Abstract
A genomic region of particular interest for sweet cherry (Prunus avium L.) breeding is a quantitative trait locus (QTL) “hotspot” on chromosome 2. QTLs for fruit size, firmness, sweetness, and flowering time are reported to map to this region. An understanding of genetic diversity, allele sources, linkage relationships, and historical recombinations is critical to enable the combining of favorable alleles at multiple loci. The objectives of this study were to characterize, visualize, and interpret the genetic structure of this previously identified QTL hotspot within North American sweet cherry breeding germplasm, using a pedigree-based haploblocking approach. Across the 29.4 cM (6.3 Mbp) region defined by single nucleotide polymorphism (SNP) information from the RosBREED cherry 6K SNP array v1, a total of 12 recombination events falling into six inter-marker regions were traced within the pedigree of elite and wild germplasm (n = 55). These recombinations defined five haploblocks containing 5–15 markers and exhibiting 7–11 haplotypes each. Over the entire QTL hotspot, 30 extended haplotypes were identified for which parental gametes could be determined. When the haploblocks and their haplotypes were used to explore genetic diversity, ancestry, and recombination patterns, and then integrated with previous QTL results for fruit size, the results indicated that favorable alleles at this QTL hotspot are under positive selection in breeding. The genetic framework provided by a haploblock approach and knowledge of haplotype-level diversity sets the stage for assigning breeding utility to these haplotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sweet cherry (Prunus avium L.) is a clonally propagated, diploid, outcrossing species for which cultivar development uses a pedigree-based breeding approach. Potential new cultivars are usually selected from segregating F1 families. Because of the long generation time of sweet cherry (typically 3 to 5 years) and the fact that sweet cherry is not native to the Americas, both the historical number of founder cultivars used in breeding and the number of breeding generations are few. For example, in North American breeding programs, most cultivar releases from four sweet cherry breeding programs were reported to be descendants from five founding clones (Choi and Kappel 2004). Three of these five founders (‘Emperor Francis’, ‘Napoleon’, and ‘Windsor’) are still maintained and therefore available for genotypic characterization. As such, sweet cherry breeding is well suited for a pedigree-based method that integrates the concept of identity-by-descent to follow mosaics of founder genomes in breeding germplasm.
One chromosomal region of particular interest for sweet cherry breeding is a quantitative trait locus (QTL) “hotspot” in the middle of chromosome 2. This region, spanning approximately 29.4 cM (6.3 Mbp), includes two fruit size QTLs (Zhang et al. 2010; Rosyara et al. 2013; Campoy et al. 2015) and a candidate fruit size gene associated with domestication (De Franceschi et al. 2013), along with QTLs for fruit firmness (Campoy et al. 2015), fruit sweetness (Y. Zhao., personal communication), and flowering time (Castède et al. 2014). An understanding of the genetic composition of germplasm for this important QTL hotspot would assist breeders in their goal of combining favorable alleles for the multiple linked loci. With knowledge of the genetic diversity and linkage relationships among desirable QTL alleles in breeding germplasm, breeders could select desirable series of QTL alleles and identify regions to target for recombination (Jaganathan et al. 2015).
Tracing the inheritance of the chromosome 2 QTL hotspot across multiple generations of breeding germplasm is enabled with genome-wide, high-density, single nucleotide polymorphism markers (SNPs). Such SNP data sets capture historical recombination events and resulting haplotype patterns in descendent generations (Peace et al. 2012). However, visualizing the inheritance of thousands of bi-allelic SNPs from multiple founders is challenging given that most sweet cherries cultivars have been shown to have a high level of heterozygosity (Wünsch and Hormaza 2002; Lacis et al. 2009; Sharma et al. 2015; Farsad and Esna-Ashari 2016). Instead, the data contained within a large number of SNPs can be simplified by identifying non-recombining multi-locus regions called “haploblocks” that consist of completely linked SNPs. Haploblocks also have the advantage of behaving as multi-allelic markers within a particular germplasm set, facilitating tracing the inheritance of all SNPs from many founders. For example, if each of four founding clones are heterozygous at a locus, up to eight haplotypes (i.e., alleles) could occur in descendent germplasm. These eight haplotypes could be distinguished with a minimum of three phased SNPs, and the eight haplotypes for the three-SNP haploblock could easily be traced through multi-generation pedigrees (Voorrips et al. 2016).
The objective of this study was to characterize, visualize, and interpret the genetic structure of a previously identified QTL hotspot on chromosome 2 within North American breeding sweet cherry germplasm, using a pedigree-based haploblocking approach. The haploblocks and haplotypes identified for this region were used to explore ancestry, historical recombinations, and patterns of selection based on previous QTL allele results for fruit size.
Materials and methods
Plant material and genotypic data
This study used a pedigree-connected set of 62 elite sweet cherry clones (cultivars, advanced breeding selections, and landraces), three wild (i.e., undomesticated) sweet cherry clones, and 463 unselected F1 seedlings from 86 crosses of the Washington State University sweet cherry breeding program (Table S1). The germplasm, spanning six generations, was considered representative of US public breeding germplasm for this crop, covering the sweet cherry Crop Reference Set and Breeding Pedigree Set established in the RosBREED project (Peace et al. 2014). Of these 528 individuals, 12 elite sweet cherry clones could not be genotyped as sources of DNA were not available. SNP genotyping was done using the cherry 6K Illumina Infinium® SNP array (Peace et al. 2012) for which physical positions of the markers (Campoy et al. 2016) were based on the peach genome v2.0 (Verde et al. 2017, updated from the peach genome v1.0, Verde et al. 2013) as a proxy for the cherry genome. Genetic positions of all markers were determined by aligning and integrating these physical positions with the sweet cherry ‘Regina’ × ‘Lapins’ SNP linkage map (Klagges et al. 2013). Of the 516 individuals SNP-genotyped, 438 were also genotyped for two simple sequence repeat markers (SSRs), CPSCT038, and BPPCT034, located within the chromosome 2 QTL hotspot as described in Zhang et al. (2010). Confirmed pedigree information, genotypic data, and genetic map information were obtained from Peace et al. (in prep.), including various parentage adjustments such as the change of the paternal parent of ‘Sweetheart’ from ‘Newstar’ (reported in Lane and MacDonald 1996) to Lapins (Table S1).
Haploblock structure and diversity for the chromosome 2 QTL hotspot
The linkage phases of SNP and SSR markers for the chromosome 2 QTL hotspot for each germplasm individual were determined with pedigree information and offspring segregation data using FlexQTL™ (version 0.99130) and VisualFlexQTL (version 0.1.0.42) software for genetic analyses in multiple pedigree-connected families (Bink et al. 2014; www.flexqtl.nl). This phasing included the identification of recombination breakpoint positions, i.e., changes in linkage phase between parent and offspring. For those founders with just one genotyped offspring, phases of the founder homologs were considered putative as recombinations inherited by their one offspring could not be determined. For those founders with just two genotyped offspring, recombinations were arbitrarily assigned between the two offspring, as the true recombinant offspring could not be determined. If the recombination could not be assigned to an interval between two consecutive markers, VisualFlexQTL estimated the recombination breakpoint to be the genetic center position of the recombined sequences (i.e., the alleles of the first and last marker of a recombined sequence originated from two grandparents). When a double recombination was detected in an interval smaller than 10 cM, a manual inspection was performed to ensure that it was not an erroneous double recombination which might have resulted from data errors such as incorrect SNP calling, phasing, or pedigree information. The breakpoint positions were further used in VisualFlexQTL to delimit the boundaries of phased markers into haploblocks within which no recombination was identified for any of the genotyped plant progenitors (Fig. S1). Progenitors were defined as the 65 elite and wild clones, and excluded their unselected offspring. Haplotypes of each haploblock were assigned using the PediHaplotyper software (Voorrips et al. 2016), based on observed SNP and SSR marker haplotypes previously phased by FlexQTL™. Inheritance patterns of haplotypes were visualized with the software Pedimap (Voorrips et al. 2012).
Integration of historic and current data
The likely locations of two previously published chromosome 2 fruit size QTLs (Rosyara et al. 2013) and the location of a candidate fruit size gene (De Franceschi et al. 2013) were connected to the new haploblock designations based on placement of BPPCT034 and PavCNR12, respectively, for two sweet cherry linkage mapping populations, NY 54 × ‘Emperor Francis’ and ‘Regina’ × ‘Lapins’. Alleles 1–3 for the PavCNR12 gene were determined in De Franceschi et al. (2013) by sequencing of the gene and ~1500 bp of the upstream region for all four parents of the two F1 mapping populations. All four parents were reported to be heterozygous for PavCNR12 (De Franceschi et al. 2013) and BPPCT034 (Rosyara et al. 2013). In the study of De Franceschi et al. (2013), offspring were genotyped for flanking SSRs, and the PavCNR12 alleles in recombinant offspring were sequenced to determine their PavCNR12 genotypes. Here, significant allele effects from these prior studies were connected to the new haplotype designations of two haploblocks, HB-E and HB-F.
Results
Recombination at the QTL hotspot
The upper and lower flanking markers of the QTL hotspot on chromosome 2 were defined by recombinations between HB-A and HB-B, and HB-F and HB-G, respectively (Fig. 1; Table 1; Table S2). Across this 29.4 cM (6.3 Mbp) QTL hotspot region, a total of 12 recombinations, which fell into six recombination regions, here called “gaps,” were traced within the pedigree of the elite and wild germplasm. All six gaps were identified from the gametes leading to EE, ‘Sweetheart’, ‘Lapins’, and ‘Stella’ (Fig. S1). The ‘Stella’ gamete that gave rise to ‘Lapins’ had a recombination between SNPs ss490549138 and ss490549172, and this recombination split HB-C from HB-D. These gaps delimited the QTL hotspot into five haploblocks (Fig. 1; Table 1). The largest gap in this QTL hotspot region was 5.5 cM (1.3 Mbp, between HB-B and HB-C) and the smallest genetic distance of a gap was 0.0 cM (0.1 Mbp, between HB-D and HB-E) (Table 1).
Haploblocks (blue) and gaps (red, no SNPs) between the haploblocks across a QTL hotspot on chromosome 2 and summary of QTLs previously reported for this region. The two flanking haploblocks and gaps are also illustrated (hatched blue and red, respectively). A candidate gene for sweet cherry fruit size, PavCNR12 (De Franceschi et al. 2013) is located in haploblock HB-E, flanked by the SSRs CPSCT038 and BPPCT034 (Zhang et al. 2010) in adjacent haploblocks
The five haploblocks spanning the QTL hotspot (HB-B to HB-F) had an average length of 4.3 cM and 0.9 Mbp in genetic and physical units, respectively (Fig. 1; Table 1; Table S2). The fruit size QTL reported by Rosyara et al. (2013) was suspected in that report to consist of two adjacent QTLs. These two QTLs together spanned three of the haploblocks, HB-D to HB-F (Fig. 1). The estimated locations of the fruit size, firmness, sweetness, and flowering time QTLs on chromosome 2 detected in other studies (Zhang et al. 2010; Castède et al. 2014; Campoy et al. 2015; Y Zhao, personal communication) also spanned three or four haploblocks of this hotspot (Fig. 1).
When progeny from the unselected F1 families were considered, an additional 200 recombination events were identified within the QTL hotspot, with 85 and 115 recombination events identified within haploblocks and gaps, respectively (Table 1). The highest observed number of recombinations within haploblocks was in the 5.5-cM HB-E where PavCNR12 was located. For example, an offspring was determined to have inherited a gamete with a recombination within HB-E between the SNPs located at 19,200,549 and 19,324,328 that flanked the candidate gene for fruit size, PavCNR12 (Fig. S1). The highest observed number of recombination within gaps was in the 5.5-cM Gap-B that overlapped with the 95% confidence interval of the fruit sweetness QTL (Fig. 1).
Haplotype prevalence and diversity
The five haploblocks that spanned the chromosome 2 QTL hotspot were defined by a total of 45 markers (43 SNPs and two SSRs), with each of the five haploblocks containing 5–15 markers and exhibiting 7–11 haplotypes (Table 1; Fig. 2). A manual examination identified three to five markers per haploblock that were sufficient to distinguish among all haplotypes (Fig. 2). Inclusion of the seven-allele SSR BPPCT034, located in HB-F, increased the number of haplotypes in that haploblock to 11 from seven without this SSR (Fig. S2).
Marker allele composition of each haplotype across the five haploblocks for the sweet cherry QTL hotspot on chromosome 2 illustrated using the smallest number of markers needed to differentiate the haplotypes. SSR alleles are recorded as fragment sizes in base pairs. Haplotypes were assigned by the PediHaplotyper software (Voorrips et al. 2016). Haplotypes containing missing marker scores were omitted from the table. The complete marker compositions are in Supplementary Fig. S2
For the middle three of the five haploblocks (i.e., HB-C, HB-D, and HB-E), the most common haplotype (“2”) was represented in 69.4, 70.0, and 64.5% of the combined elite and wild germplasm (Table S3). In contrast, for the two outer haploblocks of the hotspot (HB-B and HB-F), the most common haplotypes (“3” and “2”) were represented in only 34.3 and 48.2% of the combined elite and wild germplasm, respectively. Two sweet cherry landrace cultivars from Spain, Cristobalina and Ambrunes, contributed unique haplotypes for HB-E, and Cristobalina contributed at least one unique haplotype for three of the four other haploblocks. All wild sweet cherry individuals (MIM 17, MIM 23, and NY 54) also contributed unique haplotypes for HB-E and the two MIM clones also contributed unique haplotypes for the other four haploblocks.
Over the entire QTL hotspot, 30 extended haplotypes (haplotypes extending over the 5 haploblocks HB-B to HB-F) were identified for the 55 elite and wild individuals for which parental gametes could be determined (Table S4). The most common extended haplotype (20.4% frequency) was 3,2,2,2,2, observed in the founders ‘Napoleon’, PMR-1, and ‘Windsor’, as well as the wild accession NY 54. However, different haplotypes for both flanking haploblocks (HB-A and HB-G) for ‘Windsor’ compared to ‘Napoleon’, PMR-1, and NY 54 suggest that the extended haplotype from ‘Windsor’ might not be identical by descent. Another frequent extended haplotype (19.4% frequency) was 4,2,2,2,2, contributed by ‘Schneiders’, ‘Schmidt’, and ‘Krupnoplodnaya’, although different flanking haplotypes for ‘Schneiders’ and ‘Schmidt’ also suggest that this extended haplotype is not identical by descent among the three cultivars. Rare extended haplotypes were contributed from ‘Cristobalina’, MIM 17 and MIM 23. ‘Black Republican’ and ‘Windsor’ shared a rare extended haplotype for HB-C to HB-F, 5,5,10,9 (Table S4); however, their derived cultivars, Bing and Venus, respectively, did not inherit this extended haplotype. Only one individual, ‘Kordia’, was homozygous for an extended haplotype across this five-haploblock region.
Haplotype inheritance as mosaics of founder contribution
Inheritance of the extended haplotypes in the ‘Sweetheart’ pedigree revealed four recombinations in its four-generation pedigree (Fig. 3). Although ‘Black Republican’ only had two genotyped offspring, ‘Van’ and ‘Bing’, both inherited the same extended haplotype (dark blue) indicating that neither ‘Van’ nor ‘Bing’ resulted from a recombinant gamete in this region. It was not possible to determine if the extended haplotype ‘Van’ inherited from ‘Empress Eugenie’ (light blue, Fig. 3) had a recombination in this region because ‘Van’ was the only genotyped offspring of ‘Empress Eugenie’. ‘Stella’ inherited a recombinant gamete from its maternal parent, ‘Lambert’, with the recombination occurring between HB-E and HB-F. ‘Lapins’ inherited two further recombinant gametes, one from each parent. ‘Sweetheart’ inherited recombinant gametes from ‘Lapins’ and ‘Van’, where the recombination of the former occurred within the hotspot and the recombination of the latter defined the distal margin of the hotspot. The extended haplotypes for ‘Newstar’ are also shown (Fig. 3) to illustrate that breeding records indicating ‘Newstar’ is a parent of ‘Sweetheart’ are incorrect.
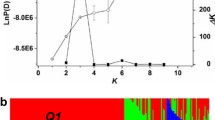
The pedigree of 'Sweetheart', where a yellow background color indicates that an individual was genotyped and gray indicates not genotyped. a Phased haplotypes across the five haploblocks spanning a QTL hotspot on chromosome 2 and two flanking haploblocks, HB-A and HB-G. Within each individual, haplotypes on the left were inherited from the mother (red line) and haplotypes on the right from the father (blue line). A transition in color within a column indicates a recombination event. Asterisk indicates this haplotype could not be deduced. For JI 2420, information from flanking haploblocks (HB-A and HB-G) was used to determine that the extended HB-B to HB-F haplotype 2, 3, 3, 3, 3 was derived from 'Napoleon'. b The IBD probabilities that a founder allele is present for the length of chromosome 2 where the vertical dimension represents the position along the chromosome. Each rectangle in an individual represents one of the two copies of chromosome 2, where each color represents a distinct founder chromosome. For founders not genotyped ('Empress Eugenie' and 'Blackheart'), the two founder homologs were assigned the same color, consistent with their offspring. The two horizontal black lines across each rectangle indicate boundaries of the five-haploblock region of a. The width of the color at each position along the chromosome indicates that the IBD probability that the corresponding founder allele is present in this individual. The images were drawn using PediMap software (Voorrips et al. 2012)
Certain founder haplotypes were more prevalent than others in this QTL hotspot. Of the 10 haplotypes of ‘Sweetheart’ across the five haploblocks, seven originated from ‘Empress Eugenie’ through the twice-recurrent parent ‘Van’ (Fig. 3). Of the other three haplotypes, two originated from ‘Black Republican’ and one from ‘Napoleon’. Haplotypes from the other two founder ancestors of ‘Sweetheart’, ‘Blackheart’ and ‘Emperor Francis’, were not inherited by ‘Sweetheart’. Considering also five other first-generation offspring cultivars of Van (Rainier, Lapins, Newstar, Olympus, and Summit; Table S1), the haplotypes derived from ‘Empress Eugenie’ for HB-D, HB-E, and HB-F were observed in four of them (all except Lapins). Even more extreme, for HB-E and HB-F, all six offspring carried ‘Empress Eugenie’ haplotypes, four being homozygous at HB-E and three for HB-F.
Integration of historic and current data
Haplotypes of HB-E and HB-F, spanning the likely location(s) of previously published fruit size QTLs, were successfully associated with previously reported alleles of PavCNR12 and BPPCT034, respectively (Fig. 4). The PavCNR12 alleles 1, 2, and 3 were associated with the HB-E haplotypes 2, 3, and 19, respectively (Fig. 4a, b). In the ‘Regina’ (2/3) × ‘Lapins’ (2/3) offspring, significant differences in offspring mean fruit weight based on their PavCNR12 genotypes, and thereby on the current HB-E genotypes, were as follows: 2/2 > 2/3 > 3/3 (Fig. 4a). In the NY 54 (2/19) × ‘Emperor Francis’ (2/3) offspring, significant differences in mean fruit weights were also identified. Mean fruit weights based on their PavCNR12 genotypes, and therefore on the current HB-E genotypes, were highest for those genotypes possessing at least one haplotype 2 (2/2 = 2/3 > 2/19 > 3/19; Fig. 4b).
Alignment of the chromosome 2 QTL hotspot haploblocks and haplotypes (blue) with the fruit size QTL candidate gene and SSR marker alleles reported by De Franceschi et al. (2013) (a, b) and Rosyara et al. (2013) (c, d) (black), respectively. a, b Mean fruit weight comparison for 'Regina' × 'Lapins' offspring and NY 54 × 'Emperor Francis' offspring, respectively, based on segregation of PavCNR12 which is located in HB-E. c, d Mean fruit weight comparison for 'Regina' × 'Lapins' offspring and NY 54 × 'Emperor Francis' offspring, respectively, based on segregation of the SSR BPPCT034 which is located in HB-F
In Rosyara et al. (2013), when the progeny from ‘Regina’ (2/6) × ‘Lapins’ (2/3) were grouped here by HB-F haplotype, significant differences in mean fruit weight were observed, where the most extreme fruit weight progeny means were 8.8 and 6.2 g for the 2/2 and 3/6 genotypes, respectively (Fig. 4c). For the NY 54 (2/13) × ‘Emperor Francis’ (2/3) progeny, individuals with genotype 2/2 for HB-F also showed the largest mean fruit weight (Fig. 4d). When HB-E and HB-F results were taken together, fruit size was highest in individuals that were homozygous for haplotype 2 for both haploblocks. Homozygous 2,2 was also the most frequent haplotype series for HB-E and HB-F in the breeding germplasm (Fig. 3).
Discussion
Diversity and ancestry
Condensing into five haploblocks the 43 bi-allelic SNP markers and two SSR markers that spanned the chromosome 2 QTL hotspot allowed a simpler visualization of the genetic structure of this genomic region across breeding germplasm. The phased linked markers revealed numerous haplotypes, as many as 11 for one of the haploblocks. The observation that the two Spanish landrace clones, ‘Cristobalina’ and ‘Ambrunes’, carried haplotypes not previously present in US breeding germplasm is consistent with prior findings that these two landraces are genetically distant from earlier US sweet cherry breeding germplasm (Cabrera et al. 2012). The Spanish landraces were also found to be genetically distant from northern and central European germplasm (Mariette et al. 2010, Wünsch and Hormaza 2002). The lack or scarcity of certain wild germplasm haplotypes for the chromosome 2 QTL hotspot in elite materials is consistent with a domestication bottleneck, a phenomenon previously documented for sweet cherry (Choi and Kappel 2004; Mariette et al. 2010; Campoy et al. 2016).
The haplotype structure facilitated detection of the available diversity which was maintained or lost in modern-bred cultivars and other selected materials. For example, when the five haploblocks were considered together in series, extended haplotypes were identified that are very common in current breeding germplasm (3,2,2,2,2 from ‘Napoleon’) or lost from advanced breeding germplasm (5,5,5,10,9 in ‘Windsor’ and 6,5,5,10,9 in ‘Black Republican’). Although the extended haplotype 2,2,2,2 for HB-C to HB-F is identical-by-state in many founders, it is possible that this shared extended haplotype is not always identical-by-descent in cases that pedigree connections do not confirm shared descent (e.g., ‘Napoleon’ and ‘Krupnoplodnaya’). This distinction has particular relevance for the interpretation of QTL studies, because if haplotypes are not identical-by-descent they are less likely to contain the same QTL allele. However, the overwhelming maintenance of cultivar Empress Eugenie’s contribution to the chromosome 2 QTL hotspot in derived selected germplasm suggests that this region has been a target of positive selection. For both HB-E and HB-F, fruit size was highest in individuals that were homozygous for haplotype 2 for both haploblocks.
Historical and new recombinations
‘Sweetheart’ was homozygous for the 2,2 haplotypes for HB-E and HB-F originating from ‘Empress Eugenie’ as a result of four recombinant gametes in this region over its four generation pedigree. The haploblocking procedure resulted in the easy identification and visualization of these historical recombinations, hereby clarifying the genetic basis of one of the outstanding characteristics (fruit size) of Sweetheart. This clarification is useful to breeders, as ‘Sweetheart’, a mainstay in the late part of the fruiting season in North America, has become a common breeding parent in recent decades.
The haploblock and haplotype information presented for the chromosome 2 QTL region also provided clarity in efficiently determining prospects of families for QTL dissection if their parents and some related germplasm are genotyped with common bridging markers. F1 families have been developed to fine-map the QTLs for fruit size, firmness, sweetness, and flowering time. Two reported mapping families are ‘Regina’ × ‘Lapins’ and ‘Black Tartarian’ × ‘Kordia’ (Klagges et al. 2013). Their parents were chosen for their differing phenotypes without prior genome-wide or QTL region-specific genetic information. Extended haplotypes at the chromosome 2 QTL hotspot for the first family are 3,2,3,3,3 and 2,3,3,2,2 for ‘Lapins’ and 4,2,2,2,2 and 2,3,3,3,6 for ‘Regina’. Because both parents are heterozygous for most haploblocks, it would be possible to genetically dissect the region by identifying recombinant offspring, as illustrated by the recombinations identified in the progeny from the F1 families in the present study (e.g., Fig. S1). In contrast, ‘Kordia’ is unusually homozygous at this region (Cabrera et al. 2012; Campoy et al. 2016) and therefore segregation for QTLs in the second family could only be identified from ‘Black Tartarian’ if that cultivar is heterozygous for QTL alleles conferring contrasting trait levels and the corresponding alleles from ‘Kordia’ are not dominant. Although homozygosity reduces efficiency for fine-mapping in the chromosome 2 QTL hotspot, the ‘Black Tartarian’ × ‘Kordia’ population provides an opportunity to more readily detect and characterize segregating ‘Kordia’ QTLs for these traits elsewhere in the genome.
Understanding linkage relationships and identifying recombinations is critical to the goal of combining favorable alleles for the multiple linked QTLs. Such is the case for fruit size and firmness where chromosome 2 fruit size and firmness QTL allele effects have been reported to be negatively associated in most of the haplotypes evaluated, although pleiotropy at a single gene cannot be ruled out (Campoy et al. 2015; C. Peace, unpublished data). Future QTL studies will likely result in improved predictions of QTL locations for these multiple traits and assignment of trait values to haplotypes.
Conclusion
The chromosome 2 QTL hotspot of sweet cherry is a region with multiple QTLs of breeding interest that can be compartmentalized as a series of discrete haploblocks and characterized by associated haplotypes. Using the haploblocks and haplotypes to explore genetic diversity, ancestry, historical recombinations, and patterns of selection for fruit size QTL alleles provided evidence that favorable alleles from a particular ancestral source have been under positive selection. With such information on genetic structure available genome-wide, breeders can efficiently target identification of recombinant individuals to achieve desired QTL allele combinations, thereby expanding selection decisions from one or a few large-effect QTLs to all known QTL-containing regions.
Change history
22 July 2017
An erratum to this article has been published.
References
Bink MCAM, Jansen J, Madduri M, Voorrips RE, Durel CE, Kouassi AB, Laurens F, Mathis F, Gessler C, Gobbin D, Rezzonico F, Patocchi A, Kellerhals M, Boudichevskaia A, Dunemann F, Peil A, Nowicka A, Lata B, Stankiewicz-Kosyl M, Jeziorek K, Pitera E, Soska A, Tomala K, Evans KM, Fernández-Fernández F, Guerra W, Korbin M, Keller S, Lewandowski M, Plocharski W, Rutkowski K, Zurawicz E, Costa F, Sansavini S, Tartarini S, Komjanc M, Mott D, Antofie A, Lateur M, Rondia A, Gianfranceschi L, van de Weg WE (2014) Bayesian QTL analyses using pedigreed families of an outcrossing species, with application to fruit firmness in apple. Theor Appl Genet 127(5):1073–1090
Cabrera A, Rosyara UR, De Franceschi P, Sebolt A, Sooriyapathirana SS, Dirlewanger E, Quero-Garcia J, Schuster M, Iezzoni AF, van der Knaap E (2012) Rosaceae conserved orthologous sequences marker polymorphism in sweet cherry germplasm and construction of a SNP map. Tree Genet Genomes 8:237–247
Campoy JA, Dantec L, Barreneche T, Dirlewanger E, Guero-Garcia J (2015) New insights into fruit firmness and weight control in sweet cherry. Plant Mol Biol Rep 33(4):783–796
Campoy JA, Lerigoleur-Balsemin E, Christmann H, Beauvieux R, Girollet N, Quero-Garcia J, Dirlewanger E, Barrenecha T (2016) Genetic diversity, linkage disequilibrium, population structure and construction of a core collection of Prunus avium L. landraces and bred cultivars. BMC Plant Biol 16:49
Castède S, Campoy JA, García JQ, Le Dantec L, Lafargue M, Barreneche T, Wenden B, Dirlewanger E (2014) Genetic determinism of phenological traits highly affected by climate change in Prunus avium: flowering data dissected into chilling and heat requirements. New Phytol 202(2):703–715
Choi C, Kappel F (2004) Inbreeding, coancestry, and founding clones of sweet cherries from North America. J Am Soc Hortic Sci 129:535–543
De Franceschi P, Stegmeir T, Cabrera A, van de Knaap E, Rosyara UR, Sebolt AM, Dondini L, Dirlewanger E, Quero-Garcia J, Campoy JA, Iezzoni AF (2013) Cell number regulator genes in Prunus provide candidate genes for the control of fruit size in sweet and sour cherry. Mol Breed 32:311–326
Farsad A, Esna-Ashari M (2016) Genetic diversity of some Iranian sweet cherry (Prunus avium) cultivars using microsatellite markers and morphological traits. Cytol Genet 50(1):9–19
Jaganathan D, Thudi M, Kale S, Azam S, Roorkiwal M, Gaur PM, Kishor PBK, Nguyen H, Sutton T, Varshney RK (2015) Genotyping-by-sequencing based intra-specific genetic map refinds a “QTL-hotspot” region for drought tolerance in chickpea. Mol Gen Genomics 290:559–571
Klagges C, Campoy JA, Quero-Garcia J, Guzman A, Mansur L, Gratacos E, Silva H, Rosyara UR, Iezzoni A, Meisel LA, Dirlewanger E (2013) Construction and comparative analyses of highly dense linkage maps of two sweet cherry intra-specific progenies of commercial cultivars. PLoS One 8(1):e54743
Lacis G, Rashal I, Ruisa S, Trajkovski V, Iezzoni AF (2009) Assessment of genetic diversity of Latvian and Swedish sweet cherry (Prunus avium L.) genetic resources collections by using SSR (microsatellite) markers. Sci Hortic 121(4):451–457
Lane WD, MacDonald RA (1996) Sweetheart sweet cherry. Can J Plant Sci 76:161–163
Mariette S, Tavaud M, Arunyawat U, Capdeville G, Millan M, Salin F (2010) Population structure and genetic bottleneck in sweet cherry estimated with SSRs and the gametophytic self-incompatibility locus. BMC Genet 11:77
Peace C, Bassil N, Main D, Ficklin S, Rosyara UR, Stegmeir T, Sebolt A, Gilmore B, Lawley C, Mockler TC, Bryant DW, Wilhelm L, Iezzoni A (2012) Development and evaluation of a genome-wide 6K SNP array for diploid sweet cherry and tetraploid sour cherry. PLoS One 7(12):e48305
Peace C, Luby J, van de Weg WE, Bink MCAM, Iezzoni A (2014) A strategy for developing representative germplasm sets for systematic QTL validation, demonstrated for apple, peach and sweet cherry. Tree Genet Genomes 10(6):1679–1694
Rosyara U, Bink M, van de Weg WE, Zhang G, Wang D, Sebolt A, Dirlewanger E, Quero-Garcia J, Schuster M, Iezzoni A (2013) Fruit size QTL identification and the prediction of parental QTL genotypes and breeding values in multiple pedigreed populations of sweet cherry. Mol Breed 32:875–887
Sharma K, Xuan H, Sedlák P (2015) Assessment of genetic diversity of Czech sweet cherry cultivars using microsatellite markers. Biochem Syst Ecol 63:6–12
Verde I, Abbott AG, Scalabrin S, Jung S, Shu S, Marroni F, Zhebentyayeva T, Dettori MT, Grimwood J, Cattonaro F, Zuccolo A, Rossini L, Jenkins J, Vendramin E, Meisel LA, Decroocq V, Sosinski B, Prochnik S, Mitros T, Policriti A, Cipriani G, Dondini L, Ficklin S, Goodstein DM, Xuan P, Del Fabbro C, Aramini V, Copetti D, Gonzalez S, Horner DS, Ralchi R, Lucas S, Mica E, Maldonado J, Lazzari B, Bielenberg D, Pirona R, Miculan M, Barakat A, Testolin R, Stella A, Tartarini S, Tonutti P, Arús P, Orellana A, Wells C, Main D, Vizzotto G, Silva H, Salamini F, Schmutz J, Morgante M, Rokhsar DS (2013) The high-quality draft of peach (Prunus persica) identifies unique patterns of genetic diversity, domestication and genome evolution. Nat Genet 45:487–494
Verde I, Jenkins J, Dondini L, Micali S, Pagliarani G, Vendramin E, Paris R, Aramini V, Gazza L, Rossini L, Bassi D, Troggio M, Shu S, Grimwood J, Tartarini S, Dettori MT, Schmutz J (2017) The peach v2.0 release: high-resolution linkage mapping and deep resequencing improve chromosome-scale assembly and contiguity. BMC Genomics 18:225
Voorrips RE, Bink MCAM, van de Weg WE (2012) Pedimap: software for the visualization of genetic and phenotypic data in pedigrees. J Hered 103(6):903–907
Voorrips RE, Bink MCAM, Kruisselbrink JW, Koehorst-van Putten HJ, van de Weg WE (2016) PediHaplotyper: software for consistent assignment of marker haplotypes in pedigrees. Mol Breed 36:119
Wünsch A, Hormaza JI (2002) Molecular characterization of sweet cherry (Prunus avium L.) genotypes using peach [Prunus persica (L.) Batsch] SSR sequences. Heredity 89:56–63
Zhang G, Sebolt A, Sooriyapathirana S, Wang D, Bink M, Olmstead J, Iezzoni A (2010) Fruit size QTL analysis of an F1 population derived from a cross between a domesticated sweet cherry cultivar and a wild forest sweet cherry. Tree Genet Genomes 6:25–36
Acknowledgements
This work was supported by USDA’s National Institute of Food and Agriculture-Specialty Crop Research Initiative project, “RosBREED: Combining Disease Resistance and Horticultural Quality in New Rosaceous Cultivars” (2014-51181-22378).
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article is available at https://doi.org/10.1007/s11032-017-0699-4.
Electronic supplementary material
Supplementary Table S1
Sweet cherry plant materials and their parentage information. (XLSX 15 kb)
Supplementary Table S2
Descriptions of locations (cM, bp) and lengths (cM, Mbp) of the haploblocks and gaps between sweet cherry haploblocks identified for a QTL hotspot on chromosome 2 in the sweet cherry breeding germplasm listed in Table S1. (XLSX 10 kb)
Supplementary Table S3
Frequencies and founder contributors of haplotypes identified for five haploblocks spanning a QTL hotspot on chromosome 2, calculated for elite and wild germplasm only. (XLSX 11 kb)
Supplementary Table S4
Haplotypes observed across five haploblocks for a QTL hotspot on chromosome 2 in U.S. elite and wild sweet cherry breeding germplasm for which parental gametes could be determined (n=55). (XLSX 14 kb)
Supplementary Fig. S1
(A) Recombination sites identified for determining haploblock boundaries of a QTL hotspot on chromosome 2, illustrated using the Segregation Indicator Pattern and Recombination Sequence parameters of FlexQTL™. Phased marker genotype data for a maternal parent (‘Van’) – paternal parent (‘Lapins’) – child (‘Sweetheart’) combination is shown. For each individual, alleles in the left column were inherited from the maternal parent and alleles on the right column were from the paternal parent. ‘0’ or ‘1’ indicates the grandparental origin of alleles (maternal or paternal, respectively). Recombination Sequences are marked by black line boxes. Recombination sites are highlighted by red lines. Six recombination sites used for delimiting the QTL hotspot’s five haploblocks were identified from elite germplasm including EE, ‘Sweetheart’, ‘Lapins’, and ‘Stella’. Examples of within-haploblock recombination are illustrated with five unselected offspring from different families. For example, offspring 4 was determined to have inherited a gamete with a recombination within HB-E between the SNPs located at 19,200,549 and 19,324,328. (B) Pedigree of individuals presented in Supplementary Fig. S1 (A). (DOCX 1054 kb)
Supplementary Fig. S2
Marker allele composition of each haplotype across five haploblocks for the sweet cherry QTL hotspot on chromosome 2. SSR alleles are recorded as fragment sizes in base pairs. The smallest subset of markers needed to differentiate the haplotypes within each haploblock are highlighted in red font. Haplotypes were assigned by the PediHaplotyper software (Voorrips et al. 2016). Haplotypes containing missing marker scores were omitted from the table. (DOCX 275 kb)
Rights and permissions
About this article
Cite this article
Cai, L., Voorrips, R.E., van de Weg, E. et al. Genetic structure of a QTL hotspot on chromosome 2 in sweet cherry indicates positive selection for favorable haplotypes. Mol Breeding 37, 85 (2017). https://doi.org/10.1007/s11032-017-0689-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-017-0689-6