Abstract
Carbonic anhydrase (CA) and phosphoenolpyruvate carboxylase (PEPC) activity were found in different Solanum lycopersicum fruit tissues, predominantly in the locular parenchyma and pericarp. The distribution of the CA and PEPC proteins in the tomato fruit tissues was examined by immunohistolocalization. CA and PEPC proteins were found in all fruit tissues examined as well as in the seeds. Three full length cDNA clones designated SlCA1, SlCA2 and SlCA3 coding for β-carbonic anhydrases (CA; EC 4.2.1.1) were identified and characterized from tomato fruit. SlCA1 and SlCA3 encode two putative cytosolic isoforms whereas SlCA2 encodes a putative plastidial isoform. Quantitative real time RT-PCR analysis revealed that accumulation of SlCA1 mRNA transcripts was detected in all examined tomato fruit tissues or organs, whereas SlCA2 gene transcripts were found in abundance in leaves. Stems also had SlCA2 transcripts, with transcript levels being higher in flowers than in stems. The SlCA3 gene transcripts were found only in the flowers and the roots. The SlPEPC1 and SlPEPC2 gene transcript levels in different fruit tissues of the tomato were also examined. Τhe possible role of CA isoforms in relation to PEPC in tomato fruit is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carbonic anhydrase (CA; EC: 4.2.1.1) is a zinc-containing enzyme that catalyzes the reversible hydration of CO2 to HCO3 − and has been found in the three domains of life: archaea, bacteria and eukarya. So far, five distinct, evolutionary unrelated CA classes, have been reported namely α-, β-, γ-, δ- and ζ-CA, which independently evolved similar catalytic mechanisms [1, 2]. The β-CA can be further subdivided into two main subclasses, the plant type and the cab-type class, named after the β-CA CAB from the archeon Methanobacterium thermoautotropicum [3]. A third sub-class of β-CAs, designated as ε-class was found in several marine cyanobacteria and chemolithoautotrophic bacteria [4, 5]. The δ- and ζ-CA classes are so far restricted to marine diatoms [6].
CA has been found in a broad range of organisms. It participates in photosynthesis and respiration through the carboxylation and decarboxylation reactions. Several other roles have been attributed to this enzyme such as pH regulation [7], ion transport, and water and electrolyte balance [8].
In plants, all known CAs belong to the α, β or γ classes with the β class being the most predominant. Ιn the chloroplast stromal compartment, HCO3 – is reported to be present at almost 100 times the concentration of CO2 [9]. Hence, in photosynthetic tissues, plastidial β-CA has the major role of converting bicarbonate into CO2 which is the substrate for Rubisco [10]. On the other hand, cytosolic β-CA could provide bicarbonate for phosphoenolpyruvate carboxylase (PEPC) either for C4 photosynthesis [11, 12] or for anaplerotic CO2 fixation in C3 plants. CAs in anaplerotic assimilation catalysed by phosphoenolpyruvate carboxylase have also been proposed [13]. Other biochemical roles, in relation to β-CA, are lipogenesis, glyconeogenesis and pyrimidine biosynthesis [14].
cDNAs encoding β-CA isoforms have been isolated and characterized so far from a number of species including Pisum sativum, [15, 16], soybean [17], alfalfa [18], Lotus japonicus [19], tobacco [20], Arabidopsis [21–23], cotton [24], Flaveria bidentis [25], Hordeum vulgare [26], Oryza sativa [27], Zea mays [28]. CA activity has also been reported in Vicia faba and Lupinus angustifolius [29]. Furthermore, the expression of several β-CA isoforms has been reported in Arabidopsis [23], F. bidentis [30] and F. pringlei [31]. Biochemical evidence has also suggested the multiplicity of β-CA isoforms in pea chloroplast [32]. Subsequent analysis of Arabidopsis thaliana β-CA1 indicated that in addition to its localization to the chloroplast it is also found close to the plasma membrane [33].
During development, tomato fruit passes through distinct stages which range from immature green to ripe red. The tomato fruit epidermis has no stomata [34] and bears a relatively thick cuticle. Carbon dioxide produced by fruit respiration could be re-fixed inside the fruit. At the mature green stage, the CO2 concentration inside the fruit was measured at about 0.29 % (i.e. a ninefold increase compared to the external ambient CO2 concentration) [35]. If any CO2 re-fixation in tomato fruits occurs, carbonic anhydrase could be the key enzyme in this process.
In this study, the presence of CA and PEPC activity and the distribution of CA and PEPC proteins were examined in tomato fruit. Furthermore, the isolation and characterization of three cDNAs encoding CA isoforms in tomato is reported. The SlCA1 and SlCA3 genes represent two full-length cDNAs encoding cytosolic β-CAs. The SlCA2 gene corresponds to a full-length cDNA encoding a plastidic β-CA. The expression pattern of the three β-CA genes was investigated, in parallel with that of the already characterized PEPC gene expression pattern, (SlPEPC1; SlPEPC2) in the fruit and other tissues of tomato plant [36].
Materials and methods
Plants of S. lycopersicum (var. Dombito) were grown in the Agricultural University of Athens greenhouse according to standard agricultural practices. Samples were taken from immature green fruit (10 mm) and mature green fruit (55 mm) which was separated into locular parenchyma with seeds, pericarp and placenta. Samples were also taken from leaves, petioles, flowers (mixed preanthesis and anthesis tissues) and roots. Tissues were cut, frozen immediately with liquid nitrogen and stored at −80 °C.
Measurement of CA and PEPC activity in tomato tissues
CA activity was assayed by measuring changes in pH during the reaction using a dye indicator method [37]. Tomato tissues were ground in liquid nitrogen and homogenized in 2.5 volumes of extraction buffer containing 50 mM MES pH 6.3, 1 mM EDTA, 0.1 mM phenylmethane-sulfonylfluoride, and 0.1 % (w/v) polyvinylpolypyrrolidone (PVPP). Supernatants were dialyzed overnight at 4 °C against 50 mM MES pH 6.3, 1 mM ZnCl2. Assays were performed at 4 °C. The buffer/indicator reaction solution used was 50 mM MES containing 0.005 % (w/v) bromothymol blue (pH 6.3, A615). The reaction buffer contained 50 mM sodium sulfate to maintain a relatively constant ionic strength in the reaction medium. The reaction was initiated by the addition of NaHCO3 at final concentrations of 25–100 mM. Measurements were taken at 615 nm using a Hitachi U-2800 Spectrophotometer (Tokyo, Japan). Uncatalyzed rates, measured in the absence of the enzyme, were subtracted from the catalyzed rates. The catalyzed rate was converted to equivalent μmol H+ consumed, after a calibration established by titrating the reaction buffer with NaOH. For the PEPC activity assay, 2 g of powdered tomato tissues were extracted with 5 ml extraction medium according to Guillet et al. [36]. The protein concentration was determined according to Bradford [38].
Protein extraction, SDS-PAGE and immunoblot analysis
Approximately 0.5 g of fruit tissues were ground in a mortar containing liquid nitrogen and the powder was mixed with 800 μl of ice-cold 50 mM TRIS–HCl (pH 6.8), 5 % (w/v) SDS, 10 % (v/v) glycerol, 5 % (v/v) 2-mercaptoethanol, and 0.002 % (w/v) bromophenol blue. The extract was centrifuged at 12,000×g for 5 min and the supernatant was used for SDS-PAGE. Approximately 20 μg of protein was loaded onto each track of the gel for detecting CA and PEPC proteins. These were analyzed in 15 % SDS-PAGE for CA and in 10 % for PEPC and were then blotted onto nitrocellulose filter (Nitrocell, Hoefer). Polyclonal antibodies were raised in rabbits against the overexpressed recombinant GmCA and GmPEPC proteins [17]. The blots were probed with the polyclonal antibodies and bound antibodies were visualized with the ProtoBlot Western Blot AP System (Promega).
Immunohistolocalization
Polyclonal antibodies raised against soybean β-CA and PEPC polypeptides [17] were used for immunolocalization of tomato CA and PEPC. Fruit tissues at the mature green stage were fixed in 4 % (v/v) paraformaldehyde and 0.25 % (v/v) glutaraldehyde in 10 mM phosphate buffer (pH 7) at 4 °C, overnight. Sections of 8-10 μm, were blocked for 1 h in TBS (10 mM Tris–HCl, 100 mM NaCl, pH 8) containing 3 % (w/v) BSA, and 0.05 % (w/v) Tween-20 at room temperature and incubated overnight with an antiserum raised against either the soybean β-CA or PEPC polypeptides. Pre-immune serum and immune serum were used at a dilution of 1:1,500. Alkaline phosphatase-conjugated anti-rabbit IgGs (Promega) [diluted to 1:5,000 in TBS containing 1 % (w/v) BSA, and 0.05 % (v/v) Tween-20] were used as secondary antibodies. Signal was detected using the alkaline phosphatase substrate, BCIP/NBT (Promega). No signal was observed when pre-immune serum was used as a negative control.
Identification and characterization of cDNAs encoding tomato β-CA gene family
Searches within public tomato EST databases revealed the presence of a number of ESTs in S. lycopersicum coding for β-CAs. The corresponding ESTs were aligned and the complete nucleotide sequence of the larger cDNA clone was determined. The deduced amino acid sequences revealed the presence of SlCA cDNAs representing three full-length open reading frames (GenBank accession nos. CAH60890, CAH60891 and CAQ30514 corresponding to SlCA1, SlCA2 and SlCA3 polypeptides, respectively).
Determination of transcripts levels using real-time qRT-PCR
Organs from S. lycopersicum plants, including locular parenchyma, pericarp and placenta, leaves, young stems, mixed flowers (anthesis and preanthesis), roots and small fruit (10 mm in diameter), were harvested and ground in liquid nitrogen. Total RNA was extracted according the method described by Brusslan and Tobin [39]. RNA samples were then treated with DNase I (Promega, Madison, WI) at 37 °C for 45 min to eliminate contaminating genomic DNA.
First-strand cDNA was reverse transcribed from 2 μg of DNase-treated total RNA. All DNase-treated total RNA samples were denatured at 65 °C for 5 min followed by quick chill on ice in a 12 μl reaction mixture containing 500 ng oligo(dT)12–18 mer, and 0.833 mM dNTPs. After the addition of 4 μl of 5× First-Strand buffer (Invitrogen, Paisley, UK), 1 μl (40 U) RNaseOUT (Invitrogen) ribonuclease inhibitor and 0.016 M DTT at final concentrations, the reaction mixture was preheated to 42 °C for 2 min before the addition of 1 μl (200 U) of SuperScript II reverse transcriptase (Invitrogen). The reaction mixture was incubated at 42 °C for 50 min, followed by heat inactivation at 70 °C for 15 min. The resulting first-strand cDNA was diluted to a final volume of 200 μl, and target cDNAs were amplified using gene-specific primers (Table 1) designed from the transcribed region of each gene using Primer Express 1.5 software (Applied Biosystems, Darmstadt, DE). Quantitative RT-PCRs were performed on the ABI PRISM 7900HT Sequence Detection System using SYBR Green master mix (Applied Biosystems), gene specific primers at a final concentration of 0.2 μM each and 1 μl of the cDNA as template. PCR cycling started with the initial polymerase activation at 95 °C for 15 s and 60 °C for 1 min [40]. The primer specificity and the formation of primer-s were monitored by dissociation curve analysis and agarose gel electrophoresis on a 4 % (w/v) gel. The expression levels of a S. lycopersicum actin gene, detected using SlACTIN-F 5′-CCACAACCACCTCAGCAGAAT-3′ and SlACTIN-R 5′-GACAACTGCTGTGGAGGCATT-3′ primers were used as internal standards to normalize small differences in cDNA template concentrations. Real-time RT-PCR was performed using an MxPro MX3005P PCR system and analyzed with MxPro v3.0 software (Stratagene). For the relative quantification of gene expression, a modification of the comparative threshold cycle method was used. In the case of different tomato tissues, relative transcript levels of the gene of interest (X) were calculated as a ratio to the expressed actin gene transcripts (U), as (1 + E)−ΔCt, where ΔCt was calculated as (C xt − C ut ) [41]. PCR efficiency (E) for each amplicon was calculated using the linear regression method, calculated with the LinRegPCR software [42]. All real-time qPCRs were performed on three biological repeats and all measurements were taken in duplicate.
Results
CA and PEPC activity in tomato fruit
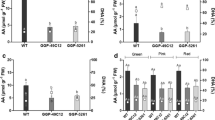
The CA activity in tomato fruit tissues (pericarp, locular parenchyma and placenta) was determined spectrophotometrically using a modified version of the pH-indicator method [37]. CA activity was found to be predominant in the locular parenchyma measuring at about 0.69 mmol H+ consumed per min and per mg of protein and at similar levels in the pericarp (0.57 mmol H+ min−1 mg−1 protein) (Fig. 1a). In contrast, CA activity reached at 0.14 mmol H+ min−1 mg−1 protein in the placenta which is four times lower than the locular parenchyma and pericarp, respectively. PEPC activity was found to be 0.58 μΜ NADH/min mg protein in the pericarp. This value was twice as high as the activity in the locular parenchyma (0.26 μM NADH min−1 mg−1 protein) (Fig. 1b). In the placenta, however, an average activity of 0.38 μM NADH/min mg protein was observed.
Carbonic anhydrase activity (a) in locular parenchyma, pericarp and placenta. The activity was determined spectrophotometrically using a pH-indicator method at pH 6.3. The activity is expressed in mmole of H+ consumed per min and per mg of protein. The PEPC activity (b) was assayed according to [36]. Bars show mean activity +SD (n = 3)
Immunohistological localization and tissue distribution of CA and PEPC proteins in tomato mature green fruit
To gain further insight into the localization of CA and PEPC proteins in the fruit tissues an immunohistolocalization approach was employed. Paraffin-embedded sections of mature green fruit including pericarp, locular parenchyma with seeds and placenta were incubated with polyclonal antibodies raised against soybean CA or PEPC recombinant proteins. The specificity of the soybean antibodies was tested against S. lycopersicum mature green fruit proteins. In each western blot polyclonal antibodies raised against soybean PEPC or β-CA recombinant proteins detected one band with a molecular mass of 110 kDa and one with a molecular mass of 24 kDa (Suppl. Fig. 1).
As shown in Fig. 2, β-CA and PEPC proteins were present in all fruit tissues examined with the signal (blue precipitate) being more intense in the seeds. In the developing seed, the accumulation of both proteins is observed in all seed parts, endosperm, integuments and embryo proper (Fig. 2c, g). Moreover, high signal was also observed in vascular bundles (Fig. 2h). Pre-immune serum was used as a negative control (Fig. 2d, insert). In this case no signal above background was detected.
Immunohistolocalization of CA (a–d) and PEPC (e–h) in mature green fruit tissues. Sections (8–10 μm) of S. lycopersicum probed with the antiserum raised against either the GmCA1 or the GmPEPC proteins. The proteins were visualized as a dark blue staining. a, b, e, f Fruit sections containing pericarp (Pe) locular parenchyma (Lp), placenta (Pl) and developing seed with integuments (In) cotyledons (Co) and embryonic root (Ro). c, g Detail of developing seed. Integuments (In) and endosperm (En). d, h Vascular bundle (Vb) from pericarp. Insert control treated with preimmune serum
Identification and characterization of cDNAs encoding tomato β-CA gene family
cDNA clones coding for tomato CA were isolated and characterized in our laboratory. The corresponding ESTs were aligned and the complete nucleotide sequence of the larger cDNA clone was determined. The deduced amino acid sequences revealed the presence of SlCA cDNAs representing three full-length open reading frames One codes for a polypeptide designated as SlCA1, which contains an open reading frame (ORF) of 268 amino acids. The second designated as SlCA2, contains an ORF of 321 amino acids. The third designated SlCA3 contains an ORF of 255 aminoacids. Topology prediction indicated localization in the cytoplasm for both SlCA1 and SlCA3 and localization in the chloroplast for SlCA2. The putative chloroplast targeting peptide of the SlCA2 exhibited high homology to chloroplast targeting peptides present in chloroplastic β-CAs from different plant species.
Comparative analysis of the deduced SlCA1, SlCA2 and SlCA3 amino acid sequences with previously characterized CA polypeptides (Suppl. Fig. 2) revealed that the three isoforms exhibit high similarity with other β-CAs. The phylogenetic relationship of SlCA1, SlCA2 and SlCA3 to other β-CAs was investigated by the construction of a dendrogram based on the respective amino acid sequences (Fig. 3).
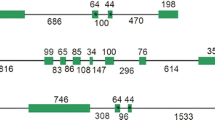
Phylogenetic relationship of SlCA1, SlCA2 and SlCA3 aminoacid sequences (underlined) to other β-type CAs. The dendrogram is a graphical representation of a multiple sequence alignment constructed with the Clustal method using the PAM 250 residue weight table. N. tabacum (P27141); A. thaliana (CA1 AT3G01500, CA2 AT5G14740, CA3 AT1G23730, CA4 AT1G70410, CA5 AT4G33580, CA6 AT1G58180); F. bidentis (CA1 AAA86939, CA2 AAO17573, CA3 AY167113); F. pringlei (CA1 AAA86992, CA2 ABC41657, CA3 ABC41658), Z. mays (CA1 B4G0V7, CA2 B6T9W3, CA3 Q41729)
The updated genome sequencing of tomato revealed that SlCA1 gene is located on chromosome 5 and its position is 5 cM while SlCA2 gene is located on chromosome 2 and its position is 93.00 cM in the Tomato-Expen 2000 Map. On the other hand, SlCA3 gene is located on chromosome 3 at 10.37 cM as annotated to Arabidopsis COSII Map (http://solgenomics.net/).
Expression of S. lycopersicum SlCA1, SlCA2, SlCA3 and SlPEPC1, SlPEPC2 in various tissues
Having established the presence of CA and PEPC proteins in the different fruit tissues of tomato, we then turned our attention towards assessing the CA and PEPC transcript levels in these tissues by performing real time quantitative PCR. Total RNA was isolated from immature green fruit (10 mm), locular parenchyma with seeds, pericarp and placenta of mature green fruit as well as from the leaves, flowers, stems and roots. Figure 4a shows the relative abundance of SlCA1, SlCA2 and SlCA3 transcripts in the various tissues and organs. Accumulation of SlCA1 gene transcript was detected in all the fruit tissues (especially in locular parenchyma), flowers and stems. On the contrary, SlCA3 and SlCA2 gene transcripts were not detected in any fruit tissues (or they were detected at very low levels). SlCA2 gene transcript was present in the leaf, stem and flower while being most abundant in the leaf tissue. SlCA3 transcript was present at significant levels in the flowers and roots. Interestingly, SlPEPC1 and SlPEPC2 gene transcripts were detected in all tissues and organs (Fig. 4b). The SlPEPC1 mRNA transcripts were 10 times higher in the fruit (10 mm) than in the leaf. SlPEPC2 mRNA transcripts were highly abundant in the fruit tissues and particularly in the locular parenchyma with seeds, the pericarp, the placenta and the fruit (10 mm) (Fig. 4b). Finally, SlPEPC2 mRNA transcript abundance reached up to 80 times that of SlPEPC1 mRNAs in the placenta (Fig. 4b).
Accumulation of SlCA1, SlCA2, SlCA3 (a) and SlPEPC1, SlPEPC2 (b) transcripts in various tissues of tomato plant. Total RNA was isolated from various organs was subjected to reverse-transcription and followed real-time quantitative polymerase chain reaction analysis. Transcript levels in the different samples were normalized to those of the constitutive gene, actin. Relative mRNA levels were calculated with respect to the levels of actin transcripts. Bars show mean +SD (n = 3)
Discussion
As tomato fruit has a relatively thick cuticle and no stomata, gas exchange should be extremely low. Carbon dioxide concentration inside mature green fruit was measured to be, on average, ninefold higher than that of the ambient air [35]. However CO2 concentration in specific tissues, like the seed tissues, should be even higher, given the high rates of respiration in developing embryos [43]. The high concentration of CO2 inside the fruit raises the question of its fate. It could be transported out of fruit, reused in the fruit [44] or both. A mechanism that could participate in CO2 re-fixation would involve the activity of a CA enzyme. A possible role for cytoplasmic β-CAs could be the conversion of CO2 to bicarbonate, which serves as a substrate for PEPC. The produced oxaloacetate through the action of PEPC could be used for amino acid synthesis [45] or converted to malate and/or citrate which are known to accumulate in tomato fruits during its development [36].
Our studies revealed CA and PEPC enzyme activity in all the fruit tissues examined. Furthermore, the immunohistolocalization studies indicated that both the β-CA and PEPC proteins were co-localized in all the fruit tissues examined with the signal being more intense in the vascular bundles and developing seeds (Fig. 2). Similar results have been observed in Medicago seeds [46]. High local CO2 concentration is expected to occur in the vascular bundles due to the high metabolic activity [47, 48]. The seed coat surrounding the developing embryo is a major barrier for gas diffusion [49]. Thus, the presence of high levels of enzymes involved in CO2 fixation in the seeds and the vascular bundles may contribute to the efficient removal of CO2 and its conversion to oxaloacetate, a precursor for amino acid synthesis.
The observed high levels of SlCA1 transcript accumulation in the tomato pericarp are in close agreement with the reported accumulation of the protein encoded by the SlCA1 gene in cherry tomato pericarp [50]. These data taken together suggested that SlCA1 may provide the bicarbonate as a substrate for SlPEPC2, which is highly expressed, thus increasing the malic acid content, reaching up to 35 μmol g−1 FW [51]. The detection of SlCA3 transcripts exclusively in the roots and flowers, suggests that this isoform may serve a special function in these tissues. Recent observations reported the presence of CA activity in tomato roots [52]. The identification of root CA isoforms is not surprising given that recent observations also reported the expression of three CA isoforms in roots of F. bidentis [31] and Arabidopsis [24]. However, these isoforms are also found in other tissues besides the roots. The accumulation of SlPEPC1 and SlPEPC2 along with SlCA1 and SlCA3 transcripts in tomato roots suggest that some of the respired CO2 may be re-fixed through a mechanism similar to that proposed in tomato fruits.
In the plastids, CA could participate in CO2 re-fixation through photosynthesis or lipogenesis [14]. Plastidial CA has been reported to facilitate the diffusion of CO2 across the chloroplast envelope [10, 53]. In this respect, the very low or undetectable levels of SlCA2 transcript in the fruit tissues, is surprising. However, the possibility of a second, not yet identified, plastidial isoform present in the fruit tissues can not be excluded. The expression of two plastidial isoforms has been detected in the leaves, siliques, stems and flowers of Arabidopsis [23].
The presence of two PEPC isoforms expressed in tomato fruit has also been reported [36]. PEPC also appears to play a role in the extension of cotton fibres and it is proposed that PEPC activity allows malate production and therefore increased turgor that is required for fibre elongation [54]. The relative levels of SlPEPC1 transcript was similar in the different fruit tissues examined, whereas the relative levels of SlPEPC2 transcripts were very high compared to those found for SlPEPC1 (Fig. 4b). It has been suggested that the SlPEPC1 isoform could play a housekeeping role [36] in tomato fruit, and SlPEPC2 could serve in the expansion of the fruit. The high PEPC activity ratio (20–40 fold) between the fruit and leaf [51] is in accordance with our qRT-PCR data (Fig. 4b) rendering PEPC a key enzyme in the CO2 metabolism of the tomato fruit.
In conclusion, we have provided data which point to the combined action of CA and PEPC in re-fixing the respired CO2 thus making a net contribution to fruit carbon economy.
Abbreviations
- ACCase:
-
Acetyl-CoA carboxylase
- BSA:
-
Bovine serum albumin
- BCIP/NTB:
-
5-Bromo-3-chloro-3-indolylphosphate nitroblue tetrazolium
- CA:
-
Carbonic anhydrase
- PEPC:
-
Phosphoenolpyruvate carboxylase
References
Tripp B, Smith K, Ferry JG (2001) Carbonic anhydrase: new insights for an ancient enzyme. J Biol Chem 276:48615–48618
Lane TW, Morel FM (2000) A biological function for cadmium in marine diatoms. Proc Natl Acad Sci USA 97:4627–4631
Smith KS, Ingram-Smith C, Ferry JG (2002) Roles of aspartate and arginine in the catalytic mechanisms of an archaeal beta-class carbonic anhydrase. J Bacteriol 184:4240–4245
So AK-C, Aspie GS, Williams EB, Sively JM, Heinhorst S, Cannon GC (2004) A novel evolutionary lineage of carbonic anhydrase (ε class) is a component of the carboxysome shell. J Bacteriol 186:623–630
Sawaya MR, Cannon GC, Heinhorst S, Tanaka S, Williams EB, Yeates TO, Kerfeld CA (2006) The structure of beta-carbonic anhydrase from the carboxysomal shell reveals a distinct subclass with one active site for the price of two. J Biol Chem 281(11):7546–7555
Park H, Song B, Morel FM (2007) Diversity of cadmium-containing carbonic anhydrase in marine diatoms and natural waters. Environ Microbiol 9:403–413
Tsuruoka S, Swenson ER, Petrovic S, Fujimura A, Schwartz GJ (2001) Role of basolateral carbonic anhydrase in proximal tubular fluid and bicarbonate absorption. Am J Physiol Renal Physiol 280:F146–F154
Dagher PC, Egnor RW, Charney AN (1993) Effect of intracellular acidification of colonic NaCl absorption. Am J Physiol 264:G569–G575
Moroney JV, Bartlett SG, Samuelsson G (2001) Carbonic anhydrases in plants and algae. Plant Cell Environ 24:141–153
Badger MR, Price GD (1994) The role of carbonic anhydrase in photosynthesis. Ann Rev Plant Physiol Plant Mol Biol 45:369–392
Hatch MD, Burnell JN (1990) Carbonic anhydrase activity in leaves and its role in the first step of C4 photosynthesis. Plant Physiol 93:825–828
Von Caemmerer S, Quinn V, Hancock N, Price GD, Furbank RT, Ludwig M (2004) Carbonic anhydrase and C4 photosynthesis: a transgenic analysis. Plant Cell Environ 27:697–703
Giordano M, Norici A, Forssen M, Eriksson M, Raven JA (2003) An anaplerotic role for mitochondrial carbonic anhydrase in Chlamydomonas reinhardtii. Plant Physiol 132:2126–2134
Ohlrogge JB, Jaworski JG (1997) Regulation of fatty acid synthesis. Ann Rev Plant Physiol Plant Mol Biol 48:109–136
Roeske CA, Ogren WL (1990) Nucleotide sequence of pea cDNA encoding chloroplast carbonic anhydrase. Nucleic Acids Res 18:3413
Majeau N, Coleman JR (1991) Isolation and characterization of a cDNA coding for pea chloroplastic carbonic anhydrase. Plant Physiol 95:264–268
Kavroulakis N, Flemetakis E, Aivalakis G, Katinakis P (2000) Carbon metabolism in developing soybean root nodules: the role of carbonic anhydrase. Mol Plant Microbe Interact 13:14–22
Coba de la Pena TC, Frugier F, McKhann HI, Bauer P, Brown S, Kondorosi A, Crespi M (1997) A carbonic anhydrase gene is induced in the nodule primordium and its cell-specific expression is controlled by the presence of Rhizobium during development. Plant J 11:407–420
Flemetakis E, Dimou M, Cotzur D, Aivalakis G, Efrose R, Kenoutis C, Udvardi M, Katinakis P (2003) A Lotus japonicus β-type carbonic anhydrase gene expression pattern suggests distinct physiological roles during nodule development. Biochem Biophys Acta 1628:186–194
Majeau N, Coleman JR (1992) Nucleotide sequence of a complementary DNA encoding tobacco chloroplastic carbonic anhydrase. Plant Physiol 100:1077–1078
Raines CA, Horsnell PR, Holder C, Lloyd JC (1992) Arabidopsis thaliana carbonic anhydrase: cDNA sequence and effect of CO2 on mRNA levels. Plant Mol Biol 20:1143–1148
Fett JP, Coleman JR (1994) Characterization and expression of 2 cDNAs encoding carbonic anhydrase in Arabidopsis thaliana. Plant Physiol 105:703–713
Fabre N, Reiter IM, Becuwe-Linka N, Genty B, Rumeau D (2007) Characterization and expression analysis of genes encoding α and β carbonic anhydrases in Arabidopsis. Plant Cell Eniron 30:617–629
Hoang CV, Chapman KD (2002) Regulation of carbonic anhydrase gene expression in cotyledons of cotton (Gossypium hirsutum L.) seedlings during post-germinative growth. Plant Mol Biol 49:449–458
Cavallaro A, Ludwig M, Burnell J (1994) The nucleotide sequence of a complementary DNA encoding Flaveria bidentis carbonic anhydrase. FEBS Lett 350:216–218
Bracey MH, Bartlett SG (1995) Sequence of a cDNA encoding carbonic anhydrase from barley. Plant Physiol 108:433–434
Suzuki S, Burnell JN (1995) Nucleotide sequence of a cDNA encoding rice chloroplastic carbonic anhydrase. Plant Physiol 107:299–300
Alexandrov NN, Brover VV, Freidin S, Troukhan ME, Tatarinova TV, Zhang H, Swaller TJ, Lu YP, Bouck J, Flavell RB, Feldmann KA (2009) Insights into corn genes derived from large-scale cDNA sequencing. Plant Mol Biol 69:179–194
Atkins C, Smith P, Mann A, Thumfort P (2001) Localization of carbonic anhydrase in legume nodules. Plant Cell Environ 24:317–326
Tetu S, Tanz SK, Vella N, Burnell J, Ludwig M (2007) The Flaveria bidentis β-carbonic anhydrase gene family encodes cytosolic and chloroplastic isoforms demonstrating distinct organ-specific expression patterns. Plant Physiol 144:1316–1327
Tanz SK, Tetu SG, Vella N, Ludwig M (2009) Loss of the transit peptide and an increase in gene expression of an ancestral chloroplastic carbonic anhydrase were instrumental in the evolution of the cytosolic C4 carbonic anhydrase in Flaveria. Plant Physiol 150:1515–1529
Rudenko NN, Ignatova LK, Kamornitskaya VB, Ivanov BN (2006) Pea leaf thylakoids contain several carbonic anhydrases. Dokl Biochem Biophys 408:155–157
Hu H, Boisson-Dernier A, Israelsson-Nordstrom M, Bohmer M, Xue S, Ries A, Godoski J, Kuhn JM, Schroeder JI (2010) Carbonic anhydrases are upstream regulators of CO2-controlled stomatal movements in guard cells. Nat Cell Biol 12:87–93
Fahn A (1982) Plant anatomy. Pergamon Press, Oxford/New York/Toronto/Sydney/Paris/Frankfurt
Czarnowski M, Starzecki W (1992) Spectral properties and CO2 exchange of tomato fruits. Photosynthetica 27:513–519
Guillet C, Just D, Bénard N, Destrac-Irvine A, Baldet P, Hernould M, Causse M, Raymond P, Rothan C (2002) A fruit-specific phosphoenolpyruvate carboxylase is related to rapid growth of tomato fruit. Planta 214:717–726
Khalifah RG (1971) The carbon dioxide hydration activity of carbonic anhydrase. I. Stop-flow kinetic studies on the native human isoenzymes B and C. J Biol Chem 246:2561–2573
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brusslan JA, Tobin EM (1992) Light-independent developmental regulation of cab gene expression in Arabidopsis thaliana seedlings. Proc Natl Acad Sci USA 89:7791–7795
Czechowski T, Bari RP, Stitt M, Scheible WR, Udvardi MK (2004) Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: unprecedented sensitivity reveals novel root- and shoot-specific genes. Plant J 38(2):366–379
Flemetakis E, Efrose RC, Ott T, Stedel C, Aivalakis G, Udvardi MK, Katinakis P (2006) Spatial and temporal organization of sucrose metabolism in Lotus japonicus nitrogen-fixing nodules suggests a role for the elusive alkaline/neutral invertase. Plant Mol Biol 62:53–69
Ramakers C, Ruijter JM, Deprez RH, Moorman AF (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66
Goffman FD, Ruckle M, Ohlrogge J, Shachar-Hill Y (2004) Carbon dioxide concentrations are very high in developing oilseeds. Plant Physiol Biochem 42:703–708
Blanke MM, Lenz F (1989) Fruit photosynthesis—a review. Plant Cell Environ 12:31–46
Andreo CS, Gonzales DH, Iglesias AA (1987) Higher plant phosphoenolpyruvate carboxylase. Structure and regulation. FEBS Lett 213:1–8
Aivalakis G, Dimou M, Flemetakis E, Plati F, Katinakis P, Drossopoulos JB (2004) Immunolocalization of carbonic anhydrase and phosphoenolpyruvate carboxylase in developing seeds of Medicago sativa. Plant Physiol Biochem 42:181–186
Geingenberger P, Langenberger S, Wilke I, Heineke D, Heldt HW, Stitt M (1993) Sucrose is metabolized by sucrose synthase and glycolysis within the phloem complex of Ricinus communis L. seedlings. Planta 190:446–453
Van Bel AJE, Knoblauch M (2000) Sieve element and companion cell: the story of comatose patient and the hyperactive nurse. Aust J Plant Physiol 27:477–487
King SP, Badger MR, Furbank RT (1998) CO2 refixation characteristics of developing canola seeds and silique wall. Aust J Plant Physiol 25:377–386
Fourobert M, Mihr C, Bebtin N, Pawlowski T, Negroni N, Sommerer N, Causse M (2007) Major proteome variation associated with cherry tomato pericarp development and ripening. Plant Physiol 143:1327–1346
Laval Martin D, Farineau J, Diamond J (1977) Light versus dark carbon metabolism in cherry tomato fruits I. Occurrence of photosynthesis. Study of the intermediates. Plant Physiol 60:872–876
Viktor Α, Cramer MD (2005) The influence of root assimilated inorganic carbon on nitrogen acquisition/assimilation and carbon partitioning. New Phytol 165:157–169
Graham D, Reed ML, Patterson BD, Hockley DG, Dwyer MR (1984) Chemical properties, distribution, and physiology of plant and algal carbonic anhydrases. Ann NY Acad Sci 429:222–237
Li XR, Wang L, Ruan YL (2010) Developmental and molecular physiological evidence for the role of phosphoenolpyruvate carboxylase in rapid cotton fibre elongation. J Exp Bot 61:287–295
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2013_2498_MOESM1_ESM.tif
Immunodetection of CA (a) and PEPC (b). Total proteins were extracted from tomato mature green fruit. Blots were probed with antiserum raised against either GmCA1 or GmPEPC proteins. Supplementary material 1 (TIFF 30,367 kb)
11033_2013_2498_MOESM2_ESM.tif
Comparison of the deduced amino acid sequences of S. lycopersicum CA (SlCA1, SlCA2 and SlCA3) with various plant CAs. Sequences shown include: N. tabacum (P27141); A.thaliana (CA1: AT3G01500, CA2: AT5G14740, CA3: AT1G23730, CA4: AT1G70410, CA5: AT4G33580, CA6: AT1G58180); F. bidentis (CA1: AAA86939, CA2: AAO17573, CA3: AY167113); F. pringlei (CA1: AAA86992, CA2: ABC41657, CA3: ABC41658), Z. mays (CA1: B4G0V7, CA2: B6T9W3, CA3: Q41729). Black-shaded boxes represent conserved amino acids, while dashes represent gaps in the alignment. Sequences were aligned using the Clustal method with PAM250 residue weight table. Supplementary material 2 (TIFF 4,943 kb)
Rights and permissions
About this article
Cite this article
Diamantopoulos, P.D., Aivalakis, G., Flemetakis, E. et al. Expression of three β-type carbonic anhydrases in tomato fruits. Mol Biol Rep 40, 4189–4196 (2013). https://doi.org/10.1007/s11033-013-2498-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-013-2498-2