Abstract
α-amy gene amplified from barley genome was cloned into MCS of pGAP9K to generate pGAP9K-α-amy which was then transformed into Pichia pastoris GS115 by electroporation. Transformants with multi-copies and high expression for the foreign gene were selected on G418 containing plate and expression analysis. The fermentation was carried out in a 50 l bioreactor with 20 l working volume, using a high-density cell culture method by continuously feeding with 50% glycerol-0.8% PTM4 to the growing culture for 54 h at 30°C. Under the control of GAP promoter (pGAP), α-amy gene was constitutively expressed. At the end of the fermentation, the α-AMY expression reached 125 mg/l, while the biomass growth was 186 as measured by absorption of 600 nm. The secreted α-AMY was purified to 97.5% by SP-Sepharose FF ion-exchange chromatography and affinity purification. The recombinant α-AMY showed activity on hydrolysis of starch.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many species of prokaryotes and eukaryotes can produce of α-amylase (α-1,4-d-glucan-glucanohydrolase, EC.3.2.1.1, α-amylase, α-AMY)which can specially digest α-1,4 indican bond to hydrolyze starch as maltose, maltotrise hydrate and other oligosaccharide [1]. α-AMY has larger sales volume each year in the word. Now the Marketable α-AMY is general the crude product from natural α-AMY producing strains, such as the Bacillus subtilis [2], which has been playing important role on bio-ethanol production, paper making, light industry and rock oil exploitation. But the purified α-AMY requiring in some trades, such as bio-pharmaceuticals and analytical chemistry, is hard up because natural α-AMY is always with lower rate to stay in the fermentation broth along other proteins from the host to make it become difficult to be purified [3]. Since it’s possible to construction engineering strain by using well productivity strain as host and setting stronger promoter to control the aim gene [4], the recombinant protein expression would be preponderantly and easily to be purified [5, 6].
The methylotrophic yeast Pichia pastoris (P. pastoris) is an efficient host for expression and secretion of heterologous proteins [7, 8]. It can grow to very high cell densities to high express and secrete recombinant proteins but only very few proteins coming from the host cell consist in the fermentation broth [5, 9]. Glyceraldehyde-3-phosphate dehydrogenase (GAP) is a key enzyme in glycolysis, its promoter constitutively expresses genes well while P. pastoris cells grown on glucose, glycerol or oleic acid [9, 10]. Here, we study of using P. pastoris’s pGAP expression system to constitutive expression of α-AMY in P. pastoris by high-density cell culture.
Materials and methods
Strains, plasmids, enzymes and reagents
The strain of P. pastoris GS115(his4) was purchased from Invitrogen. The pGAP9K [8] expression vector and α-amy gene were kept in our laboratory. Restriction and modified enzymes are the products of TaKaRa and Fermentas. PCR primers and DNA sequence were performed by Shanghai Invitrogen Company. Mediums using for yeast culture were prepared according to the operation manual of Invitrogen [5]. Peptone is the product of Bejing Aoboxing Universeen Bio-Tech CO., LTD.
Preparation of engineering strain
With cDNA of barley genome as template, α-amy gene was amplified and inosculated with the sequence of his6, a purification label for Ni, at the 3′-end by using the primers of 5′-CGCAGAATTCCAAGTCCTCTTTCAGGGGTTC-3′ and 5′-TAAGCGGCCGCTCAATGATGATGATGATGATGGCTCCGTTGTAGTGTTGCCGCGGCACC-3′, which was cloned into the MCS of pGAP9K [8] at EcoR I and Not I sites to generate pGAP9K-α-amy. The BglII linearized pGAP9K-α-amy in 10 μg was transformed into GS115 of P. pastoris by electroporation. The treated GS115 was grown on the YPD plate containing 700 μg/ml of G418, and the grown clones were moved to YPD plates with higher G418 [11] at 1,000–5,000 μg/ml. Each of the eight recombinants from the different concentrations of G418 was moved to YPD medium in a shaking flash, fermented in 30°C for 2 days, and under went SDS-PAGE test and hydrolyzing starch to compare expression levels to select well-expressed clone as engineering strain.
Fermentation condition and metabolite determination for constitutive expression
Eight 1,000 ml flasks with 200 ml YPD medium each were inoculated with 1 ml GS115 (pGAP9K-α-amy) (A 600 = 1.5) and incubated at 30°C and 250 rpm for 20 h. The culture was transferred to a 50 l fermentor with a 20 l working volume. A modified growth medium recommended by Invitrogen Corp was used. It consisted of (l) 26.7 ml of 85% H3PO4, 9.3 g of CaSO4·2H2O, 18.2 g of K2SO4, 14.9 g of MgSO4·7H2O, 4.13 g of KOH, 40 g of glycerol, 40 ml of PTM4, 20 g of peptone and 10 g of yeast extracts. The composition of PTM4 trace elements was described as mentioned above [5]. The fermentation was carried out at 30°C and 100 to 750 rpm by continuously feeding with 50% glycerol-0.8% PTM4. The dissolved oxygen (DO) level was set between 20 and 30%. pH was controlled at five by adding 7 M NH4OH. Samples were taken every 6 h to measure the biomass and α-AMY during the course of fermentation.
Purification of the expressed product
Biomass was removed from the fermentation broth by centrifugation. The supernatant was loaded on SP-Sepharose Fast Flow column (Pharmacia Biotech., NJ, USA). Then, the column was washed with an equilibrium buffer (50 mM Na2HPO4–24 mM C6H8O7). Bound proteins were eluted successively with 0.3, 0.6 and 1 M NaCl. Fractions containing α-AMY were collected, concentrated, and desalted using an ultrafiltration device (Millipore, Bedford, MA, USA), and purified again using a Ni-Agarose 6 × Tagged Protein Purification Kit (Cwbio Co., LTD.). Reverse HPLC analysis was used in the purity assay for the protein with the high performance liquid chromatograph (SHIMADZU, LC-2010CHT).
Activity analysis for the recombinant α-AMY
For qualitative analysis of the α-AMY activity, 50 μl sample and equal volume buffer (pH5.0, 0.1 mol/l C6H8O7-Na2-HPO4) were added in each of the oxford cup stood on 0.5% solubility starch containing plate. After incubation at 37°C for 12 h, the plate was dyed by 1% KI for 5 min. The result was evaluated according the diameter of clarity circle under the oxford cup [12]. And the method of DNS [13] was using for quantitative analysis the activity of the recombinant α-AMY.
Result
Screening high gene copies and well-expressed strain
A total of 120 clones were grown on the 700 μg/ml G418-containing YPD plate after 3 days incubation of the cell transformed with 10 μg plasmid DNA of pGAP9K-α-amy. PCR proved that the α-amy gene was integrated in the genome of P. pastoris (Fig. 1). The results of the clones enduring G418 concentration after being moved to higher G418-containing plates are shown in Table 1. Each eight clones between the 1,000–3,000 μg/ml of G418 concentrations were randomly selected to ferment in shaking flask, and a 50 μl supernatant was taken from each fermentation broth to test the hydrolyzing starch. The dot values of the clarity circles analyzed by software of the Gel Imaging System (Tanon, Ver. 4.00) were shown in Table 2. The fermentation supernatants from the transformants with same anti-G418 concentration were then mixed and run on the SDS-PAGE. The average expression levels of each of the eight clones from the 1,000–3,000 μg/ml of G418 concentration are shown in Fig. 2. Table 2 and Fig. 2 indicated it is positive correlations of gene copy number and foreign protein average expression level. But the expression could be obviously difference between the recombinants in the same G418 resistance (Table 2; Fig. 3). No. 5, the best expressed clone as showing in Fig. 3 were selected as engineering strain.
Peptone selection
GS115 (pGAP9K-α-amy) was inoculated into YPD medium with peptone sourced from three different companies which are the peptone of Bejing Aoboxing Universeen Bio-Tech CO., LTD., tryptone of Beijing Land Bridge Technology CO. LTD. and Tryptone of Oxoid LTD. to ferment for 2 days. The result of fermentation broth hydrolyzing starch is showing in Fig. 4. The dot values analyzed by software of the Gel Imaging System (Tanon, Ver. 4.00) indicated that the peptone from Bejing Aoboxing Universeen Bio-Tech Co., LTD. is better and it was chosen to use in the high-density fermentation.
Peptones affect on GS115(pGAP9K-α-amy) expression of α-AMY. a Fermentation broth hydrolyzed starch, b Dot values analyzed by software of the Tanon Gel Imaging System. 1 Peptone of Bejing Aoboxing Universeen Bio-Tech Co., LTD. 2 Tryptone of Beijing Land Bridge Technology Co. LTD. 3 Tryptone of Oxoid LTD
α-AMY expression by high-density cell culture
The GS115 (pGAP9K-α-amy) cells were initially grown on glycerol and then fed-batch with 50% glycerol-0.8% PTM4 for 54 h. Fig. 5 shows the α-amy gene expression during high-density cell culture of GS115 (pGAP9Kα-amy). P. pastoris cell concentration/biomass (A 600), α-AMY production, agitation, aeration and glycerol feeding during the fermentation were described in Fig. 6. At the end of fermentation, the culture volume increased from 21.6 to 33.3 l, accumulation of biomass is A 600 = 186 while 125 mg/l of α-AMY was secreted into the fermentation broth.
Purification of recombinant α-AMY
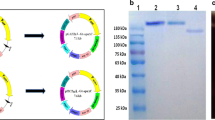
After purification by the SP-sepharose fast flow, the mostly proteins from host were excluded in the process and the foreign protein existed in the eluted buffer of 1 M NaCl with some hybrid proteins which were further excluded by the affinity purification and the purity quotient of the aim protein is 97.5% (Fig. 7).
Quantitative determination of the purified α-AMY
As shown in Fig. 8, the purified recombinant α-AMY and the α-AMY product (Shanghai Kayon Biological Technology Co. Ltd.) hydrolyzed starch as reducing sugar to react with 3.5-dinitrosalicylic acid, presenting a reddish brown color. 546 U/mg of the recombinant α-AMY had been harvested menstruated by ultraviolet Spectrophotometer (Shimadzu, UV-2550).
Discussion
Rabbani reported using E. coli to express α-AMY [14]. The expressed product stayed in the host cell in the way of inclusion body which is misfolding, poor solubility, and necessitates complicated purification methodologies [15, 16]. Wang reported expressing α-AMY in Saccharomyces cerevisiae, but the yield is very low[17]. The development of S. cerevisiae expression system was limited by the reasons of lacking strong promoter and being incapable high-density cell culture [18]. P. pastoris has been widely reported as an efficient expression system secreting very low levels of native proteins. The secreted heterologous protein comprises high ratio to hybrid protein in the medium serving as the first step in the purification of the protein [5]. And also it can grow to very high cell densities to express foreign protein [9]. The insufficiency of P. pastoris is its low expression level [7]. More attentions have been playing on the increase of foreign protein expression in P. pastoris. Generally speaking, mRNA level is parallel to the gene copy number in genome, and the amount of protein expression correlates with the mRNA level in P. pastoris [11]. Designing a resistance gene in expression vector is widely used for selecting more gene-copied recombinant of P. pastoris [11, 18]. Then, some authors reported that gene copy number does not correlate or is even contrary to foreign protein expression level [19, 20]. Our results indicated that gene copy number is positively correlated to foreign protein expression in the mass. However, the lower expressed, even unexpressed, strains may consist in the group of high gene copy numbers, which indicates that higher gene copies no mean certainly more expression. A second screening is necessary between the higher gene-copied recombinants.
High-density cell culture is beneficial for harvesting more products with unit volume of culture by cultivating the cell to high-density with high vigor in a bioreactor [9]. Using the screened strain to progress in 54 h fermentation in a bioreactor, 125 mg/l of α-AMY was secreted by the cell of A 600 = 186, which possesses about 20% of total proteins secreted by the P. pastoris and the enzyme activity is 546 U/mg. This expression yield is significantly higher than Yang used the same promoter to express α-AMY in P. pastoris [21].
References
Atsuhiko S, Hiroyuki K, Hirosuke O (1982) Physiology of α-amylase production by immobilized Bacillus amyloliquefaciens. Eur J Appl Microbiol Biotechnol 14(1):7–12
Konsoula Z, Liakopoulou-Kyriakides M (2007) Co-production of alpha-amylase and beta-galactosidase by Bacillus subtilis in complex organic substrates. Bioresour Technol 98(1):150–157
Deutch CE (2002) Characterization of a salt-tolerant extracellular a-amylase from Bacillus dipsosauri. Lett Appl Microbiol 35(1):78–84
Gasser B, Maurer M, Gach J, Kunert R, Mattanovich D (2006) Engineering of Pichia pastoris for improved production of antibody fragments. Biotechnol Bioeng 94(2):353–361
Invitrogen Corp. (1998) A manual of methods of expression of recombinant proteins in Pichia pastoris, Invitrogen Corp., San Diego
Cai X, Wang J, Wang Y, Yang Y, Gao J, Fu W, Wang J, Xu D (2010) Expression, purification and characterization of recombinant human interleukin-22 in Pichia pastoris. Mol Biol Rep 37(6):2609–2613
Cos O, Ramón R, Montesinos JL, Valero F (2006) Operational strategies, monitoring and control of heterologous protein production in the methylotrophic yeast Pichia pastoris: a review. Microb Cell Factories 5:17
Zhang AL, Luo JX, Zhang TY, Pan YW, Tan YH, Fu CY, Tu FZ (2009) Recent advances on the GAP promoter derived expression system of Pichia pastoris. Mol Biol Rep 36(6):1611–1619
Zhang AL, Zhang TY, Luo JX, Fu CY, Qu Z, Yi GH, Su DX, Tu FZ, Pan YW (2009) Inducible expression of human angiostatin by AOXI promoter in P. pastoris using high-density cell culture. Mol Biol Rep 36(8):2265–2270
Waterham HR, Digan ME, Koutz PJ, Lair SV, Cregg JM (1997) Isolation of the Pichia pastoris glyceraldehydes-3-phosphate dehydrogenase gene and regulation and use of its promoter. Gene 186:37–44
Scorer CA, Clare JJ, McCombie WR, Romanos MA, Sreekrishna K (1994) Rapid selection using G418 of high copy number transformants of Pichia pastoris for high-level foreign gene expression. Biotechnology (NY) 12(2):181–184
Worthington P, Hoang V, Perez-Pomares F, Blum P (2003) Targeted disruption of the alpha-amylase gene in the hyperthermophilic archaeon Sulfolobus solfataricus. J Bacteriol 185(2):482–488
Swamy MN, Murthy HS, Rao GS (2001) Intraoperative blood glucose levels in neurosurgical patients: an evaluation of two fluid regimens. Neurol India 49(4):371–374
Rabbani M, Mirmohammad SH, Moazen F, Rahimi M, Salehi G (2011) Cloning and expression of randomly mutated Bacillus subtilis α-amylase genes in HB101. Biotechnol Res Int. doi:10.4061/2011/305956
Butt TR, Edavettal SC, Hall JP, Mattern MR (2005) SUMO fusion technology for difficult-to-express proteins. Protein Expr Purif 43(1):1–9
Gao Y, Huang YF (2002) Advances in eukaryotic expression systems. Zhonghua Nan Ke Xue 8(4):292–298
Wang Y, Liu H, Sun T, Zhang S (1998) Cloning of alpha-amylase gene from Schwanniomyces occidentalis and expression in Saccharomyces cerevisiae. Sci China Life Sci 41(6):569–675
Gurkan C, Ellar DJ (2005) Recombinant production of bacterial toxins and their derivatives in the methylotrophic yeast Pichia pastoris. Microb Cell Fact 4:33
Sunga AJ, Tolstorukov I, Cregg JM (2008) Post transformational vector amplification in the yeast Pichia pastoris. FEMS Yeast Res 8(6):870–876
Hohenblum H, Gasser B, Maurer M, Borth N, Mattanovich D (2004) Effects of gene dosage, promoters, and substrates on unfolded protein stress of recombinant Pichia pastoris. Biotechnol Bioeng 85:367–375
Inan M, Fanders SA, Zhang W, Hotez PJ, Zhan B, Meagher MM (2007) Saturation of the secretory pathway by overexpression of ahookworm (Necator americanus) protein (Na-ASP1). Methods Mol Biol 389:65–76
Acknowledgments
This study was supported by National Nonprofit Institute Research Grant of CATAS-ITBB (No. ITBB120503) and the Science foundation of Chinese Academy of Tropical Agricultural Sciences (No. RKY0623).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, Z.W., Yin, H.X., Yi, X.P. et al. Constitutive expression of barley α-amylase in Pichia pastoris by high-density cell culture. Mol Biol Rep 39, 5805–5810 (2012). https://doi.org/10.1007/s11033-011-1390-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-1390-1