Abstract
The RNA-binding protein Arabidopsis thaliana glycine-rich RNA-binding protein 7 (AtGRP7) regulates the steady-state abundance of numerous target transcripts in A. thaliana. Here we show that the GA1 and GA2 transcripts encoding the first enzymes of the gibberellin biosynthetic pathway are expressed at reduced levels in transgenic plants ectopically over-expressing AtGRP7 (AtGRP7-ox plants). Furthermore, the levels of the bioactive phytohormone GA4 as well as of several intermediates of the GA biosynthetic pathway are reduced in AtGRP7-ox plants. The transgenic plants show a reduced length of the vegetative stem. The application of exogenous GA largely reverses the phenotype by increasing the number of vegetative internodes. AtGRP7-ox plants flower with fewer leaves than wt plants, suggesting that the floral promotive effect of AtGRP7 bypasses the effect of a reduced GA level in AtGRP7-ox plants. Upon GA treatment, AtGRP7-ox plants flower only slightly earlier than wild type plants. Thus, exogenous GA has only a small additional effect in reducing the number of leaves at the onset of flowering in AtGRP7-ox plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The RNA-binding protein Arabidopsis thaliana glycine-rich RNA-binding protein 7 (AtGRP7) is a small glycine-rich RNA-binding protein with a single RNA recognition motif at its N-terminus and a C-terminus consisting mainly of glycine repeats. The AtGRP7 transcript undergoes circadian, i.e. 24 h oscillations in steady-state abundance with a peak at the end of the daily light phase [1, 2]. Furthermore, AtGRP7 responds to cold and oxidative stress [2–4]. AtGRP7 has also been shown to participate in the defence against bacterial pathogens [5–8]. Accordingly, transcripts encoding PATHOGENESIS RELATED proteins associated with salicylic acid-mediated defence responses are constitutively elevated in transgenic plants ectopically expressing AtGRP7 (AtGRP7-ox plants) whereas defensins associated with jasmonic acid and ethylene responses are expressed at reduced levels [9, 10].
AtGRP7-ox plants flower earlier than wild type (wt) plants, particularly in short photoperiods [11]. The transition to flowering is controlled by a network of interwoven signaling pathways in response to environmental and developmental factors [12–14]. Long photoperiods promote floral transition via the photoperiodic pathway. Vernalization, the exposure to an extended period of cold, promotes flowering by down-regulating the key floral repressor FLOWERING LOCUS C (FLC). FLC is also down-regulated by endogenous regulators, collectively referred to as the autonomous pathway, independently of environmental stimuli. AtGRP7-ox plants still flower earlier in long days (LDs) than in short days (SDs) and thus retain the response to inductive photoperiods. The effect on flowering time is largely mediated by a reduced level of FLC in the AtGRP7-ox plants. Because AtGRP7-ox plants still respond to vernalization, AtGRP7 acts at least partly via the autonomous pathway that promotes flowering independent of environmental stimuli [11].
Global transcript profiling has identified a suite of target transcripts that show altered steady state abundances or splicing patterns upon ectopic expression of AtGRP7 in transgenic plants [9, 15, 16]. Among the transcripts with reduced steady-state abundance in AtGRP7-ox plants was GA2 encoding an enzyme involved in gibberellin biosynthesis.
Gibberellins (GAs) control many aspects of plant physiology including germination, transition to flowering, flower development, fertility, pollen tube growth, and pathogen responses [17–24]. In particular, GA is required for stem elongation in rosette plants to overcome the growth-repressive effect of the DELLA proteins that negatively regulate GA signaling [25]. Mutants defective in GA biosynthesis, ga1, ga2, and ga3 are extreme dwarfs. GA application in turn promotes elongation growth. Upon binding of GA, the GIBBERELLIN INSENSITIVE DWARF 1 (GID1) receptor undergoes a conformational change to interact with the DELLA proteins. The GID1-GA-DELLA complex targets the DELLA proteins for proteasomal degradation [26]. Furthermore, GAs promote floral transition via activation of the floral integrators SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1 and LEAFY at the shoot apical meristem [27]. This so-called GA pathway is predominantly active in noninductive SDs: The ga1-3 mutant, which is impaired in GA biosynthesis, does not flower in SDs but has a relatively weak late-flowering phenotype in LDs [28].
The action of the GA pathway has long been thought to be largely masked by the photoperiodic pathway under LD conditions [29]. More recently, GAs have been assigned a role in floral induction in response to inductive LDs through activation of FLOWERING LOCUS T transcription in leaves and of the SQUAMOSA PROMOTER BINDING PROMOTER LIKE genes in the meristem [20].
Greater than one hundred GA derivatives have been identified in plants. The most common bioactive components are GA1, GA3, and GA4 [17]. GA biosynthesis starts from the precursor geranylgeranyl diphosphate that is converted to ent-kaurene by ent-copalyl diphosphate synthase (GA1) and ent-kaurene synthase (GA2). Ent-kaurene is then converted to GA12 by ent-kaurene oxidase (GA3) and ent-kaurenoic acid oxidase. GA12 is converted to the bioactive GA4 through oxidation on C-20 by GA 20-oxidase and on C-3 by GA3-oxidase, respectively. In a parallel pathway, GA12 is also a substrate for GA13-oxidase to produce GA53, a precursor for GA1 in the 13-hydroxylation pathway. Whereas transcription factors that regulate the expression of genes encoding GA biosynthesis enzymes have been well described, only little is known about RNA-binding proteins that may affect the expression at the post-transcriptional level [17, 30, 31].
Here we investigate the impact of constitutive over-expression of the RNA-binding protein AtGRP7 on the expression of genes encoding GA biosynthesis enzymes. Furthermore, we characterize AtGRP7-ox plants with respect to endogenous GA levels and GA-related traits including vegetative stem growth and flowering time.
Materials and methods
Plant growth
The genotypes used were AtGRP7-ox in the Col-2 and L. er background and the corresponding wild types [11, 32]. Seeds were sown on soil, stratified at 4 °C for 2 days, and germinated and grown in short days (8 h light/16 h dark cycles) at 20 °C. To assess elongation growth, the length of the vegetative stem (from rosette to the first node producing inflorescences) and of the vegetative internodes was measured [33].
RNA isolation and RT PCR transcript analysis
AtGRP7-ox and Col-2 wt plants were grown on soil for 8 weeks in SDs and harvested at zt 8. Isolation of total RNA was performed as described [34]. For Real time PCR, total RNA was treated with DNaseI and reverse-transcribed using Superscript II (Invitrogen). The absence of amplification products from genomic DNA was confirmed in non-retro-transcribed controls. Twenty ng of cDNA was amplified in the presence of SYBR Green using an initial denaturation step of 2 min, followed by 45 cycles of 20 s at 94 °C, 30 s at 60 °C and 40 s at 68 °C (MJ research Opticon DNA Engine). CT values were determined and relative expression levels for the analyzed transcripts were calculated based on non-equal efficiencies for each primer pair [35]. Data were normalized to the PP2A transcript. Primers are listed in Table 1.
GA determination
AtGRP7-ox plants and wt plants were grown on soil in a randomized fashion for 8 weeks in SDs and harvested at zt8 (zeitgeber time 8, 8 h after lights on). Five plants were pooled per biological replicate. The analysis of GA levels was done as described previously [36]. Data are expressed as means of three biological replicates ± SE.
Determination of flowering time
Plants were grown in a randomized fashion on soil in SDs at 20 °C in Percival incubators AR66-L3 (CLF Laboratories). For GA treatment, plants were sprayed with 100 μM GA3 twice a week starting at day 10 after stratification in the middle of the light period (zt4–zt6). Mock treatment was performed by spraying with 0.1 % DMF/0.02 % Tween 20. Flowering time was determined by counting the rosette leaves once the bolt was 0.5 cm tall. Mean values ± SD were calculated [11]. For statistical analysis, ANOVA followed by a Dunnett test and a factorial ANOVA were performed.
Results and discussion
GA biosynthesis transcripts are reduced in AtGRP7-ox plants
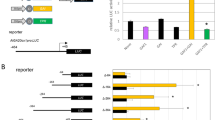
Global transcript analysis using the Affymetrix ATH1 GeneChip revealed that in transgenic plants ectopically expressing AtGRP7, the steady-state abundance of GA2 encoding ent-kaurene synthase catalyzing the second step in gibberellin biosynthesis is reduced [9]. To investigate a potential influence of AtGRP7 on GA biosynthesis in more detail, transcript levels of genes involved in GA biosynthesis were determined in independent transgenic lines. AtGRP7-ox and Col-2 wt plants were grown on soil for 8 weeks in SDs and harvested at zt 8. In independent transgenic lines, the GA2 level was 40 % of that in wt plants (Fig. 1). Furthermore, the level of GA1 encoding the first biosynthetic enzyme ent-copalyl diphosphate synthase was reduced to 60 % of that in wt plants. Transcript levels encoding enzymes involved in later steps of GA biosynthesis or GA catabolism were not strongly affected by ectopic AtGRP7 expression ([9] and data not shown). Steady-state abundance of the GA responsive transcript GASA9, a member of the GA-STIMULATED IN ARABIDOPSIS family, was also reduced in AtGRP7-ox plants compared to wt plants (Fig. 1). None of these transcripts undergoes high amplitude circadian oscillations (data not shown).
Transcript levels of GA biosynthesis and GA responsive genes in AtGRP7-ox lines. AtGRP7-ox and wt plants were grown in SD conditions and harvested at zt8. The levels of GA1, GA2, and GASA9 transcripts were measured by real time PCR. Transcript levels were normalized to PP2A and are expressed relative to wt (set to 1) based on at least two biological replicates with two technical replicates each. For statistical analysis a student′s t test was applied (*p < 0.05, **p < 0.01)
GA levels are reduced in AtGRP7-ox plants
As the levels of the transcripts encoding GA biosynthetic enzymes are slightly, but significantly reduced in AtGRP7-ox plants, we tested whether GA levels may be likewise affected. AtGRP7-ox plants and Col wt plants were grown on soil for 8 weeks in SDs and harvested at zt8. GA levels were determined as described previously [36].
The level of bioactive GA4 was strongly reduced in AtGRP7-ox plants compared to Col-2 wt plants (Table 2). The GA12, GA15, GA24 and GA9 precursors were also reduced. Levels of GAs of the 13-hydroxylation pathway were also reduced as determined for precursors GA53 and GA19 (Table 2). Levels for other GAs of the 13-hydroxylation pathway (GA44, GA20, GA1, and GA8) were below the detection limit (data not shown). The same trend was observed in an independent AtGRP7-ox line in the Landsberg background compared to L. er wt plants (data not shown). Thus, our data suggest that AtGRP7 may influence GA levels by regulating enzymes of GA biosynthesis. Whereas transcriptional regulators of GA biosynthesis have been well described, so far only one predicted RNA-binding protein has been suggested to affect GA biosynthesis at the post-transcriptional level [17, 30]. For example, a T-DNA insertion in the 3′untranslated region of the pioneer protein At4g20010 with two RNA-binding domains leads to reduced GA4 levels [31].
AtGRP7-ox plants show a dwarf phenotype that can be reverted by exogenous GA
Plants lacking sufficient levels of GA have long been known for their dwarfed phenotypes. As AtGRP7-ox plants have a somewhat stunted growth phenotype (Fig. 2a), we tested whether this correlates with the reduced GA levels. Indeed, treatment of AtGRP7-ox plants with GA3 reversed the growth phenotype (Fig. 2b). The effect was quantified by measuring the height of the vegetative stem and the number and length of vegetative internodes. In mock-treated AtGRP7-ox plants, the vegetative stem was significantly shorter than in wt plants. GA treatment increased the length of the stem in both wt and AtGRP7-ox plants. GA-treated AtGRP7-ox plants remained significantly smaller than GA-treated wt plants but reached the height of untreated wild type plants (Fig. 3a). The number of elongated vegetative internodes was lower in untreated AtGRP7-ox plants than in untreated wt plants. Upon GA treatment, the number of internodes increased considerably and was about the same in both AtGRP7-ox and wt plants (Fig. 3b). Furthermore, the length of the vegetative internodes was smaller in AtGRP7-ox plants than in wt plants (Fig. 3c). Upon GA treatment, the length of the vegetative internodes was reduced in both AtGRP7-ox and wt plants, and vegetative internodes were still shorter in AtGRP7-ox plants compared to wt plants (Fig. 3c). Thus, GA was found to promote the number of internodes that elongate upon bolting and reduce the length of the vegetative internodes, as previously observed [33]. The dwarf phenotype of AtGRP7-ox plants is reverted by GA mainly via an increase in the number of vegetative internodes.
Growth of the vegetative stem in AtGRP7-ox and wt plants upon GA treatment. AtGRP7-ox plants in the Col background were grown along with the wt plants in SD conditions and either mock-treated or sprayed with 100 μM GA3 twice a week starting at day 10 after stratification. The length of the vegetative stem below the uppermost cauline leaf (a), the number of vegetative internodes (b) and the length of the vegetative internodes was recorded for 10–17 plants per line. For statistical analysis a student′s t test was applied. A statistically significant difference between AtGRP7-ox plants and wt (mock-treated or GA3-treated as reference) with p value <0.01 is indicated with (*). The response to GA3 treatment was analyzed with factorial ANOVA
Impact of GA on flowering in AtGRP7-ox plants
We have observed previously that AtGRP7-ox plants flower after forming fewer leaves than wt plants predominantly in SDs [11]. Gibberellins have been shown to promote flowering also predominantly in SDs [28]. The accelerated flowering of AtGRP7-ox plants suggests that the reduced GA levels we observed here in the AtGRP7-ox plants do not limit floral transition. AtGRP7 has been implicated in the autonomous pathway and impacts flowering at least partly by down-regulating FLC [11]. Therefore, AtGRP7 over-expression has an effect that bypasses the reduced GA level.
To test whether exogenously applied GA would further decrease the time to flowering in plants ectopically expressing AtGRP7, independent AtGRP7-ox plants in the Col-2 and L. er background and the corresponding wt plants were grown in SDs and sprayed with GA3 twice a week starting at day 10 after stratification. The mock-treated AtGRP7-ox plants flowered with a significantly reduced leaf number compared to their corresponding wild types (p < 0.01) (Fig. 4a). GA promoted flowering both in wt and AtGRP7-ox plants (p < 0.01). AtGRP7-ox plants treated with GA still flowered with slightly fewer leaves than GA-treated wt plants. Thus, AtGRP7-ox plants are able to respond to exogenous GA but the promotive effect on flowering is much smaller than in wt plants (p < 0.01) (Fig. 4b, c).
GA accelerates flowering in AtGRP7-ox plants AtGRP7-ox plants in the Col-2 and Landsberg erecta ecotypes were grown along with the respective wt plants in SDs and sprayed with 100 μM GA3 twice a week starting at day 10 after stratification in the middle of the light period. Leaf number was recorded at 0.5 cm bolt (a). For statistical analysis a student′s t test was applied (p < 0.01). A representative analysis with factorial ANOVA evaluating the reduction of leaf number at bolting in response to GA3 treatment is shown in (b) for two independent AtGRP7-ox lines in the Col-2 background and for an AtGRP7-ox line in the L. er background (p < 0.01) (c)
Conclusion
In transgenic plants ectopically expressing AtGRP7, steady-state abundance of GA2 and GA1 catalyzing the first steps in gibberellin biosynthesis is reduced. Accordingly, significantly reduced levels of the bioactive GA4 and several intermediates were found in AtGRP7-ox plants. The stunted growth of AtGRP7-ox plants was rescued by application of exogenous GA predominantly through an increase in the number of vegetative internodes, suggesting that the phenotype of AtGRP7-ox plants at least partially is due to reduced GA levels.
Intriguingly, AtGRP7-ox plants flowered with fewer leaves than wt plants, particularly in SD conditions, despite reduced GA levels. Thus, the effect of AtGRP7 on floral transition overcomes the reduced GA levels. Treatment with exogenous GA has a small additional effect to AtGRP7 over-expression but GA treated AtGRP7-ox plants flower only slightly earlier than GA treated wt plants. Obviously, GA levels are not limiting for floral transition in AtGRP7-ox plants.
Taken together, our finding that the RNA-binding protein AtGRP7 affects GA biosynthesis and level extends previous findings that AtGRP7 affects jasmonic acid, salicylic acid and abscisic acid response genes [9, 37]. It will be interesting to further define a potential role of AtGRP7 in hormonal cross-talk and unravel the molecular underpinnings, given the nucleo-cytoplasmic shuttling of AtGRP7 and its role in determining steady-state abundance and alternative splicing of target transcripts [9, 15, 38, 39].
References
Heintzen C, Melzer S, Fischer R, Kappeler S, Apel K, Staiger D (1994) A light- and temperature-entrained circadian clock controls expression of transcripts encoding nuclear proteins with homology to RNA-binding proteins in meristematic tissue. Plant J 5:799–813
Carpenter CD, Kreps JA, Simon AE (1994) Genes encoding glycine-rich Arabidopsis thaliana proteins with RNA-binding motifs are influenced by cold treatment and an endogenous circadian rhythm. Plant Physiol 104:1015–1025
Schmidt F, Marnef A, Cheung M-K, Wilson I, Hancock J, Staiger D, Ladomery M (2010) A proteomic analysis of oligo(dT)-bound mRNP containing oxidative stress-induced Arabidopsis thaliana RNA-binding proteins ATGRP7 and ATGRP8. Mol Biol Rep 37:839–845
Schöning JC, Streitner C, Meyer IM, Gao Y, Staiger D (2008) Reciprocal regulation of glycine-rich RNA-binding proteins via an interlocked feedback loop coupling alternative splicing to nonsense-mediated decay in Arabidopsis. Nucleic Acids Res 36:6977–6987
Jeong B-r, Lin Y, Joe A, Guo M, Korneli C, Yang H, Wang P, Yu M, Cerny RL, Staiger D et al (2011) Structure function analysis of an ADP-ribosyltransferase type III effector and its RNA-binding target in plant immunity. J Biol Chem 286:43272–43281
Fu ZQ, Guo M, Jeong BR, Tian F, Elthon TE, Cerny RL, Staiger D, Alfano JR (2007) A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 447:284–288
Nicaise V, Joe A, Jeong B, Korneli C, Boutrot F, Wested I, Staiger D, Alfano JR, Zipfel C (2013) Pseudomonas HopU1 affects interaction of plant immune receptor mRNAs to the RNA-binding protein GRP7. EMBO J 32:701–712
Lee HJ, Kim JS, Yoo SJ, Kang EY, Han SH, Yang K-Y, Kim YC, McSpadden Gardener B, Kang H (2012) Different roles of glycine-rich RNA-binding protein7 in plant defense against Pectobacterium carotovorum, Botrytis cinerea, and tobacco mosaic viruses. Plant Physiol Biochem 60:46–52
Streitner C, Hennig L, Korneli C, Staiger D (2010) Global transcript profiling of transgenic plants constitutively overexpressing the RNA-binding protein AtGRP7. BMC Plant Biol 10:221
Hackmann C, Korneli C, Kutyniok M, Köster T, Wiedenlübbert M, Müller C, Staiger D (2013) Salicylic acid-dependent and –independent impact of an RNA-binding protein on plant immunity. Plant Cell Environ. doi:10.1111/pce.12188
Streitner C, Danisman S, Wehrle F, Schöning JC, Alfano JR, Staiger D (2008) The small glycine-rich RNA-binding protein AtGRP7 promotes floral transition in Arabidopsis thaliana. Plant J 56:239–250
Andres F, Coupland G (2012) The genetic basis of flowering responses to seasonal cues. Nat Rev Genet 13:627–639
Wang J-W, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138:738–749
Srikanth A, Schmid M (2011) Regulation of flowering time: all roads lead to Rome. Cell Mol Life Sci 68:2013–2037
Streitner C, Köster T, Simpson CG, Shaw P, Danisman S, Brown JWS, Staiger D (2012) An hnRNP-like RNA-binding protein affects alternative splicing by in vivo interaction with target transcripts in Arabidopsis thaliana. Nucl Acid Res 40:11240–11255
Streitner C, Simpson CG, Shaw P, Danisman S, Brown JWS, Staiger D (2013) Small changes in ambient temperature affect alternative splicing in Arabidopsis thaliana. Plant Signal Behav 8:e24638
Yamaguchi S (2008) Gibberellin metabolism and its regulation. Annu Rev Plant Biol 59:225–251
Mutasa-Gottgens E, Hedden P (2009) Gibberellin as a factor in floral regulatory networks. J Exp Bot 60:1979–1989
Plackett AR, Thomas SG, Wilson ZA, Hedden P (2011) Gibberellin control of stamen development: a fertile field. Trends Plant Sci 16:568–578
Porri A, Torti S, Romera-Branchat M, Coupland G (2012) Spatially distinct regulatory roles for gibberellins in the promotion of flowering of Arabidopsis under long photoperiods. Development 139:2198–2209
Schwechheimer C (2012) Gibberellin signalling in plants (the extended version). Front Plant Physiol 2:107
Navarro L, Bari R, Achard P, Lison P, Nemri A, Harberd NP, Jones JD (2008) DELLAs control plant immune responses by modulating the balance of jasmonic acid and salicylic acid signaling. Curr Biol 18:650–655
Pimenta Lange MJ, Lange T (2006) Gibberellin biosynthesis and the regulation of plant development. Plant Biol (Stuttg) 8:281–290
Heinrich M, Hettenhausen C, Lange T, Wünsche H, Fang J, Baldwin IT, Wu J (2013) High levels of jasmonic acid antagonize the biosynthesis of gibberellins and inhibit the growth of Nicotiana attenuata stems. Plant J 73:591–606
Hirano K, Ueguchi-Tanaka M, Matsuoka M (2008) GID1-mediated gibberellin signaling in plants. Trend Plant Sci 13:192–199
Harberd NP, Belfield E, Yasumura Y (2009) The angiosperm gibberellin-GID1-DELLA growth regulatory mechanism: how an inhibitor of an inhibitor enables flexible response to fluctuating environments. Plant Cell 21:1328–1339
Blazquez MA, Green R, Nilsson O, Sussman MR, Weigel D (1998) Gibberellins promote flowering of Arabidopsis by activating the LEAFY promoter. Plant Cell 10:791–800
Wilson RN, Heckman JW, Somerville CR (1992) Gibberellin is required for flowering in Arabidopsis thaliana under short days. Plant Physiol 100:403–408
Reeves PH, Coupland G (2000) Response of plant development to environment: control of flowering by day length and temperature. Curr Opin Plant Biol 3:37–42
Magome H, Yamaguchi S, Hanada A, Kamiya Y, Oda K (2004) dwarf and delayed-flowering 1, a novel Arabidopsis mutant deficient in gibberellin biosynthesis because of overexpression of a putative AP2 transcription factor. Plant J 37:720–729
Svensson M, Lundh D, Bergman P, Mandal A (2005) Characterisation of a T-DNA-tagged gene of Arabidopsis thaliana that regulates gibberellin metabolism and flowering time. Funct Plant Biol 32:923–932
Heintzen C, Nater M, Apel K, Staiger D (1997) AtGRP7, a nuclear RNA-binding protein as a component of a circadian-regulated negative feedback loop in Arabidopsis thaliana. Proc Natl Acad Sci USA 94:8515–8520
Rieu I, Ruiz-Rivero O, Fernandez-Garcia N, Griffiths J, Powers SJ, Gong F, Linhartova T, Eriksson S, Nilsson O, Thomas SG et al (2008) The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J 53:488–504
Staiger D, Apel K, Trepp G (1999) The Atger3 promoter confers circadian clock-regulated transcription with peak expression at the beginning of the night. Plant Mol Biol Rep 40:873–882
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucl Acid Res 29:e45
Lange T, Kappler J, Fischer A, Frisse A, Padeffke T, Schmidtke S, Pimenta Lange MJ (2005) Gibberellin biosynthesis in developing pumpkin seedlings. Plant Physiol 139:213–223
Cao S, Jiang L, Song S, Jing R, Xu G (2006) AtGRP7 is involved in the regulation of abscisic acid and stress responses in Arabidopsis. Cell Mol Biol Lett 11:526–535
Ziemienowicz A, Haasen D, Staiger D, Merkle T (2003) Arabidopsis transportin1 is the nuclear import receptor for the circadian clock-regulated RNA-binding protein AtGRP7. Plant Mol Biol 53:201–212
Lummer M, Humpert F, Steuwe C, Schüttpelz M, Sauer M, Staiger D (2011) Reversible photoswitchable DRONPA-s monitors nucleocytoplasmic transport of an RNA-binding protein in transgenic plants. Traffic 12:693–702
Acknowledgments
We thank Anja Liebrandt for technical assistance. This work was supported by the DFG through STA 653/2 and SPP1530 to D.S. and LA880/7-1 to T.L.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Löhr, B., Streitner, C., Steffen, A. et al. A glycine-rich RNA-binding protein affects gibberellin biosynthesis in Arabidopsis . Mol Biol Rep 41, 439–445 (2014). https://doi.org/10.1007/s11033-013-2878-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-013-2878-7