Abstract
Heading date (HD) is a key trait for the adaptation of rice cultivar to a specific growing region. Here, we report conventional and marker-assisted breeding strategies using genetic information related to the determination of HD, where the breeding objectives were to avoid the delayed heading common in indica × japonica hybrids, to increase the efficiency in selecting hybrid rice combinations having a suitable growth duration, and to develop cultivars with target growth duration by quantitative trait locus (QTL) pyramiding. The allelic constitution at the major HD loci was determined for a set of 109 leading Chinese rice cultivars by crossing them with HD tester lines. It was shown that the late heading in indica × japonica hybrids can be overcome by replacing the strong photoperiod-sensitivity allele Se-1 n with Se-1 e. A breeding strategy to enable the selection of hybrid combinations with suitable growth duration was proposed, based on HD genotypic information in rice. Meanwhile, a QTL analysis for HD was conducted over five years based on a recombinant inbred line population, derived from two parents Asominori (japonica) and IR24 (indica). Four QTLs, located on chromosomes 2, 3, 6, and 8, respectively, could be detected in all five years, indicating they were stably expressed QTL. According to this QTL information, and taking Asominori as an example, the HD genotypes for improving the growth duration were designed, and the best breeding selection schemes were determined by use of a genetic breeding simulation tool. Results obtained in this study demonstrate that genetic information related to HD can make a significant contribution to rice breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Heading date (HD) is one of the most important traits for rice to adapt to different areas and cropping seasons. Time taken for a cultivar to head is mainly determined by a combination of, or interaction between, basic vegetative growth period (BVG) and its photoperiod-sensitivity (PS) and temperature-sensitivity (TS). Among numerous HD genes that have been identified, Ef-1 plays a major role in controlling the BVG by accelerating the switch to reproductive growth independently of photoperiod, and partially counteracting the effects of the PS gene under long-day conditions (Tsai 1986; Kinoshita 1995; Sato et al. 1988; Nishida et al. 2002; Xu et al. 2006). Se-1 and E 1 are known as strong PS genes, proven to be the most ubiquitous genes controlling HD (Ichitani et al. 1997, 1998; Okumoto et al. 1996). Many other genes also affect HD through interaction with Se-1 or E 1 . For example, i-Se-1 can inhibit the effect of Se-1, producing an early heading type even under long-day conditions (Ohshima et al. 1993; Ohshima and Kikuchi 1994; Luo et al. 2002). Co-presence of E 1 and Se-1 produces a very strong level of PS, manifested as a very late heading type under long-day conditions (Cai et al. 1987; Luo et al. 2002).
To date, a growing number of QTL for HD has emerged from analysis of various mapping populations (Li et al. 1995; Yano et al. 1997; Lin et al. 1998, 2002; Maheswaran et al. 2000; Yamamoto et al. 2000; Yu et al. 2002), and some (Hd1, Hd3a, Hd6, Ehd1 and Ghd7) have been isolated by map-based cloning (Yano et al. 2000; Takahashi et al. 2001; Kojima et al. 2002; Doi et al. 2004; Xue et al. 2008). The coincidence of both the phenotypic effect and chromosomal location suggests that Se-1 is either identical or allelic to Hd1, Ef-1 to Ehd1, and i-Se-1 to hd2 (Yano et al. 2000; Doi et al. 2004; Ohshima et al. 1993; Ohshima and Kikuchi 1994; Luo et al. 2002; Xu et al. 2006).
HD is genetically diversified in rice, which provides a wealth of variation for rice to adapt in a vast range of growing areas worldwide. However, it also increases the breeding complexity for a specific area with optimum growth duration (Luo et al. 2001). Late heading in indica × japonica hybrids limits the exploitation of substantial inter-subspecies heterosis. Conflict between early heading and high yielding always appears in hybrid rice breeding (Deng et al. 2001). Along with a growing number of sterile and restorer lines with different HDs, breeding hybrids for certain areas was even harder than before. To overcome these difficulties, breeders have to blindly make many crosses for repeated testing in the field, which results in tremendous work and low breeding efficiency. If an optimum HD for a certain area can be designed using known genetic information, breeding will become more directed and efficient than the conventional way (Peleman and van der Voort 2003; Wang et al. 2005; Wan 2006; Wang and Pfeiffer 2007). In order to avoid the delayed heading common in indica × japonica hybrids, to increase selection efficiency in hybrid rice, and to obtain lines with an HD suited to a particular rice-growing region, in this paper we first describe the HD genetic make-up of a sample of 109 leading rice cultivars from across China, and then report use of a QTL approach to analyze the genetic determination of HD in a recombinant inbred population. On the basis of this genotypic information, more efficient strategies were devised for breeding rice cultivars with optimum HD.
Materials and methods
Materials
The following genetic stocks, cultivars, hybrids and populations were employed:
-
HD near-isogenic lines (NILs) EG0, EG1, ER, LR, T65m, and T65Ebm (Yamagata et al. 1986; Ichitani et al. 1997, 1998; Inoue et al. 1998), and two HD QTL near-isogenic lines NIL(Hd1) and NIL(Hd2) (Yamamoto et al. 1998; Lin et al. 2000) (Table 1);
-
109 leading cultivars in production that are representative of different ecological regions in China (Supplementary Table 1) and cv. Nipponbare; F1 hybrids between the 109 leading cultivars and the genetic stock lines;
-
71 recombinant inbred lines (RILs) derived from Asominori (japonica) × IR24(indica) (Tsunematsu et al. 1996); and
-
66 chromosome segment substitution lines (CSSLs) including segments of IR24 in Asominori background (Kubo et al. 1999).
Determination of HD genotype for each cultivar
Stock lines EG0 and EG1 are a pair NILs differing only at the E 1 locus (EG0 is E 1 , and EG1 is e 1 ) (Ichitani et al. 1997, 1998) (Table 1). The PS allele E 1 is dominant over the e 1 (Okumoto et al. 1992, 1996). The HD of EG1 was 19.2 days later than that of EG0 under natural long-day conditions at Nanjing (latitude 32°N), when photoperiods in May, June, July, and August were 13.7 hours light (hL)/10.3 hours dark (hD), 14.2 hL/9.8 hD, 14.0 hL/10.0 hD, and 13.2 hL/10.8 hD, respectively. If the HD of EG1 × “cultivar A” F1 hybrid is about 19.2 days later than that of EG0 × cultivar A hybrid, cultivar A was considered to carry the no-PS allele e 1 . If HDs are similar in two F1 hybrids, cultivar A should be deemed to carry the PS allele E 1 . If the HD of EG1 × cultivar A F1 hybrid is later but not 19.2 days later than that of EG0 × cultivar A F1 hybrid, cultivar A should be assumed to carry a PS allele at E 1 locus, but the PS of this allele should be weaker than E 1 allele. The weak PS allele at E 1 is named E t 1 . NILs ER and LR differ only at Se-1 (LR is Se-1 u, and ER is Se-1 e). Nipponbare carries the PS allele Se-1 n, whereas NIL(Hd1) has hd1 (synonymous with Se-1 e) (Yano et al. 1997, 2000; Lin et al. 2000; Yamamoto et al. 1998) (Table 1). Se-1 u and Se-1 n are both dominant over Se-1 e, and the PS imposed by Se-1 u is greater than that by Se-1 n (Yokoo and Kikuchi 1977). In Nanjing, the HD of ER was 22.0 days earlier than that of LR, and NIL(Hd1) was 9.5 days earlier than that of Nipponbare. The genotype of each of the 109 leading cultivars at Se-1 was determined from the HD of their F1 hybrids with ER, LR, Nipponbare, and NIL(Hd1), respectively. Similarly, the allelic state at Ef-1 was identified by comparing the F1 HD of each cultivar with NILs T65m (ef-1) and T65Ebm (Ef-1), respectively. The Hd2 genotypes was determined by comparison of the F1 HD of each cultivar with Nipponbare (Hd2) and NIL(Hd2) (hd2), respectively. For each of these heading comparisons, ten parental and ten F1 plants were grown in the field from 2000 to 2006 at Nanjing. Sowing was around May 15 and seedlings were transplanted about June 15 at a density of 13.3 cm × 26.7 cm. Crop management followed commercial rice-production practices.
HD QTL analysis for designing rice cultivars with target growth duration
The linkage map was built from the Asominori/IR24 RIL population genotyped using 375 RFLP loci that cover 1275.3 cM, with an average inter-marker interval of 3.4 cM (Tsunematsu et al. 1996). Additive HD QTLs were detected by inclusive composite interval mapping (ICIM) using QTL IciMapping v2.1 software (Li et al. 2007). In the first step of stepwise regression of ICIM, the P value for entering variables (PIN) was set at 0.01 and for removing variables (POUT) was set at 0.02 to select the significant markers; while in the second step, a threshold LOD of 2.5 was used to declare the significant QTL. The 66 CSSLs were selected from 268 BC3F1 plants by a whole-genome survey at 116 RFLP loci and nominated as CSSL1–CSSL66, which represented the whole IR24 genome (Kubo et al. 1999). The CSSLs carrying the QTL of HD were used to design rice cultivars with target growth duration. The target genotypes were designed based on breeding goals and the QTL information. To obtain the target genotypes by QTL pyramiding, various breeding schemes were compared using the software QuLine (Wang et al. 2003, 2004, 2007), and the best breeding scheme was selected.
RILs and CSSLs were planted in Nanjing from 2003 to 2007. The time of sowing and transplant and the plant-density were same as those of the leading cultivars. Also, crop management followed commercial rice-production practices.
Measurement of HD and HD heterosis
HD of each line was measured by the number of days from seeding to heading when the leading panicle emerged about 1 cm beyond the leaf sheath of its flag leaf. The HD heterosis of each hybrid was calculated as:
where F1, P 1, and P 2 represented the HD of F1 hybrid, its male parent, and female parent, respectively.
Results
HD genotypes of leading cultivars and their distribution in China
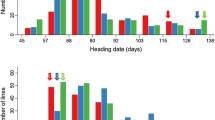
Genotypes of the 109 leading Chinese rice cultivars on the major HD gene loci E 1 , Se-1, Ef-1, and Hd2 were determined, and are presented in Supplementary Table 1. Distribution of the HD genotypes over geographic regions and cropping seasons is summarized in Table 2. This showed that the dominant allele at Ef-1 determining early heading widely distributed among cultivars adapted to a wide range of ecological regions. Japonica cultivars grown in northeast and northwest China carried either one or no PS-determining dominant allele at E 1 and Se-1, and some carried the recessive allele hd2. Most japonica cultivars from north China and central China had the PS allele at either E 1 or Se-1, while those from southwest China included either both or one of the PS alleles at these loci. Cultivars adapted to higher altitudes in southwest China carried the dominant allele Ef-1 for early heading, while those from low altitudes had the recessive allele ef-1 for late heading. In the middle-lower regions of the Yangtze River and south China, early-season indica cultivars had either one PS allele or none at E 1 and Se-1; middle-season ones had either one or two; and most late-season ones had both of the PS alleles. Some middle-season cultivars also carried the late-maturing allele ef-1 (Table 2; Fig. 1). Distribution pattern of the PS alleles demonstrated that the PS was gradually strengthened with decrease of latitude and elevation of rice growing areas (Fig. 1). These results on HD genotypes distribution in different ecological regions and cropping systems provided important information for subsequently genotype-based breeding.
Manipulation of PS genes to make indica × japonica hybrids head properly
Late heading among indica × japonica hybrids is mainly because both parents carry PS alleles at one of the two independent major PS genes (such as E 1 and Se-1) (Cai et al. 1987; Luo et al. 2002). Thus, any of three requirements has to be satisfied in selecting parents for indica × japonica hybrid rice in order to achieve proper heading. Both parents have none of the PS alleles, or the same PS-determining allelic constitution at one of the PS alleles, or independent but non-complementary PS alleles. However, the F1 hybrids of japonica Nipponbare and LR with indica rice cultivars identically and strongly exhibited late heading, indicated by their mid-parent heterosis between 4.1 and 26.2% (Table 3). This could be explained by the non-allelic complementary effects between strong PS genes in these hybrids. Japonica Nipponbare and LR carry the strong PS gene Se-1 n or Se-1 u whereas the indica cultivars have the PS-determining allele at E 1 (Table 3; Supplementary Table 1).
Therefore, in order to make indica × japonica hybrids head properly, the non-allelic complementary effects between E 1 and Se-1 n or Se-1 u need to be avoided. Either the E 1 allele in indica cultivars (such as Minghui63) should be replaced by the non-PS allele e 1 , or Se-1 n/Se-1 u in japonica parent should be replaced by the non-PS allele Se-1 e. The replacement of PS genes has proven to be effective in breeding practice using successive backcrossing with molecular marker-assisted selection based on mapping information of the target genes. According to the fine mapping of Se-1(Hd1), the Se-1 n allele in Nipponbare has been substituted by Se-1 e allele in the BC4F2, and the photoperiod sensitivity becomes weak in the resulting near isogenic line NIL(Hd1) (Lin et al. 2000). In the same way, replacement of the strong Se-1 u in LR with the weak Se-1 e has resulted in another near isogenic line ER (Yamagata et al. 1986). We crossed both the near isogenic lines with indica cultivars and the HD of these F1 hybrids was similar to that of the mid-parent value, demonstrated by their mid-parent heterosis ranging from −13.2 to 6.9% (Table 3). Thus, breeding based on prior knowledge of HD genotype should be effective for avoiding late heading in indica × japonica hybrids.
Genotype design and HD prediction in hybrid rice breeding
Early heading and high yielding are often paradoxical and the HD of hybrids is not known until they are grown out, which takes an entire season in the breeding program (Deng et al. 2001). Based on the genetic effects of major HD genes and the HD genotypes widely utilized in hybrid rice parents, we have designed genotypes of hybrids suitable for photo-temperature conditions in different cropping regions of China (Table 4). These genotypes provide direction in parental selection for hybrid breeding in a target region. Obviously, the HD of derivative hybrids is predictable, so that breeders could pay more attention to yield than heading objective. The shift of attention should help better balance early heading and high yielding and improve efficiency in hybrid rice breeding.
Here, we want to obtain a middle-season indica hybrid suitable for the Yangtze River region, and south and southwestern China. According to local climatic conditions (temperature and photoperiod), this hybrid rice should bear weak PS, a short basic vegetative phase, and moderate growth duration. These requirements could be met by type IV in Table 4, and the recommended genotypes were e 1 e 1 Se-1 e Se-1 e Ef-1_Hd2_, e 1 e 1 Se-1 n _Ef-1_hd2hd2, E 1 _Se-1 e Se-1 e Ef-1_hd2hd2, and E 1 _Se-1 n _Ef-1_hd2hd2. If E 1 _Se-1 n _Ef-1_hd2hd2 is taken as the target genotype, HD genotypes of their derivative male sterile/restorer lines should be either E 1 E 1 Se-1 e Se-1 e Ef-1Ef-1hd2hd2/e 1 e 1 Se-1 n Se-1 n ef-1ef-1hd2hd2 or E 1 E 1 Se-1 n Se-1 n Ef-1Ef-1hd2hd2/e 1 e 1 Se-1 e Se-1 e ef-1ef-1hd2hd2. For example, two elite middle-season hybrid cultivars Shanyou63 and Liangyoupeijiu are widely grown in these regions. Male sterile/restorer lines are Zhenshan 97A/Minghui 63 for Shanyou63 and PeiAi 64S/93-11 for Liangyoupeijiu. HD genotypes of the parental lines are known (Supplementary Table 1). Genetic combination of both parents resulted in an HD genotype E 1 _Se-1 n _Ef-1_hd2hd2 for Shanyou63 and E 1 _Se-1 e Se-1 e Ef-1_hd2hd2 for Liangyoupeijiu (Table 5). The resultant genotypes fit perfectly into type IV for middle-season rice in vast areas of China. These results validated the feasibility and effectiveness of genotype design and HD prediction in breeding hybrid rice.
Breeding strategies based on HD QTL information
Breeding will become more powerful and effective if HD genotype can be engineered for a cultivar to fit in a specific environment. Japonica cultivar Asominori heads at about 88 days in Nanjing, and it would be desirable for this cultivar either to have a earlier HD in order to be grown as a late double-season crop, or to have a later HD in order to be grown as a single-season intermediate maturity crop in Nanjing. We conducted a QTL analysis for HD over five years using a RIL population derived from Asominori (japonica) and IR24 (indica) (Table 6). A total of seven QTLs (located on chromosomes 2, 3, 6, 8, and 12) were identified, for which accountable phenotypic variances ranged from 5.09 to 70.43%. Four of these, qDTH-2, qDTH-3b, qDTH-6, and qDTH-8 were detected in all five years, indicating they were stably expressed for HD in multiple environments and considered to be suitable for gene design. The HDs of CSSLs containing the IR24 allele at qDTH-6 and qDTH-8 were, respectively, ~6 days and ~8 days earlier than for Asominori in Nanjing. In contrast, those CSSLs including the IR24 allele at qDTH-2 and qDTH-3b headed ~7 days and ~8 days later than Asominori. The CSSLs with the IR24 alleles at both qDTH-2 and qDTH-3b flowered ~12 days later than Asominori (Fig. 2). Thus, to achieve later heading, the IR24 alleles at qDTH-2 and qDTH-3b should be introduced into Asominori; for earlier heading, the IR24 alleles at qDTH-6 and qDTH-8 need to be transferred. Our approach was to use recurrent marker-assisted backcrossing to obtain the desired genotypes.
A designed genotype from heading date QTL pyramiding for HD. The shaded area represents genome segments from the donor parent IR24, and the unshaded area represents the genetic background of Asominori. Markers from M2-5 to M8-5 are RFLP loci were used to track each IR24 segments. M2-7, M3-3, M6-3 and M8-2 were the nearest markers of qDTH-2, qDTH-3b, qDTH-6, and qDTH-8, respectively
Two target genotypes were designed in the Asominori background: the first (TG1) was to introduce the IR24 alleles at both qDTH-6 and qDTH-8 for earlier heading, and the second (TG2) was to introduce the IR24 alleles at qDTH-2 and qDTH-3b for later heading (Fig. 2).
According to the graphical genotypes of the CSSLs, the best strategy to achieve TG1 could be marker-assisted selection among the progenies of CSSL47 × CSSL57. TG2 could be achieved by progeny selection assisted by molecular markers from either crossing CSSL16 with CSSL20 or backcrossing CSSL23 with Asominori (Fig. 2). If the two target genotypes are expected in F4, there are many schemes for marker-assisted selection. Here, we take two as examples. In scheme 1, 200 F2 individuals are obtained from a cross or backcross, and each F2 individual is allowed to produce 20 F3 progenies. 4,000 F4 individuals are produced by single-seed descent, and then target genotypes are selected by marker analysis. In scheme 2, marker analysis is applied to 200 F2 individuals, and only those carrying the target alleles are advanced to F4 where marker analysis is applied again. A simulation experiment was carried out using QuLine software (the RFLP markers used in the simulation experiment are given in supplementary Table 2). From the DNA samples per selected F4 family needed, we can draw a conclusion that the optimum approach to achieving TG1 was selection from the progenies of CSSL47 × CSSL57, coupled with marker-assisted selection of scheme 2. Similarly, for TG2, the optimum approach was scheme 2 from the progenies of CSSL16 × CSSL20 (Table 7). The time taken for CSSL31, 57, 61, 43, 45, 47, and TG1 to flower was shorter than that for Asominori whereas that for CSSL16, 20, 23, and TG2 was longer. The former lines were suitable for double-season late rice cropping and the latter for single-season middle rice in Nanjing (Fig. 2).
Discussion
Growth duration or HD is an important agronomic trait in rice, which is critical for rice to adapt to specific cultivation conditions and cropping seasons. Breeding for high yield cultivars with suitable growth duration has been a major focus. Now, with the genetic mapping and isolation of many genes and QTL controlling rice HD (Li et al. 1995; Yano et al. 1997, 2000; Lin et al. 1998, 2002; Yamamoto et al. 2000; Maheswaran et al. 2000; Yu et al. 2002; Takahashi et al. 2001; Kojima et al. 2002; Doi et al. 2004; Xue et al. 2008), the genetic basis of HD has been under elucidation (Hayama et al. 2003; Izawa et al. 2003). At the same time, the rapid development of marker technology in recent years has also encouraged the elaboration of the concept of breeding based on genotype, rather than on phenotype (Peleman and van der Voort 2003; Wang et al. 2005; Wan 2006; Wang and Pfeiffer 2007). Thus, the time is right for addressing the challenge of engineering HD in rice by design breeding.
Both major genes and QTL are involved in the determination of HD, and the HD genotypes for different ecotypes in rice have been investigated (Okumoto et al. 1992, 1996; Ichitani et al. 1997, 1998; Tsai 1985; Xu et al. 2007; Wei et al. 2008). However, the best means of using this genotypic information is so far lacking. In the work discussed in this paper, the leading rice cultivars from different ecological regions of China were genotyped by analysis of the phenotype of their hybrids with a set of HD tester lines. On the basis of this information we showed that the late heading in the indica × japonica F1 hybrid is because of interactions between alleles at E 1 and Se-1. This result was consistent with previous research, which indicated that late heading of the indica × japonica F1 hybrid is highly dependent on the identity of the two parents (Cai et al. 1987; Luo et al. 2002).
In recent years, a growing number of HD QTL have been identified (Li et al. 1995; Yano et al. 1997; Lin et al. 1998, 2002; Yamamoto et al. 2000; Maheswaran et al. 2000; Yu et al. 2002), but there is little evidence that any of these is being used as an aid to rice improvement. In fact, we can design the target genotype for the rice cultivar with required HD in different ecological regions using the QTL information and then achieve the target phenotype by pyramiding of the QTLs by molecular marker-assisted selection. Here, a total of seven QTLs were identified using recombinant inbred lines in five years under the natural long-day growing conditions of Nanjing. But the genetic location of qDTH-2, qDTH-3a1, qDTH-3a2, qDTH-3b, qDTH-6, qDTH-8, and qDTH-12 was the same as that of the previously reported QTLs Hd7, Hd16, Hd6, Hd9, Hd1 (Se1), Hd5, and Hd13, respectively (Yano et al. 1997, 2000; Yamamoto et al. 2000; Lin et al. 2002; Matsubara et al. 2008). Among the seven QTLs, qDTH-2, qDTH-3b, qDTH-6, and qDTH-8 could be detected in all five years, indicating they were stably expressed for HD in multiple environments. We believe the main effect and stably expressed QTLs could be preferentially applied for gene design. The best breeding or selection schemes could be finally determined by use of a genetic breeding simulation tool, for example QuLine (Wang et al. 2003, 2004, 2007).
The results in this report show that breeding for suitable HD in rice using genotypic information is very efficient, and represents an applied example of “design breeding” (Peleman and van der Voort 2003; Wang et al. 2005; Wan 2006; Wang and Pfeiffer 2007). As yet we have only considered the major HD genes, although it is clear that HD is also affected by other minor genes. Allelic variation in these minor or unknown HD genes probably underlies differences in HD between cultivars that share the same major HD gene allelic constitution. Using crosses with the test NILs to determine the HD alleles is simple and useful, but the results need further validation by study of DNA sequences of the HD genes. The HD genes Se-1(Hd1) and Ef-1(Ehd1) have been cloned (Yano et al. 2000; Doi et al. 2004), providing the opportunity to generate DNA-based assays to determine the identity of the allele(s) carried by any given rice plant. Although several major QTLs can be identified using recombinant inbred lines in five years under the growing conditions at Nanjing, whether they will also be expressed in other environments is not clear. In addition, the outcomes from the simulation study will, in the end, also need to be validated by field experiments. Even so, the results here showed that genetic information related to HD can make a significant contribution to rice breeding, and it has important significance in guiding design breeding for important agronomic traits of rice.
References
Cai CM, Li WM, Zhou YC (1987) Complementary genes controlling photoperiod sensitivity in hybrid rice. Rice Genet Newsl 4:90–91
Deng XJ, Zhou KD, Li RD, Wang WM, Zhu LH (2001) Genetics and mapping of growth duration of rice varieties. J Sichuan Agric Univ 19(2):172–178
Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A (2004) Ehd1 a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 118(8):926–936
Hayama R, Yokoi S, Tamaki S, Yano M, Shimamoto K (2003) Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 422:719–722
Ichitani K, Okumoto Y, Tanisaka T (1997) Photoperiod sensitivity gene of Se-1 locus found in photoperiod insensitive rice cultivars of the northern limit region of rice cultivation. Breed Sci 47:145–152
Ichitani K, Okumoto Y, Tanisaka T (1998) Genetic analyses of low photoperiod sensitivity of rice cultivars from the northern most regions of Japan. Plant Breed 117:543–547
Inoue H, Nishida H, Okumoto Y, Tanisaka T (1998) Identification of an early heading time gene found in the Taiwanese rice cultivar Taichung 65. Breed Sci 48:103–108
Izawa T, Takahashi YJ, Yano M (2003) Comparative biology comes into bloom: genomic and genetic comparison of flowering pathways in rice and Arabidopsis. Curr Opin Plant Biol 6:113–120
Kinoshita T (1995) Report of the committee on gene symbolization, nomenclature and linkage groups. Rice Genet Newsl 12:9–153
Kojima S, Takahashi Y, Kobayashi Y (2002) Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowing downstream of Hd1 under short-day conditions. Plant Cell Physiol 43(10):1096–1105
Kubo T, Nakamura K, Yoshimura A (1999) Development of a series of Indica chromosome segment substitution lines in Japonica background of rice. Rice Genet Newsl 16:104–106
Li ZK, Pinson SRM, Stansel JW, Park WD (1995) Identification of quantitative trait loci (QTLs) for heading date and plant height in cultivated rice (Oryza sativa L.). Theor Appl Genet 91:374–381
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Lin SY, Sasaki T, Yano M (1998) Mapping quantitative trait loci controlling seed dormancy and heading date in rice, Oryza Sativa L., using backcross inbred lines. Theor Appl Genet 96:997–1003
Lin HX, Yamamoto T, Sasaki T, Yano M (2000) Characterization and detection of epistatic interaction three QTLs, Hd-1, Hd-2 and Hd-3, controlling heading date of rice using nearly isogenic lines. Theor Appl Genet 101:1021–1028
Lin HX, Ashikari M, Yamanouch U, Sasaki T, Yano M (2002) Identification and characterization of a quantitative trait locus, Hd9, controlling heading date in rice. Breed Sci 52:35–41
Luo LG, Zhai HQ, Wan JM (2001) Researches on genetics of rice heading date. Jiangsu J Agric Sci 17(2):119–126
Luo LG, Zhai HQ, Wan JM (2002) Genotypic analysis of heading time for Guichao2. Chin J Rice Sci 16(2):99–105
Maheswaran M, Huang N, Sreerangasamy SR, McCouch SR (2000) Mapping quantitative trait loci associated with days to flowering and photoperiod sensitivity in rice (Oryza Sativa L.). Mol Breed 6:145–155
Matsubara K, Kono I, Hori K, Nonoue Y, Ono N, Shomura A, Mizubayashi T, Yamamoto S, Yamanouchi U, Shirasawa K, Nishio T, Yano M (2008) Novel QTLs for photoperiodic flowering revealed by using reciprocal backcross inbred lines from crosses between japonica rice cultivars. Theor Appl Genet 117:935–945
Nishida H, Inoue H, Okumoto Y, Tanisaka T (2002) A novel gene ef1-h conferring an extremely long basic vegetative growth period in rice. Crop Sci 42:348–354
Ohshima I, Kikuchi F (1994) Identification of a recessive inhibitor for photoperiod-sensitive gene, Se-1, in photoperiod-insensitive varieties of Indica type rice. Proceedings of 7th International Congress SAERAO, pp 93–100
Ohshima I, Watanabe Y, Asahic C (1993) Genetic analysis of heading time in a cross between two Indica varieties with two inhibitor genes for photoperiod sensitivity. Jpn J Breed 43:101–106
Okumoto Y, Tanisaka T, Yamagata H (1992) Heading-time genes of the rice varieties grown in the Tohoku-Hokuriku region in Japan. Jpn J Breed 42:121–135
Okumoto Y, Ichitani K, Inoue H (1996) Photoperiod insensitivity gene essential to the varieties grown in the northern limit region of paddy rice (Oryza sativa L.) cultivation. Euphytica 92:63–66
Peleman JD, van der Voort JR (2003) Breeding by design. Trends Plant Sci 8:330–334
Sato S, Sakamoto I, Shirakawa K, Nakasone S (1988) Chromosomal location of an earliness gene Ef1 of rice, Oryza sativa L. Jpn J Breed 38:385–396
Takahashi Y, Shomura A, Sasaki T, Yano M (2001) Hd6, a rice quantitative trait locus involved in photoperiod sensitivity, encodes the α subunit of protein kinase CK2. Proc Natl Acad Sci 98(14):7922–7927
Tsai KH (1985) Further observations on the Ef-1 gene for early heading. Rice Genet Newsl 2:77–78
Tsai KH (1986) Gene loci and alleles controlling the duration of basic vegetative growth of rice. In: Rice genetics. International Rice Research Institute, Manila, The Philippines, pp 339–349
Tsunematsu H, Yoshimura A, Harushima Y, Nagamura Y, Kurata N, Yano M, Sasaki T, Iwata N (1996) RFLP framework map using recombinant inbred lines in rice. Breed Sci 46:279–284
Wan JM (2006) Perspectives of molecular design breeding in crops. Acta Agron Sin 32:455–462
Wang J, Pfeiffer WH (2007) Simulation approach and its applications in plant breeding. Sci Agric Sin 40(1):1–12
Wang J, van Ginkel M, Podlich D, Ye G, Trethowan R, Pfeiffer W, Delacy I, Cooper M, Rajaram S (2003) Comparison of two breeding strategies by computer simulation. Crop Sci 43:1764–1773
Wang J, van Ginkel M, Trethowan R, Ye G, DeLacy I, Podlich D, Cooper M (2004) Simulating the effects of dominance and epistasis on selection response in the CIMMYT Wheat Breeding Program using QuCim. Crop Sci 44:2006–2018
Wang YH, Xue YB, Li JY (2005) Towards molecular breeding and improvement of rice in China. Trends Plant Sci 10:610–614
Wang J, Li H, Wan X, Pfeiffer W, Crouch J, Wan J (2007) Application of identified QTL-marker associations in rice quality improvement through a design breeding approach. Theor Appl Genet 115:87–100
Wei XJ, Jiang L, Xu JF, Zhang WW, Lu GW, Zhang YS, Wan JM (2008) Genetic analyses of heading date of japonica rice cultivars from Northeast China. Field Crops Res 107:147–154
Xu JF, Jiang L, Wei XJ, Wan JM (2006) Genotyping the heading date of male-sterile rice line II-32A. J Integr Plant Biol 48(4):440–446
Xu JF, Jiang L, Wei XJ, Zhang WW, Liu SJ, Chen LM, Luo LG, Wan JM (2007) Genotypes of heading date of middle Indica rice in the Mid-lower region of the Yangtze River. J Integr Plant Biol 49(12):1772–1781
Xue WY, Xing YZ, Weng XY, Zhao Y, Tang WJ, Wang L, Zhou HJ, Yu SB, Xu CG, Li XH, Zhang QF (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40(6):761–767
Yamagata H, Okumoto Y, Tanisaka T (1986) Analysis of genes controlling heading time in Japanese rice. In: Rice genetics. International Rice Research Institute, Manila, The Philippines, pp 351–359
Yamamoto T, Kuboki Y, Lin SY (1998) Fine mapping of quantitative trait loci Hd-1, Hd-2 and Hd-3, controlling heading date of rice, as single Mendellian factors. Theor Appl Genet 97:37–44
Yamamoto T, Lin HX, Sasaki T, Yano M (2000) Identification of heading date quantitative trait locus Hd6 and characterization of its epistatic interactions with Hd2 in rice using advanced backcross progeny. Genetics 154:885–891
Yano M, Harushima Y, Nagamura Y, Kurata N, Minobe Y, Sasaki T (1997) Identification of quantitative trait loci controlling heading date in rice using a high-density linkage map. Theor Appl Genet 95:1025–1032
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering line gene CONSTANS. Plant Cell 12:2473–2483
Yokoo M, Kikuchi F (1977) Multiple allelism of the locus controlling heading time of rice, detected using the close linkage with the blast-resistance. Jpn J Breed 27:123–130
Yu SB, Li JX, Xu CG, Tan YF, Li XH, Zhang QF (2002) Identification of quantitative trait loci and epistatic interactions for plant height and HD in rice. Theor Appl Genet 104:619–625
Acknowledgments
We are grateful to Drs K. Ichitani and M. Yano for provision of the testers EG0, EG1, ER, LR, and QTL NILs, to Dr Tsai Kuo Hai for the isogenic lines T65m and T65Ebm, to Professor A. Yoshimura, for his kindly providing us the RIL and CSSL populations and genotype data. This research is supported by grants from the 863 Program of China (2006AA100101, 2006BAD01A01-5, 2006BAD13B01), the National Natural Science Foundation of China (30871497), Jiangsu Science and Technology Development Program (BG2006301), Jiangsu Agricultural Germplasm Gene Pool Program (sx(2007)g02), and the 111 Project (B08025).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, X., Liu, L., Xu, J. et al. Breeding strategies for optimum heading date using genotypic information in rice. Mol Breeding 25, 287–298 (2010). https://doi.org/10.1007/s11032-009-9332-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-009-9332-5