Abstract
DRR17A is a stable wild-abortive cytoplasmic male sterile line with medium-slender grain type. DRR17A and its maintainer line DRR17B are highly susceptible to two of the major rice diseases, bacterial blight (BB) and blast. To improve DRR17B for resistance against BB and blast, we have introgressed a major dominant gene each conferring resistance against BB (Xa21) and blast (Pi54) into the maintainer line through marker-assisted backcross breeding using RP-Bio-Patho-2 (a near-isogenic line of Samba Mahsuri possessing Xa21 and Pi54) as the donor parent. PCR-based molecular markers tightly linked to Xa21 and Pi54 were used for foreground selection of the resistance plants at each backcross generation, while molecular markers tightly linked to the major fertility restorer genes, Rf3 and Rf4, were used for negative selection (i.e. selection of plants possessing non-fertility-restoring alleles at the two loci) at BC1 generation. After foreground selection for the target genes at each backcross generation, the ‘positive’ plants were screened with parental polymorphic markers for identifying backcross plants possessing maximum recovery of DRR17B genome. Marker-assisted backcrossing was continued till BC3 generation, and a single BC3F1 plant possessing the target genes with ~94 % recovery of recurrent parent genome was identified and selfed to generate BC3F2s. A total of six homozygous BC3F2 plants were identified and advanced. At BC3F5, six promising, stable, backcross-derived lines possessing high level of resistance against BB and blast, high yield, short plant stature, fine-grain type, have been identified; their maintenance ability and heterotic potential validated through test crosses and these lines are being converted to CMS lines through marker-assisted breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
India needs to produce at least 125 million tonnes of rice by year of 2020, to meet the demands of an increasing population. Among the various technological options available for increasing rice production, large-scale adoption of hybrid rice is the most promising one (Ahmed and Siddiq 1998). Hybrids yield 15–20 % over inbred rice varieties (Virmani 1996) and at least 70 public and private bred hybrids are available for commercial cultivation in India. However, most of the hybrids released to date in India and abroad are highly susceptible to biotic stresses like bacterial blight (BB) and blast (Khush and Jena 2009). DRR17A is an elite, highly stable wild-abortive cytoplasmic male sterile (WA-CMS) line, possessing highly desirable medium-slender grain type and developed by the Directorate of Rice Research, Hyderabad, India (AS Hariprasad, Personal Communication). The WA-CMS line is being used for developing promising three-line hybrids at DRR. However, DRR17A and its maintainer line DRR17B are highly susceptible for BB and blast and have a slightly taller plant stature which is not ideal for a good WA-CMS/maintainer line.
Bacterial blight (BB) caused by Xanthomonas oryzae pv. oryzae (Xoo) is one of the most important diseases of rice worldwide (Mew 1987) and causes yield loss ranging from 74 to 81 % (Srinivasan and Gnanamanickam 2005). Deployment of host plant resistance is the only effective strategy for management of the disease, and till date more than 38 BB resistance genes have been identified (Cheema et al. 2008; Sujatha et al. 2011; Hari et al. 2013). Some of the major resistance genes like Xa1, Xa4, xa5, Xa7, xa8, xa13, Xa21 and Xa27 have been tagged and mapped by closely linked molecular markers (Sonti 1998; Rao et al. 2002; Sundaram et al. 2014). Some of the BB resistance genes are dominant in nature (e.g. Xa21, Xa33 and Xa38). Availability of tightly linked molecular markers in rice has enabled marker-assisted breeding (MAB) for bacterial blight resistance, a reality in India and abroad (Abenes et al. 1993; Yoshimura et al. 1995; Zhang et al. 1996; Huang et al. 1997; Sanchez et al. 2000; Davierwala et al. 2001; Singh et al. 2001; Joseph et al. 2004; Leung et al. 2004; Perez et al. 2008; Sundaram et al. 2008, 2009; Basavaraj et al. 2010; Hari et al. 2011, 2013). The major dominant BB resistance gene Xa21 was originally introgressed from the wild rice, Oryza longistaminata (Ronald et al. 1992; Song et al. 1995), which conferred broad-spectrum of resistance against most of the virulent isolates existing in India. Most importantly, a very closely linked PCR-based marker, pTA248 is available for marker-assisted selection of the gene (Ronald et al. 1992). Many earlier studies have shown that through marker-assisted breeding, Xa21 can be successfully introgressed into elite rice varieties (Joseph et al. 2004; Gopalakrishnan et al. 2008; Sundaram et al. 2008, 2009; Perumalsamy et al. 2010; Pandey et al. 2013) and into hybrid rice parental lines (Chen et al. 2001; Liyong et al. 2003; Basavaraj et al. 2010; Hari et al. 2011, 2013).
Rice blast, caused by the fungus Magnaporthe grisea (anamorph Pyricularia oryzae), is a devastating disease, which leads to significant yield loss up to 70–80 % during an epidemic (Khush and Jena 2009). Even though chemicals are available for control of blast, deployment of host plant resistance is one of the best options for managing the disease (Hulbert et al. 2001). At least, 100 genes conferring resistance against blast disease and 347 quantitative trait loci (QTL) associated with blast resistance have been identified (Ballini et al. 2008; Koide et al. 2009), and 19 resistance genes have been cloned and characterized (Sharma et al. 2012). The Pi54 blast resistance gene was mapped on chromosome 11L from the Vietnamese rice genotype, Tetep that conferred the resistance against predominant races of the pathogen in India (Sharma et al. 2010). Tightly linked (Sharma et al. 2005) and functional markers (Ramkumar et al. 2010) are available for marker-assisted selection of the gene, and a few research groups in India have successfully introgressed Pi54 into varieties (Narayanan et al. 2002; Joseph et al. 2004; Sundaram et al. 2008, 2009) and parental lines (Basavaraj et al. 2010; Singh et al. 2011; Zhan et al. 2012; Hari et al. 2011, 2013).
As there is an imminent need to improve DRR17B for BB and blast resistance and with the availability of very closely linked/functional markers for both BB resistance gene, Xa21, and the blast resistance gene, Pi54, the present study was initiated.
Materials and methods
Plant material
RP-Bio-Patho-2 (NIL of Samba Mahsuri, possessing Xa21 and Pi54) was used as the donor parent for bacterial blight and blast resistance genes. DRR17B, the maintainer line of an elite wild-abortive CMS line-DRR17A, was used as the recurrent parent. In addition to these, Taichung Native 1(TN1), HR12 and Tetep were used as susceptible and resistant checks for blast screening, while the recurrent parent and donor parents were used as susceptible and resistant checks, respectively, while screening for BB resistance.
Marker-assisted selection for BB and blast resistance, fertility restoration
For targeted introgression of Xa21 and Pi54 into DRR17B, a marker-assisted backcross breeding programme was adopted. Backcrossing was performed till BC3 generation, after which the plants were advanced through pedigree method. DNA was isolated from the parents and backcross progenies by following the protocol of Zheng et al. (1995). The PCR-based STS marker pTA248 (Ronald et al. 1992), functional marker Pi54 MAS (Ramkumar et al. 2010) were used to identify the allelic status of Xa21 and Pi54 at BC1F1 and subsequent backcross generations, while the SSR markers DRCG-RF4-14/DRCG-RF4-8 and DRRM-RF3-10/DRRM-RF3-5 (Balaji et al. 2012) closely linked to the major fertility restorer genes, Rf4 and Rf3, were used at BC1F1 generation to identify plants possessing the non-fertility-restoring allele (i.e. rf4rf4 and rf3rf3) in homozygous condition. Information about the markers used for foreground selection is given in Supplementary Table 1. A set of 79 parental polymorphic SSR markers spread across the 12 chromosomes of rice were utilized for background selection at BC2F4 generation.
PCR was performed using 1 U of Taq DNA polymerase (Bangalore Genei, Bangalore, India) and 1X PCR buffer (10 mM Tris, pH 8.4, 50 mM KCl, 1.8 mM MgCl and 0.01 mg/ml gelatin), 5 Pico moles of each primer, 0.05 mM dNTPs and 50 ng template DNA in 25 µl reaction volume with a thermal profile of 94 °C for 5 min (initial denaturation), followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 30 s, extension at 72 °C for 1 min and a final extension of 7 min at 72 °C. The amplified product of pTA248 (Xa21) was electrophoretically resolved on a 1.5 % Seakem LE agarose gel (Lonza, Rockland, ME, USA); the amplicons of Pi54 MAS (Pi54) were dissolved in 2 % agarose, the amplicons of DRCG-RF4-14/DRCG-RF4-8 (Rf4) in 2.5 % agarose gel and the amplicons of RRM-RF3-10/DRRM-RF3-5 (Rf3), in 3.5 % agarose gel. Those of SSR markers used for background selection were resolved on a 3.5 % agarose gel. Seakem LE agarose gels containing 0.5 mg/ml of ethidium bromide in 0.5X TBE buffer were visualized under UV.
Screening for BB resistance
Four virulent isolates of the bacterial blight pathogen, Xanthomonas oryzae pv. oryzae (Xoo) collected from BB hot-spot locations in India, viz. DX-020 (Hyderabad, Telangana), DX-002 (Faizabad, Uttar Pradesh), DX-066 (Raipur, Chhattisgarh) and DX-049 (Maruteru, Andhra Pradesh), were used to screen the donor and recurrent parents along with backcross-derived lines of DRR17B for BB resistance under both glasshouse and field conditions. The Xoo strains were cultured and stored as described by Laha et al. (2009). The rice plants were clip-inoculated with a bacterial suspension of 108–9 cfu/ml at maximum tillering stage (45–55 days after transplanting) through the methodology of Kauffman et al. (1973). Approximately 5–10 leaves were inoculated per plant, and disease reaction was scored 14 days after inoculation. Individual plants of the segregating backcross generations were screened under field condition following the same procedure. In addition to measurement of BB lesion length, the disease score was also calculated as per IRRI standard evaluation system (IRRI-SES) scale, which is based on per cent diseased leaf area (IRRI 1996).
Screening for blast resistance
A local isolate of Magnaporthe oryzae (SPI-40) from Directorate of Rice Research (DRR), Hyderabad, Andhra Pradesh, India (Madhan Mohan 2011), was used to screen the donor and recurrent parents along with backcross-derived lines of DRR17B for blast resistance under in vivo conditions following uniform blast nursery (UBN) method at Directorate of Rice Research (DRR), Hyderabad, India. The pathogen strains were cultured and stored as described by Srinivas Prasad et al. (2011). The young seedlings at four-leaf stage were inoculated with the fungal conidial suspension at a concentration of 1 × 105 conidia/ml, and high relative humidity was maintained for disease development. Inoculated seedlings were monitored for the development of blast lesions 1 week after inoculation. The plants were scored and evaluated on a 0–9 scale as per IRRI-SES scale (IRRI 1996).
Screening for agro-morphological characteristics
Thirty-day-old seedlings of the selected BC3F5 lines along with parents were transplanted in the Rajendranagar Experimental farm of Indian Institute of Rice Research, Hyderabad, India, at a spacing of 15 × 20 cm in 12-row plots in three replications. Standard agronomic practices were followed to raise a healthy crop, which were evaluated during the wet season (June–November) in 2013. Data were recorded for the agronomic traits, viz. plant height (cm), spikelet fertility (%), productive tillers (number), panicle length (cm), grains per panicle (number) and days to 50 % flowering.
Evaluation of heterosis of the experimental hybrids developed from improved DRR17B
Four selected backcross-derived lines (viz., GP17B112-3-98-118-12-204-8, GP17B112-3-98-118-12-204-15, GP17B112-3-98-118-12-204-90 and GP17B112-3-98-118-12-204-234) and DRR17B were crossed with the elite restorer line RPHR1005 to evaluate grain yield standard heterosis of the derived hybrids during wet season 2014. The popular, high-yielding variety, BPT5204 (Samba Mahsuri), possessing medium-slender grain type served as the varietal check, while the elite hybrid, DRRH3, possessing medium-slender grain type served as the hybrid check.
Results
Introgression of BB and blast resistance genes into DRR17B background
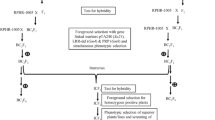
The heterozygosity of the F1s developed by crossing RP-Bio-Patho-2 (NIL of BPT5204, possessing Xa21 and Pi54) and the recurrent parent DRR17B was confirmed using pTA248, the co-dominant marker specific for Xa21, and Pi54 MAS, functional marker for Pi54. The positive F1 plants were then backcrossed with DRR17B to obtain BC1F1. A total of 261 BC1F1 plants were genotyped, of which 43 plants were double positives (i.e. Xa21 + Pi54; Fig. 1). These were then screened with Rf3- and Rf4-specific markers to identify plants devoid of the two genes (i.e. rf3rf3, rf4rf4), and a total of nine such plants were identified (Supplementary Figure 1). These were then subjected to background selection using 79 parental polymorphic SSRs to identify plants possessing maximum recurrent parent genome recovery. The best BC1F1 plant GP17B112, possessing a recovery of 73.41 % was identified and used for the next backcross generation. Backcrossing was continued up to BC3 generation, wherein a BC3F1 plant possessing Xa21 + Pi54 and 93.4 % recovery of DRR17B genome was identified and selfed to generate BC3F2s, which were then screened with gene-specific markers to identify homozygous plants. From a total of 544 BC3F2 plants, a total of 52 plants homozygous for both Xa21 and Pi54 were identified and advanced. At BC3F5, a total of six promising lines possessing grain type and agro-morphological traits similar to DRR17B were identified (viz. GP17B112-3-98-118-12-204-8, GP17B112-3-98-118-12-204-15, GP17B112-3-98-118-12-204-90, GP17B 112-3-98-118-12-204-168, GP17B112-3-98-118-12-204-234, GP17B112-3-98-118-12-204-276), and these were then screened for BB and blast resistance. The details of number of plants screened, number of plants identified to possess non-restoring alleles at Rf3 and Rf4 and percentage recurrent parent genome recovery of the best selected plant at each backcross generation are given in Supplementary Table 2.
Foreground selection for Xa21 and Pi54 among BC1F1 plants using gene-specific markers. The BC1F1 plants were screened through PCR for Xa21 gene using the marker pTA 248 (a) and for Pi54 gene using the marker Pi54 MAS (b). M marker, R recurrent parent (i.e. DRR17B) and D donor parent (i.e. RP-Bio-Patho-2). Arrows indicates plants which were heterozygous (i.e. positive) for both the genes
Evaluation of BB and blast resistance of the selected backcross-derived lines
The selected backcross-derived lines (mentioned above) were evaluated for their resistance to BB and blast under glass house condition. The resistance check Tetep harbouring the blast gene Pi54 showed a blast disease score of 1, whereas the susceptible checks, DRR 17B and HR12, showed a score of 9. The selected six backcross-derived lines showed a very high level of resistance with score of 0 (Fig. 2a). With respect to screening for bacterial blight, the donor RP-Bio-Patho-2 displayed a score of 3, whereas the recurrent parent, DRR17B, showed score of 9. The selected backcross-derived lines showed a very high level of resistance against BB with a score of 3 (Fig. 2b). The details of results of screening of the selected lines against BB and blast are given in Table 1.
Screening of selected BC3F5 plants for blast and bacterial blight resistance. a Screening against rice blast disease through uniform blast nursery (UBN) method revealed that the susceptible check HR12 and recurrent parent DRR17B were highly susceptible to blast disease, while the resistant donor (i.e. check), Tetep, along with the selected backcross-derived lines at BC3F5 generation (viz., IL-1: GP17B112-3-98-118-12-204-8, IL-2: GP17B112-3-98-118-12-204-15, IL-3: GP17B112-3-98-118-12-204-90, IL-4: GP17B 112-3-98-118-12-204-168, IL-5: GP17B112-3-98-118-12-204-234, IL-8: GP17B112-3-98-118-12-204-276) showed high level of resistance. b Screening against bacterial blight disease under glasshouse conditions revealed that the recurrent parent, DRR17B, was highly susceptible to the disease, while the donor parent, RP-Bio-Patho-2, and the selected backcross-derived lines at BC3F5 generation (viz. 1: GP17B112-3-98-118-12-204-8, 2: GP17B112-3-98-118-12-204-15, 3: GP17B112-3-98-118-12-204-90, 4: GP17B 112-3-98-118-12-204-168, 5: GP17B112-3-98-118-12-204-234) were highly resistant to bacterial blight
Selection for agro-morphological and yield characters in backcross-derived lines
The selected six BC3F5 backcross-derived lines showing good level of resistance against BB and blast disease were evaluated for agro-morphological traits. Significant difference was observed in plant height, whereas no such difference was observed in number of productive tillers, panicle length and number of grains per panicle for these lines when compared with the recurrent parent (Table 2). Further except two lines viz. GP17B112-3-98-118-12-204-168 and GP17B112-3-98-118-12-204-276 remaining lines displayed good maintenance ability when test-crossed with IR58025A and DRR17B (Table 2).
Evaluation of heterosis of hybrids derived by crossing improved versions of DRR17B lines with the elite restorer line, RPHR1005
The four elite, improved versions of DRR17B along with DRR17B were crossed with popular restorer line RPHR1005 to assess their heterotic potential. Two improved lines, viz. GP17B112-3-98-118-12-204-15 and GP17B112-3-98-118-12-204-234, showed a higher level of heterosis as compared to the crosses between DRR17B and the restorer line (Table 3). Further, all the hybrids developed from the improved DRR17B lines were resistant to BB and blast and possessed MS grain type (data not shown), indicating that the improved versions have good potential for development of high-yielding, disease-resistant rice hybrids after their conversion to CMS lines.
Discussion
DRR17B is a recently developed maintainer line of rice, possessing stable maintenance ability, with medium duration and having semi-tall plant type, and medium-slender (MS) grain type. As DRR17A and its maintainer parent, DRR17B, are highly susceptible to two important diseases, bacterial blight and blast, the present study was aimed at targeted improvement of the maintainer parent for disease resistance by introgressing a major dominant gene each conferring resistance against BB (i.e. Xa21) and blast (Pi54) through MABB, so that the hybrids developed from the improved version of the maintainer line would also be resistant.
Backcross breeding is one of the most commonly deployed methods for improvement of one or few target traits which are lacking in elite crop varieties (Stoskopf et al. 1993). This is particularly true with respect to parental lines of rice hybrids wherein targeted and precise improvement of one or few traits is usually desired, as gross changes in the genetic background of an elite maintainer or restorer line will make it unfit for use in heterosis breeding (Zhou et al. 2011). Molecular markers can aid backcross breeding, reducing the duration and the cost involved (Collard and Mackill 2008). There are many successful instances wherein marker-assisted backcross breeding has been deployed for targeted improvement of elite varieties and parental lines of rice hybrids. Marker-assisted selection (MAS) has been successfully applied for improving resistance against biotic stresses like bacterial blight, blast and BPH in rice (Narayanan et al. 2002; Joseph et al. 2004; Sundaram et al. 2008, 2009; Basavaraj et al. 2010; Sreewongchai et al. 2010; Hari et al. 2011, 2013; Singh et al. 2012; Zhan et al. 2012; Khanna et al. 2015). Breeders can easily transfer beneficial alleles into other genetic backgrounds, whenever DNA markers tightly linked to the resistance genes have been identified. There are many successful instances wherein marker-assisted backcross breeding has been deployed for targeted improvement of elite varieties and parental lines of rice hybrids. For example, Sundaram et al. (2008, 2009) improved two elite rice varieties, Samba Mahsuri and Triguna, for BB resistance by deploying Xa21, xa13 and xa5 through marker-assisted backcross breeding (MABB). Similarly, Singh et al. (2012) improved an elite basmati restorer line, PRR78, through MABB by introgressing two blast genes Piz-5 and Pi54. Later, Hari et al. (2011, 2013) improved an elite restorer line, KMR3R, and a popular maintainer line, IR58025B, for BB and blast resistance through MAS by combining two dominant resistance genes (Xa21 and Pi54). Recently, Khanna et al. (2015) developed near-isogenic lines (NILs) of Pusa Basmati-1 possessing single major blast resistance genes, Pi54, Pi1, Pita, Pi2, Pib, Pi5 and Pi9, through MABB. In the present study, we used previously reported DNA markers for bacterial blight and blast resistance along with stringent phenotype-based selection for agro-morphological characters to improve an elite maintainer line, i.e. DRR17B. This approach of combining MAS with phenotype-based selection can improve precision in breeding in the initial stages and can reduce costs significantly in the later stages of backcross breeding.
In the present study, we deployed two PCR-based functional markers, viz. pTA248 and Pi54 MAS, for foreground selection of the BB and blast resistance genes, Xa21 and Pi54, respectively. Precise foreground selection is vital for success of MABB (Hospital et al. 1997). As the two markers are functional markers, no recombination can be expected between the marker and the gene, thus ensuring 100 % selection accuracy. The F1 plants and backcross plants were again validated with another marker, RM206, which is reported to be very closely linked to Pi54 (Sharma et al. 2005). No recombination was noticed between Pi54 MAS and RM206, indicating that both the markers are highly useful for tracking the introgression of Pi54 in breeding populations. In addition to the target resistance genes, we deployed molecular markers closely linked to the major fertility restorer genes, Rf3 and Rf4, at BC1F1 generation. This permitted us to identify resistance gene(s)-positive plants which also possessed non-restoring alleles in homozygous condition at BC1F1 generation. This was necessary as the donor parent has been characterized to possess Rf4 through marker analysis (data not shown). In addition to foreground selection for the target genes and non-restoration trait, we also carried out background selection using large number (#12) of parental polymorphic SSR markers located on carrier chromosome (i.e. Chr. 11) and flanking the target resistance genes and a reasonable number of markers (# 4–6) located on non-carrier chromosomes to minimize the linkage drag, thus limiting the number of backcrosses. Earlier, Sundaram et al. (2008) used same strategy while improving the Samba Mahsuri for BB resistance through MABB. Even though the extent of recovery of the recurrent parent genome was considerably lower (Supplementary Table 2) than the expected value (i.e. 75 %) in the first backcross generation, by the third backcross generation, we were able to identify a resistance gene(s)-positive plant possessing ~93.4 % recovery of DRR17B genome, which is nearly equal to the expected value of 93.75 %. Further, only a small region of <2 Mb has been introgressed from the donor parent in the vicinity of Xa21 and Pi54 on chromosome 11 (i.e. target chromosome), and most of the non-target chromosomes have only negligible segments from the donor parent (i.e. RP-Bio-Patho-2). Further, the selected backcross plants also closely resembled DRR17B in terms of most agro-morphological traits and grain quality (Table 2) with introgression line # 2 (GP17B112-3-98-118-12-204-15) possessing maximum recovery of the recurrent parent genome on the target chromosome (Supplementary Figure 2), thus demonstrating the utility of background selection as reported earlier by Sundaram et al. (2008, 2009).
In the early generations of backcrossing, genotypic background analysis was carried out and the plants with maximum recurrent parent genome recovery were used for backcrossing up to BC3F1 generation. From BC3F2 to BC3F4 generations, strict phenotypic selections were followed and those plants which were similar to recurrent parent DRR17B in almost of the key agro-morphological traits (viz., number of days to flowering, number of tillers, grain type and gain number) were selected and advanced further. In BC3F4 generation, six stable lines which were similar to or better than DRR17B were identified. They were then subjected to genotypic background selection with parental polymorphic SSR markers spread all over the 12 chromosomes including markers flanking the gene of interest (Supplementary Figure 2). The process of intensive marker-assisted selection coupled in the initial stages of backcrossing coupled with a stringent phenotypic selection in the later stages resulted in near-complete recovery of the recurrent parent genome in all the six elite backcross-derived improved lines of DRR17B, with only small segments flanking the target genes possessing donor chromosomal introgression.
The advanced backcross-derived improved lines (ABILs) developed in the present study were screened for BB and blast resistance under controlled conditions by artificially inoculating with virulent isolates of respective pathogens. All the six improved ABILs screened showed significantly higher level of resistance to both the diseases compared to the recurrent parent, DRR17B (Table 1). Bacterial blight resistance gene, Xa21, provides a broad-spectrum resistance to majority of the pathotypes in India barring a few reports (Shanti et al. 2010; Sundaram et al. 2008). Xa21 has tremendous potential in BB resistance breeding programme and can be used either singly or in combination with other BB resistance genes (Song et al. 1995; Khush et al. 1990; Wang et al. 1996; Chen et al. 2000; Sundaram et al. 2008). This is evident from several recent reports (Joseph et al. 2004; Hari et al. 2011, 2013) where the gene had been deployed singly in hybrid rice parental lines. As no additional broad-spectrum, dominant BB resistance gene was available, when the study was initiated in 2008, we introgressed only Xa21. However, there are recent reports about identification and fine mapping of at least two broad-spectrum, wild-rice-derived BB resistance genes, Xa33 (Natrajkumar et al. 2012; Shaik et al. 2014) and Xa38 (Bhasin et al. 2012). We are in the process of introgressing Xa33 in the genetic background of improved lines of DRR17B possessing Xa21 and Pi54. Similar to the BB resistance Xa21 gene, the blast resistance gene, Pi54, is known to be a dominant and broad-spectrum and is effective throughout India (Sharma et al. 2005; Rai et al. 2011; Singh et al. 2011; Hari et al. 2013), having deployed in many Indian hybrid rice parental lines and varieties. Even though, owing to the very dynamic nature of these pathogen, it is very difficult to predict the durability of resistance conferred by a single gene (i.e. Pi54), data from All India Coordinated Rice Improvement Project (DRR Progress report, Vol. 2, 2008–2013) clearly indicate that NILs of Samba Mahsuri and Swarna possessing Pi54 are highly effective across the country. In these trials, which were carried out in >25 test centres, the Pi54-containing lines showed resistance reaction against multiple isolates with very low severity index values (2.6–2.7), which was significantly different when compared with the susceptible checks (SI values of 6.8–6.9). This clearly indicates that Pi54 gene is very effective against most of the isolates across the country. However, in order to enhance the spectrum and durability of blast resistance, we have started introgressing an additional broad-spectrum blast resistance gene, Pi2, into DRR17B.
In molecular breeding programmes, breeders generally deploy MAS for improvement of one or few target traits and carry out phenotype-based selection for improvement or retaining certainly key agro-morphological and grain quality characters, for which reliable markers are not available. This strategy was earlier adopted by Joseph et al. (2004), Gopalakrishnan et al. (2008), Sundaram et al. (2008), Singh et al.(2011) and Hari et al. (2011, 2013). Based on the experience gathered from these studies, we adopted MAS for selection of lines possessing Xa21, Pi54, rf3 and rf4 and morphology-based selection for certain key agro-morphological characters like plant height, grain type, number of productive tillers and days to flower among the backcross-derived lines. At BC3F5 generation, we identified six lines (based on visual selection carried out from BC1 generation onwards), viz. GP17B112-3-98-118-12-204-8, GP17B112-3-98-118-12-204-15, GP17B112-3-98-118-12-204-90, GP17B 112-3-98-118-12-204-168, GP17B112-3-98-118-12-204-234 and GP17B112-3-98-118-12-204-276, which were significantly shorter (75–80 cm) than the recurrent parent DRR17B (95 cm). For a CMS (or its maintainer line), possessing a short plant stature is an important morphological character (Virmani and Kumar 2004) as it assists in better seed set, provided that the restorer line is taller than the CMS lines. In addition, two ABILs (GP17B112-3-98-118-12-204-8 and GP17B112-3-98-118-12-204-90) were similar in duration to DRR17B 105 days, while some of the ABILs were significantly early in duration as compared to DRR17B (Table 2). This could be an important trait for development of hybrids in the early or mid-early category.
The ABILs were test-crossed with the WA-CMS line IR58025A to study their stability of sterility maintenance (Table 2). Except two lines, all the other ABILs showed complete sterility, and these are being crossed with DRR17A for line conversion through MAS, a strategy adopted by Hari et al. (2013). When the selected ABILs possessing stable maintenance and medium-slender grain type were crossed with a medium-slender (MS) grain type restorer, RPHR1005 (Ramesha et al. 2010), some of the newly developed hybrids were observed to possess not only medium-slender grain type, but also a higher level of heterosis as compared to the popular hybrid, DRRH3 (Table 3). Further, the experimental hybrids were resistant to BB and blast disease, indicating that high-yielding hybrids possessing MS grain type along with BB and blast resistance could be developed from the improved lines of DRR17B after their line conversion.
All the improved lines showed good level of BB and blast resistance, while the recurrent parent DRR17B was observed to be completely susceptible for both the diseases (Table 1). Among the backcross-derived lines, we made rigorous phenotype-based selections for characters which were equivalent to or better than DRR17B (i.e. more number of grains per panicle, dwarf plant type, and early or late flowering). In this process, we could recover some of the backcross-derived lines possessing lesser plant height as compared to DRR17B (Table 2) more number of gains per panicle and those possessing early flowering plant type as compared to the recurrent parent (Table 2), while retaining the premium medium-slender (i.e. fine) grain type exactly similar to DRR17B, whereas selections made for shorter height than recurrent parent ideal for maintainer line (Table 2). The improved lines also showed good maintainer ability when crossed with WA-CMS line, IR58025A, indicating their equivalence to DRR17B with respect to maintenance ability (Table 2). Interestingly, hybrids derived from two of the improved lines viz., GP17B112-3-98-118-12-204-15 and GP17B112-3-98-118-12-204-234, exhibited more grain yield heterosis when compared to the hybrids derived from crosses with DRR17B (Table 3).
In conclusion, through the present study, we have developed improved versions of the stable maintainer line DRR17B, possessing resistance against BB and blast along with MS grain type, complete sterility maintenance ability and demonstrated the heterotic potential of the improved lines.
References
Abenes MLP, Angeles ER, Khush GS, Huang N (1993) Selection of bacterial blight resistant rice plant in the F2 generation via their linkage to molecular markers. Rice Genet Newsl 10:120–123
Ahmed MI, Siddiq EA (1998) Rice. In: Banga SS, Banga SK (eds) Hybrid cultivar development. Narosa Publishing House, New Delhi, pp 221–256
Balaji SP, Vemireddy LR, Srikanth B, Dharika N, Sambasiva Rao KRS, Hemanth Kishore V, Sundaram RM, Viraktamath BC, Subhakara Rao I, Ramesha MS, Neeraja CN (2012) Fine mapping of Rf3 and Rf4 fertility restorer loci of WA-CMS of rice (Oryza sativa L.) and validation of the developed marker system for identification of restorer lines. Euphytica 187:421–435
Ballini E, Morel JB, Droc G, Price A, Courtois B, Notteghem JL, Tharreau DA (2008) A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microbe Interact 217:859–868
Basavaraj SH, Singh VK, Singh Singh A, Singh A, Anand D, Yadav S, Ellur RK, Singh D, KrishnanS Gopala, Nagarajan M, Mohapatra T, Prabhu KV, Singh AK (2010) Marker-assisted improvement of bacterial blight resistance in parental lines of Pusa RH10, a superfine grain aromatic rice hybrid. Mol Breed 26:293–305
Bhasin H, Bhatia Dharminder, Raghuvanshi Saurabh, Jagjit S, Lore Gurpreet K, Sahi Baljit Kaur, Vikal Yogesh, Singh Kuldeep (2012) New PCR-based sequence-tagged site marker for bacterial blight resistance gene Xa38 of rice. Mol Breed 30:607–611
Cheema KK, Grewal NK, Vikal Y, Sharma R, Lore JS, Das A, Bhatia D, Mahajan R, Gupta V, Bharaj TS, Singh K (2008) A novel bacterial blight resistance gene from Oryza nivara mapped to 38 Kb region on chromosome 4 and transferred to Oryza sativa L. Genet Res 90:1–11
Chen SX, Lin H, Xu CG, Zhang Q (2000) Improvement of bacterial blight resistance of ‘Minghui 63’, an elite restorer line of hybrid rice by molecular marker-assisted selection. Crop Sci 40:239–244
Chen SC, Xu G, Lin XH, Zhang Q (2001) Improving bacterial blight resistance of 6078, an elite restorer line of hybrid rice by molecular marker-aided selection. Plant Breed 120:133–137
Collard BCY, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Phil Trans R Soc B 363:557–572
Davierwala AP, Reddy APK, Lagu MD, Ranjekar PK, Gupta VS (2001) Marker assisted selection of bacterial blight resistance genes in rice. Biochem Genet 39:261–278
DRR progress report vol. 2 (2008–2013) All India coordinated rice improvement programme (ICAR). Directorate of Rice Research, Rajendranagar, Hyderabad, India
Gopalakrishnan S, Sharma RK, Anand Rajkumar K, Joseph M, Singh VP, Singh AK, Bhat KV, Singh NK, Mohapatra T (2008) Integrating marker assisted background analysis with foreground selection for identification of superior bacterial blight resistant recombinants in Basmati rice. Plant Breed 127:131–139
Hari Y, Srinivasarao K, Viraktamath BC, Hariprasad AS, Laha GS, Ilyas Ahmed M, Natarajkumar P, Ramesha MS, Neeraja CN, Balachandran SM, Shobha Rani N, Balaji Suresh P, Sujatha K, Pandey M, Ashok Reddy G, Madhav MS, Sundaram RM (2011) Marker-assisted improvement of a stable restorer line, KMR-3Rand its derived hybrid KRH2 for bacterial blight resistance and grain quality. Plant Breed 130:608–616
Hari Y, Srinivasa Rao K, Viraktamath BC, Hariprasad AS, Laha GS, Ahmed M, Nataraj Kumar P, Sujatha K, Srinivasprasad MS, Rani NS, Balachandran SM, Kemparaju S, Mohan KM, Sama VSAK, Shaik H, Balachiranjeevi CH, Pranathi K, Reddy GA, Madhav MS, Sundaram RM (2013) Marker-assisted introgression of bacterial blight and blast resistance into IR 58025B, an elite maintainer line of rice. J Plant Breed 132:586–594
Hospital F, Moreau L, Lacoudre F, Charcosset A, Gallais A (1997) More on the efficiency of marker-assisted selection. Theor Appl Genet 95:1181–1189
Huang N, Angeles ER, Domingo J, Magpantay G, Singh S, Zhang Q, Kumaravadivel N, Bennett J, Khush GS (1997) Pyramiding of bacterial resistance genes in rice: marker aided selection using RFLP and PCR. Theor Appl Genet 95:313–320
Hulbert SH, Webb CA, Smith SM, Sun Q (2001) Resistance gene complexes: evolution and utilization. Annu Rev Phytopathol 39:285–312
International Rice Research Institute (1996) Standard evaluation system for rice, 4th edn. International Rice Research Institute, Manila
Joseph M, Gopalakrishnan S, Sharma RK (2004) Combining bacterial blight resistance and basmati quality characteristics by phenotypic and molecular marker-assisted selection in rice. Mol Breed 13:377–387
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Khanna A, Sharma V, Ellur RK, Shikari AB, Gopala Krishnan S, Singh UD, Prakash G, Sharma TR, Rathour R, Variar M, Prashanthi SK, Nagarajan M, Vinod KK, Bhowmick PK, Singh NK, Prabhu KV, Singh BD, Singh AK (2015) Development and evaluation of near-isogenic lines for major blast resistance gene(s) in Basmati rice. Theor Appl Genet. doi:10.1007/s00122-015-2502-4
Khush GS, Jena KK (2009) Current status and future prospects for research on blast resistance in rice (Oryza sativa L.). In: Advances in genetics, genomics and control of rice blast disease. pp 1–10
Khush GS, Bacalango E, Ogawa T (1990) A new gene for resistance to bacterial blight from O. longistaminata. Rice Genet Newsl 7:121–122
Koide Y, Kobayashi N, Xu D, Fukuta Y (2009) Resistance genes and DNA selection markers for blast disease in rice Oryza sativa L. Jpn Agric Res 43:255–280
Laha GS, Reddy CS, Krishnaveni D, Sundaram RM, Srinivas PM, Ram T, Muralidharan K, Viraktamath BC (2009) Bacterial Blight of Rice and its Management. In: DRR Technical Bulletin No. 41. Directorate of Rice Research (ICAR), Hyderabad, India, p 37
Leung HJ, Liu WB, Bustaman M, Sridhar R, Singh K, Redona E, Quang VD, Zheng K, Bernardo M, Wang G, Leach J, Choi IR, Cruz CV (2004) Sustainable disease resistance in rice: current and future strategies. In: New directions for a diverse planet. 4th International Crop Sci Cong Sep–Oct 2004, Brisbane, Australia.Published on CDROM. Web site www.cropscience.org.au
Liyong CZ, Jie-yun Shou-jiang, Xiao-deng Z, Kang-le Z, Shi-hua C (2003) Hybrid rice resistant to bacterial leaf blight developed by marker assisted selection. Rice Sci 11:68–70
Madhan Mohan K (2011) Molecular characterization of pathogenic variability of Pyricularia grisea (Rice Blast fungus). Dissertation, Jawaharlal Nehru Technological University Hyderabad, Andhra Pradesh, India
Mew TW (1987) Current status and future prospects of research on bacterial blight of rice. Annu Rev Phytopathol 25:359–382
Narayanan NN, Baisakh N, Vera Cruz N, Gnananmanickam SS, Datta K, Datta SK (2002) Molecular breeding for the development of blast and bacterial blight resistance in rice cv. IR50. Crop Sci 42:2072–2079
Natrajkumar P, Sujatha K, Laha GS, Srinivasa Rao K, Mishra B, Viraktamath BC, Hari Y, Reddy CS, Balachandran SM, Ram T, Sheshu Madhav M, Shobha Rani N, Neeraja CN, Ashok Reddy G, Shaik H, Sundaram RM (2012) Identification and fine-mapping of Xa33, a novel gene for resistance to Xanthomonas oryzae pv. oryzae. Phytopathology 102:222–228
Pandey MK, Shobha Rani N, Sundaram RM, Laha GS, Madhav MS, Srinivasa Rao K, Injey Sudharshan, Yadla Hari, Varaprasad GS, Subba Rao LV, Kota Suneetha, Sivaranjani AKP, Viraktamath BC (2013) Improvement of two traditional Basmati rice varieties 3 for bacterial blight resistance and plant stature through 4 morphological and marker-assisted selection. Mol Breed 31:239–246
Perez ML, Redona DE, Mendioro SM, Verna Cruz MC, Leung H (2008) Introgression of Xa4, Xa7 and Xa21 for resistance to bacterial blight in thermo sensitive genetic male sterile rice (Oryza sativa L.) for the development of two line hybrids. Euphytica 164:627–636
Perumalsamy S, Bharani M, Sudha M, Nagarajan P, Arul L, Saraswathi R, Balasubramanian P, Ramalingam J (2010) Functional marker-assisted selection for bacterial leaf blight resistance genes in rice (Oryza sativa L.). Plant Breed 129:400–406
Rai AK, Kumar SP, Gupta SK, Gautam N, Singh NK, Sharma TR (2011) Functional complementation of rice blast resistance gene Pikh (Pi54) conferring resistance to diverse strains of Magnaporthe oryzae. J Plant Bioche Biotechnol 20:55–65
Ramesha MS, Viraktamath BC, Ilyas Ahmed M (2010) RPHR-1005 (IC569494; INGR09085), a Paddy (Oryza sativa) germplasm with broad spectrum of fertility restoration and with good plant type, high rate of pollen production and good combining ability BPT5204 derivative with short slender grain, lower panicle position and prominent top leaves. Indian J Plant Genet Resour 23(3):327–354
Ramkumar G, Srinivasarao K, Madhan Mohan K, Sudarshan I, Sivaranjani AKP, Gopalakrishna K, Neeraja CN, Balachandran SM, Sundaram RM, Prasad MS, Shobha Rani N, Rama Prasad AM, Viraktamath BC, Madhav MS (2010) Development and validation of functional marker targeting an In Del in the major rice blast disease resistance gene Pi54 (Pikh). Mol Breed 27:129–135
Rao KK, Lakshminarasu M, Jena KK (2002) DNA markers and marker-assisted breeding for durable resistance to bacterial blight disease in rice. Biotechnol Adv 20:33–47
Ronald PC, Albano B, Tabien R, Abenes L, Wu K, Mc Couch S, Tanksley S (1992) Genetic and physical analysis of the rice bacterial blight resistance locus, Xa21. Mol Gen Genet 235:113–120
Sanchez AC, Brar DS, Huang N, Khush GS (2000) Sequence tagged site markers-assisted selection for three bacterial blight resistance genes in rice. Crop Sci 40:792–797
Shaik H, Yugander A, Balachiranjeevi CH, Pranathi K, Anila M, Mahadevaswamy HK, Kousik BVN, Dilip Kumar T, Ashok Reddy G, Bhaskar S, Abhilash Kumar V, Harika G, Rekha G, Laha GS, Viraktamath BC, Balachandran SM, Neeraja CN, Sheshu Madhav M, Mangrauthia SK, Bhadana VP, Sundaram RM (2014) Development of durable bacterial blight resistant lines of samba mahsuri possessing Xa33, Xa21, Xa13 & Xa5. Progr Res 9:1224–1227
Shanti LM, Mohan Kumar Varma C, Premalatha P, Lalitha Devi G, Zher Usha, Freeman Wayne (2010) Understanding the bacterial blight pathogen combining pathotyping and molecular marker studies. Int J Plant Pathol 1(2):58–68
Sharma TR, Madhav MS, Singh BK, Shanker P, Jana TK, Dalal T, Pandit A, Singh A, Gaikwad K, Upreti HC, Singh NK (2005) High resolution mapping, cloning and molecular characterization of the Pik h gene of rice, which confers resistance to M. grisea. Mol Genet Genomics 274:569–578
Sharma TR, Rai AK, Gupta SK, Singh NK (2010) Broad spectrum blast resistance gene Pikh cloned from the rice line Tetep designated as Pi54. J Plant Biochem Biotechnol 19:87–89
Sharma TR, Rai AK, Gupta SK, Vijayan J, Devanna BN, Ray S (2012) Rice blast management through host resistance: retrospect and prospects. Agric Res 1:37–52
Singh S, Sidhu JS, Huang N, Vikal Y, Li Z, Brar DS, Dhaliwal HS, Khush GS (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker- assisted selection into indica rice cultivar PR-106. Theor Appl Genet 102:1011–1015
Singh AK, Gopala Krishnan S, Singh VP, Prabhu KV, Mohapatra T, Singh NK, Sharma T, Nagarajan M, Vinod KK, Singh D, Singh UD, Chander S, Atwal SS, Seth R, Singh VK, Ellur RK, Singh A, Anand D, Khanna A, Yadav S, Goel N, Singh A, Shikari AB, Singh A, Marathi B (2011) Marker assisted selection: a paradigm shift in Basmati breeding. Indian J Genet Plant Breed 71:1–9
Singh VK, Singh A, Singh SP, Ellur RK, Choudhary V, Singh D, Gopala Krishnan S, Nagarajan M, Vinod KK, Singh UD, Prashanthi SK, Agrawal PK, Bhatt JC, Mohapatra T, Prabhu KV, Sarkel S, Rathore R, Singh AK (2012) Incorporation of blast resistance into “PRR78”, an elite Basmati rice restorer line, through marker assisted backcross breeding. Field Crops Res 12:8–16
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald PC (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Sci 270:1804–1806
Sonti RV (1998) Bacterial leaf blight of rice: new insights from molecular genetics. Curr Sci 74:206–212
Sreewongchai T, Toojinda T, Thanintorn N, Kosawang C, Vanavichit A, Tharreau D, Sirithunya P (2010) Development of elite indica rice lines with wide spectrum of resistance to Thai blast isolates by pyramiding multiple resistance QTLs. Plant Breed 129:176–180
Srinivas Prasad M, Shesu Madhav M, Laha GS, Ladha Lakshmi D, Krishnaveni D, Mangrauthia SK, Balachandran SM, Sundaram RM, Arunakanthi B, Madhan Mohan K, Ratna Madhavi K, Kumar V, Virakatamath BC (2011) Rice blast disease and its management. Technical Bulletin No. 57. Directorate of Rice Research (ICAR), Hyderabad
Srinivasan B, Gnanamanickam S (2005) Identification of a new source of resistance in wild rice, Oryza rufipogon to bacterial blight of rice caused by Indian strains of Xanthomonas oryzae pv. Oryzae. Curr Sci 88:25
Stoskopf NC, Tomes DT, Christie BR (1993) Plant breeding: theory and practice. West view Press Inc., Oxford, San Francisco
Sujatha K, Natarajkumar P, Laha GS, Mishra B, Srinivasarao K, Viraktamath BC, Kirti PB, Hari Y, Balachandran SM, Rajendrakumar P, Ram T, Hajira SK, Madhav MS, Neeraja CN, Sundaram RM (2011) Inheritance of bacterial blight resistance in the rice cultivar Ajaya and high-resolution mapping of a major QTL associated with resistance. Genet Res Camb 93:397–408
Sundaram RM, Vishnupriya MR, Biradar SK, Laha GS, Reddy GA, Rani NS, Sarma NP, Sonti RV (2008) Marker assisted introgression of bacterial blight resistance in Samba Mahsuri, an elite indica rice variety. Euphytica 160:411–422
Sundaram RM, Vishnupriya MR, Laha GS, Shobha Rani N, SrinivasRao P, Balachandaran SM, Ashok Reddy G, Sarma NP, Shonti RV (2009) Introduction of bacterial blight resistance into Triguna, a high yielding, mid-early duration rice variety. Biotechnol J 4:400–407
Sundaram RM, Chatterjee S, Oliva R, Laha GS, Leach JE, Sonti RV, Cruz CV (2014) Update on bacterial blight of rice: fourth international conference on bacterial blight rice 7: 12
Virmani SS (1996) Hybrid rice. Adv Agron 57:377–462
Virmani SS, Kumar I (2004) Development and use of hybrid rice technology to increase rice productivity in the tropics. Int Rice Res Notes 29:10–20
Wang GL, Song WY, Ruan DL, Sideris S, Ronald PC (1996) The cloned Xa21 confers resistance to multiple Xanthomonas oryzae pv. Oryzae isolates in transgenic plants. Mol Plant Microbiol Interact 9:850–855
Yoshimura S, Yoshimura A, Iwata N, Mc Couch SR, Abenes MN, Baraoidan MR, Mew TW, Nelson RJ (1995) Tagging and combining bacterial blight resistance genes using RAPD and RFLP markers. Mol Breed 1:375–387
Zhan XD, Zhou HP, Chai RY, Zhuang JY, Cheng SH, Li CY (2012) Breeding of R8012, a Rice restorer line resistant to blast and bacterial blight through marker-assisted selection. Rice Sci 19(1):29–35
Zhang G, Angeles ER, Abenes MLP, Khush GS, Huang N (1996) RAPD and RFLP mapping of the bacterial blight resistance gene xa13 in rice. Theor Appl Genet 93:65–70
Zheng K, Subudhi PK, Domingo J, Maopanty G, Huang N (1995) Rapid DNA isolation for marker assisted selection in rice breeding. Rice Genet Newsl 12:255–258
Zhou YL, Uzokwe VNE, Zhang CH, Cheng LR, Wang L, Chen K, Gao X, Sun Q, ChenY ZhuJJ, Zhang LH, Ali Q, Xu JL, Li ZK (2011) Improvement of bacterial blight resistance of hybrid rice in China using the Xa23 gene derived from wild rice (Oryza rufipogon). Field Crop Res 30:637–644
Acknowledgments
The authors would like to acknowledge the funding support provided by the Department of Biotechnology (DBT), Government of India, for execution of the research study through the Grant # BT/PR11705/AGR/02/646/2008. The authors also thank Project Director, ICAR-Indian Institute of Rice Research, for providing all the necessary facilities.
Author information
Authors and Affiliations
Corresponding author
Additional information
CH Balachiranjeevi, S. Bhaskar Naik, V. Abhilash and R. M. Sundaram have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Balachiranjeevi, C., Bhaskar, N.S., Abhilash, V. et al. Marker-assisted introgression of bacterial blight and blast resistance into DRR17B, an elite, fine-grain type maintainer line of rice. Mol Breeding 35, 151 (2015). https://doi.org/10.1007/s11032-015-0348-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0348-8