Abstract
Fe participates in several important reactions in plant metabolism. However, Fe homeostasis in plants is not completely understood, and molecular studies on Fe-excess stress are scarce. Rice (Oryza sativa L. ssp. indica) is largely cultivated in submerged conditions, where the extremely reductive environment can lead to severe Fe overload. In this work, we used representational difference analysis (RDA) to isolate sequences up-regulated in rice shoots after exposure to Fe-excess. We isolated 24 sequences which have putative functions in distinct cellular processes, such as transcription regulation (OsWRKY80), stress response (OsGAP1, DEAD-BOX RNA helicase), proteolysis (oryzain-α, rhomboid protein), photosynthesis (chlorophyll a/b binding protein), sugar metabolism (β glucosidase) and electron transport (NADH ubiquinone oxireductase). We show that the putative WRKY transcription factor OsWRKY80 is up-regulated in rice leaves, stems and roots after Fe-excess treatment. This up-regulation is also observed after dark-induced senescence and drought stress, indicating that OsWRKY80 could be a general stress-responsive gene. To our knowledge, this is the first report of an Fe-excess-induced transcription factor in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron (Fe) is a transition metal essential for virtually all living organisms. Because of its singular capacity to gain or lose electrons, it has a crucial participation in metabolic processes such as oxy-reductive reactions of photosynthesis, respiration and nitrogen assimilation. Fe is very abundant in the soil, but is normally chelated to organic matter in oxidized and insoluble forms, which are not readily available to absorption by plants [1]. Fe-deficiency leads to chlorosis, decreased abundance of photosynthetic proteins and senescence [2–4], being a yield-limiting factor with major implications for field crop production in many agricultural regions of the world. Under Fe-deficiency, plants make use of ingenious Fe-uptake strategies [5]. Rice (Oryza sativa) plants are especially susceptible to low Fe supply, differently from other cultivated grass species, such as barley [6].
On the other hand, Fe toxicity is very common in waterlogged and irrigated rice. High concentrations of ferrous (Fe2+) ions, found in soils under anaerobic and acidic conditions, lead to Fe toxicity due to excessive Fe uptake, which can result in yield reductions from 12% to 100%, depending on the soil characteristics and the rice cultivar [7]. Fe toxicity has been reported worldwide, including China, India, Indonesia, Thailand, Malaysia, Philippines, Sri Lanka, Burundi, Colombia, several countries of the West African region and Brazil [7]. The symptoms of Fe toxicity in rice are bronzing or yellowing of older leaves, Fe deposition in spots on young leaves, Fe plaque formation on roots, increased lipid peroxidation in shoots, decreased growth rate and grain production [8] and even plant death [7, 9]. Two distinct types of Fe toxicity are observed, directly or indirectly related to excessive Fe absorption. The direct Fe toxicity is caused by excessive concentrations of the metal in plant tissues, while the indirect toxicity is caused by Fe plaque formation in roots, impairing the absorption of other minerals [7].
Fe can be very dangerous when in excess inside the cell. Free Fe can act as a potent generator of reactive oxygen species (ROS), especially the hydroxyl radical, through the Fenton reaction [10]. This radical is extremely toxic to cell metabolism, leading to oxidation of biological macromolecules such as lipids, proteins and nucleic acids, causing membrane leakage and cell death [11, 12]. Physiological effects of Fe-excess in plants, including evidence of oxidative stress, were described in Nicotiana plumbaginifolia [13] and rice [8, 14]. Increased cytochrome b6/f content, leading to higher susceptibility of the photosystem II (PSII) to photoinhibition and consequent lower photosynthetic rates was described in pea plants [15] submitted to Fe-excess. Around 80% of plant Fe is located in chloroplasts [16], and therefore they are the first organelles to be damaged by Fe-excess. Ferritin and nicotianamine are pointed out as candidate molecules to perform Fe-buffering functions, being important for maintaining Fe homeostasis when plants are under Fe-excess stress [17–19]. Recently, our group brought up a new insight about Fe-excess responses, using resistant and susceptible cultivars to show that the capacity to keep Fe bound to low mass compounds (MW < 3 kDa) could be associated to Fe-excess tolerance [20]. Although Fe metabolism and homeostasis issues in plants have received increasing attention in the last years, especially mechanisms of Fe uptake by roots, the molecular aspects of Fe-excess responses remain largely unknown.
In this work, we used the cDNA-RDA technique to isolate sequences up-regulated by Fe-excess in shoots of rice plants. We found several sequences belonging to gene families already described as stress- and senescence-related. Among them, we describe OsWRKY80, a transcription factor (TF) of the WRKY superfamily. This family of proteins comprises structurally related members that share a common domain with about 60 amino acids, which include the signature heptapeptide WRKYGQK as well as a zinc-binding motif [21]. WRKY TFs are mainly found in plants, although their presence is not specific to this kingdom [22, 23]. These transcription factors bind to the conserved W box motif [24–26] and are related to functions as diverse as plant defense [27, 28], development of trichomes and seed coat [29], senescence [4, 25, 30] and response to abiotic stresses such as high salinity, heat, drought, water deficit and cold [31–34].
We demonstrate that OsWRKY80 is up-regulated by Fe-excess treatment in rice. Drought and dark-induced senescence also increase the steady state level of OsWRKY80 transcripts. This is the first attempt to uncover the Fe-excess response at the molecular level.
Materials and methods
Plant material
Rice seeds (Oryza sativa L. ssp. indica) from BR-IRGA 409 cultivar were provided by Instituto Rio-Grandense do Arroz (IRGA). This cultivar was previously shown to be susceptible to Fe toxicity [35, 36]. Seeds were germinated for 4 days in an incubator (28°C, first 2 days in the dark and last 2 days in the light) on Petri dishes lined with filter paper soaked in distilled water. After germination, plants were grown in hydroponic conditions using the nutrient solution described by Ogo et al. [37] until the four-leaf stage (~30 days). For Fe-excess treatments, 500 ppm of Fe was added to the nutrient solution. For generation of RDA representations, shoots were harvested after 3, 6 and 9 days and pooled. For semi-quantitative and quantitative RT-PCR analyses, plants were kept in the same conditions as above and harvested after 3, 6 and 9 days. All solutions were replaced every 3 days. Plants were grown in a room at 28 ± 1°C under white light with a photoperiod of 16/8 h light/dark cycle (irradiance of approximately 100 μmol m−2 s−1 at plant tops).
Abiotic stress and dark-induced senescence treatments
For the abiotic stress treatments, plants grown in the same conditions as above were submitted to control, drought (no water supply), cold (4°C) and high salinity (100 mM NaCl) treatments for 12 and 24 h, and all leaves from each plant were sampled. The senescence experiments were performed as described by Kusaba et al. [38] with minor modifications. Fully expanded leaves from plants at the four-leaf stage were excised, fragmented (0.5 cm2 of leaf area) and incubated in 3 mM MES (2,4-morpholino-ethane sulfonic acid) buffer, pH 5.8, at 27°C in the darkness. For phytohormone treatments, 50 μM abscisic acid (ABA) or 50 μM 6-benzylaminopurine (BAP) was added to the MES solution. Samples were harvested after 0, 1, 3, 5 and 7 days of treatment.
RNA extraction and cDNA synthesis
Each RNA extraction was performed with pooled material from at least three plants. Total RNA was extracted using Concert (Invitrogen® Life Technologies) reagent. RNA quality was assessed by denaturing agarose gel electrophoresis and spectrophotometric analysis. First-strand cDNA synthesis was performed after DNAse treatment with reverse transcriptase (M-MLV, Invitrogen® Life Technologies) using 1 μg of RNA. For the cDNA-RDA experiments, RNA samples from three-time points were pooled (300 ng from each sampling time: 3, 6 and 9 days, for each treatment) and cDNA was then prepared using the SMART PCR cDNA Synthesis Kit (Clontech® Laboratories), according to the manufacturer’s instructions. For the senescence experiments, eight leaf fragments were pooled for each RNA extraction. First-strand cDNA synthesis was performed as mentioned above. Three true biological replicates were used for each time point and treatment.
Representational difference analysis
The cDNA representational difference analysis (cDNA-RDA) was performed essentially as described by Pastorian et al. [39] and optimized by Dutra et al. [40]. The shoot cDNA pool from plants maintained in the control treatment was digested with the restriction enzyme Sau3AI (Amersham Pharmacia® Biotech). The resulting product was purified using the GFX kit (Amersham Pharmacia® Biotech) and ligated to adaptors (RBam12: GATCCTCGGTGA and RBam24: AGCACTCTCCAGCCTCTCTCACCGAG). To provide sufficient amounts of starting material for RDA, the cDNA was diluted and amplified using RBam24 as a primer (25 cycles, denaturation at 95°C for 45 s and annealing and extension at 70°C for 3 min). The final PCR product was purified using the GFX kit, resulting in driver cDNA populations from shoots of plants under the control treatment. To generate the Fe-excess tester population, the same procedure was performed with the cDNA pool from Fe-excess treated plant shoots, using JBam adaptors instead of RBam (JBam12: GATCCGTTCATG and JBam24: ACCGACGTCGACTATCCATGAACG).
In order to isolate Fe-excess-induced sequences, the first differential product (DP1) was obtained by mixing control driver and Fe-excess tester in a 50:1 ratio (5 μg driver: 100 ng tester), in a final volume of 5 μl, followed by hybridization at 67°C for 24 h and amplification by PCR with JBam24 as primer. To minimize cDNA linearly amplified, the product of the first seven cycles of amplification was used as template in a new reaction, diluting the unwanted sequences. After purification, DP1 products were digested with Sau3AI to remove JBam adaptors before ligation to NBam adaptors (NBam12: GATCCTCCCTCG and NBam24: AGGCAACTGTGCTATCCGAGGGAG). The second hybridization was performed at a ratio of 500:1 (5 μg driver:10 ng DP1). The second differential product (DP2) was obtained by the same procedure as for DP1, but using NBam24 as primer instead of JBam24.
Cloning and sequence analysis
The DP2 product was purified and cloned into pCR2.1-TOPO Vector (TOPO TA Cloning Kit, Invitrogen® Life Technologies) and transformed in Escherichia coli XL1 Blue competent cells by electroporation. Individual bacterial clones were grown in 96-well plates. Plasmid DNA was prepared from 192 selected clones and samples were sequenced in the ACTGene Laboratory (Centro de Biotecnologia, UFRGS, Porto Alegre, RS, Brazil) using the automatic sequencer ABI-PRISM 3100 Genetic Analyzer (Applied Biosystems®). The vector sequence was removed manually and the resulting sequences were then matched to rice full-length cDNA clones using the Rice Pipeline Database (http://cdna.01.dna.affrc.go.jp/PIPE/), a unification tool that dynamically integrates data from various rice databases [41].
Semi-quantitative and quantitative RT-PCR analysis
All RNA samples for expression analyses were generated from a new experiment, which was conducted under the same conditions as the experiment which originated the samples for cDNA-RDA. For semi-quantitative RT-PCR analysis, the synthesized first-strand cDNA from each time point was diluted five times and amplified using different sets of gene-specific primers. PCR amplifications started with an initial denaturation step of 5 min at 95°C, followed by cycling of 30 s at 95°C, 30 s at 60°C and 30 s at 72°C in a final volume of 50 μl. The optimal number of PCR cycles was determined for each gene to ensure that amplification occurred in the exponential phase. Obtained PCR products were subjected to agarose gel electrophoresis and stained with ethidium bromide. The signal intensity of the stained bands was photographed using a digital imager (Kodak DC120 Zoom Digital Camera) and analyzed using the Kodak Digital Science 1D Image Analysis Software, Version 3.0. Amplification of OsUBQ5 [42] transcripts was used as cDNA loading control.
For quantitative RT-PCR analysis, the synthesized first-strand cDNA from each time point was diluted 100 times. Reactions were carried out in an Applied Biosystems® 7500 real-time cycler. PCR amplifications included an initial denaturation step of 5 min at 94°C, followed by 40 cycles of 10 s at 94°C, 15 s at 60°C, 15 s at 72°C and 35 s at 60°C, after which samples were held for 2 min at 40°C for annealing and then heated from 55 to 99°C with a ramp of 0.1°C/s to produce the denaturing curve of the amplified products. qRT-PCRs were carried out in 20 μl final volume composed of 10 μl of each reverse transcription sample diluted 100 times, 2 μl of 10× PCR buffer, 1.2 μl of 50 mM MgCl2, 0.1 μl of 10 mM dNTPs, 0.4 μl of 10 μM primer pairs, 4.25 μl of water, 2.0 μl of SYBR green (1:10,000 Molecular Probe), and 0.05 μl of Platinum Taq DNA polymerase (5 U/μl) (Invitrogen®). Obtained data were analyzed by the comparative CT (threshold cycle) method [43]. The PCR efficiency from the exponential phase (Eff) was calculated for each individual amplification plot using the LinReg software [44]. In each plate, the average of PCR efficiency for each amplicon was determined and used in further calculations. C t values for all genes were normalized to the C t value of UBQ5 [42] using the equation \( Q_{{0\,{\text{target\,gene}}}} /Q_{{0\,{\text{UBQ5}}}} = [ ( {\text{Eff}}_{\text{UBQ5}} )^{{C_{\text{t}} {\text{UBQ5}}}}/({\text{Eff}}_{\text{target\,gene}} )^{{C_{\text{t}} \,{\text{target\,gene}}}} ] \) [43], where Q 0 is the initial amount of transcripts. All analyses were performed using three biological replicates for each time point and treatment.
Primers for semi-quantitative and quantitative RT-PCR were designed to amplify 100–200 bp of the 3′-UTR of the genes and to have similar T m values (60 ± 2°C). Supplementary Table 1 contains the sequences of all gene-specific primers used in this work.
Statistical analyses
When appropriate, data were subjected to ANOVA and means were compared by the Duncan or Student’s t test using the SPSS Base 12.0 for Windows (SPSS Inc., Chicago, IL, USA).
Results and discussion
Isolation of Fe-excess responsive cDNA clones by RDA
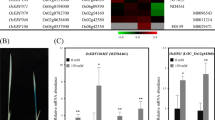
After 9 days of Fe-excess treatment, plants showed clear bronzing in leaves, a symptom of Fe-excess toxicity (Fig. 1a). Under these conditions, our group has already characterized the Fe-excess response of BR-IRGA 409 plants [45]: this work shows that Fe over accumulates in shoots under Fe-excess when compared to control, showing 2- to 3-fold higher Fe concentrations after 3 days and 8- to 10-fold after 9 days of treatment. It is also demonstrated that ferritin transcripts accumulate in response to Fe-excess treatment [45]. As we were interested in sequences that are relevant to the Fe-excess response, we performed the cDNA-RDA using two cDNA pools generated from RNAs of shoots extracted after 3, 6 and 9 days of control or Fe-excess treatments. Figure 1b shows all cDNA representations and differential products obtained. The first hybridization was performed with a 50:1 ratio of control: Fe-excess cDNAs. After PCR amplification using primers specific for the Fe-excess cDNAs, the DP1 was obtained. Slight differences could be noted when comparing Fe-excess cDNA and DP1 cDNAs (Fig. 1b, lanes 1 and 3), although no discrete bands were observed. The second hybridization was then performed with a 500:1 ratio using control: DP1 cDNAs. PCR amplification using primers specific for DP1 cDNAs was performed in order to obtain the DP2. Discrete bands could be observed (Fig. 1b, lane 4), indicating that the DP2 was correctly subtracted and enriched for differentially expressed sequences.
Fe-excess stress symptoms and differential products obtained in RDA experiments. a Plants after 9 days of exposure to control and Fe-excess treatments. Leaves from three different plants are shown for each treatment. Bar indicates 1 cm. b Fe-excess rice shoot cDNAs were subtracted from control condition rice shoot cDNAs by representational difference analysis after the first (DP1) and second (DP2) rounds of hybridization. DP1 and DP2 were fractionated by electrophoresis in 1% agarose gel and stained with ethidium bromide. M 100 bp DNA Ladder, 1 tester Fe-excess condition, 2 driver control condition, 3 differential product 1 (1:50), 4 differential product 2 (1:500)
Randomly chosen 192 clones from DP2 were isolated and sequenced. The sequence analysis showed 24 unique cDNAs. These cDNAs were functionally categorized according to the description of the most similar clone found in the Rice Pipeline Database. Four cDNAs could not have a putative function assigned, although they have matched expressed sequences in the database. Supplementary Table 2 shows the description of the cDNAs, functional categorization and best E-value found for each different sequence.
Up-regulated genes
The genes found are included in functional categories related to transcription regulation, stress response, proteolysis and photosynthesis (Supplementary Table 2). Those which are members of gene families already described as related to stress response or senescence were indicated in the last column of Supplementary Table 2.
To ensure that our experimental conditions were indeed leading to Fe-excess, we used a primer set that amplifies both rice ferritin genes (OsFER1 and OsFER2) in quantitative RT-PCR analysis. Induction of ferritin expression under Fe-excess is well documented in plants [46], including rice [18]. OsFER1 and OsFER2 genes are known to be very similar and thus their differential detection is difficult to achieve with qPCR [18]. Figure 2a clearly demonstrates that OsFER transcripts were induced by 1.5-fold after 3 days of Fe-excess treatment, further increasing to 3-fold after 6 and 9 days.
Gene expression analysis by quantitative RT-PCR. Shoots from plants under control condition (CC) or Fe-excess treatment (Fe+) were harvested after 3, 6 and 9 days. a Expression level of OsFER. b Expression level of oryzain-α. c Expression level of C2 protein (OsGAP1). d Expression level of OsWRKY80. Expression levels were calculated relative to OsUBQ5 expression. The expression levels are presented as fold-induction relative to control. Mean ± SE of three biological replicates are shown. Statistical differences by the Student’s t test are shown by one (P ≤ 0.05) or two asterisks (P ≤ 0.01)
To confirm the RDA results, we also performed quantitative RT-PCR (qPCR) analysis of three genes (OsGAP1, oryzain-α and OsWRKY80) with cDNAs from 3, 6 and 9 days of control or Fe-excess treatments separately (instead of the pooled samples used in the cDNA-RDA). Both oryzain-α and OsGAP1 were already characterized as related to stress in rice [47, 48] and OsWRKY80 was previously described as responsive to pathogen attack [49].
Expression of oryzain-α increased by 2.4- and 4.2-fold after 6 and 9 days of treatment, respectively (Fig. 2b). Oryzain-α is a cysteine proteinase first described as up-regulated during germination [50]. This gene was recently found to be up-regulated by various stresses in rice leaves, and it was suggested to be involved in the final steps of leaf senescence [47]. Proteinases are also related to senescence in Gladiolus flowers [51] and in nodule senescence of the legume Astragalus sinicus [52]. A cysteine proteinase was also suggested be involved in autophagy in rice [53], a process related to leaf senescence [54]. Around 7% of the 2,500 genes expressed in Arabidopsis senescent leaves are proteases [55], and cysteine-proteases constitute one of the most representative classes [56].
The C2 domain protein OsGAP1 was also up-regulated in all time points tested (Fig. 2c). Transcripts were 6-fold increased by the third day of Fe-excess treatment, and were 1.9- and 3.3-fold higher when compared to control plants in the sixth and ninth days (Fig. 2c). Recently, this gene product was described as coding a C2-containing GTPase-activating protein, named OsGAP1 [48]. OsGAP1 is up-regulated by wounding and interacts with a G-protein called OsYchF1, suggesting a role in signal transduction. OsGAP1 confers protection against Xanthomonas oryzae and Pseudomonas syringae when overexpressed in rice, indicating its involvement in defense responses [48]. Moreover, a C2 domain-containing protein has been described as responsive to cadmium and copper toxicity and is associated with leaf senescence in H. vulgare [57].
The amount of OsWRKY80 transcripts increased only after 6 days of treatment, further accumulating after 9 days, by 1.6- and 3-fold, respectively (Fig. 2d). WRKY proteins are a super family of TFs that are mainly found in higher plants, with 72 members in Arabidopsis [58] and up to 103 members in rice [59]. WRKY proteins were already described as involved in responses to stress such as high salinity in rice shoots [33], water deficit in grapevine [32], sucrose starvation in rice suspension cells [60], cold and drought in barley [31], and senescence in Arabidopsis [25, 61, 62], rice [4] and wheat [30], fungus infection in ramie (Boehmeria nivea) [63], as well as in other plant stresses ([59] and references therein). OsWRKY80 was first named by Xie et al. [23], although other authors named the same rice locus as OsWRKY13 [64]. Since OsWRKY13 from Xie et al. [23] nomenclature have already been characterized [65, 66], we decided to follow their nomenclature [23], avoiding ambiguities.
To further confirm the up-regulation of the genes present in our RDA library, we conducted semi-quantitative RT-PCR analysis of other five genes with cDNAs from 3, 6 and 9 days of control or Fe-excess treatments. The intensity of each band was obtained and used to compare the amount of product after PCR. All tested genes showed increased expression after Fe-excess treatment (DEAD-BOX RNA helicase, rhomboid protein, chlorophyll a/b-binding protein, β glucosidase and NADH ubiquinone oxireductase), demonstrating the quality of our subtractive library (Fig. 3).
Semi-quantitative RT–PCR expression analysis of selected genes identified by RDA in shoots of rice plants exposed or not to Fe-excess for 3, 6 and 9 days. Band intensity of each PCR product was normalized in relation to the optical density obtained for the OsUBQ5 gene. Numbers below each lane represent a percentage in relation to the highest normalized optical density. Numbers in parenthesis shown after gene names indicate the number of cycles used to detect the PCR product in the exponential phase. Control: control condition; +Fe: Fe-excess condition. Agarose gels (1%) were stained with ethidium bromide
OsWRKY80 expression in different plant organs
Among the sequences identified by our RDA experiment, we were mostly interested in the OsWRKY80 gene, possibly the first rice transcription factor found to be involved with Fe-excess responses. As the cDNA-RDA was performed with total shoots, we decided to further characterize the mRNA expression pattern of OsWRKY80 in different plant organs under Fe-excess. Rice plants at the four-leaf stage (about 30 days) were submitted to control or Fe-excess treatment for 6 days and leaves, stems and roots were harvested. The OsWRKY80 gene was expressed in all the analyzed plant organs under control conditions (Fig. 4), as already observed by Ramamoorthy et al. [59] using semi-quantitative RT-PCR. The relative expression levels of OsWRKY80 in leaves under control treatment is higher, with average expression levels in culms and roots corresponding to 47.9% an 53.4% of the expression in leaves. Fe-excess treatment resulted in increased OsWRKY80 expression in all organs tested (Fig. 4). Therefore, OsWRKY80 response to Fe-excess occurs throughout the plant.
OsWRKY80 expression in different rice plant organs. Samples were harvested after 6 days under control condition (CC) or Fe-excess treatment (Fe+) for quantitative RT-PCR. The expression levels are presented as fold-induction relative to control. Means ± SE of three biological replicates are shown. Statistical differences in Student’s t test are shown by one (P ≤ 0.05) or two asterisks (P ≤ 0.01)
Drought stress up-regulates OsWRKY80
We submitted rice plants to three different abiotic stresses: drought (no water supply), cold (4°C) and high salinity (100 mM NaCl). Leaves were harvested after 12 and 24 h and OsWRKY80 expression was addressed. After 12 h, a nearly 2-fold increase was observed in leaves of drought-stressed plants, while no change was observed in cold- and high salinity-treated plants (Fig. 5). The increase in OsWRKY80 expression is even higher after 24 h, reaching 4.4-fold when comparing drought-treated expression level to control level (Fig. 5). Therefore, OsWRKY80 seems to be up-regulated by drought stress in rice leaves. In cold-treated plants, there was no change in OsWRKY80 mRNA accumulation in leaves, while high salinity-treated plants showed a slight decrease in this organ (Fig. 5).
OsWRKY80 expression under drought (no water supply), cold (4°C) and high salinity (100 μM NaCl) stresses for 12 and 24 h. Samples from three plants were harvested after 6 days of Fe-excess or control treatment for quantitative RT-PCR. The expression levels are presented as fold-induction relative to control. Means ± SE of three biological replicates are shown. Statistical differences in Student’s t test in comparison to control are shown by one (P ≤ 0.05) or two asterisks (P ≤ 0.01)
Dark-induced senescence induces OsWRKY80 expression
To test whether OsWRKY80 is up-regulated during senescence in rice leaves, we evaluated the expression of this gene under dark-induced senescence and compared it to OsSGR [67] expression, a senescence marker. Detached leaves were submitted to dark under three conditions: no hormonal treatment, ABA-treated and BAP-treated [38]. ABA is a senescence inductor, while BAP is a senescence inhibitor [38]. Under normal dark-induced senescence, OsWRKY80 up-regulation was observed after 7 days, when the leaves showed a well-established senescence process (Fig. 6a, c). Under ABA-enhanced dark-induced senescence, up-regulation was observed earlier, after 3 days of treatment, with a further increase after 5 days (Fig. 6a). In BAP-treated leaves, these increases were not observed, but only a slight OsWRKY80 up-regulation occurred after 5 days (Fig. 6a). OsSGR was used as a control for all treatments, as it is known to be up-regulated by dark-induced senescence, with ABA acting synergistically to up-regulate its expression and BAP antagonistically to delay its up-regulation during the senescence process [67]. Dark induction of OsSGR expression started after 3 days of treatment, and expression increased with time. ABA treatment was able to anticipate OsSGR induction to the first day of treatment, while the BAP treatment partially delayed its increase in expression (Fig. 6b). Therefore, we could observe that OsWRKY80 is up-regulated by dark-induced senescence and this up-regulation is anticipated by ABA and delayed by BAP treatments, a pattern similar to OsSGR regulation. Interestingly, both dark and dark + ABA treatments resulted in earlier increase of expression (in relation to the control treatment) of the OsSGR gene than of the OsWRKY80 gene. It is possible that the onset of senescence, with the consequent Fe release during the disorganization of the photosynthetic apparatus, may have a role in the induction of OsWRKY80 expression.
OsWRKY80 and OsSGR mRNA expression during dark-induced senescence. Leaves were kept in the dark, dark + 50 μM ABA or dark + 50 μM BAP. Quantitative RT-PCR was carried out with eight leaf fragments per sample. Means ± SE of three biological replicates are shown. a Expression level of OsWRKY80, relative to OsUBQ5. b Expression level of OsSGR, relative to OsUBQ5. Different letters indicate that means are different by the Duncan test (P ≤ 0.05). c Detached leaves from each treatment and time of harvesting. ABA abscisic acid, BAP 6-benzylaminopurine
Possible OsWRKY80 functions
In the last years, Fe regulation of genes involved in its uptake and storage has been widely studied, but the focus is mainly on the identification of regulatory proteins that function in the Fe-deficiency-signaling pathway (for a review, see [68]). However, genes involved in Fe-excess responsive pathways are still largely unknown. Recently, Duc et al. [69] described the nuclear factor time for coffee (TIC) as a regulator of Fe-excess responsive genes in Arabidopsis. Our findings indicate that OsWRKY80 is a transcription factor regulated by Fe-excess, and thus one of the first candidate genes to act as regulator of Fe-excess responses in plants.
Transcription factors related to abiotic and biotic stresses have been widely described [70–74]. Abiotic and biotic stresses can trigger and affect senescence to various extents [75], and the WRKY family is one of the two most representative transcription factor groups which are up-regulated during senescence in Arabidopsis leaves [76]. We showed that OsWRKY80 is regulated by drought stress and dark-induced senescence. These results indicate that OsWRKY80 could be classified as a senescence-associated gene (SAG). Several WRKY genes have been described as SAGs [25, 61, 62] and WRKY proteins were also implied in drought response in rice [34], barley [31], soybean [70] and in the desert plant creosote bush (Larrea tridentata), in which LtWRKY21 acts through an ABA-dependent pathway [77].
The up-regulation observed in dark-induced senescence is inhibited by BAP treatment and is synergistically increased by the addition of ABA, demonstrating a regulation pattern typical of a SAG [67, 72]. The increased expression in leaves under drought stress and by ABA treatment further suggests that OsWRKY80 up-regulation is mediated by this hormone. ABA is a classical stress regulator related to abiotic stresses as drought, cold and salinity. In rice seedlings, ABA was shown to induce stress-related proteins [78], and stress-responsive transcription factors from the WRKY family are regulated by ABA in rice [23], barley [79], Arabidopsis [80], Brassica napus [81] and the resurrection plant Boea hygrometrica [82]. Moreover, the ABA pathway was suggested to act as a linker between transcription factors responsive to abiotic stress and senescence in rice [72] and soybean [73].
Ramamoorthy et al. [59] analyzed the expression pattern of 103 WRKY mRNAs in different tissues and in response to various stresses. Expression of OsWRKY80 (referred as 09g30400) was not altered when plants were submitted to cold, drought and high salinity. From these three stress conditions, only drought was able to increase OsWRKY80 expression in our experiments. Ramamoorthy et al. [59] analyzed plants under drought stress for as long as 3 h, while our experiments showed significant increase of OsWRKY80 expression only after a longer time (12 h), becoming much more evident after 24 h. Treatments with ABA, gibberellic acid (GA3), auxin (IAA), methyl jasmonate (MeJA) and salicylic acid (SA) for up to 16 h were considered unable to up-regulate the expression of OsWRKY80 mRNA [59]. Among these, we tested ABA, which in combination with dark treatment was able to induce earlier expression of OsWRKY80. The apparent discrepancy can be explained by the different techniques used: semi-quantitative RT-PCR [59] and quantitative RT-PCR (our work), which allows easier detection of changes in mRNA expression levels.
WRKY transcription factors are mainly involved in plant innate responses to biotic stress, such as pathogen attack and through signaling of the endogenous molecule salicylic acid (SA) [27, 28]. SA is required to induce systemic acquired resistance (SAR), in which pathogen-related proteins accumulate [83], and is normally antagonized by jasmonic acid (JA). An Arabidopsis WRKY gene, AtWRKY70, was described as a convergence point determining the balance between SA- and JA-dependent defense pathways, and many others were involved in biotic stress response (for a review, see [28]). Although Ramamoorthy et al. [59] did not observe any alteration in the expression of OsWRKY80 by SA or JA hormones, Guo et al. [49] showed that OsWRKY80 (referred as OsiWRKY) mRNA is induced upon Xanthomonas oryzae attack and in response to wounding, although in his experiments the control plants also exhibited a slight up-regulation. Fe has already been proposed to be a mediator of pathogen attack signaling in wheat [84]. After Blumeria graminis attack, Fe is secreted to the apoplast and induces an oxidative burst mediated by H2O2. The cytoplasmatic Fe-deficiency in concert with the production of H2O2 induce pathogenesis-related genes [84]. Fe-excess could be partially mimicking this high apoplastic Fe concentration when inducing OsWRKY80. Once more data are available, it may become possible to elaborate a model where OsWRKY80 has a role in both abiotic and biotic stresses as well as in senescence.
In summary, we described a new Fe-excess up-regulated transcription factor from the WRKY family, which is also regulated by dark-induced senescence and drought stress, responses that are possibly mediated by ABA. The OsWRKY80 is the first transcription factor shown to be induced by Fe-excess in plants.
Abbreviations
- ABA:
-
Abscisic acid
- BAP:
-
6-Benzylaminopurine
- CAB:
-
Chlorophyll a/b binding protein
- DP:
-
Differential product
- LRR:
-
Leucine-rich repeat receptor
- MES:
-
2,4-Morpholino-ethane sulfonic acid
- PSII:
-
Photosystem II
- RDA:
-
Representational difference analysis
- ROS:
-
Reactive oxygen species
- RuBisCO:
-
Ribulose 1,5-bisphosphate carboxylase/oxygenase
- SAG:
-
Senescence-associated gene
- SGR:
-
Staygreen
- TF:
-
Transcription factor
References
Guerinot ML, Yi Y (1994) Iron: nutritious, noxious, and not readily available. Plant Physiol 104:815–820
Spiller SC, Terry N (1980) Limiting factors of photosynthesis II: iron stress diminishes photochemical capacity by reducing the number of photosynthetic units. Plant Physiol 65:121–125
Sperotto RA, Ricachenevsky FK, Fett JP (2007) Iron deficiency in rice shoots: identification of novel induced genes using RDA and possible relation to leaf senescence. Plant Cell Rep 26:1399–1411
Sperotto RA, Boff T, Duarte GL, Fett JP (2008) Increased senescence-associated gene expression and lipid peroxidation induced by iron deficiency in rice roots. Plant Cell Rep 27:183–195
Kim SA, Guerinot ML (2007) Mining iron: iron uptake and transport in plants. FEBS Lett 581:2273–2280
Takahashi M, Nakanishi H, Kawasaki S, Nishizawa NK, Mori S (2001) Enhanced tolerance of rice to low iron availability in alkaline soils using barley nicotianamine aminotransferase genes. Nature Biotechnol 19:466–469
Sahrawat KL (2004) Iron toxicity in wetland rice and the role of other nutrients. J Plant Nutr 27:1471–1504
Fang WC, Wang JW, Lin CC, Kao CH (2001) Iron induction of lipid peroxidation and effects on antioxidative enzymes activities in rice leaves. Plant Growth Reg 35:75–80
Ponnamperuma FN, Bradfield R, Peech M (1955) Physiological disease of rice attributable to iron toxicity. Nature 175:265
Becana M, Moran JF, Iturbe-Ormaetxe I (1998) Iron dependent oxygen free radical generation in plants subjected in environmental stress: toxicity and antioxidant protection. Plant Soil 201:137–147
Cadenas E (1989) Biochemistry of oxygen toxicity. Ann Rev Biochem 58:79–110
Halliwell B, Gutteridge JMC (1992) Biologically relevant metal ion-dependent hydroxyl radical generation. FEBS Lett 307:108–112
Kampfenkel K, Van Montagu M, Inzé D (1995) Effects of iron-excess on Nicotiana plumbaginifolia. Implications to oxidative stress. Plant Physiol 107:725–735
Yamauchi M, Peng XX (1995) Iron toxicity and stress-induced ethylene production in rice leaves. Plant Soil 173:21–28
Suh H, Kim CS, Lee J, Jung J (2002) Photodynamic effect of iron-excess on photosystem II function in pea plants. Photochem Photobiol 75:513–518
Mengel K, Kirkby EA (1982) Iron. In: Mengel K, Kirkby EA (eds) Principles of plant nutrition, 3rd edn. Der Bund Press, Bern, Germany, pp 473–489
Ravet K, Touraine B, Boucherez J, Briat JF, Gaymard F, Cellier F (2009) Ferritins control interaction between iron homeostasis and oxidative stress in Arabidopsis. Plant J 57:400–412
Stein RJ, Ricachenevsky FK, Fett JP (2009) Differential regulation of the two rice ferritin genes (OsFER1 and OsFER2). Plant Sci 177:563–569
Pich A, Manteuffel R, Hillmer S, Scholz G, Schmidt W (2001) Fe homeostasis in plant cells: does nicotianamine play multiple roles in the regulation of cytoplasmic Fe concentration? Planta 213:967–976
Stein RJ, Duarte GL, Spohr MG, Lopes SIG, Fett JP (2009) Distinct physiological responses of two rice cultivars subjected to iron toxicity under field conditions. Ann Appl Biol 154:269–277
Duan MR, Nan J, Liang YH, Mao P, Lu L, Li L, Wei C, Lai L, Li Y, Su XD (2007) DNA binding mechanism revealed by high resolution crystal structure of Arabidopsis thaliana WRKY1 protein. Nucl Acids Res 35:1145–1154
Ulker B, Somssich IE (2004) WRKY transcription factors: from DNA binding towards biological function. Curr Opin Plant Biol 7:491–498
Xie Z, Zhang ZL, Zou X, Huang J, Ruas P, Thompson D, Shen QJ (2005) Annotations and functional analyses of the rice WRKY gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol 137:176–189
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206
Miao Y, Laun T, Zimmermann P, Zentgraf U (2004) Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol Biol 55:853–867
Ciolkowski I, Wanke D, Birkenbihl RP, Somssich IE (2008) Studies on DNA-binding selectivity of WRKY transcription factors lend structural clues into WRKY-domain function. Plant Mol Biol 68:81–92
Eulgem T, Somssich IE (2007) Networks of WRKY transcription factors in defense signaling. Curr Opin Plant Biol 10:366–371
Pandey SP, Somssich IE (2009) The role of WRKY transcription factors in plant immunity. Plant Physiol 150:1648–1655
Johnson CS, Kolevski B, Smyth DR (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375
Gregersen PL, Holm PB (2007) Transcriptome analysis of senescence in the flag leaf of wheat (Triticum aestivum L.). Plant Biotech J 5:192–206
Marè C, Mazzucotelli E, Crosatti C, Francia E, Stanca AM, Cattivelli L (2004) Hv-WRKY38: a new transcription factor involved in cold- and drought-response in barley. Plant Mol Biol 55:399–416
Cramer GR, Ergul A, Grimplet J, Tillett RL, Tattersall EA, Bohlman MC, Vincent D, Sonderegger J, Evans J, Osborne C, Quilici D, Schlauch KA, Schooley DA, Cushman JC (2007) Water and salinity stress in grapevines: early and late changes in transcript and metabolite profiles. Funct Integrat Genom 7:111–134
Zhou J, Wang X, Jiao Y, Qin Y, Liu X, He K, Chen C, Ma L, Wang J, Xiong L, Zhang Q, Fan L, Deng XW (2007) Global genome expression analysis of rice in response to drought and high-salinity stresses in shoot, flag leaf, and panicle. Plant Mol Biol 63:591–608
Wu X, Shiroto Y, Kishitani S, Ito Y, Toriyama K (2009) Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep 28:21–30
Nava G, Bohnen H (2002) Oxidação de ferro em raízes de dois cultivares de arroz em solução de solo inundado. R Bras Cienc Solo 26:325–332
Silveira VC, Oliveira AP, Sperotto RA, Espindola LS, Amaral L, Dias JF, Cunha JB, Fett JP (2007) Influence of iron on mineral status of two rice (Oryza sativa L.) cultivars. Braz J Plant Physiol 19:127–139
Ogo Y, Itai RN, Nakanishi H, Inoue H, Kobayashi T, Suzuki M, Takahashi M, Mori S, Nishizawa NK (2006) Isolation and characterization of IRO2, a novel iron-regulated bHLH transcription factor in graminaceous plants. J Exp Bot 57:2867–2878
Kusaba M, Ito H, Morita R, Iida S, Sato Y, Fujimoto M, Kawasaki S, Tanaka R, Hirochika H, Nishimura M, Tanaka A (2007) Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence. Plant Cell 19:1362–1375
Pastorian K, Hawell L III, Byus CV (2000) Optimization of cDNA representational difference analysis for the identification of differentially expressed mRNAs. Anal Biochem 283:89–98
Dutra V, Nakazato L, Broetto L, Schrank IS, Vainstein MH, Schrank A (2004) Application of representational difference analysis to identify sequence tags expressed by Metarhizium anisopliae during the infection process of the tick Boophilus microplus cuticle. Res Microbiol 155:245–251
Yazaki N, Kojima K, Suzuki K, Kishimoto N, Kikuchi S (2004) The Rice PIPELINE: a unification tool for plant functional genomics. Nucl Acids Res 32:383–387
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345:646–651
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408
Ramakers C, Ruijter JM, Deprez RH, Moorman AF (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66
Silveira VC, Fadanelli C, Sperotto RA, Stein RJ, Basso LA, Santos DS, Vaz I Jr, Dias JF, Fett JF (2009) Role of ferritin in the rice tolerance to iron overload. Sci Agric 66:549–555
Briat JF, Cellier F, Gaymard F (2006) Ferritins and iron accumulation in plant tissues. In: Barton LL, Abadia J (eds) Iron nutrition in plants and rhyzospheric microorganisms. Springer, Verlag, pp 341–357
Fu Y, Zhao W, Peng Y (2007) Induced expression of oryzain alpha gene encoding a cysteine proteinase under stress conditions. J Plant Res 120:465–469
Cheung MY, Zeng NY, Tong SW, Li WY, Xue Y, Zhao KJ, Wang C, Zhang Q, Fu Y, Sun Z, Sun SS, Lam HM (2008) Constitutive expression of a rice GTPase-activating protein induces defense responses. New Phytol 179:530–545
Guo ZJ, Kan YC, Chen XJ, Li DB, Wang DW (2004) Characterization of a rice WRKY gene whose expression is induced upon pathogen attack and mechanical wounding. Acta Botanica Sinica 46:955–964
Watanabe H, Abe K, Emori Y, Hosoyama H, Arai S (1991) Molecular cloning and gibberellin-induced expression of multiple cysteine proteinases of rice seeds (oryzains). J Biol Chem 266:16897–16902
Azeez A, Sane AP, Bhatnagar D, Nath P (2007) Enhanced expression of serine proteases during floral senescence in Gladiolus. Phytochem 68:1352–1357
Li Y, Zhou L, Li Y, Chen D, Tan X, Lei L, Zhou J (2008) A nodule-specific plant cysteine proteinase, AsNODF32, is involved in nodule senescence and nitrogen fixation activity of the green manure legume Astragalus sinicus. New Phytol 180:185–192
Su W, Ma H, Liu C, Wu J, Yang J (2006) Identification and characterization of two rice autophagy associated genes, OsAtg8 and OsAtg4. Mol Biol Rep 33:73–78
Xiong Y, Contento AL, Bassham DC (2005) AtATG18a is required for the formation of autophagosomes during nutrient stress and senescence in Arabidopsis thaliana. Plant J 42:535–546
Gepstein S (2004) Leaf senescence—not just a ‘wear and tear’ phenomenon. Genome Biol 5:212
Guo G, Cai Z, Gan S (2004) Transcriptome of Arabidopsis leaf senescence. Plant Cell Environm 27:521–547
Ouelhadj A, Kuschk P, Humbeck K (2006) Heavy metal stress and leaf senescence induce the barley gene HvC2d1 encoding a calcium-dependent novel C2 domain-like protein. New Phytol 170:261–273
Dong J, Chen C, Chen Z (2003) Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol Biol 51:21–37
Ramamoorthy R, Jiang SY, Kumar N, Venkatesh PN, Ramachandran S (2008) A comprehensive transcriptional profiling of the WRKY gene family in rice under various abiotic and phytohormone treatments. Plant Cell Physiol 49:865–879
Wang HJ, Wan AR, Hsu CM, Lee KW, Yu SM, Jauh GY (2007) Transcriptomic adaptations in rice suspension cells under sucrose starvation. Plant Mol Biol 63:441–463
Robatzek S, Somssich IE (2002) Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev 16:1139–1149
Ulker B, Shahid Mukhtar M, Somssich IE (2007) The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta 226:125–137
Wang XX, Wang B, Liu LJ, Cui XP, Yang JY, Wang H, Jiang H, Luo BB, Long Z, Dou WX, Zhang N, Peng DX (2009) Isolation of high quality RNA and construction of a suppression subtractive hybridization library from ramie (Boehmeria nivea L. Gaud.). Mol Biol Rep 37:2099–2103
Berri S, Abbruscato P, Faivre-Rampant O, Brasileiro AC, Fumasoni I, Satoh K, Kikuchi S, Mizzi L, Morandini P, Pè ME, Piffanelli P (2009) Characterization of WRKY co-regulatory networks in rice and Arabidopsis. BMC Plant Biol 9:120
Qiu D, Xiao J, Ding X, Xiong M, Cai M, Cao Y, Li X, Xu C, Wang S (2007) OsWRKY13 mediates rice disease resistance by regulating defense-related genes in salicylate- and jasmonate-dependent signaling. Mol Plant Microbe Interact 20:492–499
Qiu D, Xiao J, Xie W, Cheng H, Li X, Wang S (2009) Exploring transcriptional signalling mediated by OsWRKY13, a potential regulator of multiple physiological processes in rice. BMC Plant Biol 9:74
Park SY, Yu JW, Park JS, Li J, Yoo SC, Lee NY, Lee SK, Jeong SW, Seo HS, Koh HJ, Jeon JS, Park YI, Paek NC (2007) The senescence-induced staygreen protein regulates chlorophyll degradation. Plant Cell 19:1649–1664
Walker EL, Connolly EL (2008) Time to pump iron: iron-deficiency-signaling mechanisms of higher plants. Curr Opin Plant Biol 11:530–535
Duc C, Cellier F, Lobreaux S, Briat JF, Gaymard F (2009) Regulation of iron homeostasis in Arabidopsis thaliana by the clock regulator time for coffee. J Biol Chem 284:36271–36281
Zhou QY, Tian AG, Zou HF, Xie ZM, Lei G, Huang J, Wang CM, Wang HW, Zhang JS, Chen SY (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol J 6:486–503
Yang Y, Wu J, Zhu K, Liu L, Chen F, Yu D (2009) Identification and characterization of two chrysanthemum (Dendronthema × moriforlium) DREB genes, belonging to the AP2/EREBP family. Mol Biol Rep 36:71–81
Sperotto RA, Ricachenevsky FK, Duarte GL, Boff T, Lopes KL, Sperb ER, Grusak MA, Fett JP (2009) Identification of up-regulated genes in flag leaves during rice grain filling and characterization of OsNAC5, a new ABA-dependent transcription factor. Planta 230:985–1002
Pinheiro GL, Marques CS, Costa MD, Reis PA, Alves MS, Carvalho CM, Fietto LG, Fontes EP (2009) Complete inventory of soybean NAC transcription factors: sequence conservation and expression analysis uncover their distinct roles in stress response. Gene 444:10–23
Agarwal P, Agarwal PK, Joshi AJ, Sopory SK, Reddy MK (2010) Overexpression of PgDREB2A transcription factor enhances abiotic stress tolerance and activates downstream stress-responsive genes. Mol Biol Rep 37:1125–1135
Buchanan-Wollaston V, Earl S, Harrison E, Mathas E, Navabpour S, Page T, Pink D (2003) The molecular analysis of leaf senescence–a genomic approach. Plant Biotechnol J 1:3–22
Balazadeh S, Riaño-Pachón DM, Mueller-Roeber B (2008) Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Biol 10(Suppl 1):63–75
Zou X, Seemann JR, Neuman D, Shen QJ (2004) A WRKY gene from creosote bush encodes an activator of the abscisic acid signaling pathway. J Biol Chem 279:55770–55779
Rakwal R, Komatsu S (2004) Abscisic acid promoted changes in the protein profiles of rice seedling by proteome analysis. Mol Biol Rep 31:217–230
Xie Z, Zhang ZL, Hanzlik S, Cook E, Shen QJ (2007) Salicylic acid inhibits gibberellin-induced alpha-amylase expression and seed germination via a pathway involving an abscisic-acid-inducible WRKY gene. Plant Mol Biol 64:293–303
Jiang W, Yu D (2009) Arabidopsis WRKY2 transcription factor mediates seed germination and postgermination arrest of development by abscisic acid. BMC Plant Biol 9:96
Yang B, Jiang Y, Rahman MH, Deyholos MK, Kav NN (2009) Identification and expression analysis of WRKY transcription factor genes in canola (Brassica napus L.) in response to fungal pathogens and hormone treatments. BMC Plant Biol 9:68
Wang Z, Zhu Y, Wang L, Liu X, Liu Y, Phillips J, Deng X (2009) A WRKY transcription factor participates in dehydration tolerance in Boea hygrometrica by binding to the W-box elements of the galactinol synthase (BhGolS1) promoter. Planta 230:1155–1166
Durrant WE, Dong X (2004) Systemic acquired resistance. Annu Rev Phytopathol 42:185–209
Liu G, Greenshields DL, Sammynaiken R, Hirji RN, Selvaraj G, Wei Y (2007) Targeted alterations in iron homeostasis underlie plant defense responses. J Cell Sci 15:596–605
Acknowledgments
This research was supported by CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, Brazil, grant 481131/2004-3 to JPF and scholarships to FKR and PKM) and HarvestPlus (Agreement number 6005-05). RAS was recipient of a scholarship from CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Brazil). The authors thank IRGA (Instituto Rio-Grandense do Arroz) for technical support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Supplementary Table 1
Gene-specific PCR primers used for semi-quantitative and quantitative RT-PCR (DOC 39 kb)
Supplementary Table 2
Rice sequences up-regulated by Fe-excess treatment (DOC 69 kb)
Rights and permissions
About this article
Cite this article
Ricachenevsky, F.K., Sperotto, R.A., Menguer, P.K. et al. Identification of Fe-excess-induced genes in rice shoots reveals a WRKY transcription factor responsive to Fe, drought and senescence. Mol Biol Rep 37, 3735–3745 (2010). https://doi.org/10.1007/s11033-010-0027-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0027-0