Abstract
Pathologic complete response after neoadjuvant systemic treatment appears to be a valid surrogate for better overall survival in breast cancer patients. Currently, together with standard clinicopathologic assessment, novel molecular biomarkers are being exhaustively tested in order to look into the heterogeneity of breast cancer. The aim of our study was to examine an association between 23-gene real-time-PCR expression assay including ABCB1, ABCC1, BAX, BBC3, BCL2, CASP3, CYP2D6, ERCC1, FOXC1, GAPDH, IGF1R, IRF1, MAP2, MAPK 8, MAPK9, MKI67, MMP9, NCOA3, PARP1, PIK3CA, TGFB3, TOP2A, and YWHAZ receptor status of breast cancer core biopsies sampled before neoadjuvant chemotherapy (anthracycline and taxanes) and pathologic response. Core-needle biopsies were collected from 42 female patients with inoperable locally advanced breast cancer or resectable tumors suitable for downstaging, before any treatment. Expressions of 23 genes were determined by means of TagMan low density arrays. Analysis of variance was used to select genes with discriminatory potential between receptor subtypes. We introduced a correction for false discovery rates (presented as q values) due to multiple hypothesis testing. Statistical analysis showed that seven genes out of a 23-gene real-time-PCR expression assay differed significantly in relation to pathologic response regardless of breast cancer subtypes. Among these genes, we identified: BAX (p = 0.0146), CYP2D6 (p = 0.0063), ERCC1 (p = 0.0231), FOXC1 (p = 0.0048), IRF1 (p = 0.0022), MAP2 (p = 0.0011), and MKI67 (p = 0.0332). The assessment of core biopsy gene profiles and receptor-based subtypes, before neoadjuvant therapy seems to predict response or resistance and to define new signaling pathways to provide more powerful classifiers in breast cancer, hence the need for further research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathologic complete response (pCR) after neoadjuvant systemic treatment appears to be a valid surrogate for better overall survival in breast cancer patients [1]. A vast amount of data suggests that pCR may be a more sensitive and straightforward way to document treatment effects than disease-free survival (DFS) [1–3]. Currently, together with standard, established clinicopathologic assessment, novel molecular biomarkers are being exhaustively tested in order to look into the heterogeneity of breast cancer [4, 5]. The assessment of core biopsy gene profiles and receptor based subtypes, before neoadjuvant therapy seems to predict response or resistance and to define new signaling pathways to provide more powerful classifiers in breast cancer [6].
Aim
The aim of our study was to examine an association between 23-gene real-time-PCR expression assay including ABCB1, ABCC1, BAX, BBC3, BCL2, CASP3, CYP2D6, ERCC1, FOXC1, GAPDH, IGF1R, IRF1, MAP2, MAPK 8, MAPK9, MKI67, MMP9, NCOA3, PARP1, PIK3CA, TGFB3, TOP2A, and YWHAZ (Table 1), receptor status of breast cancer core biopsies sampled before neoadjuvant systemic treatment and pathologic response in the subsequent mastectomy or breast conservation specimens.
Patients and methods
The study was conducted under Institutional Review Board protocol # RNN/159/10/KE/07/09/2010, Medical University of Lodz and all patients gave written informed consent.
Before any treatment, ultrasound guided 14-gauge core needle biopsies using an ultra automatic biopsy instrument (Pro-Mag™, Angiotech) were collected from 42 female patients with inoperable locally advanced breast cancer or resectable tumors suitable for downstaging, and from two healthy controls at the Cancer Center between September 2010 and April 2011. Four to five specimens per lesion were obtained, half of which were frozen immediately at −80°C, for subsequent RNA extraction, cDNA generation and custom-designed TaqMan® gene expression assay. The other samples were paraffin embedded and reviewed by dedicated breast pathologists in the Department of Pathology. Estrogen receptor (ER) and progesterone receptor (PR) status were determined by immunohistochemistry (IHC) using the Allred score. Human epidermal growth factor receptor 2 (HER2) status was evaluated by immunohistochemistry or by fluorescence in situ hybridization. HER2-positive tumors were defined as 3+ receptor overexpression on IHC staining and/or gene amplification found on fluorescent in situ hybridization. TNM clinical staging was assessed by mammography, ultrasound of the breast, axilla, and abdomen, and chest X-ray. In selected cases, MRI of the breast was performed. The following preoperative chemotherapy regimens were used: AT (doxorubicin 50 mg/m2, docetaxel 75 mg/m2) in 29 patients, AC (doxorubicin 60 mg/m2, cyclophosphamide 600 mg/m2) in 13 patients. Upon completion of chemotherapy (six cycles), dedicated breast surgeons performed mastectomy or breast conservation, with axillary dissection or sentinel node biopsy. Pathologic response in the mastectomy or breast conservation specimens was assessed by dedicated breast pathologists. The pCR was defined as postoperative microscopic absence of invasive or in situ carcinoma in breast tissue, and axillary lymph nodes after neoadjuvant systemic treatment. A near complete response with only minimal residual disease was described as scattered tumor cells in the primary tumor site or lymph node or minimal cellularity or small clusters in the surgical specimen, with >90% loss of tumor cells. Pathologic no response (pNR) was defined as no change or some minor alteration to individual malignant cells, but no reduction in overall cellularity. Partial pathologic response (pPR) reduction in overall cellularity, not exhibiting the changes listed for pCR, near-pCR or pNR. pCR and near-pCR were key points in statistical analysis.
Total RNA extraction and cDNA generation
Total RNA was extracted from samples according to the manufacturer’s RNeasy mini kits protocol (Qiagen, Hilden, Germany). In the initial step, RLT buffer (containing β-mercaptoethanol) was added to Eppendorf tubes containing the frozen samples which were homogenized using a Qiagen homogenizer (TissueRuptor) and centrifuged for 3 min at 14,000 rpm. Following the manufacturer’s protocol, a DNase digestion was performed and RNA was quantified using PicoDrop spectrophotometer (Picodrop, Saffon Walden, Cambridgeshire, UK). The quality of RNA samples was analyzed by measuring the ratio of absorptions at 260/280 nm. The purified total RNA was immediately used for cDNA synthesis or stored at −80°C.
Generation of cDNA was performed with High Capacity cDNA Reverse Transcription Kits (Applied Biosystems Inc., Foster City, CA, USA) following the reverse transcription protocols of the manufacturer. 500 ng of DNase-treated total RNA was used as starting material, to which was added 2× RT master mix containing 2 μl of 10× RT buffer, 0.8 μl of 25× dNTP mix (100 mM), 2 μl of 10× RT random primers, 1 μl MultiScribe™ Reverse Transcriptase and 1 μl RNase inhibitor per each 20 μl reaction. Reverse transcription was performed in conditions optimized for use with this kit (25°C for 10 min, 37°C for 120 min, 85°C for 5 min). The samples were kept frozen at −20°C.
Custom-designed TaqMan® gene expression assays
Gene expression was measured using custom-made TaqMan low density arrays (Applied Biosystems Inc., Foster City, CA). The assay comprised of probes selected to measure expression of 23 a priori selected genes: ABCB1, ABCC1, BAX, BBC3, BCL2, CASP3, CYP2D6, ERCC1, FOXC1, GAPDH, IGF1R, IRF1, MAP2, MAPK 8, MAPK9, MKI67, MMP9, NCOA3, PARP1, PIK3CA, TGFB3, TOP2A, and YWHAZ. Lists of genes that represent various biological pathways were assembled from gene datasets [7, 8].
The microfluidic cards consisted of eight ports with 23 different TaqMan primer pair/probe sets arrayed in duplicate in a 384-well microplate. Each well contained a gene-specific forward and reverse primer, as well as a gene-specific probe, which is labeled at the 5′ position with 6-FAM (reporter dye) and at the 3′ position with minor groove binder/non-fluorescent quencher.
RT reactions were performed after adding 500 ng cDNA mixed with 2× TaqMan Universal PCR Master Mix (Applied Biosystems Inc., Foster City, CA, USA), loaded on the TLDA card, and analyzed by PCR on the 7900HT instrument using Applied Biosystems Sequence Detection System 2.0 software according to the manufacturer’s instructions. Target gene expression data from samples was normalized using 18S RNA to compensate for variability in the amount of RNA and for exclusion of general transcriptional effects.
Molecular analyses were performed in the Department of Molecular Biology, Central Laboratory.
Statistical analysis
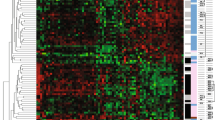
Expression ratios were computed for each gene by dividing the values obtained for each gene in each of the study patients by those from healthy tissue samples. Gene expression data underwent standard filtering procedures. After logarithmic transformation, ratios deviating more than three standard deviations were treated as outliers. Following that stage, data were standardized by dividing their difference from group mean by respective standard deviations. This provided a homogenous group of variables with similar scales and ranges of values. Univariate comparisons of gene profile between receptor subtypes were performed using analysis of variance (ANOVA) with p values verified by false discovery rates (FDR) to correct for multiple hypotheses testing. Genes that showed different expression depending on receptor subtype in ANOVA entered post hoc analyses with Tukey’s HSD test to determine which subgroup deviated from the remainder in expression values. Statistical computations were performed in Statistica 9.0 PL (Statsoft, Tulsa, OK, USA). Hierarchical clustering of gene expression ratios was used to visualize expression patterns in the analyzed groups. The GenePattern online suite (http://genepattern.broadinstitute.org/gp/pages/index.jsf) was used for this purpose. Q values representing FDRs were computed in R using the q values package. A p value less than 0.05 was considered as statistically significant. For comparisons of gene expression profiles, a q value less than 0.05 for ANOVA comparisons was necessary to deem a particular result as significant rather than an incidental one due to multiple hypothesis testing.
Results
Patients enrolled in the study were aged between 32- and 80-years-old, mean age 55.6 years. Histopathological tumor types were: invasive ductal breast cancer (37 patients), invasive lobular cancer (four patients), and adenoid cystic cancer (one patient). Tumor grades were: one, seven and 34 patients with grades Gx, G2, and G3, respectively. Tumor stage was: IIA in three patients, IIB in six, IIIA in 12, IIIB in 17, IIIC in three and IV in one patient (breast cancer with isolated metastasis to the uterus, removed with clear margins). Receptor status was: ER positive in 21 patients, ER negative in 21 patients, PR positive in 17 patients, PR negative in 25 patients, HER2 positive in eight patients, HER2 negative in 34 patients. Surrogates of intrinsic subtypes were: Luminal A (12 patients), Luminal B HER2 negative (five patients), Luminal B HER2 positive (four patients), Triple Negative (17 patients), HER2 positive (four patients).
After neoadjuvant chemotherapy or hormonal therapy, pCR was achieved in 12% of patients (5/42), near-pCR in 19% (8/42), partial pathologic response in 38% (16/42) and no response or progression in 31% of patients (13/42).
Statistical analysis showed that seven genes out of a 23-gene real-time-PCR expression assay differed significantly in relation to pathologic response regardless of breast cancer subtypes. Among these genes, we identified: BAX (p = 0.0146), CYP2D6 (p = 0.0063), ERCC1 (p = 0.0231), FOXC1 (p = 0.0048), IRF1 (p = 0.0022), MAP2 (p = 0.0011), and MKI67 (p = 0.0332). Expression levels of these seven genes were compared between pathologic response groups, and results of these comparisons are shown in Fig. 1a–d.
P values of the remaining genes in the 23-gene realtime-PCR expression assay (ABCB1, ABCC1, BBC3, BCL2 CASP3, IGF1R, MAPK8, MAPK9, MMP9, NCOA3, PARP1, PIK3CA, TGFB3, TOP2A, and YWHAZ) did not reach a level of statistical significance (Table 2).
Discussion and conclusions
Oncologists are in pursuit of novel biomarkers which might supplement the standard, well established receptor subtype-classification in order to improve stratification of breast cancer patients for neo- and adjuvant systemic therapies [9]. Preoperative chemo- or endocrine treatment provides an attractive clinical model to study multiple molecular pathways that determine outcomes in terms of drug complete, partial, or no response. In the current study, we have performed an analysis of tumor gene expression profile using a 23-gene realtime-PCR expression assay in which seven genes such as BAX2, CYP2D6, ERCC1, FOXC1, IRF1, MAP2, and MKI67 differed significantly in discriminating breast cancer samples according to pathologic response. Biological functions of these genes included apoptosis, proliferation, immunity, DNA repair, drug metabolism, and yet-to-be-identified mechanisms which may influence the sensitivity of cancer cells to systemic treatment (Table 1). These pathways are thought to be possible points of therapeutic intervention and have been a focus of intensive drug discovery efforts [10, 11].
It is widely accepted that triple negative tumors are more chemosensitive compared with Luminal A subtypes. This is consistent with our own study in which higher rates of pCR were observed in patients with triple negative receptor status (p = 0.036). Additionally, near-pCR was noted in some Luminal B HER2 positive and Luminal B HER2 negative patients (Table 3). Of the seven genes: BAX (p = 0.0146), CYP2D6 (p = 0.0063), ERCC1 (p = 0.0231), FOXC1 (p = 0.0048), IRF1 (p = 0.0022), MAP2 (p = 0.0011) and MKI67 (p = 0.0332), that showed pathologic response-specific expression profiles in the current study, two of them, FOXC1 and IRF1 were of specific interest. In our previous study FOXC1 and IRF1 mRNA levels were significantly elevated in core biopsies from triple negative breast cancers, compared with Luminal A subtype (unpublished data). Ray and Giuliano et al. have convincingly shown that FOXC1 may be a pivotal prognostic biomarker of basal-like breast cancer, but they did not correlate this gene with pathologic response as we have now done [12, 13].
In the studies conducted by Rody et al. [14], ER negative tumors showing a high expression of immune function metagenes seemed to respond better to neoadjuvant chemotherapy. Cavalli et al. [15] have observed that low IRF1 mRNA expression was associated with poor clinical outcome and correlated with risk of recurrence and death. We have found that high mRNA levels of the immune function gene are associated with pCR which could be a surrogate for good prognosis. Similarly, Teschendorff et al. have demonstrated that immune signatures correlate well with good prognosis in ER negative disease [16].
Our study should be considered to be preliminary in view of its small sample size and a limited observation time. For these reasons, pathologic responses were the key outcomes rather than survival associated endpoints. We did not attempt to predict response to individual drugs, because diverse types of systemic treatment were used (although mainly anthracycline- or taxane-based regimens). On the other hand, the current standard of care for breast cancer patients is combined therapy, so our findings may be more relevant in terms of clinical practice, although our ability to decode drug-specific response pathways was limited. Other studies elsewhere are addressing this problem, such as the correlation of ERCC1, involved in DNA repair, and response to platinum based drug therapy in triple negative breast cancer, or MAP, stabilizing microtubules, and response to paclitaxel [17–20].
To generate integrated prognostic and predictive models that contain both genomic and clinical features is a challenging task. Fluctuations in a single gene expression in biological samples affecting functional change are examples of potential obstacles. Although, opponents of gene signatures claim that we are lost in a sea of genomic data, surely any tool which can better inform oncologists and patients about targeted therapies should be introduced into clinical practice at the earliest opportunity, hence the need for further research [9, 21–26]. Currently, we are collecting more samples and continuing the follow-up of the original group. In addition, we plan to delineate gene signatures predictive for specific pathologic response to endocrine treatment. A prospective validation in a secondary cohort will be made possible in cooperative studies with other departments. On the basis of our current and future research on gene signatures, we would like to enrich clinicopathologic data to more accurately identify a population of patients most likely to benefit from treatment.
References
Von Minckwitz G, Loibl S, Maisch A, Untch M (2011) Lessons from the neoadjuvant setting on how best to choose adjuvant therapies. Breast 20(Suppl 3):S142–S145
Marchio C, Sapino A (2011) The pathologic complete response open question in primary therapy. J Natl Cancer Inst Monogr 43:86–90
Arun BK, Dhinghra K, Valero V, Kau SW, Broglio K et al (2011) Phase III randomized trial of dose intensive neoadjuvant chemotherapy with or without G-CSF in locally advanced breast cancer: long-term results. Oncologist 16:1527–1534
Ena G (2011) Neoadjuvant therapy in breast cancer. Rev Med Brux 32:243–246
Vaklavas C, Forero-Tores A (2011) How do I treat triple negative disease? Curr Treat Options Oncol 12:369–388
Gonzalez-Angulo AM, Hennessy BT, Meric-Bernstam F, Sahin A, Liu W et al (2011) Functional proteomics can define prognosis and predict pathologic complete response in patients with breast cancer. Clin Proteomics 8(1):11
http://www.ncbi.nlm.nih.gov. Accessed 27 June 2010
http://www.genecards.org/cgi-bin/carddisp.pl. Accessed 13 Nov 2011
Sotiriou C, Pusztai L (2009) Gene-expression signatures in breast cancer. N Engl J Med 360(8):790–800
Hoeflich K (2011) PI3 K + MEK inhibition. Ann Oncol 22(suppl 2):ii23. doi:10.1093/annonc/mdr077
Fumagalli D, Desmedt C, Ignatiadis M, Loi S, Piccart M, Sotiriou C (2011) Gene profiling assay and application: the predictive role in primary therapy. J Natl Cancer Inst Monogr 43:124–127
Ray PS, Wang J, Qu Y, Sim MS, Shamonki JM et al (2010) FOXC1 is a potential prognostic biomarker with functional significance in basal- like breast cancer. Cancer Res 70(10):3870–3876. doi:10.1158/0008-5472
Ray PS, Bagaria SP, Wang J, Shamonki JM, Ye X et al (2011) Basal- like breast cancer defined by FOXC1 expression offers superior prognostic value: a retrospective immunohistochemical study. Ann Surg Oncol 18:3839–3847
Rody A, Holtrich U, Pusztai L, Liedtke C, Gaetje R et al (2009) T-cell metagene predicts a favorable prognosis in estrogen receptor- negative and HER2 positive breast cancers. Breast Cancer Res 11:R15
Cavalli LR, Riqqins RB, Wang A, Clarke R, Haddad BR (2010) Frequent loss of heterozygosity at the interferon regulatory factor -1 gene locus in breast cancer. Breast Cancer Res Treat 121:227–231
Teschendorff AE, Gomez S, Arenas A, El-Ashry D, Schmidt M et al (2010) Improved prognostic classification of breast cancer defined by antagonistic activation patterns of immune response pathway modules. BMC 10:604
Kim D, Jung W, Koo JS (2011) The expression of ERCC1, RRM1, and BRCA1 in breast cancer according to the immunohistochemical phenotypes. J Korean Med Sci 26:352–359
Chen X, Wu J, Lu H, Huang O, Shen K (2011) Measuring β-tubulin III, Bcl-2 and ERCC1 improves pathological complete remission predictive accuracy in breast cancer. Cancer Sci 103(2):262–268. doi:10.1111/j.1349-7006.2011.02135.x
Shao YY, Kuo KT, Hu FC, Lu JS, Huang CS et al (2010) Predictive and prognostic values of tau and ERCC1 in advanced breast cancer patients treated with paclitaxel and cisplatin. Jpn J Clin Oncol 40:286–293
Silver DP, Richardson AL, Eklund AC, Wang ZC, Szallasi Z et al (2010) Efficacy of neoadjuvant cis-platin in triple negative breast cancer. J Clin Oncol 28:1145–1153
Iwamoto T, Pusztai L (2010) Predicting prognosis of breast cancer with gene signatures: are we lost in a sea of data? Genome Med 2:81
Tordai A, Wang J, Andre F, Liedtke C, Yan K et al (2008) Evaluation of biological pathways involved in chemotherapy response in breast cancer. Breast Cancer Res 10(2):R37. doi:10.1186/bcr2088
Fan C, Prat A, Parker JS, Liu Y, Carey A et al (2011) Building prognostic models for breast cancer patients using clinical variables and hundreds of gene expression signatures. BMC Medical Genomics 4:3
Nielsen TO, Parker JS, Leung S, Voduc D, Ebbert M et al (2011) A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer. Clin Cancer Res 16:5222–5232
Froehlich H (2011) Network based consensus gene signatures for biomarker discovery in breast cancer. Plos one 6(10):e25364. doi:10.1371/journal.pone.0025364
Gabrovska PN, Smith RA, Tiang T, Weinstein SR, Haupt LM, Griffiths LR (2011) Development of an eight gene expression profile implicating human breast tumours of all grades. Mol Biol Rep (in press)
Acknowledgment
This work was supported by the grant of the Ministry of Science, Poland, NN402350838.
Conflict of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kolacinska, A., Fendler, W., Szemraj, J. et al. Gene expression and pathologic response to neoadjuvant chemotherapy in breast cancer. Mol Biol Rep 39, 7435–7441 (2012). https://doi.org/10.1007/s11033-012-1576-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1576-1