Abstract
High-affinity ammonium transporters (AMT1) are responsible for ammonium (NH4 +) acquisition and/or perception in the micromolar range, and their expressions can be differentially regulated by nitrogen (N) availability. The present study characterised the functions of the rice (Oryza sativa) OsAMT1.3 transporter to understand its contribution to NH4 + acquisition and plant adaptation to environments with low N availability. Transgenic rice plants were obtained to study the activity of the OsAMT1.3 promoter (P OsAMT1.3 :GFP:GUS) and the overexpression of the OsAMT1.3 gene (UBIL:OsAMT1.3:3xHA) in plants. The OsAMT1.3 promoter activity was induced strongly in the absence of N and occurred primarily in the zones of lateral root emission and root tips. Anatomical sections of the segment of root tips and the middle third showed a differential pattern of OsAMT1.3 activity. Analysis of the OsAMT1.1–1.3 transporter expression profiles indicated that overexpression of OsAMT1.3 positively affected OsAMT1.2 expression. When subjected to a low N supply, plants overexpressing OsAMT1.3 showed lower K M and C min values. Additionally, these lines showed longer roots with a higher area, volume, and number of tips. The data suggested that OsAMT1.3 is involved in the ability of rice plants to adapt to low NH4 + supplies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrogen (N) is an essential element for plants and is the most limiting element for crop productivity and cereal grain quality (Bu et al. 2011). The current high rice productivities became possible partly because of the intensive use of fertilisers, primarily N fertilisers. Approximately 110 million tons of N fertilisers are applied annually worldwide, at high cost (FAO 2012). Additionally, this excess application may be associated with severe environmental damage (Mulvaney et al. 2008).
Plants absorb N preferentially as ammonium (NH4 +) and nitrate (NO3 −). Under anaerobiosis, NH4 + is the main form of N available to plants (Funayama et al. 2013). However, excess NH4 + absorption can be toxic (Britto et al. 2001), and the absorption and metabolism of this nutrient is highly regulated in plants (Sonoda et al. 2003a, b).
Plants take up NH4 + through high-affinity (HATS) and low-affinity (LATS) transport systems, depending on the nutrient concentration in the external medium (Wang et al. 1993). Recent studies have characterised the high-affinity ammonium transporters (AMT1) because they act at low soil solution N concentrations and may be involved in NH4 + uptake efficiency in plants (Ranathunge et al. 2014; Lima et al. 2010; Gu et al. 2013).
Genome sequencing identified several ammonium transporters belonging to the AMT1 family in various species, such as Oryza sativa (OsAMT1.1–1.3) (Suenaga et al. 2003; Sonoda et al. 2003a, b), Lycopersicon esculentum (LeAMT1.1–1.3) (von Wirén et al. 2000), Arabidopsis thaliana (AtAMT1.1–1.5, AtAMT2) (Kaiser et al. 2002; Sohlenkamp et al. 2002), and Triticum aestivum (TaAMT1.1–1.3) (Jahn et al. 2004) profiles. OsAMT1.1 is an NH4 +-responsive gene, expressed in the roots and shoots (Ranathunge et al. 2014), whereas the OsAMT1.2 and OsAMT1.3 genes are specifically expressed in the roots (Sonoda et al. 2003a, b). OsAMT1.2 responds positively to resupply with increasing doses of ammonium, whereas OsAMT1.3 shows increased expression at N concentrations below 0.15 mM and is repressed at higher concentrations (Gaur et al. 2012).
The OsAMT1.2 and OsAMT1.3 gene expression profiles in the roots suggest that both are key components of the uptake system. However, they play different roles in N utilisation. OsAMT1.3 appears to act as an N sensor, whereas OsAMT1.2 acts as an NH4 + transporter (assimilator) at low concentrations. The observation that OsAMT1.3 was repressed not only in the presence of NH4 + but also in the presence of NO3 − has led to the hypothesis that it may serve as an N sensor (Yao et al. 2008).
Sonoda et al. (2003a, b) suggested that NH4 + transporters are regulated by the glutamine concentrations inside the roots and not by the NH4 + concentrations in the external solution (i.e., the control over the NH4 + uptake is internal). However, the regulation of the AMT1 family of transporters varies with the genotype and NH4 + availability (Gaur et al. 2012).
Plant varieties with increased N uptake efficiency in soils containing low N concentrations must be selected for sustainable agriculture (Glass et al. 2002). Therefore, the molecular and physiological responses of plants under low N concentrations should be well understood. Some researchers have suggested that the OsAMT1.3 transporter may function to signal the presence of N in the soil, in addition to its potential involvement as a transporter in the uptake of reduced NH4 + levels in the soil (Sonoda et al. 2003a, b; Gaur et al. 2012).
The present study aimed to evaluate the contribution of the OsAMT1.3 transporter in increasing the NH4 + uptake efficiency and to characterise its role in the mechanisms by which plants adapt to environments with low N availability.
Materials and methods
Plant material and growth conditions
Transgenic rice plants of the variety Nipponbare (Oryza sativa L. subsp. Japonica) were used in this study. The experiments were performed in a climatic chamber (light/dark cycle, 12/12 h; 28/26 °C; light intensity, 500 µmol m−2 s−1; relative humidity, 70 %). The rice seeds were surface-sterilised with sodium hypochlorite (2 %) for 10 min and germinated in distilled water. Five days after germination (DAG), plants were transferred to Hoagland solution (Hoagland and Arnon 1950) with different N regimes. The solutions were changed every 3 days, and the pH was maintained at 5.8.
Gene constructs
Transgenic rice lines were generated by expressing OsAMT1.3 under the control of the maize ubiquitin 1 promoter (UBIL:OsAMT1.3:3xHA) using the MultiSite Gateway cloning kit (Life Technologies, Carlsbad, CA, USA). The amplified fragment of OsAMT1.3 was cloned into the molecular cloning site of the pH7m34GW vector, with the recombination sites necessary for cloning the Gateway vectors (Table S1). To generate the green fluorescent protein (GFP) and β-glucuronidase (GUS) OsAMT1.3 construct, a 1500-bp fragment upstream of the translation initiation site was amplified from genomic DNA in two subsequent PCRs with hybrid primers (Table S2) and cloned into the molecular cloning site of the pGWFS7 vector (Karimi et al. 2002) to drive GFP expression and GUS activity. The resulting constructs were transformed into Escherichia coli.
Genetic transformation and in vitro development of transgenic plants
Rice plants were transformed according to the Agrobacterium-mediated transformation of embryogenic calli with the OsAMT1.3 binary construct. The subsequent regeneration of transgenic lines and the separation of early events of independent stable transformations within the callus material have been described previously (Toki et al. 2006). Transformed calli were selected via hygromycin resistance conferred by a UBIL promoter-driven hph gene. Several T1 transformants were generated and confirmed by hygromycin resistance. The transgenic lines selected were grown in a greenhouse, and homozygous lines of the T3 generation were selected by segregation analysis for hygromycin resistance.
Localisation studies
Transgenic rice plants (L#4) were grown for 14 days in a Hoagland solution with two different N treatments: a constant N supply (2.0 mM NH4 +) as a control or N deficiency. The roots were harvested, infiltrated with staining solution containing 5-bromo-4-chloro-3-indolyl-β-d-glucuronide (X-gluc), and maintained at 37 °C for 3 h (Jefferson et al. 1987). Root segments (15 mm) from the tips of the roots and middle third were infiltrated with a 2.5 % glutaraldehyde solution in 0.01 M phosphate buffer. The samples were embedded in Leica® synthetic resin (hydroxyethyl methacrylate), according to the manufacturer’s instructions, and sectioned using a Leica rotary microtome. The 2-µm sections were observed on an Axioplan light microscope (Zeiss) equipped with AxioVision software.
OsAMT1.3 promoter activity
Transgenic rice plants (L#4) were grown in a Hoagland solution with low N availability until 10 DAG (0.5 mM NO3 −) to reduce OsAMT1.3 expression after germination and reduce the interference of this dose in the subsequent treatments. The plants were then subjected to three different N regimes over 14 days: without N (control), 2.0 mM NO3 −, and 2.0 mM NH4 +. Roots were harvested at 0, 3, 7, and 14 days after treatment and stored at −80 °C for subsequent use.
Root samples were ground in liquid N2; homogenised in three volumes of 50 mM Tris–HCl pH 8.0 buffer containing 1 mM EDTA, 1.5 % polyvinylpolypyrrolidone (PVPP), 10 mM dithiothreitol (DTT), 30 % glycerol, and 1 mM phenylmethylsulfonyl fluoride (PMSF); and then centrifuged at 14,000×g for 30 min. The supernatant was used to determine the enzyme activities. The protein concentration was determined according to Bradford (1976). The GUS activity was determined spectrophotometrically using the enzyme substrate p-nitrophenyl-β-d-glucuronide (PNPG), according to Aich et al. (2001). The activity was expressed as ∆OD405 in mg−1 protein h−1.
OsAMT1.1–1.3 NH4 + transporter expression
Transgenic rice plants (L#2 and L#8), which overexpressed OsAMT1.3, and WT plants were grown in a Hoagland solution with 0.5 mM NO3 −, to reduce the natural OsAMT1.3 expression after germination. At 30 DAG, the plants were treated with a solution containing 0.5 mM NH4 +. Plants were harvested at 2 and 6 h following treatment, and root samples were stored at −80 °C for subsequent use.
Total RNA was extracted according to GAO et al. (2001) in NTES buffer (0.2 M Tris–HCl pH 8.0, 25 mM EDTA, 0.3 M NaCl, 2 % SDS). The total RNA was quantified using a Qubit 2.0 fluorometer (Life Technologies), according to the manufacturer’s instructions. The total RNA was treated with DNaseI (Life Technologies) and used for cDNA synthesis using a High-Capacity RNA-to-cDNA™ kit (Life Technologies) and oligo(dT) primers, according to the manufacturer’s instructions. qRT-PCR was performed using the Power SYBR® Green PCR Master Mix kit and a StepOne real-time PCR system (Applied Biosystems, Carlsbad, CA, USA). The PCR program consisted of 95 °C for 15 s and 60 °C for 60 s. Two qRT-PCR determinations were performed for each cDNA sample. The threshold cycle (C t) values for each sample were normalised with the O. sativa elongation factor (eEF1-α) as a housekeeping gene. The relative quantity was calculated using the 2−ΔΔCT method (Livak and Schmittgen 2001). The primers designed by Duan et al. (2007) for the ammonium transporter genes (OsAMT1.1, 1.2, and 1.3) were used.
NH4 + uptake kinetics of rice lines overexpressing the OsAMT1.3 gene
Transgenic rice plants (L#2 and L#8) were grown as previously described. At 27 DAG, the plants were submitted to N starvation for 72 h, followed by resupply with 0.2 mM NH4 +. Samples of nutrient solution (1.0 mL) were collected at intervals of 30 min until N exhaustion. At the end of the experiment, shoots and roots were harvested (Table S3). The kinetic parameters, V max and K M , were measured by depletion of NH4 + in the uptake solution over time according to Claassen and Barber (1974). The ammonium content of the nutrient solution was determined according to Felker (1977). The integrated analysis of NH4 + uptake was calculated as α = V max/K M (Marschner 1995). In the final samples of nutrient solution, the concentration at which net uptake of ions ceases before the ions are completely depleted was measured (C min) (Marschner 1995). Root parameters (root length, surface area, projected area, volume, and number of tips) were determined using the Winrhizo 4.1 software (Regent Instruments, Quebec, Canada).
Statistical analysis
A completely randomised experimental design was utilised, with four replicates in all experiments. Analysis of variance was performed by applying the F test, the averages were compared using a Scott–Knott test at p ≤ 0.05, and the standard error was calculated.
Results
OsAMT1.3 tissue-specific localisation and quantification
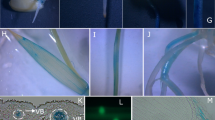
To identify the sites of action of the OsAMT1.3 transporter and study its possible involvement in the adaptation of plants to low N supplies, rice plants (L#4) were grown under high NH4 + supply or N starvation for 14 days. The roots were collected and infiltrated with a solution containing the β-glucuronidase (X-gluc) substrate (Fig. S1). Under treatment by 2.0 mM of NH4 +, no promoter activity was observed (Fig. 1a), whereas under N starvation, an intense blue stain was observed (Fig. 1b). Two segments of roots (black boxes) were selected for further analyses: 15 mm from the tip and the middle third (Fig. 1c, d). In these segments, intense GUS and GFP activities were observed close to the epidermis, at the zones of lateral root emission and at the tips of the lateral roots (Fig. 1e–h). Histological sections of the selected segments are shown in Fig. 2a–f.
Rice roots (L#4) grown with 2.0 mM NH4 + (a) and under N starvation (scale bar 10 mm) (b). Black boxes indicate the root segments selected for the anatomical sections. Middle third (c) and tip (d) (scale bar 2.0 mm). An intense blue stain (GUS) and fluorescence (GFP) were observed in the lateral root emission zone (scale bar 100 μm) (e, f) and in the tips (scale bar 400 μm) (g, h)
Cross sections obtained from the tips and from the middle third of the root exhibited different patterns of staining (Fig. 2). Cross section from the tips showed OsAMT1.3 activity in the exodermis, sclerenchyma, cortex, and stele (Fig. 2a, b), while in cross and longitudinal sections from the middle third region, with longer lateral roots, a higher OsAMT1.3 activity was observed in the sites of lateral root emission and at the exodermis (Fig. 2c–f; Fig. S2).
In addition to the histochemical assays (Figs. 1, 2), the in vitro GUS activity was also quantified under treatment with NH4 + or NO3 − and under N deficiency for 14 days. High GUS activity was observed in plants (L#4) grown without N at 3 days following the treatment (Fig. 3). This high activity remained until 7 days and then decreased after 14 days. High OsAMT1.3 activity was observed in all periods analysed under N deprivation. No changes were noted in the GUS activity in plants grown under a constant N supply. These results indicated that the OsAMT1.3 ammonium transporter is induced strongly by N deficiency and repressed under NO3 − or NH4 + supply.
OsAMT1.3 positively affects OsAMT1.2 expression
Rice plants overexpressing OsAMT1.3 were obtained to examine the effect of its expression on the NH4 + uptake and root growth. L#2 and L#8 were selected for further study because they showed high OsAMT1.3 expression levels. Rice lines showed increased levels of OsAMT1.3 expression under constant N supply (Fig. 4a). In addition, OsAMT1.2 expression showed the same pattern as OsAMT1.3 (Fig. 4a, b; Fig. S3). The OsAMT1.2 expression levels were higher at 2 h than at 6 h (Fig. 4c, d), and the lines did not present significant changes in their OsAMT1.1 expression levels at 2 and 6 h (Fig. 4c).
Uptake kinetics and root parameters under low NH4 + supply and N deficient
The overexpression of the OsAMT1.3 did not alter the V max significantly in the rice lines; however, the K M values were 26.4 and 52.4 % lower for L#2 and L#8, respectively, than for the WT (Table 1). The integrate analysis of NH4 + uptake using the α value indicated that the rice lines showed kinetic parameters more favourable to the uptake of NH4 + at low concentrations than the WT. Furthermore, a significant reduction in C min values of 4.2 and 17 % was observed for L#2 and L#8, respectively (Table 1). When plants were grown in N-deficient medium or 0.2 mM NH4 + for 14 days, several root parameters were modified in the rice lines. Greater root length, projection area, surface area, root volume, and number of tips were observed for L#2 and L#8 in both treatments (Table 2).
Despite the higher values observed for the root parameters in 0.2 mM NH4 +, the differences between the lines and WT were more evident without N. L#2 and L#8 showed greater increases in the root length, approximately 32 and 48 %, respectively, without N, while with 0.2 mM NH4 +, the increases were 12 and 20 % for L#2 and L#8, respectively. In addition, marked increases in the number of tips were observed for L#2 and L#8 (44 and 77 %, respectively) during N starvation, while under the treatment with 0.2 mM NH4 + a minor increase was observed: 4 and 7 %, for L#2 and L#8, respectively.
Discussion
In this study, transgenic rice plants (L#4) were submitted to N deprivation or 2.0 mM NH4 + for 14 days to identify whether the promoter activity is regulated by N and the sites of action of OsAMT1.3. The OsAMT1.3 activity was restricted to the roots under N deficiency (Fig. 1), supporting the data found by Sonoda et al. (2003a, b) and Yao et al. (2008). The plants subjected to N deprivation showed higher OsAMT1.3 promoter activity, mainly in the root emission zone and roots tips (Fig. 1e–h). Furthermore, root sections at the segment from the tips and middle third exhibited different patterns of OsAMT1.3 promoter activity (Fig. 2). In the tips, the OsAMT1.3 activity was uniformly distributed at the exodermis, sclerenchyma, cortex, and stele (Fig. 2a, b), while in the middle third segment, it was observed only at the cortex and exodermis (Fig. 2c–f). The ammonium taken up by the AMTs is readily assimilated by cytosolic GS1 and NADH-GOGAT isozymes in the surface cell layers of the roots, epidermis, and exodermis (Hirose et al. 1997; Ishiyama et al. 1998). The OsAMT1.3 promoter activity observed mainly in the exodermis (Fig. 2c, d) is related to the sites of primary ammonium assimilation, indicating a role of the encoded protein in the ammonium uptake. The OsAMT1.3 promoter activity at root emission zones also suggested a role in signalling events that result in lateral root emission under N deficiency (Fig. 2e, f; Fig. S2). Yao et al. (2008) observed that OsAMT1.3 was expressed preferentially at the apex of the lateral and seminal roots. These data support the idea that OsAMT1.3 acts as a sensor for nutrients present in the soil, changing the plant metabolism through the activation of signal transduction pathways (Gojon et al. 2011).
In A. thaliana with a quadruple knockout of ammonium transporter genes (amt1.1, amt1.2, amt1.3, and amt2.1), no lateral root formation was observed, and the lateral root formation decreased significantly in a atamt1.3 mutant under conditions of low N, supplied as NH4 + and NO3 − (Lima et al. 2010). These authors suggested that AtAMT1.3 might be involved in triggering lateral root formation in Arabidopsis. The AtAMT1.3 ammonium transporter does not show high sequence similarity with OsAMT1.3 (Li et al. 2009); however, OsAMT1.3 could also act as a signal for the emission of lateral roots under N deficiency in rice (Fig. 2c–f).
The OsAMT1.3 activity in plants grown in nutrient solution without N was higher at 3 and 7 days after treatment. When these plants were transferred to solutions containing N as NH4 + or NO3 − ions, no changes were observed in the OsAMT1.3 promoter activity (Fig. 3). This result is consistent with the data of Sonoda et al. (2003a, b), where rice plants submitted to N starvation showed upregulation of OsAMT1.3 expression, while in plants submitted to N supply as NH4 + or NO3 −, the opposite behaviour was observed: downregulation of OsAMT1.3 expression. Gaur et al. (2012) reported that the repression of OsAMT1.3 through an increase in N might not be a universal mechanism, but may depend on the genotype and on the N level required by a given genotype. These authors observed repression of OsAMT1.3 with increasing NH4 + concentrations in the solution, up to 1.0 mM, for the rice cultivar Kalanamak 3119, whereas the cultivar Pusa Basmati showed the opposite behaviour, increasing the OsAMT1.3 expression under higher NH4 + levels. The Nipponbare rice variety used in our study requires low N supply and also showed high OsAMT1.3 expression under N deficiency (Fig. 3). OsAMT1.3 expression is believed to be useful as a biomarker to determine the optimal N supply for rice varieties adapted to low and high N environments. These differences in induction of the high-affinity AMT genes might be attributed to differences in N perception and signalling (Gaur et al. 2012).
Thus, overexpression of members of the AMT1 family might be useful to improve N uptake from soils with low NH4 + concentrations (Ranathunge et al. 2014). However, some members of the AMT1 family may not be directly involved in the acquisition of NH4 + from the external solution, acting instead as sensors of the intracellular NH4 + status (Hoque et al. 2006). Thus, we developed rice plants overexpressing OsAMT1.3 to identify whether the gene product is involved in ammonium uptake or signalling.
The relative expression of the high-affinity ammonium transporter genes (OsAMT1.1–1.3) was performed at 2 and 6 h with a constant N supply (Fig. 4). L#2 and L#8 showed high OsAMT1.3 expression (Fig. 4a). In addition, a strong, positive correlation was observed between the OsAMT1.3 and OsAMT1.2 expressions at 2 and 6 h (Fig. 4a, b; Fig. S3). No significant change was observed in the expression of the OsAMT1.1. These data support the hypothesis that the overexpression of OsAMT1.3 could alter the expression of other members involved in high-affinity ammonium transport, such as OsAMT1.2. At 6 h, a decrease in the expression of OsAMT1.2 was observed, possibly indicating negative feedback regulation by glutamine (Sonoda et al. 2003a, b).
Rice lines overexpressing OsAMT1.3 showed lower K M and C min values than WT plants when supplied with 0.2 mM NH4 + (Table 1). This result suggested that OsAMT1.3 overexpression, associated with a higher OsAMT1.2 expression (Fig. 4), resulted in increased uptake efficiency by these plants, considering that OsAMT1.2 is involved in NH4 + uptake from soil solutions (at concentrations <200 nM) and in the retrieval of NH4 + in the vascular system (Sonoda et al. 2003a, b). The AMT1 family might show different K M values at low concentrations of NH4 +, as observed in Arabidopsis and maize plants (Gazzarrini et al. 1999 and Gu et al. 2013). This suggested that the higher expression of OsAMT1.3 and OsAMT1.2 changed the K M of the NH4 + high-affinity transport system, resulting in increased uptake efficiency.
The combination of low K M and C min values associated with increased root growth is a desirable characteristic in crop plants because it equates to increased N uptake efficiency (Barber 1995). In addition to improved kinetic parameters, the rice lines showed longer roots with more tips, indicating that OsAMT1.3 expression might contribute to an increase in lateral root emission under N deficiency or low N supply (Table 2). A greater difference was observed for root length and number of tips during N starvation, indicating a contribution of the natural OsAMT1.3 expression to the root parameters.
Our results showed that overexpression of OsAMT1.3 was associated with the natural expression of OsAMT1.2, and promoted the uptake of NH4 + at low concentrations and changes to the root morphology of the lines. However, further studies should be carried out to isolate the role of the OsAMT1.3 using knockout plants.
References
Aich S, Delbaere LTJ, Chen R (2001) Expression and purification of Escherichia coli beta-glucuronidase. Protein Expr Purif 22:75–81. doi:10.1006/prep.2001.1401
Barber SA (1995) Soil nutrient bioavailability: a mechanistic approach, 2nd edn. Wiley, New York
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Britto DT, Siddiqi MY, Glass ADM, Kronzucker HJ (2001) Futile transmembrane NH4 + cycling: a cellular hypothesis to explain ammonium toxicity in plants. PNAS 98:4255–4258. doi:10.1073/pnas.061034698
Bu Y, Takano T, Nemoto K, Liu S (2011) Research progress of ammonium transporter in rice plants. Genomics 2:19–23. doi:10.5376/gab.2011.02.0003
Claassen N, Barber SA (1974) A method for characterizing the relation between nutrient concentration and flux into roots of intact plants. Plant Physiol 54:564–568. doi:10.1104/pp.54.4.564
Duan YH, Zhang YL, Ye LT, Fan XR, Xu GH, Shen QR (2007) Responses of rice cultivars with different nitrogen use efficiency to partial nitrate nutrition. Ann Bot 99:1153–1160. doi:10.1093/aob/mcm051
Felker P (1977) Microdetermination of nitrogen in seed protein extracts with the salicylate–dichloroisocyanurate color reaction. Anal Chem 49:1080. doi:10.1021/ac50015a053
Food and Agriculture Organization of the United Nations (FAO) (2012) Current world fertilizer trends and outlook to 2016. ftp://ftp.fao.org/ag/agp/docs/cwfto16.pdf. Accessed 09 Jan 2014
Funayama K, Kojima S, Tabuchi-Kobayashi M, Sawa Y, Nakayama Y, Hayakawa T, Yamaya T (2013) Cytosolic glutamine synthetase1;2 is responsible for the primary assimilation of ammonium in rice roots. Plant Cell Physiol 54:934–943. doi:10.1093/pcp/pct046
Gao J, Liu J, Li B, Li Z (2001) Isolation and purification of functional total RNA from blue-grained wheat endosperm tissues containing high levels of starches and flavonoids. Plant Mol Biol Rep 19:185–186. doi:10.1007/BF02772163
Gaur VS, Singh US, Gupta AK, Kumar A (2012) Understanding the differential nitrogen sensing mechanism in rice genotypes through expression analysis of high and low affinity ammonium transporter genes. Mol Biol Rep 39:2233–2241. doi:10.1007/s11033-011-0972-2
Gazzarrini S, Lejay L, Gojon A, Ninnemann O, Frommer WB, von Wíren N (1999) Three functional transporters for constitutive, diurnally regulated, and starvation-induced uptake of ammonium into Arabidopsis roots. Plant Cell 11:937–947. doi:10.1105/tpc.11.5.937
Glass ADM, Britto DT, Kaiser BN, Kinghorn JR, Kronzucker HJ, Kumar A, Okamoto M, Rawat S, Siddiqi MY, Unkles SE, Vidmar JJ (2002) The regulation of nitrate and ammonium transport systems in plants. J Exp Bot 53:855–864. doi:10.1093/jexbot/53.370.855
Gojon A, Krouk G, Perrine-Walker F, Laugier E (2011) Nitrate transceptor(s) in plants. J Exp Bot 62:2299–2308. doi:10.1093/jxb/erq419
Gu R, Duan F, An X, Zhang F, von Wirén N, Yuan L (2013) Characterization of AMT-mediated high-affinity ammonium uptake in roots of maize (Zea mays L.). Plant Cell Physiol 54:1515–1524. doi:10.1093/pcp/pct099
Hirose N, Hayakawa T, Yamaya T (1997) Inducible accumulation of mRNA for NADH-dependent glutamate synthase in rice roots in response to ammonium ions. Plant Cell Physiol 38:1295–1297. doi:10.1093/oxfordjournals.pcp.a029120
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. California Agricultural Experiment Station, Berkeley, p 347
Hoque MS, Masle J, Udvardi MK, Ryan PR, Upadhyaya NM (2006) Over-expression of the rice OsAMT1-1 gene increases ammonium uptake and content, but impairs growth and development of plants under high ammonium nutrition. Funct Plant Biol 33:153–163. doi:10.1071/FP05165
Ishiyama K, Hayakawa T, Yamaya T (1998) Expression of NADH-dependent glutamate synthase protein in the epidermis and exodermis of rice roots in response to the supply of ammonium ions. Planta 204:288–294. doi:10.1007/s004250050258
Jahn TP, Møller ALB, Zeuthen T, Holm LM, Klaerke DA, Mohsin B, Kühlbrandt W, Schjoerring JK (2004) Aquaporin homologues in plants and mammals transport ammonia. FEBS Lett 574:31–36. doi:10.1016/j.febslet.2004.08.004
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kaiser BN, Rawat SR, Siddiqi MY, Masle J, Glass ADM (2002) Functional analysis of an Arabidopsis T-DNA “knockout” of the high-affinity NH4 + transporter AtAMT1;1. Plant Physiol 130:1263–1275. doi:10.1104/pp.102.010843
Karimi M, Inzé D, Depicker A (2002) Gateway vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7:193–195. doi:10.1016/S1360-1385(02)02251-3
Li B-Z, Merrick M, Li S, Li H, Zhu S, Shi W, Su Y (2009) Molecular basis and regulation of ammonium transporter in rice. Gene 16:314–322. doi:10.1016/S1672-6308(08)60096-7
Lima JE, Kojima S, Takahashi H, von Wíren N (2010) Ammonium triggers lateral root branching in Arabidopsis in an ammonium transporter1;3-dependent manner. Plant Cell 22:3621–3633. doi:10.1105/tpc.110.076216
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Marschner H (1995) Mineral nutrition of higher plants, 2nd edn. Academic Press, London
Mulvaney RL, Khan SA, Ellsworth TR (2008) Synthetic nitrogen fertilizers deplete soil nitrogen: a global dilemma for sustainable cereal production. J Environ Qual 38:2295–2314. doi:10.2134/jeq2008.0527
Ranathunge K, El-Kereamy A, Gidda S, Bi Y-M, Rothstein SJ (2014) OsAMT1;1 transgenic rice plants with enhanced NH4 + permeability show superior growth and higher yield under optimal and suboptimal NH4 + conditions. J Exp Bot. doi:10.1093/jxb/ert458
Sohlenkamp C, Wood CC, Roeb GW, Udvardi MK (2002) Characterization of Arabidopsis AtAMT2, a high-affinity ammonium transporter of the plasma membrane. Am Soc Plant Biol 130:1788–1796. doi:10.1104/pp.008599
Sonoda Y, Ikeda A, Saiki S, von Wirén N, Yamaya T, Yamaguchi J (2003a) Distinct expression and function of three ammonium transporter genes (OsAMT1;1-1;3) in rice. Plant Cell Physiol 44:726–734. doi:10.1093/pcp/pcg083
Sonoda Y, Ikeda A, Saiki S, Yamaya T, Yamaguchi J (2003b) Feedback regulation of the ammonium transporter gene family AMT1 by glutamine in rice. Plant Cell Physiol 44:1396–1402. doi:10.1093/pcp/pcg169
Suenaga A, Moriya K, Sonoda Y, Ikeda A, von Wirén N, Hayakawa T, Yamaguchi J, Yamaya T (2003) Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants. Plant Cell Physiol 44:206–211. doi:10.1093/pcp/pcg017
Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S, Tanaka H (2006) Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J 47:969–976. doi:10.1111/j.1365-313X.2006.02836.x
von Wirén N, Lauter FR, Ninnemann O, Gillissen B, Walch-Liu P, Engels C, Jost W, Frommer WB (2000) Differential regulation of three functional ammonium transporter genes by nitrogen in root hairs and by light in leaves of tomato. Plant J 21:167–175. doi:10.1046/j.1365-313x.2000.00665.x
Wang MY, Siddiqi MY, Ruth TJ, Glass ADM (1993) Ammonium uptake by rice roots—kinetics of 13NH4 + influx across the plasmalemma. Plant Physiol 103:1259–1267. doi:10.1104/pp.103.4.1259
Yao S-G, Sonoda Y, Tsutsui T, Nakamura H, Ichikawa H, Ikeda A, Yamagushi J (2008) Promoter analysis of OsAMT1;2 and OsAMT1;3 implies their distinct roles in nitrogen utilization in rice. Breed Sci 58:201–207. doi:10.1270/jsbbs.58.201
Acknowledgments
This study was supported by the National Council for Scientific and Technological Development (Conselho Nacional de Desenvolvimento Científico e Tecnológico-CNPq), the Research Support Foundation of the State of Rio de Janeiro (Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro-FAPERJ), and the Coordination for the Improvement of Higher Education Personnel (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior-CAPES).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ferreira, L.M., de Souza, V.M., Tavares, O.C.H. et al. OsAMT1.3 expression alters rice ammonium uptake kinetics and root morphology. Plant Biotechnol Rep 9, 221–229 (2015). https://doi.org/10.1007/s11816-015-0359-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-015-0359-2