308–3317. https://doi.org/10.1128/AAC.00535-10
Nolan T, Hands RE, Bustin SA (2006) Quantification of mRNA using real-time RT-PCR. Nat Protoc 1(3):1559–1582. https://doi.org/10.1038/nprot.2006.236
Xu YY, Li H, Li XG, Lin J, Wang ZH, Yang QS et al (2015) Systematic selection and validation of appropriate reference genes for gene expression studies by quantitative real-time PCR in pear. Acta Physiol Plant 37(2):40. https://doi.org/10.1007/s11738-015-1784-0
Udvardi MK, Czechowski T, Scheible WR (2008) Eleven golden rules of quantitative RT-PCR. Plant Cell 20(7):1736–1737. https://doi.org/10.1105/tpc.108.061143
Joseph JT, Poolakkalody NJ, Shah JM (2018) Plant reference genes for development and stress response studies. J Biosci 43(1):173–187
Leal MF, Astur DC, Debieux P, Arliani GG, Silveira Franciozi CE, Loyola LC, Andreoli CV, Smith MC, Pochini Ade C, Ejnisman B, Cohen M (2015) Identification of suitable reference genes for investigating gene expression in anterior cruciate ligament injury by using reverse transcription-quantitative PCR. PLoS One 10(7):e0133323. https://doi.org/10.1371/journal.pone.0133323
Gutierrez L, Mauriat M, Guénin S, Pelloux J, Lefebvre JF, Louvet R, Rusterucci C, Moritz T, Guerineau F, Bellini C, Van Wuytswinkel O. (2008). The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol J 6(6):609–618. https://doi.org/10.1111/j.1467-7652.2008.00346.x
Mascia T, Santovito E, Gallitelli D, Cillo F (2010) Evaluation of reference genes for quantitative reverse-transcription polymerase chain reaction normalization in infected tomato plants. Mol Plant Pathol 11(6):805–816. https://doi.org/10.1111/j.1364-3703.2010.00646.x
Feng K, Liu JX, Xing GM, Sun S, Li S, Duan AQ, Wang F, Li MY, Xu ZS, Xiong AS (2019) Selection of appropriate reference genes for RT-qPCR analysis under abiotic stress and hormone treatment in celery. Peer J 7:e7925. https://doi.org/10.7717/peerj.7925
Li MY, Song X, Wang F, Xiong AS (2016) Suitable reference genes for accurate gene expression analysis in parsley (Petroselinum crispum) for abiotic stresses and hormone stimuli. Front Plant Sci 7:1481. https://doi.org/10.3389/fpls.2016.01481
Yang J, Yang X, Kuang Z, Li B, Lu X, Cao X, Kang J (2020) Selection of suitable reference genes for qRT-PCR expression analysis of Codonopsis pilosula under different experimental conditions. Mol Biol Rep 47(6):4169–4181. https://doi.org/10.1007/s11033-020-05501-8
Schlotter YM, Veenhof EZ, Brinkhof B, Rutten VP, Spee B, Willemse T, Penning LC (2009) A GeNorm algorithm-based selection of reference genes for quantitative real-time PCR in skin biopsies of healthy dogs and dogs with atopic dermatitis. Vet Immunol Immunopathol 129(1–2):115–118. https://doi.org/10.1016/j.vetimm.2008.12.004
Zhang X, Xu ZC, Xu J, Ji AJ, Luo HM, Song JY, Sun C, Hu YL, Chen SL (2016) Selection and validation of reference genes for normalization of quantitative real-time reverse transcription PCR analysis in Poria cocos (Schw.) Wolf (Fuling). Chin Med 11:8. https://doi.org/10.1186/s13020-016-0079-8
Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP (2004) Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper-Excel-based tool using pair-wise correlations. Biotechnol Lett 26:509–515. https://doi.org/10.1023/b:bile.0000019559.84-305.47
Xie F, Xiao P, Chen D, Xu L, Zhang B (2012) miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol Biol, Advance online publication. https://doi.org/10.1007/s11103-012-9885-2
Kudo T, Sasaki Y, Terashima S, Matsuda-Imai N, Takano T, Saito M, Kanno M, Ozaki S, Suwabe K, Suzuki G, Watanabe M, Matsuoka M, Takayama S, Yano K (2016) Identification of reference genes for quantitative expression analysis using large-scale RNA-seq data of Arabidopsis thaliana and model crop plants. Genes Genet Syst 91(2):111–125. https://doi.org/10.1266/ggs.15-00065
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):RESEARCH0034. https://doi.org/10.1186/gb-2002-3-7-research0034
Dekkers BJ, Willems L, Bassel GW, van Bolderen-Veldkamp RP, Ligterink W, Hilhorst HW, Bentsink L (2012) Identification of reference genes for RT-qPCR expression analysis in Arabidopsis and tomato seeds. Plant Cell Physiol 53(1):28–37. https://doi.org/10.1093/pcp/p-cr113
Andersen CL, Jensen JL, Ørntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64(15):5245–5250. https://doi.org/10.1158/0008-5472.CAN-04-0496
Kumar V, Sharma R, Trivedi P, Vyas G, Khandelwal V (2011) Traditional and novel references towards systematic normalization of qRT-PCR data in plants. Aust J Crop Sci 5(11):1455–1468. https://doi.org/10.1016/j.fcr.2011.04.016
Li J, Han X, Wang C, Qi W, Zhang W, Tang L, Zhao X (2017) Validation of suitable reference genes for RT-qPCR data in Achyranthes bidentata Blume under different experimental conditions. Front Plant Sci 8:776. https://doi.org/10.3389/fpls.2017.00776
Niu K, Shi Y, Ma H (2017) Selection of candidate reference genes for gene expression analysis in Kentucky bluegrass (Poa pratensis L.) under abiotic stress. Front Plant Sci 8:193. https://doi.org/10.3389/fpls.2017.00193
Zhou W, Wang S, Yang L, Sun Y, Zhang Q, Li B, Wang B, Li L, Wang D, Wang Z (2019) Reference genes for qRT-PCR normalisation in different tissues, developmental stages, and stress conditions of Hypericum perforatum. Peer J 7:e7133. https://doi.org/10.7717/peerj.7133
Niu X, Qi J, Chen M, Zhang G, Tao A, Fang P, Xu J, Onyedinma SA, Su J (2015) Reference genes selection for transcript normalization in kenaf (Hibiscus cannabinus L.) under salinity and drought stress. Peer J 3:e1347. https://doi.org/10.7717/peerj.1347
Hu R, Fan C, Li H, Zhang Q, Fu YF (2009) Evaluation of putative reference genes for gene expression normalization in soybean by quantitative real-time RT-PCR. BMC Mol Biol 10:93. https://doi.org/10.1186/1471-2199-10-93
Reid KE, Olsson N, Schlosser J, Peng F, Lund ST (2006) An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol 6:27. https://doi.org/10.1186/1471-2229-6-27
Duan M, Wang J, Zhang X, Yang H, Wang H, Qiu Y, Song J, Guo Y, Li X (2017) Identification of optimal reference genes for expression analysis in radish (Raphanus sativus L.) and its relatives based on expression stability. Front Plant Sci 8:1605. https://doi.org/10.3389/fpls.2017.01605
Mafra V, Kubo KS, Alves-Ferreira M, Ribeiro-Alves M, Stuart RM, Boava LP, Rodrigues CM, Machado MA (2012) Reference genes for accurate transcript normalization in citrus genotypes under different experimental conditions. PLoS One 7(2):e31263. https://doi.org/10.1371/journ-al.pone.0031263
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139(1):5–17. https://doi.org/10.1104/pp.105.063743
Xia W, Liu Z, Yang Y, Xiao Y, Mason AS, Zhao S, Ma ZL (2014) Selection of reference genes for quantitative real-time PCR in Cocos nucifera during abiotic stress. Botany 92(3):179–186. https://doi.org/10.1139/cjb-2013-0212
Rosic NN, Pernice M, Rodriguez-Lanetty M, Hoegh-Guldberg O (2011) Validation of housekeeping genes for gene expression studies in Symbiodinium exposed to thermal and light stress. Mar Biotechnol 13(3):355–365. https://doi.org/10.1007/s10126-010-9308-9
Bansal R, Mittapelly P, Cassone BJ, Mamidala P, Redinbaugh MG, Michel A (2015) Recommended reference genes for quantitative PCR analysis in soybean have variable stabilities during diverse biotic stresses. PLoS One 10(8):e0134890. https://doi.org/10.1371/journal.pone.0134890
Acknowledgments
This work was supported by the National Key Technologies R & D Program for Modernization of Traditional Chinese Medicine (2017YFC1701300) and the Youth Innovation Team of Shaanxi Universities.
Funding
Dr. Xiaoyan Cao was funded by The National Key Technologies R & D Program for Modernization of Traditional Chinese Medicine (2017YFC1701300) and The Youth Innovation Team of Shaanxi Universities.
Author information
Authors and Affiliations
Contributions
Conceptualization: XYC, WW. Data curation: WW, YC, RC. Formal analysis: YC, RC, WW. Funding acquisition: ZW, XC. Methodology: WW, SH, YC, RC. Software: SH, WW. Supervision: SH. Writing – original draft: WW. Writing–review and editing: ZW, XC.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have declared that no competing interests exist.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, W., Hu, S., Cao, Y. et al. Selection and evaluation of reference genes for qRT-PCR of Scutellaria baicalensis Georgi under different experimental conditions. Mol Biol Rep 48, 1115–1126 (2021). https://doi.org/10.1007/s11033-021-06153-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06153-y