Abstract
The Bloom syndrome (BS) is an autosomic recessive disorder comprising a wide range of abnormalities, including stunted growth, immunodeficiency, sun sensitivity and increased frequency of various types of cancer. Bloom syndrome cells display a high level of genetic instability, including a 10-fold increase in the sister chromatid exchanges (SCE) level. Bloom syndrome arises through mutations in both alleles of the BLM gene, which was identified as a member of the RecQ helicase family. In this study, we screened a Tunisian family with three BS patients. Cytogenetic analysis showed several chromosomal aberrations, and an approximately 14-fold elevated SCE frequency in BS cells. A significant increase in SCE frequency was observed in some family members but not reaching the BS patients values, leading to suggest that this could be due to the heterozygous profile. Microsatellite genotyping using four fluorescent dye-labeled microsatellite markers revealed evidence of linkage to BLM locus and the healthy members, sharing higher SCE frequency, showed heterozygous haplotypes as expected. Additionally, the direct BLM gene sequencing identified a novel homozygous frameshift mutation c.3617–3619delAA (p.K1207fsX9) in BS patients and a heterozygous BLM mutation in the family members with higher SCE frequency. Our findings suggest that this latter mutation likely leads to a reduced BLM activity explaining the homologous recombination repair defect and, therefore, the increase in SCE. Based on the present data, the screening of this mutation could contribute to the rapid diagnosis of BS. The genetic confirmation of the mutation in BLM gene provides crucial information for genetic counseling and prenatal diagnosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Bloom syndrome (BS, MIM# 210900) is a rare human autosomic recessive disorder associated with growth retardation, immunodeficiency, sunlight sensitivity and an increased risk of malignancy at a young age [1–4]. Besides these clinical manifestations, the BS cells exhibit a number of cytogenetic abnormalities, including chromosome breakage, quadriradial chromatid interchanges, and especially a 10-fold elevated sister chromatid exchanges (SCEs) level, which is the hallmark of BS [2].
Bloom syndrome is caused by inactivating mutations in both copies of the BLM gene which is located on chromosome 15 at 15q26.1 and encodes a 1,417 amino acid protein (159 kDa), homologous to the RecQ subfamily of DExH box-containing DNA and RNA helicases [5]. BLM protein consists of seven domains: The amino terminus poly-aspartate domain (Poly D1), poly-serine domain (Poly S), poly-aspartate domain (Poly D2), DEAH helicase domain (DEAH) [6], RecQ helicase C-terminal domain (RecQCt) [6], helicase and RNase D C-terminal domain (HRDC) [7], and nuclear localization signals (NLS) [8] (Fig. 1).
Schematic depiction of distribution of the wild-type BLM protein and the identified mutations. BLM protein consists of seven domains, from the amino terminus poly-aspartate domain (Poly D1), poly-serine domain (Poly S), poly-aspartate domain (Poly D2), DEAH helicase domain (DEAH), RecQ helicase C-terminal domain (RecQCt), helicase and RNase D C-terminal domain (HRDC), and nuclear localization signals (NLS). IR intervening region. All currently known mutations are presented in boxes according http://bioinf.uta.fi/BLMbase/
Like other RecQ family helicases, BLM interacts physically and functionally with numerous proteins in the cell to perform its diverse functions [9–13]. BLM helicase activity is necessary for genomic instability correction. Thus, several in vitro and in vivo studies demonstrated that BLM unwinds the canonical Watson–Crick duplex, and recognizes and disrupts alternative DNA structures including the Holliday junctions, the triple helices and the highly stable G-quadruplex [14–19].
Up to date, a number of nonsense or frameshift mutations and missense mutations are found in the BLM gene of BS patients, according to the previous reports [20–22]. Frameshift mutations are found along the entire length of BLM gene, whereas all missense mutations affect the DEAH helicase or the RecQ-CT domains (Fig. 1).
In the present study, we performed clinical, cytogenetic, microsatellite genotyping and mutational analyses on a Tunisian consanguineous family with BS. We identified several chromosomal aberrations (CA) and an approximately 14 fold-elevated SCE frequency in BS cells associated with a novel homozygous frameshift mutation c.3617–3619delAA (p.K1207fsX9). Heterozygous BLM mutation was found to be associated with an SCE increase in the healthy family members, a finding which has not been reported elsewhere.
Materials and methods
Studied subjects
A consanguineous family (Is.) (12 members) from Sidi Bouzid (South of Tunisia) with three affected males (Fig. 2) referred to the Dermatology Department, Hedi Chaker Hospital, Sfax, Tunisia. The age of the affected individuals ranged from 27 to 38 years. For the present study, fifty (50) control individuals were also tested. A questionnaire on clinical information and family history was drawn up. Informed consent was obtained from all the family members and control individuals in accordance with the ethic committee of the University Hospital of Sfax, Tunisia.
CA and SCE analysis
Separate lymphocyte cultures were set up for CA and SCEs assays. A detailed description of CA and SCE assays was reported in our previous studies [22, 23].
Microsatellite genotyping
Four (4) fluorescent dye-labeled microsatellite markers (D15S996, D15S130, D15S127 and D15S158) spanning the BLM locus (15q26.1) were selected on the basis of their map position and heterozygosity coefficient. The distances between the two close markers (D15S996 and D15S127) and the BLM gene are 39 and 273 Kb, respectively. The markers were genotyped for the family members after DNA extraction according to the phenol–chloroform protocol [24]. True Allele PCR Premix (Applied Biosystems, Foster City, CA, USA) was used for PCR reactions according to the manufacturer’s instructions. Fluorescent labelled alleles were analysed on an ABI PRISM 3100-Avant automated Genetic Analyser (Applied Biosystems). The genotypes were determined using GenScan software (Applied Biosystems). The construction of haplotypes in genotyped members was done regardless of the individual’s affection status.
PCR amplification of the BLM gene
Mutational analysis was performed by PCR amplification of each of the 21 encoding exons of BLM gene and the intron–exon boundaries using appropriate primers chosen so that at least 80 bp of flanking intronic sequences were readable. Primers were shown in our previous study [22]. Intronic sequences containing known polymorphic sites were avoided to prevent false-negatives resulting from the amplification of single allele. PCR amplification of each exon was performed in a thermal cycler (Gene Amp PCR system 9700, Applied Biosystem) [22].
Sequencing of the BLM gene and bioinformatics analysis
PCR product was purified using exonuclease before sequencing. Each exon was sequenced on both strands. The region containing putative novel variations was sequenced twice on both strands to exclude that they were PCR artefacts. Direct sequencing of PCR products was performed in an ABI PRISM 3100-Avant automated DNA sequencer using the Big Dye Terminator Cycle Sequencing reaction kit v1.1 (Applied Biosystems). The identified mutation of the BLM gene was investigated by direct sequencing in 50 Tunisian healthy individuals. The BLAST homology searches were performed using NCBI website (http://www.ncbi.nlm.nih.gov/).
Results
A consanguineous family (12 members) from Sidi Bouzid (South of Tunisia) with three affected males (Fig. 2) referred to the Dermatology Department, Hedi Chaker Hospital, Sfax, Tunisia.
Clinical characteristics of BS patients
The physical and biological data collected during the follow-up are summarized in Table 1. The medical history revealed a proportionate dwarfism, small stature, reduced head circumference, telangiectatic erythema with sun sensitivity (see supplementary Figure). According to the questionnaire data, the BS patients had a smoking habit of more than 10 years. However, fortunately, none of the patients had cancer at their most recent follow-up visit when they were between 5 and 38 years old.
Cytogenetic analysis
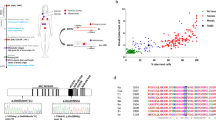
The results indicated multiple CA in the BS cells such as chromatid breaks, chromosome breaks, acentrics, dicentrics, gaps, fragile sites, and especially telomeric associations, which are characteristic of BS cells (Fig. 3). Furthermore, an approximately 14-fold increase in SCE levels was found in BS patients compared to the control group (7.34 ± 2.31) (p < 0.001) (Table 2). The mean frequencies of SCE per cell were 96.39 ± 18.40, 113.40 ± 22.32, and 110.50 ± 12.52 in BS patients Is.1, Is.2 and Is.3, respectively (Table 2). Interestingly, an unexpected heterogeneity was observed in the mean frequency of SCE within the healthy family members. So, compared with control group, a significant increase was found in the mother (I.1) (10.61 ± 2.10 SCE/cell), in the healthy brother (II.5) (10.49 ± 3.47 SCE/cell) (Fig. 3), and in the two daughters (III.1 and III.2) of the patient Is.1 (11.40 ± 5.23 SCE/cell and 10.67 ± 3.76 SCE/cell) (p < 0.05).
Metaphases observed with CA assay (a, b) and SCE assay (c, d). CA are represented with the following red symbols: Chromatid breaks, stars; acentric fragment, bold arrow; dicentric chromosome, square; fragiles sites, circles; telomeric associations, triangles. Three of 168 and 22 SCE are represented by black arrows in cells from a BS patient (c) and a healthy brother (d), respectively
Microsatellite genotyping
According to the heterogeneity of SCE frequencies among the healthy family and since four of the nine members shared a significantly higher SCE frequency, we hypothesized that these members might be heterozygous for BLM mutation. Before BLM gene sequencing, we performed a haplotype analysis on all the family members using four fluorescent dye-labeled polymorphic microsatellite markers covering all BLM locus. This analysis revealed evidence of linkage to BLM mapping in chromosome 15q. Bloom syndrome patients showed a homozygous haplotype for alleles 194, 145, and 84 bp of D15S996, D15S127, and D15S158 microsatellite markers, respectively (Fig. 4). As expected by SCE data, heterozygous haplotypes were observed in members I.1, II.5, III.1 and III.2 (Fig. 4).
Microsatellite genotyping and mutation analyses. a Pedigree of the family showing the segregation of BLM haplotypes and the inheritance of the c.3617–3619delAA mutation. The distances between the two markers D15S996 and D15S127 and the BLM gene are 273 and 39 Kb, respectively. The part of haplotypes not boxed is non-informative for all patients and their relatives. +: Presence of c.3617–3619AA. b Sequence chromatograms showing the presence of the new homozygous c.3617–3619delAA mutation in exon 19 in the studied patients (proband) (at the middle, nucleotides deletions are indicated by an arrow), the heterozygous mutation profile in some healthy family members (at the right) and its absence in a control (at the left). Mutation profiles in family members are represented with the following symbols: homozygous BLM mutation, black squares; heterozygous BLM mutation, half squares and half circles in black; homozygous wild-type; white circles
BLM gene mutational analysis
To further strengthen this hypothesis and confirm the haplotypes data, we performed BLM gene sequencing for all family members. The entire BLM gene coding sequence of (exon 2–22) was, therefore, analysed, including all intron–exon junctions and at least 80 bp of intronic sequence flank each exon. A novel homozygous mutation c.3617–3619delAA was identified in BS patients as predicted by the consanguinity of their parents and the markers haplotype (Fig. 4). This mutation occurred within exon 19 of the BLM gene and predicted to generate a premature termination codon (p.K1207fsX9). The healthy members (I.1, II.5, III.1 and III.2) sharing higher SCE frequencies were found to be heterozygous for the BLM mutation; while, it was absent in the rest of family members and the 50 healthy controls (Fig. 4).
Discussion
Our study reports clinical, cytogenetic, microsatellite and mutational analyses on three patients, belonging to a consanguineous Tunisian family, with typical BS clinical features. Cytogenetic analysis by SCE and CA assays confirmed the suspected diagnosis of the patients. Indeed, in line with previous reports, BS cells display chromosome breaks, spontaneous symmetric quadriradial interchanges and an increase in SCE levels [2, 22, 25]. However, in our study, the SCE mean frequencies in BS patients seemed to be higher than those reported in previous study [2, 22, 25]. It was an approximately 14-fold higher than control frequency especially, in patients Is.2 and Is.3. This finding could be due to the smoking effect, according to our previous study [23] that reported an increase of SCE frequency in higher smokers (smoking habit >10 years).
In somatic cells, SCEs are mediated by homologous recombination (HR) [26]. It has been reported in Sonoda study that HR uses nascent sister chromatid to repair potentially lethal DNA lesions accompanying replication, which might explain the lethality or tumorigenic potential associated with defects in HR or HR-associated proteins [26]. In BS cells, the BLM protein unwinds DNA structures mimicking replication forks and HR intermediates in combination with topoisomerase IIIα, RMI1 and RMI2 (BTR complex) to dissolve double Holliday junctions [17, 27–30].
In addition, it has been reported that BLM interacts with RAD51 and can efficiently disrupt D-loops and might, therefore, act to prevent inappropriate template usage during HR [31–33]. BLM might suppress SCEs through the dismantling of the D-loops to promote synthesis-dependent strand annealing, a pathway of HR that prevents crossovers formation and thus SCEs [32, 33]. Therefore, the BS cells defective for BLM protein could engender an elevated frequency of homologous and non-homologous tri and quadriradial chromosomes and SCEs.
Interestingly, in this study, an unexpected heterogeneity in SCE frequency in the healthy family members was observed. So, a significant increase in SCE frequency was noted in some family members, but did not lead the BS patient’s values. This finding has not been observed elsewhere and, therefore, leads us to suggest that this could be due to the heterozygous profile of BLM gene since these members were non-smokers and without any indication of previous occupational/environmental genotoxic exposure or other agents suspicious of genotoxicity which could increase their SCE rates.
To confirm our hypothesis, we performed a haplotype analysis on all the family members using microsatellite markers covering the BLM gene. This analysis revealed evidence for linkage to BLM mapping to chromosome 15, and, therefore a homozygous haplotype was observed in BS patients, which supported our hypothesis that the healthy members sharing higher SCE frequency might be heterozygous for BLM gene. To strengthen this suggestion, we investigated a direct BLM gene sequencing.
Mutational analysis of BLM gene revealed a novel homozygous mutation c.3617–3619delAA in exon 19 of BS patients; however, as expected by haplotype analysis, a heterozygous BLM mutation was found in the healthy family members sharing higher SCE frequency. This suggests that this heterozygous mutation might reduce BLM activities explaining the HR defects and, therefore, the increase in SCE. This is in disagreement with Ellis’ study showing normal SCE rates in BS cells with at least one non mutant BLM allele [34].
The c.3617–3619delAA mutation seemed to be specific to the Tunisian population, consistent with the considerable mutations diversity affecting the BLM gene (http://bioinf.uta.fi/BLMbase/). The c.3617–3619delAA mutation introduced a premature stop codon (p.K1207fsX9) in the intervening region 5 (IR5) situated between RecQCt domain and HRDC domain. As a result, this small deletion likely leads to a truncated protein containing an intact DEAH helicase and the RecQ-CT domains located in the first 1,077 amino acids [6] but lacking the HRDC domain situated between amino acids 1,212–1,292 [7] and NLS domain located between amino acids 1,334–1,349 [8] in the C-terminal region of BLM protein (see Fig. 1).
The C-terminus may confer on BLM protein the ability to recognize and bind abnormal DNA structures such as quadruplexes [35]. It has been reported in Wu’s studies that C-terminus deletions have a stronger negative effect on genomic stability than those occurring in N-terminus domain and that the mutant proteins can be incorporated into normal BLM-containing complexes and inhibit their function or regulation [36]. Helicase and RNase D C-terminal domain residing in the C-terminal region of BLM protein [7], is a critical determinant of the dissolution function of double Holliday junctions by the BLM–Topoisomerase IIIa complex [37]. In addition, Killoran (2006) reported that the HRDC domain could independently bind DNA and, consequently, the RecQ helicases containing mutations in the HRDC domain have altered DNA structure specific binding and unwinding properties [38]. Recently, Amor Guéret group (2008) identified a homozygous frameshift mutation (p.Ser1196fsX3) in the IR5 domain (Fig. 1), leading to a truncated BLM protein upstream to the HRDC domain; however, no data are available for the correlation between heterozygous mutation and cell phenotype [21].
Overall, in this study, we described a higher chromosomal instability in Tunisian BS patients associated with a novel homozygous fameshift mutation leading to the deletion of a BLM protein upstream to the HRDC domain. Additionally, we established that the heterozygous profile of this mutation seemed to be associated with BLM defects and higher SCE frequency in healthy family members. This identification may allow a genetic counselling in the relatives of this family and an antenatal diagnosis which is now possible.
References
German J (1974) Bloom’s syndrome. II. In: German J (ed) Chromosomes and cancer. Wiley, New Yok, pp 601–617
German J (1993) Bloom syndrome: a mendelian prototype of somatic mutational disease. Medicine 72:393–406
Diaz A, Vogiatzi MG, Sanz MM, German J (2006) Evaluation of short stature, carbohydrate metabolism and other endocrinopathies in Bloom’s syndrome. Horm Res 66(3):111–117
Amor-Gueret M (2006) Bloom syndrome, genomic instability and cancer: the SOS-like hypothesis. Cancer Lett 236(1):1–12
Ellis NA, Groden J, Ye TZ, Straughen J, Lennon DJ, Ciocci S, Proytcheva M, German J (1995) The Bloom’s syndrome gene product is homologous to RecQ helicases. Cell 83:655–666
Bahr A, De Graeve F, Kedinger C, Chatton B (1998) Point mutations causing Bloom’s syndrome abolish ATPase and DNA helicase activities of the BLM protein. Oncogene 17:2565–2571
Morozov V, Mushegian AR, Koonin EV, Bork P (1997) A putative nucleic acid-binding domain in Bloom’s and Werner’s syndrome helicases. Trends Biochem Sci 22:417–418
Kaneko H, Orii KO, Matsui E, Shimozawa N, Fukao T, Matsumoto T, Shimamoto A, Furuichi Y, Hayakawa S, Kasahara K, Kondo N (1997) BLM (the causative gene of Bloom syndrome) protein translocation into the nucleus by a nuclear localization signal. Biochem Biophys Res Commun 240:348–353
Hickson ID (2003) RecQ helicases: caretakers of the genome. Nat Rev Cancer 3:169–178
Temime-Smaali N, Guittat L, Wenner T, Bayart E, Douarre C, Gomez D, Giraud-Panis MJ, Londono-Vallejo A, Gilson E, Amor-Guéret M, Riou JF (2008) Topoisomerase IIIa is required for normal proliferation and telomere stability in alternative lengthening of telomeres. EMBO J 27:1513–1524
Grierson PM, Lillard K, Behbehani GK, Combs KA, Bhattacharyya S, Acharya S, Groden J (2012) BLM helicase facilitates RNA polymerase I-mediated ribosomal RNA transcription. Hum Mol Genet 21(5):1172–1183
Machwe A, Karale R, Xu X, Liu Y, Orren DK (2011) The Werner and Bloom syndrome proteins help resolve replication blockage by converting (regressed) holliday junctions to functional replication forks. Biochemistry 50(32):6774–6788
Bachrati CZ, Hickson ID (2008) RecQ helicases: guardian angels of the DNA replication fork. Chromosoma 117:219–233
Neff NF, Ellis NA, Ye TZ, Noonan J, Huang K, Sanz M, Proytcheva M (1999) The DNA helicase activity of BLM is necessary for the correction of the genomic instability of bloom syndrome cells. Mol Biol Cell 10:665–676
Brosh RM Jr, Majumdar A, Desai S, Hickson ID, Bohr VA, Seidman MM (2001) Unwinding of a DNA triple helix by the Werner and Bloom syndrome helicases. J Biol Chem 276:3024–3030
Mohaghegh P, Karow JK, Brosh JR Jr, Bohr VA, Hickson ID (2001) The Bloom’s and Werner’s syndrome proteins are DNA structure-specific helicases. Nucleic Acids Res 29:2843–2849
Wu L, Hickson ID (2003) The Bloom’s syndrome helicase suppresses crossing over during homologous recombination. Nature 426:870–874
Brown AD, Claybon AB, Bishop AJ (2011) A conditional mouse model for measuring the frequency of homologous recombination events in vivo in the absence of essential genes. Mol Cell Biol 31(17):3593–3602
Cheok CF, Wu L, Garcia PL, Janscak P, Hickson ID (2005) The Bloom’s syndrome helicase promotes the annealing of complementary single-stranded DNA. Nucleic Acids Res 33:3932–3941
German J, Sanz MM, Ciocci S, Ye TZ, Ellis NA (2007) Syndrome-causing mutations of the BLM gene in persons in the Bloom’s Syndrome Registry. Hum Mutat 28(8):743–753
Amor-Gueret M, Dubois-d’Enghien C, Lauge A, Onclercq-Delic R, Barakat A, Chadli E, Bousfiha AA, Benjelloun M, Flori E, Doray B, Laugel V, Lourenço MT, Gonçalves R, Sousa S, Couturier J, Stoppa-Lyonnet D (2008) Three new BLM gene mutations associated with Bloom syndrome. Genet Test 2:257–261
Ben Salah G, Hadj Salem I, Masmoudi A, Ben Rhouma B, Turki H, Fakhfakh F, Ayadi H, Kamoun H (2013) Chromosomal instability associated with a novel BLM frameshift mutation (c.1980–1982delAA) in two unrelated Tunisian families with Bloom syndrome. J Eur Acad Dermatol Venereol. Oct 1. doi: 10.1111/jdv.12279
Ben Salah G, Kamoun H, Rebai A, Ben Youssef A, Ayadi H, Belghith-Mahfoudh N, Fourati A, Ayadi H, Fakhfakh F (2011) Sister chromatid exchange (SCE) and high-frequency cells (HFC) in peripheral blood lymphocytes of healthy Tunisian smokers. Mutat Res 719:1–6
Lewin HA, Stewart-Haynes JA (1992) A simple method for DNA extraction from leukocytes for use in PCR. Biotechniques 13(4):522–524
Chaganti RSK, Schonberg S, German J (1974) A manyfold increase in sister chromatid exchanges in Bloom’s syndrome lymphocytes. Proc Natl Acad Sci USA 71(11):4508–4512
Sonoda E, Sasaki MS, Morrison C, Yamaguchi-Iwai Y, Takata M, Takeda S (1999) Sister chromatid exchanges are mediated by homologous recombination in vertebrate cells. Mol Cell Biol 19:5166–5169
Srivastava V, Modi P, Tripathi V, Mudgal R, De S, Sengupta S (2009) BLM helicase stimulates the ATPase and chromatin-remodeling activities of RAD54. J Cell Sci 122:3093–3103
Mankouri HW, Hickson ID (2007) The RecQ helicase-topoisomerase III-Rmi1 complex: a DNA structure-specific ‘dissolvasome’. Trends Biochem Sci 32(12):538–546
Singh TR, Ali AM, Busygina V, Raynard S, Fan Q, Du CH, Andreassen PR, Sung P, Meetei AR (2008) BLAP18/RMI2, a novel OBfold-containing protein, is an essential component of the Bloom helicase-double Holliday junction dissolvasome. Genes Dev 22:2856–2868
Ralf C, Hickson ID, Wu L (2006) The Bloom’s syndrome helicase can promote the regression of a model replication fork. J Biol Chem 281:22839–22846
Van Brabant AJ, Ye T, Sanz M, German IJ, Ellis NA, Holloman WK (2000) Binding and melting of D-loops by the Bloom syndrome helicase. Biochemistry 39:14617–14625
Bachrati CZ, Borts RH, Hickson ID (2006) Mobile, D-loops are a preferred substrate for the Bloom’s syndrome helicase. Nucleic Acids Res 34:2269–2279
Bugreev DV, Yu X, Egelman EH, Mazin AV (2007) Novel pro- and antirecombination activities of the Bloom’s syndrome helicase. Genes Dev 21:3085–3094
Ellis NA, Proytcheva M, Sanz MM, Ye TZ, German J (1999) Transfection of BLM into cultured bloom syndrome cells reduces the sister-chromatid exchange rate toward normal. Am J Hum Genet 65:1368–1374
Sun H, Karow JK, Hickson ID, Maizels N (1998) The Bloom’s syndrome helicase unwinds G4 DNA. J Biol Chem 273:27587–27592
Yankiwski V, Noonan J, Neff N (2001) The C-terminal domain of the Bloom syndrome DNA helicase is essential for genomic stability. BMC Cell Biol 2:11
Wu L, Chan K, Ralf C, Bernstein D, Garcia P, Bohr V, Vindigni A, Janscak P, Keck J, Hickson I (2005) The HRDC domain of BLM is required for the dissolution of double Holliday junctions. EMBO J 24:2679–2687
Killoran MP, Keck JL (2006) Sit down, relax and unwind: structural insights into RecQ helicase mechanisms. Nucleic Acids Res 34:4098–4105
Acknowledgments
We would like to thank all the volunteers for their cooperation. This work was supported by the Ministry of Higher Education and Scientific Research in Tunisia.
Conflict of interest
We declare that we have no any actual or potential conflict of interest, including financial, personal or any relationships with other people or organizations concerning this work.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2014_3624_MOESM1_ESM.tif
Photographs of Bloom’s syndrome patients. The patient Is.1, 38 years old, showing narrow face, small mandibles and achromic patches of the lips and nose. The patient Is.2, 36 years old, showing erythematous lesions of the nose and lips (TIFF 189 kb)
Rights and permissions
About this article
Cite this article
Ben Salah, G., Hadj Salem, I., Masmoudi, A. et al. A novel frameshift mutation in BLM gene associated with high sister chromatid exchanges (SCE) in heterozygous family members. Mol Biol Rep 41, 7373–7380 (2014). https://doi.org/10.1007/s11033-014-3624-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3624-5