Abstract
Winter hardiness is a quantitative trait and the lack of it limits geographic distribution of ryegrass. Improving winter hardiness is an important breeding goal in ryegrass breeding programs. An understanding of the genetic basis for the component traits of winter hardiness would allow more efficient selection. A three-generation interspecific population of an annual × perennial ryegrass consisting of 152 progenies was used to map quantitative trait loci (QTL) that control winter hardiness-related traits including fall growth (FG), freezing tolerance (FT), and winter survival (WS) over 2 years. A total of 39 QTL were identified for the three traits from both the female parental (MFA) and the male parental (MFB) maps, of which 13 were for FG, 6 for FT, and 20 for WS. The proportion of phenotypic variation explained by individual QTL ranged from 10.4 to 22.1%. Both FG and FT were positively correlated with WS. Common QTL were detected between FG, FT, and WS. The QTL associated with WS on linkage groups (LGs) 4 and 5, and the QTL for FT on LG 5 were consistently identified over years and maps. These consistent QTL might serve as potential tools for marker-assisted selection to improve ryegrass winter hardiness.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Perennial ryegrass (Lolium perenne L.) is used extensively as a forage grass due to its high nutritive values and persistence (Wilkins 1991), and as a turf species in golf course fairways and athletic fields due to its fast establishment, strong seedling vigor, and good tolerance to both traffic and low mowing. However, its poor ability to survive in severe winter conditions (Gusta 1980) limits its distribution in the far northern areas of the United States; therefore, improving winter hardiness is an important goal in perennial ryegrass breeding programs.
Winter hardiness is a complex quantitative trait affected by a number of interacting factors including responses to low temperatures, rapidly fluctuating temperatures, photoperiod, and vernalization requirement (Humphreys 1989). The complexity of winter hardiness makes it difficult to evaluate reliably and then improve it based on phenotypic selection. The best method to evaluate winter hardiness of a cultivar is to rate field survival during a test winter which is severe enough to kill most non-hardy plants, but causes variable degrees of injury to those with variable hardiness (Sulc et al. 1991). However, such a winter environment is often not available. Moreover, assessing winter hardiness is hampered by various environmental factors and the existence of genotype and environment interactions. One approach is to break down winter hardiness into more basic component traits and use these components to predict winter hardiness. An understanding of the genetic basis of these component traits and identification of molecular markers that tightly linked to these traits permit more efficient and reliable selection.
Freezing tolerance (FT) has been identified as a major component of winter hardiness (Lorenzetti et al. 1971; Hides 1979; Humphreys and Eagles 1988). Significant positive correlations between FT and field winter survival (WS) have been reported in perennial ryegrass (Humphreys 1989; Waldron et al. 1998), barley (Hordeum vulgare L.) (Hayes et al. 1993), wheat (Triticum aestivum L.) (Fowler et al. 1981), and alfalfa (Medicago sativa L.) (Brouwer et al. 1998). Both positive and negative correlations have been reported between fall growth (FG) and winter hardiness in alfalfa depending on the cultivars or populations tested (Perry et al. 1987; Schwab et al. 1996; Brouwer et al. 1998; Brummer et al. 2000). Negative correlation between FG and winter hardiness might have resulted from the fact that short days and cool temperature in the fall reduce plant growth and induce accumulation of carbohydrate, which may increase the ability of plants to survive in severe winter conditions.
Freezing tolerance can be measured by first cold-acclimating plants followed by freezing and recovery. The relative FT was expressed as LT50, the temperature at which 50% plants are killed after freezing (Fuller and Eagles 1978; Gusta 1980; Humphreys and Eagles 1988; Waldron et al. 1998). However, this method requires a lot of plant materials and a large area that can be set to different testing temperatures which is not feasible for testing a large mapping population. An alternative method that has been used extensively in various laboratories to evaluate FT is to subject plant tissue to freeze–thaw stresses at various temperatures and then measure the electrolyte leakage. With this method, FT was determined by LT50, the temperature that results in 50% of the total electrolyte leakage (Stone et al. 1993; Teutonico et al. 1995; Jaglo-Ottosen et al. 1998; Brouwer et al. 2000; Zhu et al. 2000) or electrical conductivity at a single freezing temperature (Dexter et al. 1930, 1932; Brouwer et al. 2000; Yamada et al. 2004).
Development of high-density linkage maps based on molecular markers makes it possible to detect quantitative trait loci (QTL) responsible for quantitative traits. QTL mapping for the component traits of winter hardiness has been studied in various crop species such as wheat (Sutka 1994; Galiba et al. 1995, 1997), barley (Hayes et al. 1993; Pan et al. 1994; Francia et al. 2004), oil seed brassica (Brassica napus L.) (Teutonico et al. 1995), alfalfa (Brouwer et al. 2000), and lentil (Lens culinaris Medik.) (Kahraman et al. 2004), which identified genomic regions that were associated with winter hardiness-related traits.
Molecular marker-based genetic maps of perennial ryegrass have been constructed by using the p150/112 reference mapping population (Jones et al. 2002a, b) and were used to map QTL that were related to winter hardiness and other agronomic traits (Yamada et al. 2004). A single QTL for electrical conductivity was detected on linkage group (LG) 4 in the p150/112 population of perennial ryegrass. Recently, genetic linkage maps based on an interspecific ryegrass population derived from the cross between perennial ryegrass and annual ryegrass have been constructed by using 81 RAPD (rapid amplified polymorphism of DNA), 235 AFLP (amplified fragment length polymorphism), 16 RFLP (restricted fragment length polymorphism), 106 SSR (simple sequence repeat), 2 isozymes, and 2 morphological markers (Warnke et al. 2004). These genetic linkage maps were recently enhanced by 120 additional RFLP markers from barley, oat, and rice, which allows comparative genetic mapping and reveals substantial conserved synteny between ryegrass and the Triticeae species, oat, and rice (Sim et al. 2005). As a result, a total of more than 200 codominant markers as well as about 300 dominant markers have been mapped in this interspecific ryegrass population, permitting dissection of genetic loci of quantitative traits that segregate in the population. QTL for resistance to the fungal disease of gray leaf spot have been identified by using the enhanced genetic maps in this population (Curley et al. 2005).
Our objective in this study was to use the enhanced genetic linkage maps of ryegrass to locate QTL that are associated with winter hardiness in the ryegrass population. The specific traits measured were FG, FT, and WS. The identification of genomic regions controlling FG, FT, and WS would improve our understanding of the genetics of these traits and provide an aid to breeders for improving winter hardiness of ryegrass.
Materials and methods
Plant materials
An annual ryegrass plant from the cultivar ‘Floregon’, which has poor winter hardiness was crossed to a perennial ryegrass plant from the cultivar ‘Manhattan’, which has intermediate to good winter hardiness to create the 1st F1 population. At the same time, a second F1 population was created in a similar manner but with two different parental plants each selected from L. multiflorum and L. perenne L., respectively. From the 1st F1 population, one plant named MFA was chosen randomly to cross with another plant named MFB that was randomly chosen from the 2nd F1 population to develop a three-generation population (a pseudo F2 population). All of the seed used to develop the progeny population was obtained from the plant MFA, making MFA as the female parent and MFB as the male parent. The progeny population used for mapping includes 152 individuals. More details on the population development were described by Warnke et al. (2004). The population has been maintained in a research greenhouse at 20–21°C at Iowa State University (Ames, IA, USA). Twelve single-tiller ramets were propagated for each of the 152 genotypes and their parents and were maintained in the same greenhouse. On 15 May 2003, four ramets of each genotype were transplanted to the field within each replication in an α-lattice design with three replications at the Iowa State University Horticulture Research Station near Gilbert, IA, USA. Each replication included 26 blocks and each block included 6 entries. Spacing between ramets within each genotype was 30 cm. Each genotype was separated by a 60-cm space in between. Plants were mowed at 6.4 cm above the soil surface on 17 July, 14 August, and 22 September in 2003, respectively. The experiment was repeated at a different site within the same research station in 2004 except that in 2004, the plants were transplanted to the field on 3 June and mowed on 22 July and 21 August 2004, respectively.
Trait assessment and analysis
FG was measured in centimeters for the vertical height of regrowth after mowing in the fall on 14 November 2003 and 12 November 2004, respectively. A total of 12 tillers from each genotype for one replication (3 tillers per ramet with 4 ramets in each replication) were measured and the heights of the 12 tillers were averaged as the height for the genotype in the replication.
Freezing tolerance was assessed by subjecting plant tissue to freeze–thaw stresses at various temperatures and then measuring the ion leakage at different freezing temperatures in 2003. The LT50 (°C) was determined by fitting the curve of ion leakage for each genotype. This protocol was based on Stone et al. (1993) with some modifications. One ramet of each genotype was removed from naturally hardened field plants on 30 November 2003 and stored at a 4°C walk-in cold room to avoid dehardening. Individual tillers of similar size were separated from the ramet and trimmed to 2.5-cm long segments containing 2 cm of the stem and 0.5 cm of the root, respectively. All the preparation work was done at the 4°C cold room. Trimmed tillers were placed in test tubes (15 × 125 mm) containing 100 μl deionized water which were then placed in a programmable freezer (Model 8.5–3.1; ScienTemp Corp. Adrian, MI, USA) equilibrated at −2°C. Controls were kept at 4°C. Each tube contained two tillers. After 30 min, the temperature was lowered to−3°C and held for 30 min and then lowered to −4°C and held for 30 min again. Further cooling below −4°C was at a rate of 2°C/h until −10°C, then 4°C/h for the rest and held for 15 min at each predetermined freezing temperature. One tube of each genotype was removed at each freezing temperature (−6, −10, −14, −18, −20, −24, −28, and −32°C), respectively, and thawed first on ice overnight at the 4°C cold room, and then removed from ice and kept at 4°C for a minimum of 2 h to allow slow thawing. Seven millimeter of deionized distilled water was added to the samples at room temperature. Samples were then infiltrated twice for 4 min each at 0.1 atmosphere (10 kPa) by using a vacuum pump and shaken horizontally for 1 h at 250 rpm. Temperature-specific electrical conductivity was measured for each sample with an YSI model 3403 conductance meter. Total conductivity for each sample was determined after autoclaving the samples at 121°C for 20 min. Percentage of ion leakage at each freezing temperature was calculated as the ratio of the temperature-specific conductivity to the total conductivity and was plotted as a function of freezing temperatures. The temperature value corresponding to the midpoint between the maximum and minimum (control) ion leakage obtained for each genotype was defined as the LT50 value. The LT50 values of the three replications were used for further analysis.
Because the method used for measuring FT is very time consuming, in 2004 we used another proven procedure (Dexter et al. 1932; Brouwer et al. 2000; Yamada et al. 2004) to evaluate FT by subjecting plant tissue to a freeze–thaw stress at a single freezing temperature followed by measuring electrical conductivity. One ramet of each genotype was removed from naturally hardened field plants on 15 December 2004. Material storage and sample preparation were the same as described in 2003. Three prepared individual tillers from each genotype were placed into test tubes (15 × 125 mm) containing 100-μl deionized water. The tubes were then submerged into a glycol-containing bath previously equilibrated at −1.5°C while the control was kept at 4°C. The temperature was held at −1.5°C for 2 h, and then ice chips were added to tubes to initiate the formation of ice-nucleation. The temperature continued to hold at −1.5°C overnight and was then lowered manually to −2°C at a rate of 0.5°C/30 min until to −4°C. After that, the cooling rate was 1°C/h until to −9°C, and then 2°C/h to −15°C. The temperature was held at −15°C for 30 min. Then the tubes were taken out and thawed on ice at 4°C cold room overnight and removed from ice and kept at 4°C for a minimum of 2 h to allow slow thawing. The subsequent steps were the same as described for 2003. Percentage of electrical conductivity was used to evaluate FT. The average electrical conductivity of the three replications was used for further analysis. This method allowed us to process the sample at a faster pace, yet the results obtained with the two methods in 2003 and 2004 are comparable.
Winter survival was visually scored on a scale of 0–5 with 0 being no survival and 5 being full survival following the winter of 2003 and 2004. The data were collected on 10 April 2004 and 9 April 2005, respectively.
Analysis of variance of the phenotypic data was performed by the GLM procedure of the Statistical Analysis System (version 8.0, SAS Institute Inc. (2004), Cary, NC, USA). The MIXED procedure was used to estimate variance components with all effects in the model considered random. Broad-sense heritabilities (H) were calculated as H = σ 2g /(σ 2g + σ 2e /r), where σ 2 g is the genetic variance and σ 2 e is the error variance divided by r, the number of replications for each genotype. Pearson correlation coefficients were used to evaluate correlations for pair-wise phenotypic traits by using the CORR procedure of SAS. The correlations were calculated on a genotype-mean basis and calculated separately for each year.
QTL analysis
The genetic markers from the two parental maps, MFA and MFB, constructed separately by Warnke et al. (2004), including RAPD, RFLP, AFLP, SSR, isozyme, and morphological marker data were combined with the ryegrass RFLP marker data generated recently (Sim et al. 2005) to produce a composite marker dataset for QTL analysis of the present study. Two parental maps, MFA and MFB, were constructed by using the CP (cross pollination) type option in Joinmap® (Van Ooijen and Voorrips 2001). Interval mapping analysis was employed to detect putative QTL implemented by MapQTL® (Van Ooijen et al. 2002). Mean trait values from three replications were used to identify QTL from both female (MFA) and male (MFB) maps. The logarithm of odds (LOD) score of 3.0 (Liu 1998) was chosen as the threshold for declaring putative QTL. QTL locations were determined by the peak of LOD score and the one-LOD support region was used as a support interval for the location of a QTL on the linkage maps.
Results
Phenotypic distributions, heritabilities, and correlations
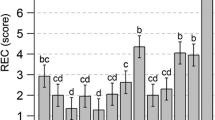
Variation was observed in all measured traits for the two parents, ‘MFA’ and ‘MFB’ as well as the mapping population (Table 1; Fig. 1). Both parents had higher values in FG and WS in 2004 than in 2003. In both years, MFA was consistently associated with a higher FG and WS compared with MFB. MFA also showed more FT than MFB in both 2003 and 2004. The mean values of MFA and MFB for FG, WS, and FT fell within the range of progenies, and all the three traits exhibited nearly normal distributions with transgressive segregants (Fig. 1).
Progeny frequency distributions for the annual × perennial ryegrass population for fall growth (a), winter survival (b) measured in 2003 and 2004. Freezing tolerance was measured as LT50 in 2003 (c), and as electrical conductivity in 2004 (d). The means of the parents (MFA and MFB) are indicated by arrows
Fall growth and WS had intermediate heritabilities in both years ranging from 0.77 to 0.89, whereas FT had a relatively lower heritability of 0.60 measured in 2003 and 0.51 in 2004.
Significant phenotypic correlations were detected among all three traits over 2 years except for between FT and FG. The strongest correlation was found between FG and WS in 2003 (r = 0.57) and the lowest correlation was that of FT and WS in 2003 with r = 0.15 (yet significant at P = 0.01). FT was positively correlated with WS consistently. Furthermore, FG had positive correlations with WS in both years. All the three traits measured in 2003 were positively correlated with those measured in 2004.
QTL detection
Fall growth
A total of 13 QTL associated with FG was identified from both the female MFA and the male MFB maps in the mapping population (Tables 3, 4; Fig. 2). Of which, nine QTL were from MFA map and four from MFB map. In the MFA map, four QTL associated with FG were detected in 2003, two in 2004, and three in the 2-year combined analysis. The nine QTL were located on four LGs with one each located on each of the LGs 1 and 7, and three located on the LG 3, and four on the LG 4 in the MFA map (Table 3, Fig. 2a). On the MFB map, three QTL associated with FG were found in 2003, and one in the 2-year combined analysis (Table 4). The four QTL were located on the LGs 1, 3, and 4, respectively, with two located on the LG 4 (Table 4, Fig. 2b). No QTL for FG were detected in 2004 from MFB map. The proportion of phenotypic variation explained by individual QTL for FG ranged from 10.4 to 21.6% (Tables 3, 4).
Location of putative QTL for fall growth (FG), winter survival (WS), and freezing tolerance measured in both 2003 and 2004 on the female parental (MFA) map (a) and the male parental (MFB) map (b) of the annual × perennial ryegrass. QTL positions for the three traits detected by simple interval mapping are shown on the left of the linkage groups. The number on the top indicates the linkage group. Black, hatched, and white bars represent interval regions of putative QTL for FG, WS, and FT, respectively. The name of QTL was given by the form of q-trait-year-female/male map linkage group. When there were more than one QTL for the same trait in the same map and same year, then different numbers were added as suffix at the end of QTL name to distinguish the QTL. For QTL detected on the traits by combining a 2-year data, the name was given by the form of q-trait-female/male map linkage group
Winter survival
A total of 20 QTL for WS was detected from both the female MFA and the male MFB maps (Tables 3, 4; Fig. 2). Nine of them were located on the LGs 1, 3, 4, and 5 of the MFA map and seven were on the LGs 2, 3, 4, and 5 of the MFB map. Four QTL associated with WS were identified in 2003, two in 2004, and three in the 2-year combined analysis on the MFA map (Table 3). On the MFB map, four QTL for WS were detected in 2003, three in 2004, and four in the 2-year combined analysis (Table 4). Individual QTL for WS explained 12.0–30.3% of the total phenotypic variance. The QTL on the LGs 4 and 5 of the MFA map were consistently identified in both years. The QTL on the LGs 2, 4, and 5 of the MFB map were also consistent in both years. The QTL on the LGs 4 and 5 were coincident between MFA and MFB maps.
Freezing tolerance
A total of six QTL associated with FT were identified from both the female MFA and the male MFB maps in 2003 and 2004 (Tables 3, 4; Fig. 2). Two QTL for FT were identified on the LGs 3 and 5 in 2003, and one on the LG 5 in 2004 on both the MFA and the MFB maps. The QTL for FT on the LG 5 was consistent in both years on both MFA and MFB maps. Individual QTL accounted for phenotypic variation ranging from 12.7 to 22.1%. All QTL for FT identified were consistent between maps.
Discussion
We measured three traits, FG, FT, and WS for 152 progeny individuals of the interspecific ryegrass population and its parents in both 2003 and 2004. Means of the parents for all measured traits fell within the range of progeny frequency distributions, suggesting transgressive segregations occurred in both directions in the progenies (Table 1; Fig. 1). Parent MFA was more winter hardy than MFB, as indicated by LT50 (°C) and electrical conductivity, although both parents derived from the crosses between the perennial ryegrass cultivar ‘Manhattan’ and the annual ryegrass cultivar ‘Floregon.’ The heterozygosity of both grandparents ‘Manhattan’ and ‘Floregon’ allowed their hybrids to combine different alleles. As a result, parents used for constructing the mapping population differed in winter hardiness.
Phenotypic correlation was observed between FT and WS (Table 2), which was consistent with previous studies (Fowler et al. 1981; Humphreys 1989; Brouwer et al. 1998; Waldron et al. 1998). The coincident QTL found between FT and WS provide evidence for the observed phenotypic correlation. One overlapped QTL for FT and WS was located on the LG 5 (Tables 3, 4; Fig. 2). Positive correlation was also found between FG and WS in our study, which was in agreement with the results reported by Brummer et al. (2000) in alfalfa where the taller plants tended to have less winter damage. The common QTL between FG and WS accounted for their phenotypic relationships. For example, the QTL for FG on LG 4 from both MFA and MFB, qFG-03-f4 and qFG-03-m4, coincided with those for WS. These genetic regions common for FG and WS, and FT and WS were consistent with their phenotypic correlations, and these common QTL associations may provide genetic information useful in predicting winter hardiness by using FG and FT. However, we also found some QTL associated with WS were not associated with FG and FT, and vice versa. Therefore, indirect selection to improve winter hardiness based on correlated traits FG and FT will not be effective for all loci affecting WS.
The inconsistency of the QTL detected for FG over the 2 years might have resulted from the specific environmental components or could be the result of the different magnitudes of errors in the 2 years. Certain environmental conditions in 2003 may favor expression and detection of some QTL, for example, qFG-03-f1, qFG-03-f7, and qFG-03-m3. The QTL for FT located on LG 5 was consistently identified in both years, and this QTL was perfectly overlapped on both MFA and MFB map (Tables 3, 4; Fig. 2), although we used different methods to evaluate the FT in 2003 and 2004. This result demonstrated that both methods worked equally well for measuring FT.
In our study, one consistent QTL for WS over 2 years was mapped to LG 4 on both MFA and MFB maps (qWS-03-f4 and qWS-04-f4, and qWS-03-m4 and qWS-04-m4). Yamada et al. (2004) reported that the QTL for FT measured by electrical conductivity was also located on the upper part of LG 4 in the p150/112 reference population of perennial ryegrass. Comparative mapping between ryegrass and Triticeae reveals that the one end of LG 4 of ryegrass is the counterpart of the Triticeae chromosome 5 due to translocation (Sim et al. 2005). QTL for FT and vernalization response have been located in wheat (Galiba et al. 1995, 1997) and barley (Hayes et al. 1993; Pan et al. 1994; Francia et al. 2004). Two QTL controlling vernalization and FT are located separately on chromosome 5 in wheat and barley (Galiba et al. 1995; Francia et al. 2004). Jensen et al. (2005) reported that the wheat vernalization gene VRN1 was mapped to LG 4 of perennial ryegrass and co-located with the QTL for heading date in perennial ryegrass. Thus, the QTL we identified for WS on LG 4 may correspond to the orthologue of the vernalization gene of wheat and barley. The LG 5 of ryegrass is also the counterpart of chromosome 5 in the Triticeae (Sim et al. 2005); therefore, the other consistent QTL we detected in the present study for WS and FT over 2 years from both MFA and MFB maps on LG 5 are possibly the orthologues of the QTL for FT detected in wheat and barely. We did not detect QTL for FT but for WS on LG 4 in the study, while Yamada et al. (2004) identified one QTL for FT on LG 4 but no QTL for WS. This could result from the different mapping population we used and the interaction of different alleles and environments.
The QTL that show stable expression over environments are desirable for marker-assisted selection (MAS). In the present study, coincident QTL for WS were identified on LGs 4 and 5 in both years from both MFA and MFB maps. The QTL for FT on LG 5 was also consistently detected in both years from both MFA and MFB maps. These QTL could serve as potential markers for MAS to improve winter hardiness in future ryegrass-breeding programs. The identification of QTL affecting winter hardiness-related traits provides new knowledge about the genetics of winter hardiness of ryegrass.
Abbreviations
- FG:
-
Fall growth
- FT:
-
Freezing tolerance
- LG:
-
Linkage group
- LOD:
-
Logarithm of odds
- QTL:
-
Quantitative trait loci
- WS:
-
Winter survival
References
Brouwer DJ, Duke SH, Osborn TC (1998) Comparison of seedlings and cuttings for evaluating winter hardiness in alfalfa. Crop Sci 38:1704–1707
Brouwer DJ, Duke SH, Osborn TC (2000) Mapping genetic factors associated with winter hardiness, fall growth and freezing injury in autotetraploid alfalfa. Crop Sci 40:1387–1396
Brummer EC, Shah MM, Luth D (2000) Reexamining the relationship between fall dormancy and winter hardiness in alfalfa. Crop Sci 40:971–977
Curley J, Sim SC, arnke S, Leong S, Barker R, Jung G (2005) QTL mapping of resistance to gray leaf spot in ryegrass. Theor Appl Genet 111:1107–1117
Dexter ST, Tottingham WE, Graber LF (1930) Preliminary results in measuring the hardiness of plants. Plant Physiol 5:215–223
Dexter ST, Tottingham WE, Graber LF (1932) Investigations of the hardiness of plants by measurement of electrical conductivity. Plant Physiol 7:63–78
Fowler DB, Gusta LV, Tyler NJ (1981) Selection for winterhardiness in wheat III. Screening methods. Crop Sci 21:896–901
Francia E, Rizza F, Cattivelli L, Stanca AM, Galiba G, Toth B, Hayes PM, Skinner JS, Pecchioni N (2004) Two loci on chromosome 5H determine low-temperature tolerance in a ‘Nure’ (winter) × ‘Tremois’ (spring) barley map. Theor Appl Genet 108:670–680
Fuller MP, Eagles CF (1978) A seedling test for cold hardiness in Lolium perenne L. J Agric Sci Camb 91:217–222
Galiba G, Quarrie SA, Sutka J, Morounov A, Snape JW (1995) RFLP mapping of the vernalization (Vrn1) and frost resistance (Fr1) genes on chromosome 5A of wheat. Theor Appl Genet 90:1174–1179
Galiba G, Kerepesi I, Snape JW, Sutka J (1997) Location of a gene regulating cold-induced carbohydrate production on chromosome 5A of wheat. Theor Appl Genet 95:265–270
Gusta LV (1980) Freezing resistance of perennial turfgrasses. HortScience 15:494–496
Hayes PM, Blake T, Chen THH, Tragoonrung S, Chen F, Pan A, Liu B (1993) Quantitative trait loci on barley (Hordeum vulgare L.) chromosome 7 associated with components of winter hardiness. Genome 36:66–71
Hides DH (1979) Winter hardiness in Lolium multiflorum Lam. III. Selection for improved cold tolerance and its effect on agronomic performance. J Br Grassl Soc 33:175–179
Humphreys MO (1989) Assessment of perennial ryegrass for breeding. II Components of winter hardiness. Euphytica 41:99–106
Humphreys MO, Eagles CF (1988) Assessment of perennial ryegrass for breeding. I freezing tolerance. Euphytica 38:75–84
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
Jensen LB, JRAndersen U, Frei Y, Xing C, Taylor PB, Holm T, Lubberstedt (2005) QTL mapping of vernalization response in perennial ryegrass (Lolium perenne L.) reveals co-location with an orthologue of wheat VRN1. Theor Appl Genet 110:527–536
Jones ES, Dupal MD, Dumsday JL, Hughes LJ, Forster JW (2002a) An SSR-based genetic linkage map for perennial ryegrass (Lolium perenne L.). Theor Appl Genet 105:577–584
Jones ES, Mahoney NL, Hayward MD, Armstead IP, Jones JG, Humphreys MO, King IP, Kishida T, Yamada T, Balfourier F, Charmet G, Forster JW (2002b) An enhanced molecular marker based genetic map of perennial ryegrass (Lolium perenne) reveals comparative relationships with other Poaceae genomes. Genome 45:282–295
Kahraman A, Kusmenoglu I, Aydin N, Aydogan A, Erskine W, Muehlbauer FJ (2004) QTL mapping of winter hardiness genes in lentil. Crop Sci 44:13–22
Liu BH (1998) Statistical genomics: linkage, mapping and QTL analysis. CRC press, Boca Raton
Lorenzetti F, Tyler BF, Cooper JP, Breese EL (1971) Cold tolerance and winter hardiness in Lolium perenne L. I. development of screening techniques for cold tolerance and survey of geographical variation. J agric Sci Camb 76:199–209
McCouch SR, Cho YG, Yano M, Paul E, Blinstrub M (1997) Report on QTL nomenclature. Rice Genet Newslett 14:11–13
Pan A, Hayes PM, Chen F, Chen THH, Blake T, Wright S, Karsai I, Bedo Z (1994) Genetic analysis of the components of winter hardiness in barley (Hordeum vulgare L.). Theor Appl Genet 89:900–910
Perry MC, McIntosh MS, Wiebold WJ, Welterlen M (1987) Genetic analysis of cold hardiness and dormancy in alfalfa. Genome 29:144–149
SAS Institute (2000) SAS language and procedure: usage. Version 8, 1st edn, SAS Institute, Cary, NC
Schwab PM, Barnes DK, Sheaffer CC (1996) The relationship between field winter injury and fall growth score for 251 alfalfa cultivars. Crop Sci 36:418–426
Sim S, Chang T, Curley J, Warnke SE, Barker RE, Jung G (2005) Chromosomal rearrangements differentiating the ryegrass genome from the Triticeae, oat, and rice genomes using common heterologous RFLP probes. Theor Appl Genet 110:1011–1019
Stone JM, Palta JP, Bamberg JB, Weiss LS, Harbage JF (1993) Inheritance of freezing resistance in tuber-bearing Solanum species: evidence for independent genetic control of nonacclimated freezing tolerance and cold acclimation capacity. Proc Natl Acad Sci USA 90:7869–7873
Sulc RM, Albrecht KA, Duck SH (1991) Leakage of intracellular substance as an indicator of freezing injury in alfalfa. Crop Sci 31:430–435
Sutka J (1994) Genetic control of frost tolerance in wheat (Triticum aestivum L.). Euphytica 77:277–282
Teutonico RA, Yandell B, Satagopan JM, Ferreira ME, Palta JP, Osborne TC (1995) Genetic analysis and mapping of genes controlling freezing tolerance in oilseed Brassica. Mol Breed 1:329–339
Van Ooijen JW, Voorrips RE (2001) JoinMap® 3.0: software for the calculation of genetic linkage maps. Plant Research International, Wageningen
Van Ooijen JW, Boer MP, Jansen RC, Maliepaard C (2002) MapQTL® 4.0, Software for the calculation of QTL positions on genetic maps. Plant Research International, Wageningen
Waldron BL, Ehlike NJ, Vellekson DL, White DB (1998) Controlled freezing as an indirect selection method for field winterhardiness in turf-type perennial ryegrass. Crop Sci 38:811–816
Warnke SE, Baker RE, Jung G, Sim SC, Rouf Mian MA, Saha MC, Brilmam LA, Dupal MP, Forster JW (2004) Genetic linkage mapping of an annual × perennial ryegrass population. Theor Appl Genet 109:294–304
Wilkins PW (1991) Breeding perennial ryegrass for agriculture. Euphytica 52:201–214
Yamada T, Jones ES, Cogan NOI, Vecchies AC, Nomura T, Hisano H, Shimamoto Y, Smith KF, Hayward MD, Forster JW (2004) QTL analysis of morphological, developmental, and winter hardiness-associated traits in perennial ryegrass. Crop Sci 44:925–935
Zhu B, Choi DW, Fenton R, Close TJ (2000) Expression of the barley dehydrin multigene family and the development of freezing tolerance. Mol Gen Genet 264:145–153
Acknowledgements
We wish to thank Dr. Steven Larson for critically reading the manuscript and Lori Evans for technical assistance in maintaining the plants in the greenhouse.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xiong, Y., Fei, S.z., Arora, R. et al. Identification of quantitative trait loci controlling winter hardiness in an annual × perennial ryegrass interspecific hybrid population. Mol Breeding 19, 125–136 (2007). https://doi.org/10.1007/s11032-006-9050-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-006-9050-1