Abstract
The allelic diversity of a collection of 4046 rice accessions was assessed using 15 neutral SSR markers distributed throughout the genome. A total of 482 alleles were detected; the average allelic richness was 32.1 alleles per locus. Using a heuristic approach, an allele-mining set was successfully developed on the basis of SSR marker data. 162 accessions of the allele-mining set, accounting for about 4.0% of the entire collection, captured all of the alleles (482) retained in the entire collection, which showed 100% coverage of alleles with minimum redundancy. As a result of validation of this heuristic approach using another 14 SSR markers associated with starch, 70% of the total alleles and 83% of the restricted alleles (allele frequency > 0.05%) were captured in this allele-mining set. The results showed that the heuristic approach meets the condition as an allele-mining set even when applied to another specific set of markers related to starch synthesis in the same entire and allele-mining set. The newly developed methodology for developing allele-mining sets can be used in other crop species. By retaining all alleles of the entire collection, this allele-mining set will be useful for future studies on introducing unused useful alleles into elite rice varieties by breeders in the post-genomic era.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
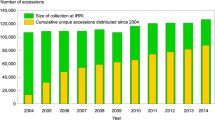
Rice (Oryza sativa L.), as a model cereal species, is one of the most important crops in the world and provides the main energy resource for more than half the world’s population (Yu et al. 2002). The survival of mankind in the future will depend to a great extent on the quantity and diversity of germplasm collections. Therefore, many countries and organizations have established hundreds of genebanks and have conserved millions of crop germplasm resources. For instance, the International Rice Genebank at the International Rice Research Institute (IRRI) maintains a collection of more than 108,925 rice accessions (http://www.cgiar.org/pdf/newsroom_svalbard_irri_shipment.pdf); there are also many other large rice collections in countries such as China and India. With the rapid increase in the number of accessions contained in crop germplasm collections, many genebanks face the problems of redundant resources and the cost of maintaining these collections, which may be an obstacle for their full exploitation, evaluation and utilization (Holden 1984). For the convenience of management, research and application, Frankel and Brown (1984) proposed the concept of the core set. The design of the core set should include the maximum possible genetic diversity contained in the entire collection with a minimum of repetitiveness. The information obtained from such a core set can aid in the judicious use of the entire collection. To date, most core sets have been developed on the basis of passport data giving the geographical origin, morphological and phenotypic traits, and biochemical or molecular markers in many crops (Perry et al. 1991; Joe and Orlando 1996; Hokanson et al. 1998; Ortiz et al. 1998; Huaman et al. 1999; Parsons et al. 1999; Chavarriaga-Aguirre et al. 1999; Marita et al. 2000; Upadhyaya and Ortiz 2001; Chandra et al. 2002). But most traits of crop varieties are quantitatively under the control of multiple genes that are easily influenced by environmental conditions, while the molecular markers reflect changes that have occurred at the DNA level but not necessarily expressed in the phenotype of the organism (Tanksley and McCouch 1997; Li et al. 2004).
A good core set should minimize redundant entries and should be sufficiently large to provide reliable conclusions for the entire collection (Brown 1989). To establish a core set, the sampling proportion and variation representation of the entire collection are important in the construction of the core set in order to retain the greatest degree of genetic diversity in it. There are many different methodologies available to build sampling strategies. These methods include simple random sampling and stratified random sampling (Peeters and Martinelli 1989; Crossa et al. 1995; Charmet and Balfourier 1995; Rincon et al. 1996; Chandra et al. 2002; Franco et al. 2003), and other sophisticated methods. For stratified random sampling, Brown (1989) proposed three allocation methods including constant (C strategy), proportional (P strategy), and logarithmic proportional (L strategy). Franco et al. (2005) proposed to use Gower’s distance between accessions within each cluster (D method) as the allocation criteria. Li et al. (2004) developed a core collection using the adjusted unbiased prediction (AUP) method based on the predicted genotypic value of rice. The clustering algorithm has now also been used as an important tool to reduce redundancy and select core sets within groups in germplasm research (van Hintum et al. 1995; Zewdie et al. 2004; Upadhyaya et al. 2006; Mosjidis and Klingler 2006), Hu et al. (2000) developed an stepwise clustering method for sampling; the least distance stepwise sampling (LDSS) has been proved to be a valid method for eliminating the influence of different clustering methods (Wang et al. 2007). A method for determining sample sizes based on genetic distances was introduced by Franco et al. (2005). Jansen and van Hintum (2007) further developed a novel sampling method for obtaining a core set using genetic distances.
Recently, molecular genetic markers have been widely used to characterize genebank collections (Bretting and Widrlechner 1995; van Hintum and van Treuren 2002). Schoen and Brown (1993) addressed the issue of how to use genetic markers to sample collections of wild crops while maximizing allelic richness. The H strategy seeks to maximize the total number of alleles in the core collection by sampling accessions from groups in proportion to their within-group genetic diversity, while the M (maximization) strategy maximizes the number of observed alleles at each marker locus. Bataillon et al. (1996) found by computer simulation that the M strategy was more effective for retaining widespread and low-frequency neutral alleles than the other sampling strategies. Gouesnard et al. (2001) developed the MSTRAT algorithm by implementing the M strategy for selecting accessions. These different approaches have been compared by Franco et al. (2006). McKhann et al. (2004), Ronfort et al. (2006) and Cunff et al. (2008) developed a nested genetic core collection using the M strategy. Kim et al. (2007) developed PowerCore software: a program applying the advanced M strategy with a heuristic search for establishing a core set.
Many genebanks all over the world contain untapped resources of distinct alleles which will remain hidden unless efforts are initiated to screen these alleles for their potential use and function; the process is known as “allele mining”, which will contribute to discovering and exploiting the hidden diversity for many complex traits (Varshney et al. 2005). The deployment of an allele-mining set, a kind of mini core set for finding new alleles from selected entries using genomic tools, has been an area of much interest for researchers, especially those working in the field of allele mining. A representative set of rice for allele mining, as the core set described, should best represent the diversity present in the entire genebank. With the rice genome sequence available (Collard et al. 2008), allele mining provides the avenue for the validation of specific gene(s) responsible for a particular trait and mining of the most favorable alleles from the rice genebank. Thus, in the post-genomics era, allele mining in a large collection of accessions will contribute to genomics research for crop improvement (Varshney et al. 2005). These developments will be a boon for plant breeders who are trying to increase yields and create new varieties which are resistant to diseases, pests, drought and salinity and/or with improved nutritional quality (Latha et al. 2004).
The objective of this study was to develop an allele-mining set, a kind of mini core set, using a heuristic approach and SSR genotype data from an entire collection of 4046 rice accessions conserved in the National Genebank of Rural Development Administration, Republic of Korea (RDA-genebank) and to evaluate the allele-mining set by applying another set of SSR markers related to starch synthesis. And the availability of the allele-mining set was also tested.
Materials and methods
Plant materials
The RDA-genebank holds 25,604 accessions of rice (Oryza sativa L.) from 60 countries (http://genebank.rda.go.kr/). From this collection, 4,046 accessions (approximately 15.8% of the total collection), including the introduced varieties, breeding lines and varieties, weedy accessions, and the Korean landraces, were selected based on the passport data in this study (Table 1). The IRRI set, a super mini-set for the DNA polymorphism test for developing a DNA bank at the IRRI genebank, was included to separate one from the others. Of these, 1,065 accessions originated from 71 countries and 2,981 accessions were from the Republic of Korea. A description of the entire rice collection used in this study is shown in Table 1.
SSR genotyping
The 29 SSR markers, including 15 neutral SSRs and 14 SSRs associated with starch synthesis in rice, were analyzed in this study. All these SSR markers were obtained from GRAMENE (http://www.gramene.org/). Markers were chosen according to their location on the rice genetic map, which gives good coverage of the whole genome map, and their suitability for high throughput genotyping. A three-primer system (Schuelke 2000), including a universal M13 oligonucleotide (TGTAAAACGACGGCCAGT) labeled with one of the fluorescent dyes 6-FAM, NED, and HEX, allowing PCR products to be triplexed during electrophoresis, a special forward primer composed by the concatenation of the M13 oligonucleotide, and the specific forward primer, were used for SSR PCR amplification. DNA amplifications were performed using an MJ Research PTC-100 96 Plus thermal cycler. PCR reactions were carried out in a volume of 15 μL containing 10 ng of total DNA, 10× PCR buffer, 0.25 mM of each dNTP, 8 pmol of each primer, and 1 U of Taq polymerase using a touchdown procedure. Information on primer sequences and PCR amplification conditions for each set of primers are available at http://www.gramene.org/. SSR alleles were resolved on the ABI PRISM 3100 DNA sequencer (Applied Biosystems, Foster City, CA, USA) using GENESCAN 3.7 software and sized precisely using GeneScan 500 ROX (6-carbon-X-rhodamine) molecular size standards (35–500 bp) with GENOTYPER 3.7 software (Applied Biosystems).
Development of an allele-mining set
The advanced M strategy by a modified heuristic algorithm implemented in the PowerCore software by Kim et al. (2007) was used to develop the allele-mining set. The PowerCore software maximizes the number of alleles with the least redundancy in the SSR data set (Kim et al. 2007), In the PowerCore software, the A* algorithm, a heuristic algorithm that finds the optimum path from the initial to the final stages, was used:
Here, with g(n) as the number of accessions inserted into the frequency table and h(n) as the maximum number of empty cells within each column, this algorithm expands the paths that have the lowest value for g(n) + h(n), where g(n) is the cost for the path from the initial state to the current node and h(n) serves as an estimate of the cost for the cheapest path from that node to the designated node. When expanding each of the steps, the sum of g(n) and h(n) will be evaluated and the accession with the lowest value will be chosen. If h(n) is admissible without overestimating the costs of reaching the goal, then A* will always find an optimal solution (http://genebank.rda.go.kr/PowerCore/) (Kim et al. 2007).
The efficiency of the sampling strategy was assessed by comparing the total number of alleles captured using a modified heuristic algorithm in samples of increasing size to the number of alleles captured in random sampling and stratified random sampling according to geographic region and variety type from the same entire collection (Brown 1995; Qiu et al. 2003; Yan et al. 2007). Fifty independent samplings were made in each case (q = 50).
Validating the core collection
Use of the allele-mining set may improve the efficiency of germplasm evaluation by reducing the number of accessions evaluated to increase the probability of finding genes of interest. To see the effectiveness of the allele-mining set in this study, another set of SSR markers was tested on the entire and the allele-mining set. The set of 14 SSR markers was selected according to their association with starch synthesis. The coverage of alleles was compared between the entire accessions and the allele-mining set constructed by 15 neutral SSR markers.
Data analysis
The total number of alleles per locus, the number of rare alleles per locus (i.e. alleles with frequency lower than 5%), the number of unique alleles per locus (alleles occurring in only one accession), Shannon and Weaver diversity index (I) (1949), Nei’s gene diversity index (H) (Nei 1973), and the polymorphism information content (PIC) per locus were calculated for the entire and the core accessions using PowerCore (Kim et al. 2007) and Powermaker 3.25 software (Liu and Muse 2005) based on Rogers’ distance (Rogers 1972). All the indices were calculated independently in both the entire and the allele-mining set to determine whether the diversity for each locus was retained in the allele-mining set. Frequency distributions for each locus were determined using Microsoft Excel 2007 software. Statistical analysis was conducted using the univariate and correlation procedures of SPSS 14.0 (http://www.spss.com/) and Statistica 7.0 (http://www.statsoft.com/) statistical software.
Nei’s gene diversity (H) was calculated based on the formula
where n i is the allele frequency at the ith locus, n is the number of alleles at this locus and N is the total number of accessions.
The Shannon–Weaver diversity index (I) as presented was estimated using
where p i is the frequency of the phenotypic class.
The PIC for each marker was calculated based on the formula
where P is the relative frequency of the jth pattern for SSR marker i (Botstein et al. 1980).
Results
Allele mining of 4046 rice accessions
The allelic diversity of a collection of 4046 rice accessions was assessed using 15 neutral SSRs distributed throughout the genome and the resulting statistics are summarized in Table 2. A total of 482 alleles were detected ranging from 15 (RM246) to 61 (RM206) with an average allelic richness of 32.1 alleles per locus. The total number of rare alleles (398) represented about 82.6% of the total number of alleles, showing that most alleles are at low frequency (Supplementary Fig. 1). The mean Shannon–Weaver diversity index and Nei’s gene diversity were 2.716 and 0.932, respectively. The PIC ranged from 0.7338 to 0.9333 with an average of 0.8448 (Table 2).
Development of an allele-mining set
The 482 alleles detected at 15 neutral SSR loci were used to develop the allele-mining set using the PowerCore software (http://genebank.rda.go.kr/PowerCore). The basis of developing an allele-mining set using PowerCore is the nominalization of variables, leading to a decrease in the number of accessions in the allele-mining set, which was considered necessary in performing the heuristic search through its evaluation function using the given data (Kim et al. 2007). Figure 1 showed that, in this case, the sampling efficiency (i.e. the ability to capture allelic diversity) implementing a modified heuristic algorithm was always better than other strategies. Furthermore, the relative efficiency of this advanced M (maximization) strategy was highest for smaller allele-mining set samples; for instance, the heuristic approach outperformed a stratified random sampling by about 50% when sample sizes were in the range of 50–150 (i.e. an allele-mining set size is 1.2–3.7% of the entire sample size).
Number of alleles captured with respect to accession sample size in three sampling strategies generated using PowerCore software. HCC, SRC and RCC represent the total number of alleles captured using a modified heuristic algorithm, stratified random sampling and random sampling method, respectively. Redundancy curves obtained using PowerCore software (fifty independent samplings)
It was found that the allele-mining set (162 accessions) (Supplementary Table 1), accounting for about 4.0% of the entire collection, captured all of the alleles (482) of the markers presented in the entire collection, which showed 100% coverage of alleles with minimum redundancy (Table 2). Compared with other conventional sampling methods, the heuristic approach showed the highest capturing efficiency (Table 5).
Genetic diversity of the allele-mining set
To fully realize the advantages of an allele-mining set, the allele-mining set should include most of the genetic diversity in the entire collection and be closely correlated with the entire collection (Yan et al. 2007). The allele-mining set of this study represented all SSR alleles of the entire rice collection. As shown in Table 2, the correlation coefficients (r) of mean diversity index between the allele-mining set and the entire collection were highly significant (Table 3; Supplementary Fig. 2). In this allele-mining set, all the alleles were covered and highly significant correlations were recorded for all parameters studied, which indicated that the allele-mining set effectively represented the genetic diversity of the entire collection. We also compared allele frequencies of the SSR markers in the allele-mining set with the frequencies observed in the entire collection. The frequency of alleles between them was very significantly correlated (r = 0.87; P < 0.01) (Fig. 2), indicating that not only were the same alleles represented but also similar frequencies were represented.
Validation of the allele-mining set
The construction of a so-called “allele-mining set” from a large germplasm collection is a situation where allelic richness is a relevant measure of diversity (Schoen and Brown 1993; Bataillon et al. 1996), because as many alleles as possible should be retained in the allele-mining set, where they would be available for phenotypic screening and breeding programs. To validate the heuristic approach, the same accessions in this study were assessed in a set of 14 additional SSR markers related to starch synthesis between the same entire and allele-mining sets. These markers are different from the markers used to build the allele-mining set. Statistics describing the allelic diversity of these 4046 accessions for 14 additional SSR markers are summarized in Table 4. 214 alleles were detected with the 14 SSR markers in the entire collection. The number of alleles per locus ranged from 4 to 34 with an average of 15.3. Compared with the neutral SSRs, the SSRs related to quality had smaller polymorphism (Fig. 3) and the distributions of frequency of alleles per locus were different between them (Fig. 4). For these 14 markers, PIC ranged from 0.3423 to 0.8923, with an average of 0.6249. Table 5 summarizes the total number of alleles detected for the two types of markers. For the second set of 14 markers, 70% of the alleles observed in the 4046 accessions were captured in the allele-mining set. For association studies, Malysheva-Otto et al. (2006) thought that rare alleles occurring in more than 0.5% of investigated accessions should be referred to as widespread or often occurring alleles, since markers with low allele frequencies would need to have a very strong effect to be detected. There are 29 unique alleles in the second set of SSRs (only 3 unique alleles were kept in the allele-mining set), but the allele-mining set represented 83% diversity of the 176 restricted alleles (frequencies > 0.05%, corresponding to two out of the entire accessions of the set) retained in the entire accessions (Cunff et al. 2008). Even if not all alleles of useful genes were captured, the heuristic approach also does better than other sampling strategies (Table 5). Given the nature of the allele-mining set, it is impossible to guarantee the complete capture of all alleles for each gene (McKhann et al. 2004). However, the allele-mining set using the heuristic approach here eliminated the redundancy in the rice collection and succeeded in capturing most of the alleles in some genes of interest.
Discussion
Studies on allelic diversity have been proved to be fruitful in understanding the genetic basis of complex traits (Szalma et al. 2005). The sequencing of the complete genome of rice makes it possible to access all the genes of this species and increases the chances of exploiting the natural genetic diversity through association genetics (Varshney et al. 2005; Collard et al. 2008). However, our basic knowledge of the extent of allelic variation within the species is still not sufficient. Considering the huge numbers of accessions that are held collectively by genebanks, germplasm collections are thought to harbor a wealth of undisclosed allelic variants. Mining alleles will improve the efficiency of conservation and use of genetic resources (Kresovich et al. 2002; Varshney et al. 2005). The allelic richness of 32.1 observed in our study was much higher than previously reported by Garris et al. (2005) (mean 11.8) using 169 SSRs and 234 rice accessions, and Ebana et al. (2008) (mean 7.7) using 23 SSRs and 236 Japanese rice landrace accessions, indicating higher levels of allelic diversity. We also compared the alleles within different populations: the allelic richness was weedy>introduced>landrace>bred>IRRI accessions (Supplementary Table 2). After comparing allelic richness with the respective index of genetic diversity, we found that allelic richness was significantly associated with the genetic diversity index, the correlation coefficients (r) between allelic richness and Shannon–Weaver diversity index, Nei’s gene diversity and PIC were 0.903, 0.748 and 0.560, respectively. Furthermore, the high proportion (82.6%) of rare alleles found in our sample indicated that, conversely, there exist many informative alleles to be mined in the rice collection (Table 2).
To devise plant breeding strategies for crop improvement, a breeder would ideally like to know the relative value of all alleles for genes of interest in the primary germplasm, an unlikely prospect. However, information can be gathered by establishing the allele-mining set (Varshney et al. 2005). So the development of an allele-mining set, which represents the genetic diversity of a crop with minimal redundancy and increases utility of the collection as a whole, is especially important as the funding for germplasm collections decreases (Marita et al. 2000). Many core sets were successfully developed after Frankel proposed the theory of the core set in 1984, but the selection of an appropriate sampling strategy is still important in the construction of a core set. In this paper, we successfully developed an allele-mining set by a heuristic approach with least redundancy in rice. The heuristic method implemented in PowerCore software captured 100% of the allelic diversity existing in the entire collection (Kim et al. 2007). There are now many methods for developing an allele-mining set in different plants; Franco et al. (2006) demonstrated that one of the best strategies is the M (for maximization) strategy developed by Schoen and Brown (1993), maximizing allelic richness at each marker locus. Kim et al. (2007) also found that MSTRAT was the best method compared with the other conventional methods in the rice accessions, but the coverage rate was only 88.9% for SSRs. In this paper, the allele-mining set developed using PowerCore showed the highest diversity and coverage compared to those allele-mining sets developed using other sampling strategies (Table 5). The basis for the development of an allele-mining set using PowerCore is the nominalization of categorical variables, a step efficiently decreasing the number of accessions selected while capturing the maximum variation and minimizing redundancy. This lies in its capability to select entries without the comparison of relative characteristics within accessions. Instead it fills all diversity cells with the least number of entries taking into account all the possible combinations of alleles that exist through an advanced maximization strategy (M strategy) (Kim et al. 2007).
Developing an allele-mining set has been proposed as a means of increasing the use of germplasm more economically (Frankel 1984). Brown et al. (1987) recommended that the number of collections in the core set should account for 5–10% of the base collection, and that the core set should represent at least 70% of the genetic diversity in the base collection. Diwan et al. (1995) indicated that core set sampling should always be greater than 10%. Van Hintum (1995) suggested that the sampling proportion should depend on the particular objective of the core set and should vary between 5 and 20% of the base collection. In establishing a core set of rice germplasm, Li et al. (2000) found that 5% of the base collection represented 96% of the phenotypic variation, Yan et al. (2007) represented approximately 10% of the 18,412 accessions with 88% certainty, Ebana et al. (2008) established that a 20% core set represented 87.5% diversity using SSR markers. Therefore, ascertaining the best threshold value for group numbers in an allele-mining set has not yet been fully resolved. The current study showed that the heuristic method implemented in PowerCore software successfully captured all of the alleles existing in the entire collection, with a threshold value of only about 4% having the highest capturing efficiency (Table 5). Agrama et al. (2009) established that the 12% mini-core represented 100% diversity on the basis of 26 phenotypic traits and 70 SSR markers using the same software. From the above results, we found that in order to retain maximum genetic diversity in the core set, with the increase of the SSR markers, especially the allele number, the size of the core set will also increase correspondingly. Therefore, the 208 accessions of the allele-mining set were developed if we used all 29 SSRs to construct the core set.
The allele-mining set here included 100% of the 482 observed SSR alleles; among them, germplasm from Korea predominated in the allele-mining set (76 entries) due to a large number of germplasm lines (2981 accessions), followed by germplasm from the IRRI (16 entries) where many entries were acquired from the IRRI collection, followed by China having 12 entries. All germplasm types, such as introduced accessions, breeding lines, weedy types, and the Korean landraces, were included in the allele-mining set. The entire accessions originated from 72 countries, but only 29 different geographical origins were represented in the allele-mining set (Table 1); this might be because the definition of the true geographical origin of rice is sometimes difficult due to many human migration events. This could be explained by differences in allelic richness between germplasm from different geographical areas and by the status of the accessions (Supplementary Table 1). For instance, landrace accessions have more alleles than bred accessions. From Table 1 and Supplementary Table 1, we also found no relationship between allelic richness and sample number in the allele-mining set. Some were under or over-represented compared to the total sample; for example, the total number of alleles in IRRI was 205, 16 accessions were sampled from a total of 55 accessions, a high proportion (30.19%) of accessions were kept in the allele-mining set, but the correlation coefficient (r = 0.983) of the mean allele number per sample between the entire and the allele-mining set was significant at the P = 0.01 level, giving a reasonable explanation for the phenomenon. The result was very valuable for sampling in constructing an allele-mining set, because the higher the mean allele number per sample in the group, the more accessions in the allele-mining set. In addition, the IRRI accessions are a super mini-set for DNA polymorphism at the IRRI genebank, and the high proportion in the allele-mining set showed indirectly that our heuristic approach is reasonable, feasible and reliable.
In order to assess the robustness of the allele-mining set, the genetic diversity index is used in genetic studies as a convenient measure of both allelic richness and allelic evenness. Although significant correlation coefficients (r = 0.725–0.860) were found between the entire and the allele-mining set (Table 3), the total genetic diversity revealed by Shannon–Weaver diversity index (I) and Nei’s gene diversity (H) was higher in the entire collection than in the allele-mining set while PIC in the allele-mining set was higher than in the entire collection, due to the fact that they were of unequal size. So sometimes the use of indices such as I and H may be disputed (Hennink and Zeven 1991). Hennink and Zeven (1991) proposed relative indices, defined as H′ = H mean/H max and I′ = I mean/I max, respectively. By comparison, we found that H′, I′ and PIC′ (=PICmean/PICmax) in the allele-mining set was similar to the entire set (Supplementary Fig. 3), indicating that H′, I′ and PIC′ indices of genetic diversity can be better used as parameters evaluating the quality of the allele-mining set.
The property of starch in rice is a very important determinant for rice quality. The method used to build the allele-mining set was validated by a second set of independent markers associated with starch synthesis in rice on a larger sample of accessions. As shown in Fig. 3, the 14 SSR markers related to starch synthesis generally showed lower diversity indices (allelic richness and all genetic diversity indices) than the 15 neutral SSRs. This could be explained by the fact that SSRs related to starch are probably more conserved than the DNA segments containing neutral SSRs. So, with the lower allelic richness and fewer rare alleles, the use of such a set of SSR markers to validate the method may have diminished the effectiveness of the validation of the allele-mining set (Balfourier et al. 2007). Maximizing the diversity of a first set of markers (15 neutral SSRs) at the same time should maximize useful gene diversity, here expressed by a second set of markers. A complete cross-validation of the method required the total sample of 4046 accessions to be tested with the second set of markers (Table 5). Cunff et al. (2008) thought that estimating the unlinked diversity within the entire collection would have been very fastidious; here the allele-mining set represented 70% of the total alleles and 83% of the restricted alleles (alleles with frequencies higher than 0.05%) observed in the 4046 accessions. This meets the accepted standard of an allele-mining set (10% of the base accessions representing more than 70% of the genetic diversity). Moreover, the allele coverage per locus is higher than with other sampling strategies, with the highest efficiency for small size cores. The results showed that the allele-mining set based on 15 neutral SSRs minimized redundancy and successfully captured the majority of the target gene alleles. Therefore, this heuristic approach can be used as an allele-mining set to uncover the loci with useful alleles and greatly facilitate the identification of useful genes, and can incorporate them into advanced breeding materials for testing and further selection (Tanksley and McCouch 1997).
In conclusion, an allele-mining set of 162 accessions (only about 4% of the entire collection) was successfully developed by a heuristic approach based on SSR markers using PowerCore software. This allele-mining set captured all of the alleles present in the entire collection and will be useful for future studies of rice gene mining to introduce unused useful alleles into elite rice varieties by breeders. Moreover, the newly presented methodology for an allele-mining set with the least allelic redundancy and maximum allelic diversity from a large germplasm collection of rice can be used in other crop species in the post-genomic era.
References
Agrama HA, Yan WG, Lee F, Fjellstrom R, Chen MH, Jia M, McClung A (2009) Genetic assessment of a mini-core subset developed from the USDA Rice Genebank. Crop Sci 49:1336–1346
Balfourier F, Roussel V, Strelchenko P, Exbrayat-Vinson F, Sourdille P, Boutet G, Koenig J, Ravel C, Mitrofanova O, Beckert M, Charmet G (2007) A worldwide bread wheat core collection arrayed in a 384-well plate. Theor Appl Genet 114:1265–1275
Bataillon TL, David JL, Schoen DJ (1996) Neutral genetic markers and conservation genetics: simulated germplasm collections. Genetics 144:409–417
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Bretting PK, Widrlechner MP (1995) Genetic markers and plant genetic resource management. Plant Breed Rev 31:11–86
Brown AHD (1989) Core collections: a practical approach to genetic resources management. Genome 31:818–824
Brown AHD (1995) The core collection at the crossroads. In: Hodgkin T, Brown AHD, van Hintum TJL (eds) Core collections of plant genetic resources. Wiley, Chichester, pp 3–19
Brown AHD, Grace JP, Speer SS (1987) Designation of a core collection of perennial glycine. Soybean Genet Newsletter 14:59–70
Chandra S, Huaman Z, Hari Krishna S, Ortiz R (2002) Optimal sampling strategy and core collection size of Andean tetraploid potato based on isozyme data—a simulation study. Theor Appl Genet 104:1325–1334
Charmet G, Balfourier F (1995) The use of geo statistics for sampling a core collection of perennial ryegrass population. Genet Resour Crop Evol 42:303–309
Chavarriaga-Aguirre P, Maya MM, Tohme J, Duque MC, Iglesias C, Bonierbale MW, Kresovich S, Kochert G (1999) Using microsatellites, isozymes and AFLPs to evaluate genetic diversity and redundancy in the cassava core collection and to assess the usefulness of DNA-based markers to maintain germplasm collections. Mol Breed 5:263–273
Collard BCY, Cruz CMV, McNally KL, Virk PS, Mackill DJ (2008) Rice molecular breeding laboratories in the genomics era: current status and future considerations. Int J Plant Genomics 2008:1–25
Crossa J, Basford K, Taba S, DeLacy I, Silva E (1995) Three-mode analysis of maize using morphological and agronomic attributes measured in multilocation Trials. Crop Sci 35:1483–1491
Cunff LL, Fournier-Level A, Laucou V, Vezzulli S, Lacombe T, Adam-Blondon AF, Boursiquot JM, Patrice T (2008) Construction of nested genetic core collections to optimize the exploitation of natural diversity in Vitis vinifera L. subsp. Sativa. BMC Plant Biol 8:31
Diwan N, McIntosh MS, Bauchan GR (1995) Methods of developing a core collection of annual Medicago species. Theor Appl Genet 90:755–761
Ebana K, Kojima Y, Fukuoka S, Nagamine T, Kawase M (2008) Development of mini core collection of Japanese rice landrace. Breed Sci 58:281–291
Franco J, Crossa J, Taba S, Shands H (2003) A multivariate method for classifying cultivars and studying group × environment × trait interaction. Crop Sci 43:1249–1258
Franco J, Crossa J, Taba S, Shands H (2005) A sampling strategy for conserving genetic diversity when forming core subsets. Crop Sci 45:1035–1044
Franco J, Crossa J, Warburton ML, Taba S (2006) Sampling strategies for conserving maize diversity when forming core subsets using genetic markers. Crop Sci 46:854–864
Frankel OH (1984) Genetic perspectives of germplasm conservation. In: Arber W, limensee K, Peacock WJ, Starlinger P (eds) Genetic manipulation: impact on man and society. Cambridge University Press, Cambridge, pp 161–171
Frankel OH, Brown AHD (1984) Plant genetic resources today: a critical appraisal. In: Holden JHW, Williams JT (eds) Crop genetic resources: conservation and evaluation. Allen & Unwin Ltd, London, pp 249–257
Garris AJ, Tai TH, Coburn J, Kresovich S, McCouch S (2005) Genetic structure and diversity in Oryza sativa L. Genetics 169:1631–1638
Gouesnard B, Bataillon TM, Decoux G, Rozale C, Schoen DJ, David JL (2001) Mstrat: an algorithm for building germplasm core collections by maximizing allelic or phenotypic richness. J Hered 92:93–94
Hennink S, Zeven AC (1991) The interpretation of Nei and Shannon-Weaver within population variation indices. Euphytica 51:235–240
Hokanson SC, Szewc-McFadden AK, Lamboy WF, McFerson JR (1998) Microsatellite (SSR) markers reveal genetic identities, genetic diversity and relationships in a Malus domestica borkh core subset collection. Theor Appl Genet 97:671–683
Holden JHW (1984) The second ten years. In: Holden JHW, Williams JT (eds) Crop genetic resources: conservation and evaluation. Allen and Unwin, Winchester, pp 277–285
Hu J, Zhu J, Xu HM (2000) Methods of constructing core collections by stepwise clustering with three sampling strategies based on the genotypic values of crops. Theor Appl Genet 101:264–268
Huaman Z, Aguilar C, Ortiz R (1999) Selecting a Peruvian sweet potato core collection on the basis of morphological, ecogeographical, and disease and pest reaction data. Theor Appl Genet 98:840–844
Jansen J, van Hintum ThJL (2007) Genetic distance sampling: a novel sampling method for obtaining core collections using genetic distances with an application to cultivated lettuce. Theor Appl Genet 114:421–428
Joe T, Orlando GD (1996) AFLP analysis of gene pools of a wild bean core collection. Crop Sci 36:1375–1384
Kim KW, Chung HK, Cho GT, Ma KH, Chandrabalan D, Gwag JG, Kim TS, Cho EG, Park YJ (2007) PowerCore: a program applying the advanced M strategy with a heuristic search for establishing allele mining sets. Bioinformatics 23:2155–2162
Kresovich S, Luongo AJ, Schloss SJ (2002) Mining the gold: finding allelic variants for improved crop conservation and use. In: Engels JMM, Rao VR, Brown AHD, Jackson MT (eds) Managing plant genetic diversity. CABI, Wallingford, pp 379–386
Latha R, Rubia L, Bennett J, Swaminathan MS (2004) Allele mining for stress tolerance genes in Oryza species and related germplasm. Mol Biotechnol 27:101–108
Li ZC, Zhang HL, Zeng YW, Yang ZY, Shen SQ, Sun CQ, Wang XK (2000) Study on sampling schemes of core collection of local varieties of rice in Yunnan, China. Sci Agri Sin 33:1–7
Li CT, Shi CH, Wu JG, Xu HM, Zhang HZ, Ren YL (2004) Methods of developing core collections based on the predicted genotypic value of rice (Oryza sativa L.). Theor Appl Genet 108:1172–1176
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Malysheva-Otto LV, Ganal MW, Röder MS (2006) Analysis of molecular diversity, population structure and linkage disequilibrium in a worldwide survey of cultivated barley germplasm (Hordeum vulgare L.). BMC Genet 7:6
Marita JM, Rodriguez JM, Nienhuis J (2000) Development of an algorithm identifying maximally diverse core collections. Genet Resour Crop Evol 47:515–526
McKhann HI, Camilleri C, Berard A, Bataillon T, David JL, Reboud X, Corre VL, Caloustian C, Gut IG, Brunel D (2004) Nested core collections maximizing genetic diversity in Arabidopsis thaliana. Plant J 38:193–202
Mosjidis JA, Klingler KA (2006) Genetic diversity in the core subset of the US red clover germplasm. Crop Sci 46:758–762
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Ortiz R, Ruiz-Tapia EN, Mujica-Sanchez A (1998) Sampling strategy for a core collection of Peruvian quinoa germplasm. Theor Appl Genet 96:475–483
Parsons BJ, Newbury HJ, Jackson MT, Ford-Lloyd BV (1999) The genetic structure and conservation of aus, aman and boro rices from Bangladesh. Genet Resour Crop Evol 46:587–598
Peeters JP, Martinelli JA (1989) Hierarchical cluster analyses as a tool to manage variation in germplasm collections. Theor Appl Genet 78:42–48
Perry MC, Mclntosh MS, Stoner AK (1991) Geographical patterns of variation in the USDA soybean germplasm collection: II. allozyme frequencies. Crop Sci 31:1356–1360
Qiu LJ, Cao YS, Chang RZ, Zhou XA, Wang GX, Sun JY, Xie H, Zhang B, Li XH, Xu ZY (2003) Establishment of Chinese soybean (G. max) core collection. I. Sampling strategy. Sci Agri Sin 36:1442–1449
Rincon F, Johnson B, Crossa J, Taba S (1996) Cluster analysis, and approach to sampling variability in maize accessions. Maydica 41:307–316
Rogers JS (1972) Measures of genetic similarity and genetic distance. Stud Genet VII Univ Tex Publ 7213:145–153
Ronfort J, Bataillon T, Santoni S, Delalande M, David JL, Prosperi JM (2006) Microsatellite diversity and broad scale geographic structure in a model legume: building a set of nested core collection for studying naturally occurring variation in Medicago truncatula. BMC Plant Biol 6:28
Schoen DJ, Brown AHD (1993) Conservation of allelic richness in wild crop relatives is aided by assessment of genetic markers. Proc Natl Acad Sci USA 90:10623–10627
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Szalma SJ, Buckler ES, Snook ME, McMullen MD (2005) Association analysis of candidate genes for maysin and chlorogenic acid accumulation in maize silks. Theor Appl Genet 110:1324–1333
Tanksley SD, McCouch SR (1997) Seed bank and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Upadhyaya HD, Ortiz R (2001) A mini core subset for capturing diversity and promoting utilization of chickpea genetic resources in crop improvement. Theor Appl Genet 102:1292–1298
Upadhyaya HD, Gowda CLL, Pundir RPS, Reddy VG, Singh S (2006) Development of core subset of finger millet germplasm using geographical origin and data on 14 quantitative traits. Genet Resour Crop Evol 53:679–685
van Hintum TJL (1995) Hierarchical approaches to the analysis of genetic diversity in crop plants. In: Hodgkin T, Brown AHD, van Hintum TJL (eds) Core collections of plant genetic resources. Wiley, Chichester, pp 23–34
van Hintum ThJL, van Treuren R (2002) Molecular markers: tools to improve genebank efficiency. Cell Mol Biol Lett 7:737–744
van Hintum ThJL, von Bothmer R, Visser DL (1995) Sampling strategies for composing a core collection of cultivated barley (Hordeum vulgare s. Iat.) collected in China. Hereditas 122:7–15
Varshney RK, Andreas GA, Sorrells ME (2005) Genomics-assisted breeding for crop improvement. Trends Plant Sci 10:621–630
Wang JC, Hu J, Xu HM, Zhang S (2007) A strategy on constructing core collections by least distance stepwise sampling. Theor Appl Genet 115:1–8
Yan WG, Ruter JN, Bryant RJ, Bockelman HE, Fjellstrom RG, Chen MH, Tai TH, McClung AM (2007) Development and evaluation of a core subset of the USDA rice germplasm collection. Crop Sci 47:869–876
Yu J, Hu S, Wang J, Wong GK, Li S et al (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296:79–92
Zewdie Y, Tong NK, Bosland P (2004) Establishing a core collection of capsicum using a cluster analysis with enlightened selection of accessions. Genet Resour Crop Evol 51:147–151
Acknowledgments
This study was supported by Biogreen 21 project (Grant 20080401034058) of the Rural Development Administration (RDA) and a grant (Code 200803101010415) from the National Academy of Agricultural Science, RDA, Republic of Korea. This research was also supported by the 2008 KU Brain Pool of Konkuk University for Dr. Zhao Weiguo.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, W., Cho, GT., Ma, KH. et al. Development of an allele-mining set in rice using a heuristic algorithm and SSR genotype data with least redundancy for the post-genomic era. Mol Breeding 26, 639–651 (2010). https://doi.org/10.1007/s11032-010-9400-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-010-9400-x