Abstract
Cystinuria is an autosomal recessive defect in reabsorptive transport of cystine and the dibasic amino acids ornithine, arginine, and lysine from renal tubule and small intestine. Mutations in two genes: SLC3A1, encoding the heavy chain rbAT of the renal cystine transport system and SLC7A9, the gene of its light chain b0, + AT have a crucial role in the diseases. In our previous studies from Iranian populations with Cystinuria totally six and eleven novel mutations respectively identified in SLC3A1 and SLC7A9 genes. In this study, we conducted an in silico functional analysis to explore the possible association between these genetic mutations and Cystinuria. MutationTaster, PolyPhen-2, PANTHER, FATHMM. PhDSNP and MutPred was applied to predict the degree of pathogenicity for the missense mutations. Furthermore, Residue Interaction Network (RIN) and Intron variant analyses was performed using Cytoscape and Human Slicing Finder softwares. These genetic variants can provide a better understanding of genotype–phenotype relationships in patients with Cystinuria. In the future, the findings may also facilitate the development of new molecular diagnostic markers for the diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transporters are embedded proteins within membranes that control the uptake of different solutes [1]. They are divided into solute carrier (SLC) and ATP-binding cassette (ABC) transporters [2]. The SLC transporters control the uptake of endogenous compounds essential for cell survival, including sugars, amino acids, digested peptides, nucleotides, and inorganic ions [3,4,5].
The solute carrier (SLC) transporter superfamily with 55 families are encoded by a total of at least 362 putatively functional protein coding genes [6]. Because of key physiological roles of SLC transporters, defects in functionally specific SLC transporters can cause many Mendelian diseases or monogenic disorders [7]. More than 80 SLC genes have been involved in monogenic disorders. For example, mutations in SLC25A19 and SLC5A2 respectively was lead to Amish lethal microcephaly and familial renal glucosuria [8,9,10]. Cystinuria is another disease due to pathogenic variants in the SLC3A1 or SLC7A9 genes [11, 12].
Here, we focus on the Cystinuria, which is an inherited autosomal recessive disorder of renal reabsorption of cystine, arginine, lysine, and ornithine [13]. The protein products of SLC3A1 (rBAT) and SLC7A9 (b0, + AT) form the heterodimeric amino-acid transporter system b0, +, which is responsible for the uptake of cystine and dibasic amino acids in the renal tubular and intestinal epithelial cells [14, 15]. In Cystinuria, mutation in the two genes resulted to increased urinary excretion of cystine and finally formation of kidney stones [16]. Patients with two SLC3A1 mutations are classified as type I Cystinuria, whereas patients with two SLC7A9 mutations are classified as non-type I Cystinuria [17, 18]. Over 100 SLC3A1 mutations have been recognized, and all, except one (dupE5–E9), were limited to patients with type I Cystinuria [17, 19, 20]. At least 66 SLC7A9 mutations were identified and these mutations were found in both type I and type non-I patients [21]. From our previous studies in Iranian patients with Cystinuria, we identified four missense mutations, one intron variant and one polymorphism in SLC3A1 as well as three missense mutations, one frame shift, four intron variant and three polymorphisms in SLC7A9 [22,23,24,25,26].
Bioinformatics prediction tools can be applied in a cost efficient manner to calculate effects of specific mutations on the protein structure and function for selecting SNPs likely contribute to an individual’s disease susceptibility. Recently, several computational methods have been developed to screen functional SNPs out of large pools of disease-sensitive SNPs related to the BRCA1, ATM, PON1, ADIPOR1 and SLC genes [27,28,29].
In the current study, we used different softwares and publicly available bioinformatics tools to comprehensively analyze various mutation identified in the SLC3A1 and SLC7A9 genes of Iranian populations with Cystinuria from our previous studies [22,23,24,25,26]. Since missense mutations of the genes are associated with more abnormalities, we aimed to study the effect of mutations on protein stability. Moreover, the pathogenic effects of the intron variants using bioinformatics tools were predicted. Subsequently, the 3D modeled protein structures of the mutants were compared with the native protein to evaluate structural deviations and topological similarities.
Methods
Bioinformatic pathogenicity predictions
The degree of pathogenicity for the missense mutations identified in SLC3A1 and SLC7A9 genes was predicted using the MutationTaster (http://www.mutationtaster.org/) [30], Polymorphism Phenotyping v2 (PolyPhen-2) (http://genetics.bwh.harvard.edu/pph2) [31], Protein Analysis Through Evolutionary Relationships (PANTHER) (http://www.pantherdb.org) [32], and Functional Analysis Through Hidden Markov Models (FATHMM) (http://fathmm.biocompute.org.uk/index.html) [33]. The Mutation Prediction Predictor of Human Deleterious Single Nucleotide Polymorphisms (PhDSNP) (http://snps.biofold.org/phd-snp/phd-snp.html) [34] and (MutPred) (http://mutpred.mutdb.org) [35] were applied to estimate its functional effects.
3D structure preparation
The 3D modelled structure of the SLC3A1 and SLC7A9 proteins for wild and mutant type prepared using Homology modeling in SWISS-MODEL webserver (https://swissmodel.expasy.org/) [36,37,38,39] were applied for structural analysis.
Exploration of residue interaction networks
Cytoscape with two plugins StructureViz [40] and RINalyzer [41] was used for analysis of residue network interaction of wild type and mutated structures [42].
Sequence alignment
Sequence alignment and visualization of conserved amino acids were prepared using the cobalt constraint-based multiple protein alignment tool (https://www.ncbi.nlm.nih.gov/tools/cobalt/re_cobalt.cgi) [43] and the universal protein resource (UniProt) (http://www.uniprot.org/align/) [44] with default parameters.
Intron variant analysis
To in silico evaluate the possible effects of the identified intron variants on gene splicing, Human Splicing Finder (http://www.umd.be/HSF/, Marseille, France) softwares were used [45]. In this tool, analysis of intron sequences for putative branch points and calculation of the breakage of exonic splicing enhancers (ESE) or creation of exonic splicing silencers (ESS) was performed.
Results and discussion
In previous studies from these authors [22,23,24,25,26], some variants were identified in SLC3A1 and SLC7A9 genes including missense, polymorphism, and intron variants summarized in Table 1. Totally, six and eleven novel mutations respectively identified in SLC3A1 and SLC7A9 genes. Wass et al found 57 different mutations in UK population [46]. Similarly, they used computational methods to discover the functional and structural consequences of the nsSNPs [29]. In the current research work, the novel variants that have not been reported so far including c.1136+2/3delT in SLC3A1 and c.177G/A, c.478+14insA, c.272−273insA, c.478+10T/C, c.604+66A/G, c.993G/A in SLC7A9 were identified. As shown in Table 1, c.177G>A, c.411T>C and c.993G>A mutations in SLC7A9 as well as c.114A/C mutation in SLC3A1 lead to polymorphism/synonymous for Thr 59, Cys 137, Ala 331 and Gly 38 residues, respectively. The c.235+22T/G, c.478+14insA, c.478+10T>C, c.604+66C>G mutations in SLC7A9 and c.1136+2/3delT mutation in SLC3A1 were in the intronic region or Untranslated/No coding region.
Intron variant mutations
Intron variant analysis indicated that only c.478+10T/C mutation was not created/changed a significant splicing motif in SLC7A9 and probably no impact on splicing (Table 2). Furthermore, c.235+22T/G and c.478+14insA intron variants in SLC7A9 had not any effect on splicing but they can created and also changed ESS and ESE motif sites in the intronic regions (Int3 for c.235+22T/G and Int4 for c.478+14insA). However, both of variants c.1136+2/3delT and c.604+66C>G mutations in SLC3A1 and SLC7A9 genes most probably affected splicing respectively through alteration in WT donor sites and exonic ESE sites (Table 2). These molecular events probably make an alternative splicing process.
Missense mutations
Only the missense mutations change the amino acid sequence of the SLC transporter proteins. The protein prediction analysis for the pathogenic effects of these missense mutations on SLC3A1 and SLC7A9 proteins were calculated using six bioinformatics programs that use different prediction algorithms: PolyPhen-2, PANTHER, FATHMM, PhD-SNP, MutPred and MutationTaster (Tables 3, 4). All of these programs predicted the variants p.R362C, p.M67K/T, p.T216M in SLC3A1 and p.G105R, p.R333W in SLC7A9 to be damaging/deleterious/disease causing. While the p.V142A variant in SLC7A9 were benign/Neutral Polymorphism/Polymorphism.
The residue interaction analysis for these mutations were presented in Figs. 1 and 2. In SLC7A9 protein, Gly 105 in the wild type had contacts with Met 101, Ileu 107, Pro 108, and Ala 109 but Arg 105 is in the connection with these residues as well as Tyr 104. Trp 333 in the mutant type of the protein, lost its contact to Ser 342 and Ala 331, while Arg in wild type connected to them as well as Tyr 329, Gly 325 and Val 330. Moreover, Val 142 which was in the connection with Cys 144, Lys 145 and Ala 142, in the mutant type, had contacts with the same residues plus Cys 137. In SLC3A1 protein, Arg 362 from the mutant type had residue interaction network similar to wild type as well as additional connection to Glu 404 and Gln 403. In M467T mutation from the mentioned protein, the mutant type had lost its connection to residues Gly645, Asp 628 and obtained connection to Leu 597. In the same way, M467K missense mutation, the mutated protein had lost its contacts to residues Leu 468, Leu 555, Gly 645, Asp 628 and expanded its network connection to other residues such as Asn 466, Leu 469 and Phe 470. In the last mutation T216M, the mutant type of the SLC3A1 protein had contacts with residues as the same as wild type and also Leu 285, His 215 and Phe 280.
Point mutations
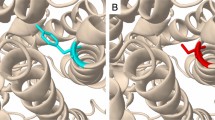
In point mutation, c.272−273insA, the SLC7A9 protein changed through Nonsense mediated mRNA Decay (NMD). The length of SLC7A9 protein sequence was decreased from 478 amino acid to 120. In this mutation, insertion “A” between nucleotide C272 and C273, lead to frame shift mutation and changing Lys 92 to Gln in the protein structure. The residue interaction analysis indicated the residues that were interacted to the mutant and wild type protein is also changed (Fig. 3). Lys 92 of SLC7A9 protein interacts with seven amino acids in the same chain including Ileu 90, Ser 93, Gly 94, Gly 95, Pro 98, Glu 102 and Thr 242. While Gln 92 had contact with only four residues of the protein: Ileu 90, Arg 94, Gly 95 and Ser 98. Therefore the major changes in the length of protein and contact network will be definitely pathogenic. The MutationTaster determined that c.272−273insA mutation was “disease causing”.
The residue 92 interaction network in wild type and mutated SLC7A9 protein structures. a Lys 92 has contacts with Ileu 90, Ser 93, Gly 94, Gly 95, Pro 98, Glu 102 and Thr 242. b Gln 92 is in connection with Ileu 90, Arg 94, Gly 95 and Ser 98. The residues 93–120 was colored blue for showing the frameshift mutation. (Color figure online)
Sequence alignment
The multiple sequence alignment obtained by cobalt constraint based multiple protein alignment tool indicated that Arg 362, Met 467, Thr 216 in SLC3A1 protein and Gly 105 and Arg 333 in SLC7A9 protein, are in a highly conserved region, whereas Val 142 in SLC7A9 protein is not conserved (Fig. 4). The other alignments were not shown. The substitution of the conserved residues which mainly contribute to the protein structure and function, confirm the deleterious effect of the mentioned mutations in SLC3A1 and SLC7A9 genes and also the benign effect of p.V142A in SLC7A9 predicted previously using different bioinformatics programs.
Conclusion
The present study offers that various computational tools were able to distinguish disease-causing mutations from benign polymorphisms. Four deleterious mutation (R362C, T216M, M467K/T) in the coding region of SLC3A1 were identified. Only missense mutation V142A had a benign effect on the protein structure and function of SLC7A9. The intron variants c.604+66C>G and c.1136+2/3delT respectively in SLC7A9 and SLC3A1 genes probably affected the splicing process. Overall, the present computational study will provide an insight into the genetic association of some novel deleterious mutations in SLC3A1 and SLC7A9 genes with Cystinuria.
References
Sahi J, Lai Y, Lee C, Lyubimov AV (2011) Solute carrier (SLC) family transporters. Encyclopedia of drug metabolism and Interactions. Wiley, Hoboken
Nakanishi T (2007) Drug transporters as targets for cancer chemotherapy. Cancer Genomics Proteomics 4(3):241–254
Forrest LR, Rudnick G (2009) The rocking bundle: a mechanism for ion-coupled solute flux by symmetrical transporters. Physiology (Bethesda Md) 24:377–386
Saier MH Jr, Yen MR, Noto K, Tamang DG, Elkan C (2009) The transporter classification database: recent advances. Nucleic Acids Res 37(Database issue):D274–D278
Forrest LR, Kramer R, Ziegler C (2011) The structural basis of secondary active transport mechanisms. Biochim Biophys Acta 1807(2):167–188
He L, Vasiliou K, Nebert DW (2009) Analysis and update of the human solute carrier (SLC) gene superfamily. Hum Genomics 3(2):195–206
Lin L, Yee SW, Kim RB, Giacomini KM (2015) SLC transporters as therapeutic targets: emerging opportunities. Nat Rev Drug Discov 14(8):543–560
Kelley RI, Robinson D, Puffenberger EG, Strauss KA, Morton DH (2002) Amish lethal microcephaly: a new metabolic disorder with severe congenital microcephaly and 2-ketoglutaric aciduria. Am J Med Genet 112(4):318–326
Lindhurst MJ, Fiermonte G, Song S, Struys E, De Leonardis F, Schwartzberg PL et al (2006) Knockout of Slc25a19 causes mitochondrial thiamine pyrophosphate depletion, embryonic lethality, CNS malformations, and anemia. Proc Natl Acad Sci USA 103(43):15927–15932
Rosenberg MJ, Agarwala R, Bouffard G, Davis J, Fiermonte G, Hilliard MS et al (2002) Mutant deoxynucleotide carrier is associated with congenital microcephaly. Nat Genet 32(1):175–179
Palacin M, Goodyer P, Nunes V, Gasparini P (2001) Cystinuria, 8th edn. The McGraw-Hill Companies, New York
Yuen YP, Lam CW, Lai CK, Tong SF, Li PS, Tam S et al (2006) Heterogeneous mutations in the SLC3A1 and SLC7A9 genes in Chinese patients with Cystinuria. Kidney Int 69(1):123–128
Gaildrat P, Lebbah S, Tebani A, Sudrie-Arnaud B, Tostivint I, Bollee G et al. (2017) Clinical and molecular characterization of Cystinuria in a French cohort: relevance of assessing large-scale rearrangements and splicing variants. Mol Med 5(4):373–389
Palacin M, Nunes V, Font-Llitjos M, Jimenez-Vidal M, Fort J, Gasol E et al (2005) The genetics of heteromeric amino acid transporters. Physiology (Bethesda Md) 20:112–124
Reig N, Chillaron J, Bartoccioni P, Fernandez E, Bendahan A, Zorzano A et al (2002) The light subunit of system b(o,+) is fully functional in the absence of the heavy subunit. EMBO J 21(18):4906–4914
Chillaron J, Font-Llitjos M, Fort J, Zorzano A, Goldfarb DS, Nunes V et al (2010) Pathophysiology and treatment of Cystinuria. Nat Rev Nephrol 6(7):424–434
Calonge MJ, Gasparini P, Chillaron J, Chillon M, Gallucci M, Rousaud F et al (1994) Cystinuria caused by mutations in rBAT, a gene involved in the transport of cystine. Nat Genet 6(4):420–425
Pras E, Kochba I, Lubetzky A, Pras M, Sidi Y, Kastner DL (1998) Biochemical and clinical studies in Libyan Jewish Cystinuria patients and their relatives. Am J Med Genet 80(2):173–176
Calonge MJ, Volpini V, Bisceglia L, Rousaud F, de Sanctis L, Beccia E et al (1995) Genetic heterogeneity in Cystinuria: the SLC3A1 gene is linked to type I but not to type III Cystinuria. Proc Natl Acad Sci USA 92(21):9667–9671
Harnevik L, Fjellstedt E, Molbaek A, Tiselius HG, Denneberg T, Soderkvist P (2001) Identification of 12 novel mutations in the SLC3A1 gene in Swedish Cystinuria patients. Hum Mutat 18(6):516–525
Font-Llitjos M, Jimenez-Vidal M, Bisceglia L, Di Perna M, de Sanctis L, Rousaud F et al (2005) New insights into Cystinuria: 40 new mutations, genotype-phenotype correlation, and digenic inheritance causing partial phenotype. J Med Genet 42(1):58–68
Koulivand L, Mohammadi M, Ezatpour B, Kheirollahi M (2015) Cystinuria in a patient with a novel mutation in SLC7A9 gene. Iran J Kidney Dis 9(1):63–66
Koulivand L, Mohammadi M, Ezatpour B, Salehi R, Markazi S, Dashti S et al (2015) Mutation analysis of SLC3A1 and SLC7A9 genes in patients with Cystinuria. Urolithiasis 43(5):447–453
Fazaeli S, Ashouri S, Kheirollahi M, Mohammadi M, Fazilati M (2017) A novel mutation in SLC7A9 gene in Cystinuria. Iran J Kidney Dis 11(2):138–141
Kheirollahi M, Pourreza MR, Khorvash F, Kazemi M, Amini G (2017) A report of a novel mutation in human prostacyclin receptor gene in patients affected with migraine. Iran J Psychiatry 12(3):219–222
Markazi S, Kheirollahi M, Doosti A, Mohammadi M, Koulivand L (2016) A novel mutation in SLC3A1 gene in patients with Cystinuria. Iran J Kidney Dis 10(1):44–47
Baynes C, Healey CS, Pooley KA, Scollen S, Luben RN, Thompson DJ et al (2007) Common variants in the ATM, BRCA1, BRCA2, CHEK2 and TP53 cancer susceptibility genes are unlikely to increase breast cancer risk. Breast Cancer Res 9(2):R27
Solayman M, Saleh MA, Paul S, Khalil MI, Gan SH (2017) In silico analysis of nonsynonymous single nucleotide polymorphisms of the human adiponectin receptor 2 (ADIPOR2) gene. Comput Biol Chem 68:175–185
Martell HJ, Wong KA, Martin JF, Kassam Z, Thomas K, Wass MN (2017) Associating mutations causing Cystinuria with disease severity with the aim of providing precision medicine. BMC Genom 18(Suppl 5):550
Schwarz JM, Rodelsperger C, Schuelke M, Seelow D (2010) MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 7(8):575–576
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Thomas PD, Campbell MJ, Kejariwal A, Mi H, Karlak B, Daverman R et al (2003) PANTHER: a library of protein families and subfamilies indexed by function. Genome Res 13(9):2129–2141
Shihab HA, Gough J, Cooper DN, Stenson PD, Barker GL, Edwards KJ et al (2013) Predicting the functional, molecular, and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum Mut 34(1):57–65
Capriotti E, Calabrese R, Casadio R (2006) Predicting the insurgence of human genetic diseases associated to single point protein mutations with support vector machines and evolutionary information. Bioinformatics 22(22):2729–2734
Li B, Krishnan VG, Mort ME, Xin F, Kamati KK, Cooper DN et al (2009) Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25(21):2744–2750
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22(2):195–201
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42(Web Server issue):W252–W258
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30(Suppl 1):S162–S173
Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009) The SWISS-MODEL repository and associated resources. Nucleic Acids Res 37(Database issue):D387–D392
Morris JH, Huang CC, Babbitt PC, Ferrin TE (2007) structureViz: linking Cytoscape and Chimera UCSF. Bioinformatics 23(17):2345–2347
Doncheva NT, Klein K, Domingues FS, Albrecht M (2011) Analyzing and visualizing residue networks of protein structures. Trends Biochem Sci 36(4):179–182
Saito R, Smoot ME, Ono K, Ruscheinski J, Wang P-L, Lotia S et al (2012) A travel guide to Cytoscape plugins. Nat Methods 9(11):1069–1076
Papadopoulos JS, Agarwala R (2007) COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23(9):1073–1079
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S et al (2004) UniProt: the universal protein knowledgebase. Nucleic Acids Res 32(Database issue):D115–D119
Desmet FO, Hamroun D, Lalande M, Collod-Beroud G, Claustres M, Beroud C (2009) Human splicing finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res 37(9):e67
Wong KA, Wass M, Thomas K (2016) The role of protein modelling in predicting the disease severity of Cystinuria. Eur Urol 69(3):543–544
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Mahdavi, M., Koulivand, L., Khorrami, M. et al. In silico analysis of SLC3A1 and SLC7A9 mutations in Iranian patients with Cystinuria. Mol Biol Rep 45, 1165–1173 (2018). https://doi.org/10.1007/s11033-018-4269-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-018-4269-6