Abstract
Ninety-five α-gliadin open reading frames were cloned and sequenced from the somatic hybrid wheat introgression line II-12 and its parents Triticum aestivum cv. Jinan177 (JN177) and Agropyron elongatum. Novel α-gliadin genes were found to originate via point mutation, unequal crossover or slippage of a parental gene, demonstrating that new genes could be rapidly created through somatic hybridization in a manner similar to that previously shown for high-molecular-weight glutenin subunits (HMW-GS) genes. The data reveal the composition and origin of the α-gliadin gene in II-12, showing that: (1) most were homologous to those of JN177; (2) a few were derived direct from A. elongatum; and (3) some new genes were created de novo. A particular quality attribute of interest was the presence or absence of celiac disease (CD) epitopes, which were found to be four times more common among α-gliadin genes from the parent wheat JN177 than in those from A. elongatum. Although four types of CD epitopes were found in introgression line II-12, the number of genes encoded CD epitopes was lower than in JN177 due to the occurrence of pseudogenes. We discuss the benefit of these α-gliadins to wheat breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat grain is a major source of energy and nutrition in the human diet. Mature wheat grain consists of two genetically different organs, the embryo and the endosperm. The latter comprises about 80% of the dry matter and 72% of the protein (Shewry et al. 2003). The majority of the protein in endosperm is gluten, which determines the bread-making quality of wheat flour (Gu et al. 2004). Gluten is classically divided into alcohol-soluble (gliadin) and alcohol-insoluble (glutenin) fractions, which are further separated by electrophoresis. Gliadin consists of monomeric proteins, which are traditionally separated into four groups, named α-, β-, γ-, and ω-gliadins, by polyacrylamide electrophoresis at low pH (Shewry et al. 2003; Wieser 2007). It is known that α- and β-gliadins are closely related to sequence and structure, so that both are usually referred to as α-gliadin. Among them, α-gliadin is most abundant, comprising 15–30% of the whole proteins of wheat seed, with average molecular weight of 31 kDa. Therefore, α-gliadin is the most consumed storage protein by humans (Gu et al. 2004; Herpen et al. 2006; Wang et al. 2007).
Genetically, the main gliadin loci are located on the chromosomes 1 (Gli-1) and 6 (Gli-2) homologous groups (Payne et al. 1982; Sobko 1984; Pogna et al. 1993; Felix et al. 1996; Shewry et al. 2003). The α-gliadins are positioned on the Gli-2 loci (Gli-A2, Gli-B2, and Gli-D2; Payne 1987; Shewry et al. 2003). The number of α-gliadin genes is highly variable, ranging from 25 to 150 copies per haploid genome of wheat and its ancestral species (Herberd et al. 1985; Okita et al. 1985; Anderson et al. 1997). These differences are probably caused by duplication and deletion of chromosome segments (D’Ovidio et al. 1991).
The common structure of the α-gliadin consists of a 20-amino-acid signal peptide (S) followed by a repetitive N-terminal domain (R) based on one or more motifs that are rich in proline and glutamine residues and a longer nonrepetitive C-terminal domain (NR1 and NR2), separated by two polyglutamine repeats (Q1 and Q2) (Anderson and Greene 1997; Shewry et al. 2003). With a few exceptions, α-gliadin contains six conserved cysteine residues in the nonrepetitive domains, which form three interchain disulfide bonds (Shewry et al. 2003). Some α-gliadins contain additional cysteine residues, which allow the formation of interchain disulfide bonds and have positive effect on pasta quality (Shewry and Tatham 1997; Anderson et al. 2001). Changes in positions of cysteine residues can also affect the pattern of intra- and intermolecular disulfide bond formation (Masci et al. 2002).
Celiac disease (CD) is a common enteropathy, occurring in 0.5–1% of the general population, induced by ingestion of wheat gluten proteins and related prolamins from oat, rye, and barley. Among these proteins, α-gliadin and some glutenins contain several peptides that constitute the main toxic components in CD (Van de Wal et al. 1998; Koning 2003). There is genetic diversity in α-gliadin proteins with toxic epitopes (Vader et al. 2003; Herpen et al. 2006).
Agropyron elongatum (Host) Nevishi (Th. Ponticum Podp) is an important wild source for wheat improvement. Its seeds have a high content of protein, and the plant shows high level of resistance to abiotic stress and many diseases (Xia et al. 2003). Asymmetric somatic hybridization between protoplast of common wheat (Triticum aestivum L cv. JN177) and ultraviolet (UV)-irradiated protoplast of A. elongatum generated fertile introgression lines with superior agronomic traits, in particular bread-making quality (Xia et al. 2003). In the present paper the α-gliadin sequences from a somatic hybrid line F7 II-12 with high flour quality and biparents JN177 and A. elongatum are characterized. This study complements earlier work on the glutenin proteins (Zhao et al. 2003; Liu et al. 2007; Chen et al. 2007).
Materials and methods
Plant materials
Agropyron elongatum (StStStStEeEeEbEbExEx, 2n = 70), Triticum aestivum cv. JN177 (AABBDD, 2n = 42), and the introgression self-fertilized line II-12 originated from a single fusion cell in the intergeneric asymmetric somatic hybridization between protoplasts of JN177 and UV-irradiated A. elongatum were used in the experiments. II-12 is genetically stable from the F2 generation (Feng et al. 2004).
Cloning and sequencing of α-gliadin gene ORFs
Seedlings of A. elongatum, JN177, and hybrid introgression line F7 II-12 were grown in darkness for 14 days at 25°C. Genomic DNA was extracted from these seedlings by the cetyl trimethylammonium bromide (CTAB) method according to Murray and Thompson (1980). A pair of primers (P1: 5′-ATG AAG ACC TTT CTC ATC CT-3′ and P2: 5′-TCA GTT AGT ACC GAA GAT GCC-3′) was used to amplify the coding region of α-gliadin genes in a public database. A high-fidelity polymerase LA Taq with GC buffer (TaKaRa) was used in genomic polymerase chain reaction (PCR). The amplification regime consisted of a denaturation step (94°C/3 min), followed by 30 cycles of 94°C/40 s, 55°C/1 min, and 72°C/2 min, with a final extension at 72°C for 10 min. The purified PCR products were cloned into the vector pMD 18-T (TaKaRa) and introduced by standard methods (Sambrook et al. 1989) into E. coli DH10B competent cells. PCR amplification was carried out to identify the positive clones. The selected clones were sequenced by a commercial company (Yingjun, Shanghai, China). At least two independent sequences of each clone were sequenced.
Sequence analysis
Sequence analyses were performed with the help of MEGA (version 3.1, Kumar et al. 2004), programs hosted by the National Centre for Biotechnology Information (NCBI) and the European Bioinformatics Institute (EBI).
Analysis of synonymous and nonsynonymous substitution
The obtained nucleotide sequences were aligned codon-by-codon using Clustal W. We analyzed general selection patterns at the molecular level using DnaSp 4.00 (Rozas et al. 2003). Insertions or deletions that cause a frameshift were treated as nonsynonymous substitutions. The number of synonymous (Ks) and nonsynonymous substitutions (Ka) per site was calculated from pairwise comparisons with incorporation of the Jukes-Cantor correction, as described by Nei and Gojobori (1986).
Results
α-Gliadin gene cloning from both parents and introgression line II-12
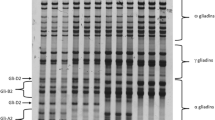
We cloned all the products from introgression line II-12 and the two parents and obtained inserts ranging in size from 0.83 to 0.95 kb (Fig. 1). Preliminary sequencing confirmed the identity of isolated clones to known gliadins. Full sequencing was carried out on 95 open reading frames of α-gliadin genes obtained from introgression line II-12 and the two parents (Table 1). The 95 sequences share identity of >86% with one another (Fig. 2), and an average of >91% with previously reported α-gliadin gene sequences of wheat and its related grass species.
Sequence alignments indicated that most of the α-gliadin genes were derived from the wheat parent, but a few trace to the donor A. elongatum or variation from one of the parent (Fig. 2). Only 22 of these sequences were full-ORF α-gliadin genes (Table 1). Many of the α-gliadin genes contained one or more internal stop codon(s) (56/73) or a frameshift mutation (17/73). Predominately, the stop codons of pseudogenes were located on positions where the full-ORF genes contain a glutamine residue codon according to their deduced amino acid sequences. A major base transition is from C to T, altering a glutamine codon (CAG or CAA) into a stop codon (TAG or TAA).
The number of synonymous (Ks) and nonsynonymous (Ka) substitutions per site was calculated based on pairwise comparison among the obtained full-ORF genes and the pseudogenes (Fig. 3a–c). It is indicated that the ratio of Ks/Ka was no obviously different in A. elongatum, whereas in JN177 and II-12 this value was significantly different between full ORFs and pseudogenes.
Relative numbers of synonymous substitutions (Ka) and nonsynonymous substitutions (Ks) per site for pairwise comparisons among full-ORF α-gliadins and pseudogene sequences. A dotted line represents a Ka/Ks ratio of 1. Linear trendlines with the intercept set to zero are shown both for full-ORF sequences and pseudogene sequences. (a) JN177; (b) A. elongatum; (c) introgression line II-12
Characteristics of deduced amino acid sequences of α-gliadin genes
Sequence alignment of the deduced amino acid sequences of the 95 amplicons showed that they possessed highly similar structures (Figs. 4 and 5). Each allele encodes a typical structure consisting of a signal peptide with 20 amino acid residues followed by a repetitive domain, a polyglutamine domain, an N-terminal unique domain, a second polyglutamine domain, and finally a C-terminal unique domain. Signal peptide and repetitive domains were more conserved than other regions of the genes. The size of each sequence varied, as a result of polymorphism mainly due to the number of glutamines in the two polyglutamine domains. Differences of most amino acids could be attributed to single nucleotide base changes, sequence changes involving complete codons, and frameshifts.
Deduced amino acid sequence alignment of 22 full-ORF α-gliadin genes, showing the disruption of epitopes glia-α2 (PQPQLPYPQ), glia-α9 (PFPQPQLPY), glia-α20 (FRPQQPYPQ), and glia-α (QGSFQPSQQ) in all A. elongatum. S, signal peptide; R, repetitive domain; Q1, N-terminal polyglutamine domain; U1, N-terminal unique domain; Q2, C-terminal polyglutamine domain; U2, C-terminal unique domain. Box 1, glia-α2 and glia-α9 epitodes; box 2, glia-α20; box 3, glia-α
Classification of the deduced amino acid sequences based on the distribution of cysteine. S, signal peptide; R, repetitive domain; Q1, N-terminal polyglutamine domain; U1, N-terminal unique domain; Q2, C-terminal polyglutamine domain; U2, C-terminal unique domain; *, cysteine involved in intermolecular S–S bonds,  , cysteine involved in intramolecular S–S bonds; 1–6, conserved cysteine residue position
, cysteine involved in intramolecular S–S bonds; 1–6, conserved cysteine residue position
Several α-gliadins showed same band/spot in polyacrylamide gel electrophoresis (PAGE) analyses because they possessed almost the same number of deduced amino acid (Fig. 4), although they could be separated by single nucleotide polymorphisms (SNPs) at the DNA level.
The repetitive domain comprised of amino acid numbers ranged from 94 to 109. The first three-codon motif in this region was separated from the followed repeat motifs in amino acid composition. Usually, the first three-codon was VRV, but diversity was also found in these sequences, such as VTV (EU018357) and VIV (EU018332) in full-ORF sequences (Fig. 4), and VRL, VRI, VRF, and FRI in pseudogenes. Each of the repeat motifs in this region, composed of 3–9 codons, was divided into two parts: the conserved front-end three codons followed by a variable-length glutamine-rich region. The repeat motifs were encoded by DNA patterns CCA TA/TT CCA/G CAR. CAR represents a 0–6 glutamine-rich region.
According to the number and position of the cysteine residues in the two unique regions, all the α-gliadins cloned in this study were classified into seven types. Most of them typically contain six conserved cysteine residues resulting in three intramolecular disulfide bonds (Fig. 5). Four were in the N-terminal unique domain and two in the C-terminal unique domain. Figure 4 also shows other six types with odd number of cysteine residues. Clone EU018285 lost a cysteine residue through a cysteine–glutamine change at cysteine residue position 3 (Fig. 5). Clones EU018286, EU018288, EU018291, EU018306, EU018311, and EU018341 made a cysteine–arginine substitution change at position 4. Clone EU018317 lost the fifth cysteine residue because of the encoding DNA deletion. Clone EU018290 lost the sixth cysteine residue because of a cysteine changed to a glycine. Clones EU018273 and EU018262, EU018295, EU018296, EU018298, EU018299, EU018337, EU018339, and EU018362 had an additional cysteine in the front-end of the C-terminal unique domain created by a serine-to-cysteine substitution.
Analysis of CD toxic epitopes
Four CD epitopes (glia-α2, glia-α9, glia-α20, and glia-α) reported previously have been screened in all the obtained full-ORF genes and pseudogenes (Fig. 4, Table 2). The result indicated that three of the four were absent in all genes from A. elongatum except for glia-α. Both the full-ORF genes and pseudogenes, as a whole in JN177 and the hybrid II-2, contained all the four epitopes, but the CD numbers were lower in II-2 than in JN177 (Table 2). Each epitope had definite position in the α-gliadin protein (Fig. 4). Glia-α was usually present in the C-terminal unique domain (U2), whereas glia-α2, glia-α9, and glia-α20 were found in all repetitive domain (R). A close look at these α-gliadin genes indicated that a SNP, which resulted in an amino acid change in a particular epitope, was present in most full-ORF genes obtained in this study; for example, Fig. 4 shows that the glia-α20 epitope in all of the full-ORF genes from A. elongatum were disrupted by the second amino acid of the epitope containing P (proline) instead of R (arginine). In parts of full-ORF sequences from wheat and hybrid, the glia-α20 epitope was disrupted due to the fifth amino acid S (serine) of the epitope being replaced by P (proline). These substitutions were also found in other epitopes. In pseudogenes, the deletion of an amino acid in epitope was presented another type of epitope disruption (data not shown).
Coding-sequences-based phylogeny of the α-gliadin gene family
Twenty-two α-gliadin full-ORF genes obtained in this paper and another eight from Triticum retrieved from the NCBI were used to construct the phylogenetic tree according to their deduced amino acid sequences (Fig. 6). The results indicated that all the cloned sequences clustered into two main groups. Most gliadins from the hybrid were closely related to those from the parent wheat and Triticum, whereas only two from the hybrid (EU018355 and EU018359) and three from Titicum (DQ296195, K03074, and M16496) were more homologous with those from A. elongatum.
Phylogenetic tree based on the deduced amino acid sequences of 22 α-gliadin genes with full ORF obtained in this study, and the eight genes from wheat and related grass according to data of NCBI. Sequences were compared from the start codon to the stop codon. EU018291–EU018299 from T. aestivum cv. JN177; EU018331–EU018335 from A. elongatum; EU018355–EU018362 from introgression line II-12; DQ14035 and DQ296195 from T. turgidum subsp. durum (durum wheat); M16496 from T. urartu; DQ246447, K03074, M01192, X01130, X17361 from T. aestivum
Discussion
Composition and origin of the α-gliadin family in introgression line II-12
Shewry et al. (1995) suggested that the prolamin superfamily of the Triticeae (S-rich, S-poor, and HMW prolamins) are derived from a single ancestral protein. Comparisons of the S-rich prolamins and HMW-GS prolamins showed that all of them contain three conserved nonrepetitive regions designated A, B, and C. Insertion of variable regions and repeated sequences between these conserved regions have generated S-rich prolamins and HMW prolamins in the present day. The S-poor prolamins are considered to have evolved from the S-rich prolamins by deletion of the most domains of regions A, B, and C.
The composition, variation, and origin of gliadins in the introgression line II-12 and the two parents indicated that: (1) most of the α-gliadin genes in the hybrid are homologous to those of common wheat (JN177; Fig. 2); (2) a few novel α-gliadin genes come from the introgression of donor to receptor, e.g., clones EU018355 and EU018359 of the hybrid wheat displayed a high similarity with clones EU018335 and EU018334 of A. elongatum, but lower similarity with all clones of JN177 and those from Triticum; (3) some genes presenting in the hybrid could be created via point mutation of parent wheat, e.g., clone EU018362 from the hybrid shares 99.9% homology (only a T to G change at position 519) with EU018298 from the parent wheat; (4) allelic variation at α-gliadin genes of JN177 or A. elongatum could occur in the same way as has been documented for the HMW-GS and low-molecular-weight glutenin subunits (LMW-GS) genes (Liu et al. 2007; Chen et al. 2007); for example the sequence alignments indicated that clone EU018292 was very similar to EU018360 except the deletion of a tri-base repeat motif and eight SNPs. Two polyglutamine domains of α-gliadin genes were encoded by microsatellite-like sequences, which were known to be hypervariable, and the repetitive codon number variation accounted for most of the differences in protein sizes among α-gliadins.
Nonsynonymous and synonymous mutations between the somatic hybrid and its biparents
From our results the occurrence of stop codons at identical positions in different sequences indicated that pseudogene duplication had occurred. It was noted that most stop codon positions were shared between the II-2 and JN177 genomes, which implied that hybrid pseudogenes were mainly from the parent wheat, which was consistent with the alignment and phylogenetic analysis. The results of the synonymous and nonsynonymous substitutions in the obtained gene sequences indicated that the pseudogenes contained more nonsynonymous substitutions than did the full-ORF genes. This is consistent with reduced selection pressure on the pseudogenes, which were not active as storage proteins. Among them, the ratio of nonsynonymous to synonymous substitutions was the highest in pseudogenes of II-2, whereas full-ORF genes were the highest in A. elongatum among the hybrid and parents.
Contribution of α-gliadin to wheat breeding
The number and location of cysteine residue in α-gliadin are strongly correlated with flour quality (Shewry et al. 2003). Most α-gliadins cloned in this paper contained six cysteine residues with conserved positions in the two unique domains. These cysteine residues could form three intramolecular disulfide bonds, resulting in a smaller and more compactly folded globular protein (Miiller and Wieser 1995; Khatkar et al. 2002). Changes in the position of cysteine residues might affect the pattern of disulfide bond formation, resulting in a failure of two cysteine residues in a protein. These two cysteine residues would then contribute to intermolecular disulfide bond formation (Masci et al. 2002). Kasarda et al. (1984) reported that α-gliadins with an odd number of cysteine residue tended to join the disulfide crosslinked gluten matrix, with a positive effect on flour quality; whereas Porceddu et al. (1998) proposed that the additional residues in gliadins with an odd number of cysteines might induce the polypeptide to act as a chain terminator, with negative effects in terms of pasta quality. In this study, a few hybrid α-gliadins contained odd numbers of cysteines. The analysis of gliadin genes in the introgression line II-12 and the two parents also indicates that these lines may be a possible source of gliadins with a reduced frequency of CD epitopes, a quality attribute that may become a significant breeding target as the consumption of wheat increases.
References
Anderson OD, Greene FC (1997) The α-gliadin gene family. II. DNA and protein sequence variation, subfamily structure, and origins of pseudogenes. Theor Appl Genet 95:59–65
Anderson OD, Litts JC, Greene FC (1997) The α-gliadin gene family. I characterization of ten new wheat α-gliadin genomic clones, evidence for limited sequence conservation of flanking DNA, and Southern analysis of gene family. Theor Appl Genet 95:50–58
Anderson OD, Hsia CC, Adalsteins AE, Lew EJL, Kasarda DD (2001) Identification of several new classes of low-molecular-weight wheat gliadin related proteins and genes. Theor Appl Genet 103:307–315
Chen F, Luo Z, Zhang Z, Xia G (2007) Variation and potential value in wheat breeding of low-molecular-weight glutenin subunit genes cloned by genomic and RT-PCR in a derivative of somatic introgression between common wheat and Agropyron elongatum. Mol Breed 21(2):141–152
D’Ovidio R, Lafiandra D, Tanzarella OA, Anderson OA, Greene FC (1991) Molecular characterization of bread wheat mutants lacking the entire cluster of chromosome 6A-controlled gliadin components. J Cereal Sci 14:125–129
Felix I, Martinant JP, Bernard M, Bernard S, Branlard G (1996) Genetic characterisation of storage proteins in a set of F1-derived haploid lines in bread wheat. Theor Appl Genet 92:340–346
Feng DS, Xia GM, Zhao SY, Chen FG (2004) Two quality-associated HMW glutenin subunits in a somatic hybrid line between Triticum aestivum and Agropyron elongatum. Theor Appl Genet 110:136–144
Gu YQ, Crossman C, Kong XY, Luo MC, You FM, Coleman-Derr D, Dubcovsky J, Anderson OD (2004) Genomic organization of the complex α-gliadin gene loci in wheat. Theor Appl Genet 109:648–657
Herberd NP, Bartels D, Thompson RD (1985) Analysis of the gliadin multigene loci in bread wheat using nullisomic-tetrasomic lines. Mol Gen Genet 198:234–242
Herpen TWJM, Goryunova SV, Schoot J, Mitreva M, Salentijn E, Vorst O, Schenk MF, Veelen PA, Koning F, Soest LJM, Vosman B, Bosch D, Hamer RJ, Gilissen LJWJ, Smulders MJM (2006) Alpha-gliadin genes from the A, B, and D genomes of wheat contain different sets of celiac disease epitopes. BMC Genomics 7:1–13
Kasarda DD, Okita TW, Bemardin JE, Baecker PA, Nimmo CC, Lew EJ, Dietler MD, Greene FC (1984) Nucleic acid (cDNA) and amino acid sequences of alpha-type gliadins from wheat (Triticum aestivum). Proc Natl Acad Sci 81:712–716
Khatkar BS, Fido RJ, Tatham AS, Schofield JD (2002) Functional properties of wheat gliadins. 11. Effects on dynamic rheological properties of wheat gluten. J Cereal Sci 35:307–313
Koning F (2003) The molecular basis of celiac disease. J Mol Recognit 16:333–336
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Liu S, Zhao S, Chen F, Xia G (2007) Generation of novel high quality HMW-GS genes in two introgression lines of Triticum aestivum/Agropyron elongatum. BMC Evol Biol 7:76–83
Masci S, Rovelli L, Kasarda DD, Vensel WH, Lafiandra D (2002) Characterisation and chromosomal localisation of C-type low-molecular-weight glutenin subunits in the bread wheat cultivar Chinese Spring. Theor Appl Genet 104:422–428
Miiller S, Wieser H (1995) The location of disulphide bonds in a-type gliadins. J Cereal Sci 22:21–27
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Okita TW, Cheesbrough V, Reeves CD (1985) Evolution and heterogeneity of the α/β-type and γ-type gliadin DNA sequences. J Biol Chem 260:8203–8213
Payne PI (1987) Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Annu Rev Plant Physiol 38:141–143
Payne PI, Holt LM, Lawrence GJ, Law CN (1982) The genetics of gliadin and glutenin, the major storage proteins of the wheat endosperm. Qualitas Plantarum Plant Foods Hum Nutr 31:229–241
Pogna NE, Metakovsky EV, Redaelli R, Raineri F, Dachkevitch T (1993) Recombination mapping of Gli-5, a new gliadin-coding locus on chromosomes 1A and 1B in common wheat. Theor Appl Genet 87:113–121
Porceddu E, Turchetta T, Masci S, D’Ovidio R, Lafiandra D, Kasarda DD, Impiglia A, Nachit MM (1998) Variation in endosperm protein composition and technological quality properties in durum wheat. Euphytica 100:197–205
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Shewry PR, Tatham AS (1997) Biotechology of wheat quality. J Sci Food Agric 73:397–406
Shewry PR, Napier JA, Tatham AS (1995) Seed storage proteins: structures and biosynthesis. Plant Cell 7:945–956
Shewry PR, Halford NG, Lafiandra D (2003) Genetics of wheat gluten proteins. Adv Genet 49:111–184
Sobko TI (1984) Identification of a new locus which controls the synthesis of alcohol-soluble endosperm proteins in winter bread wheat. J Agric Sci (Kiev) 320:78–80
Vader LW, Stepniak DT, Bunnik EM, Kooy YM, de Haan W, Drijfhout JW, van Veelen PA, Koning F (2003) Characterization of cereal toxicity for celiac disease patients based on protein homology in grains. Gastroenterology 125:1105–1113
Van de Wal Y, Kooy Y, van Veelen P, Pena S, Mearin L, Molberg O, Lundin L, Mutis T, Benckhuijsen W, Drijfhout JW, Koning F (1998) Small intestinal T cells of celiac disease patients recognize a natural pepsin fragment of gliadin. Proc Natl Acad Sci 95:10050–10054
Wang H, Wei Y, Yan Z, Zheng Y (2007) Isolation and analysis of α-gliadin gene coding sequences from Tritcum durum. Agric Sci China 6(1):25–32
Wieser H (2007) Chemistry of gluten proteins. Food Microbiol 24:115–119
Xia GM, Xiang FN, Zhou AF, Wang H, He SX, Chen HM (2003) Asymmetric somatic hybridization between wheat (Triticum aestivum L.) and Agropyron elongatum (Host) Nevski. Theor Appl Genet 107:299–305
Zhao TJ, Quan TY, Xia GM, Chen HM (2003) Glutenin and SDS sedimentation analysis of the F5 somatic hybrids between Triticum aestivum and Agropyron elongatum. J Shandong Univ (Nat Sci) 38(3):112–116. (in Chinese with English abstract)
Acknowledgements
We are grateful to Prof. Rudi Appels (Centre for Comparative Genomics, Murdoch University, Perth WA 6150, Australia) for many significant suggestions and correction. This work was supported by the National 863 High Technology Research and Development Project 2006AA10Z173 and 2006011001020, Key and Doctor-Site Foundation of Ministry of Education in China, Natural Science Foundation of Shandong Province No. Y2007D48.
Author information
Authors and Affiliations
Corresponding author
Additional information
Fanguo Chen, Chunhui Xu, and Mengzhu Chen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chen, F., Xu, C., Chen, M. et al. A new α-gliadin gene family for wheat breeding: somatic introgression line II-12 derived from Triticum aestivum and Agropyron elongatum . Mol Breeding 22, 675–685 (2008). https://doi.org/10.1007/s11032-008-9208-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-008-9208-0