Abstract
To develop a salt-tolerant upland rice cultivar (Oryza sativa L.), OsNHX1, a vacuolar-type Na+/H+ antiporter gene from rice was transferred into the genome of an upland rice cultivar (IRAT109), using an Agrobacterium-mediated method. Seven independent transgenic calli lines were identified by polymerase chain reaction (PCR) analysis. These 35S::OsNHX1 transgenic plants displayed a little accelerated growth during seedling stage but showed delayed flowering time and a slight growth retardation phenotype during late vegetative stage, suggesting that the OsNHX1 has a novel function in plant development. Northern and western blot analyses showed that the expression levels of OsNHX1 mRNA and protein in the leaves of three independent transgenic plant lines were significantly higher than in the leaves of wild type (WT) plants. T2 generation plants exhibited increased salt tolerance, showing delayed appearance and development of damage or death caused by salt stress, as well as improved recovery upon removal from this condition. Several physiological traits, such as increased Na+ content, and decreased osmotic potential in transgenic plants grown in high saline concentrations, further indicated that the transgenic plants had enhanced salt tolerance. Our results suggest the potential use of these transgenic plants for further agricultural applications in saline soil.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
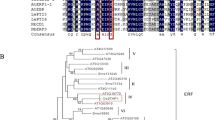
Na+/H+ antiporters, which catalyze the exchange of Na+ for H+ across membranes, are ubiquitous membrane proteins that play important roles in cellular pH and Na+ homeostasis throughout the biological kingdom. In plants, Na+/H+ antiporters are located in both plasma (Shi et al. 2002) and vacuolar (Apse et al. 1999) membranes, removing Na+ from the cytosol or compartmentalizing it in vacuoles for maintenance of a low Na+ concentration. In particular, the vacuolar Na+/H+ antiporter has been investigated and proposed to play an important role in salt tolerance (Blumwald 2000). Recently, with the molecular identification and biochemical characterization of the genes encoding these vacuolar membrane Na+/H+ antiporters, they have been isolated from many plants, including Arabidopsis thaliana (Apse et al. 1999), Oryza sativa (Fukuda et al. 1999, 2004b), Atriplex gmelili (Hamada et al. 2001), Brassica napus (Wang et al. 2003), Beta vulgaris (Xia et al. 2002), Gossypium hirsutum (Wu et al. 2004), and Hordeum vulgare (Fukuda et al. 2004a). These proteins are homologous and have a similar function. However, it is still unclear whether fine variations in activity among these orthologues may contribute to differences in physiological regulation.
Manipulating the vacuolar Na+/H+ antiporter to improve Na+ homeostasis is recognized as an attractive strategy in plants. Recently, AtNHX1 has been over-expressed in several dicotyledonous plants, including Arabidopsis (Apse et al. 1999), tomato (Lycopersion esculentum) (Zhang and Blumwald 2001), and Brassica napus (Zhang et al. 2001). These transgenic plants displayed robust salt tolerance and could grow normally and produce fruit and seeds under highly saline conditions (200 mM NaCl). More recently, AtNHX1 also has been introduced into crop plants such as wheat (Xue et al. 2004) and maize (Yin et al. 2004), improving salt tolerance. Moreover, over-expression of GhNHX1 in tobacco plants also enhanced salt tolerance (Wu et al. 2004). The examples above clearly demonstrate the feasibility of engineering salt tolerance into crop plants.
Rice is an important crop and mostly grown under continuously flooded conditions in lowlands. However, an ever-increasing water shortage has become the most serious constraint to rice production and yield stability in many rice-growing areas. Therefore, the non-flooded cultivation of upland rice is an alternative to lowland rice farming that could reduce the demand for irrigation water by 50–70% (Wang et al. 2002). However, upland rice, like lowland rice, has low salt tolerance. If salt tolerance were to be conferred upon upland rice through genetic engineering, the production of upland rice would be increased in saline soil and the plants would grow in a wider area. Thus, the persistent problem of food shortage in Asia and Africa could perhaps be alleviated to some extent through these methods. Recently, OsNHX1 has been over-expressed in rice (Fukuda et al. 2004b) and perennial ryegrass (Wu et al. 2005b). The AgNHX1 (Ohta et al. 2002) and bacterial nhaA (Wu et al. 2005a) genes have been introduced into rice as well, resulting in transgenic plants that displayed enhanced salt tolerance. However, this is the first report on improving the salt tolerance of upland rice by over-expressing a vacuolar Na+/H+ antiporter.
In the present study we produced transgenic upland rice plants over-expressing the OsNHX1, using an Agrobacterium-mediated transformation method, and investigated whether the salt tolerance in these plants could be improved by enhancing the level of OsNHX1 mRNA and protein. As the T2 generation plants were more salt tolerant, the exhibition of damage, or death, caused by high saline concentration was delayed or diminished in these plants. Furthermore, they also displayed improved recovery upon their removed from salt stress. In addition to being the first report that OsNHX1 has been over-expressed in upland rice, this is also the first description of the effects of OsNHX1 on the plant growth phenotype.
Materials and methods
Agrobacterium-mediated transformation and growth of transgenic plants
Mature seeds of upland rice Oryza sativa L. (cv IRAT109) were de-hulled, surface sterilized with 70% (v/v) ethanol for 5 min and exchanged with 0.1% (w/v) mercuric chloride (HgCl2) plus 0.1% (v/v) Tween-20 for 13–15 min. Following a rinse with sterile distilled water, the seeds were inoculated on callus induction medium ND2 (N6 medium (Zhu et al. 1975) plus 2.0 mg l−1 2,4-dichlorophenoxyacetic acid (2,4-D), 300 g l−1 casamino acid, 500 mg l−1 proline, 500 mg l−1 glutamine, 30 mg l−1 sucrose, 7 mg l−1 agar, pH 5.8) for callus induction. Cultures were transferred at 25°C in the dark.
Embryogenic calli were initiated from mature embryos and sub-cultured on ND2 medium 2–3 times at 21-day intervals and kept in the dark at 25°C. The embryonic calli were pre-cultured on fresh ND2 medium for 5 days, then infected with Agrobacterium tumefaciens LBA4404 containing the vector p3301/OsNHX1(Wu et al. 2005b) for 20–30 min; thereafter, they were transferred on ND2-AS (ND2 plus 100 μmol l−1 acetosyringone and 10 g l−1 glucose, pH 5.2) medium for 2–3 days in the dark at 25°C. We added 5 mg l−1, 10 mg l−1 and 20 mg l−1 phosphinotricine (PPT) in ND2 medium as selective media and transferred the co-cultivation calli on them at 2-week intervals. After 6 weeks of selection culture at 25°C in the dark, PPT-resistant calli were produced and regenerated into plants by being cultivated in regenerative medium, RE1-CH [Murashige and Skoog (MS) medium plus 20 mg l−1 PPT, 2 mg l−1 6-benzylaminopurine (6-BA), 0.25 mg l−1 naphthalene acetic acid (NAA), 30 mg l−1 sucrose, 8 mg l−1 agar, pH 5.8] for 15 days, then in RE2-CH (RE1-CH medium with NAA adjusted to 0.5 mg l−1) for an additional 15 days. Transgenic plants of the T0 and T1 generation were transplanted into soil and grown in a greenhouse under natural light and controlled temperature (28 ± 2°C daytime, 23 ± 2°C night) and relative humidity (50–60%). T2 generation plants were used in the analysis.
Polymerase chain reaction analysis
A polymerase chain reaction (PCR) strategy was used to identify the transgenic plants. The primers: P1 (5′-gcggtctgcaccatcgtaa-3′) and P2 (5′-gtaccggcaggctgaagtcca-3′), from the bar coding region, were used to amplify a 460 bp fragment. To avoid the interference of endogenous OsNHX1, we based the forward primer on the 35S promoter sequence. The primers for exogenous OsNHX1, P3 (5′-tcattgcgataaaggaaaggc-3′) and P4 (5′-acgaacaggttgatggacacc-3′) were used to amplify a total 453 bp DNA fragment (340 bp from the CaMV35S promoter region and 113 bp of the OsNHX1). The following PCR program was used: 94°C for 3 min, 40 cycles of 94°C for 30 s, 57°C (for 35S plus OsNHX1 fragments) or 58°C (for the bar gene) for 30 s, 72°C for 30 s, and 72°C for 10 min.
Northern and western blot analysis
Three-week-old seedlings of wild-type (WT) and transgenic plants cultured in vermiculite irrigated with half-strength MS solution were used for northern and western blot analyses. Before isolating tonoplast fractions, we treated seedlings of both WT and transgenic plants for 12 h with half-strength MS containing 0.25 mM NaCl. Total RNA (25 μg) isolated from leaf tissue was separated by electrophoresis on a 0.8% (w/v) agarose gel. The RNA was transferred onto Hybond N+ membrane (Boehringer Mannheim, Germany) and hybridized with a 536 bp fragment of OsNHX1 cDNA probe labeled with [32P] dCTP (Amersham Biosciences, USA) using a random primer labeling kit (TakaRa Biotechnology, Ltd., Dalian, China). The 536bp OsNHX1 DNA probe was amplified by PCR using the primers P5 (5′-gagcaccttccttggagtatttg-3′) and P6 (5′-gcaatcgacacagctcctctcat-3′). After hybridization for 18–20 h at 65°C, the membrane was washed once with 2× standard saline citrate (SSC) plus 0.1% sodium dodecyl sulfate (SDS) at 65°C for 30 min, then washed with 1× SSC plus 0.1% SDS at 65°C for 20 min. The membrane was exposed to X-ray film at −80°C for 7 days.
Polyclonal antibodies against the COOH-terminus of OsNHX1 (Fukuda et al. 1999, 2004b) were generously supplied by Dr. Fukuda. For western analysis, tonoplasts were isolated from the leaves of upland rice seedlings, as previously described (Fukuda et al. 1998, 2004b). Proteins were separated by 12% sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE). Blots were then incubated with a horseradish peroxidase (HRP)-goat anti-rabbit IgG (Sigma-Alrich Biotech Co., Missouri, USA) secondary antibody. Immunoreactive bands were detected by super enhanced chemiluminescence (ECL) (Applygene Tech Co. Ltd., Beijing, China).
Seed germination in salt solution and seedling growth in water
The WT and T2 transgenic upland rice seeds (lines L1, L3, and L5) were sterilized with 2.5% sodium hypochlorite for 15 min, washed thoroughly with water, then soaked in 9 cm Petri dishes with or without 100 mM NaCl for 5 days (2 days in the dark and 3 days in light). Each Petri dish contained two sheets of Whatman No. 1 filter paper, 36 seeds, and 20–30 ml solution. The seeds were germinated, and the resulting seedlings were grown in a growth chamber at 25°C with a light intensity of 100 μmol m−2 s−1 and 56–60% relative humidity. After 5 days salt treatment, seedlings were transferred to distilled water for 15 days under a growth cycle (12 h light, 12 h dark); during this period the water was exchanged every day. Growth was monitored photographically.
Salt-tolerant assay in agar and measurement of ion content
Evaluation of growth performance was carried out for T2 plants under salt-stress conditions. Seeds of these T2 and WT plants were surface sterilized with 70% ethanol for 5 min and with 0.1% HgCl2 for 12–15 min, rinsed with sterile water thoroughly, then germinated in the dark at 25°C on MS agar supplemented with (for T2 transgenic plants) or without (for WT plants) 20 mg l−1 PPT, and allowed to grow for 5 days. PPT-resistant transgenic and WT plants were then transferred to 9 cm plates with 25 ml MS agar. These MS media were supplemented with different concentrations of NaCl (0 mM, 50 mM, 100 mM, 150 mM, and 200 mM). The young seedlings were grown under identical conditions in water in Petri dishes. After 18 days of culture, representative plants were chosen and photographed. For ion content measurements, the fully expanded leaves and roots were separated (growing in flasks) and rinsed quickly with distilled water to wash off possible surface Na+ contamination. The roots were washed at least five times then oven-dried for 8 h at 80°C. We re-weighed the dried materials to obtain the dry weight, and the ion content was determined by atomic absorption spectrophotometry (Z-5000, Hitachi Instrument, Japan).
Plant growth and salt-stress treatment in soil
Seeds were germinated and grown in MS agar for 15 days then transferred into soil in trays (30 × 18 × 8 cm), with each tray containing three WT and six transgenic plants (two lines). The seedlings were grown for an additional 2 weeks before they were exposed to salt stress. Each experiment was repeated three to four times. Plants grown in different trays were watered with 900 ml solution containing different concentrations of NaCl (0 mM, 50 mM, 100 mM, and 200 mM) once and later irrigated with water. Seedlings were grown in a growth chamber at 25°C under a light intensity of 100–120 mol m−2 s−1, 12/12 h day/night period, and 56–60% relative humidity.
Osmotic potential measurements
Four-week-old seedlings of WT and transgenic plants were treated with different concentrations of NaCl (0 mM, 50 mM, 100 mM, and 200 mM) for 24 h. The fully expanded leaves were collected and stored at −20°C for 24 h. Juice was extracted from the leaves through a pillar by centrifugation at 1,000 r.p.m. for 10 min, and the osmotic potential of leaf juice was measured with an osmometer (O30 M, Gonotec, West Germany).
Results
Production and molecular analysis of transgenic upland rice plants
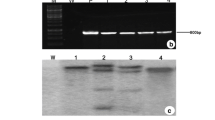
Mature embryo-derived calli of the upland rice cultivar IRAT109 were transformed with Agrobacterium strain LBA4404 harboring the p3301/OsNHX1 binary plasmid, and OsNHX1 was successfully introduced into the upland rice genome following the protocol of Hiei et al. 1994 and Huang et al. 2000 with some improvements as described in the Materials and methods section. More than 150 plantlets from seven independent transgenic calli lines carrying the additional OsNHX1 gene were identified by PCR amplification (Fig. 1A, B). Arrows indicate the target bar gene (460 bp) and 35S + OsNHX1 (453 bp) PCR products. The lower bands in each lane represent primer dimers. Three independent T2 generation lines from these transgenic plants were used in the experiments to follow. Northern blot analysis showed that these transgenic lines had higher levels of OsNHX1 transcript than did WT plants, with the transgenic lines 1–16 and 3–12 displaying higher levels of OsNHX1 transcript than transgenic lines 5–14 displayed (Fig. 1C). Immunoblots of tonoplast (vacuolar membrane) fractions isolated from the leaves of WT and transgenic plants also indicated that transgenic plants had enhanced expression of the vacuolar Na+/H+ antiporter (Fig. 1D). The growth performance of the transgenic plants was similar but slightly higher than WT seedlings under normal conditions. However, 35S::OsNHX1 transgenic plants displayed a clear retardation phenotype and a more prolonged vegetative stage than did the WT during the late vegetative growth phase (Fig. 2). On average, WT plants flowered and ripened 10 days earlier than did the transgenic plants under normal conditions.
PCR, northern-, and western-blot analyses of transgenic upland rice plants. A, B PCR products were amplified from the genomic DNA of independent T0 transgenic lines (1–7). A The bar (460 bp) gene was identified (arrow); B a PCR product (453 bp) between the 35S promoter and OsNHX1 was amplified (arrow); the lower band in each lane represents unincorporated PCR primers; marker III (M) (4,500 bp, 3,000 bp, 2,000 bp, 1,200 bp, 800 bp, 500 bp, 200 bp). p plasmid p3301/OsNHX1, ck wild type IRAT109. C Northern-blot analysis of OsNHX1 gene expression in wild-type and three independent lines (T2 generation). Both wild type and transgenic plants germinated and grew in vermiculite irrigated with half-strength MS solution. WT wild-type plants, 1 lines 1–16, 3 lines 3–12, 5 lines 5–14. D Western blot analysis of protein from the leaves of 3-week-old plants (seedlings of WT and transgenic plants treated with 25 mM NaCl for 12 h before isolation of tonoplast membrane proteins). Tonoplast proteins (12 μg) were probed with antibodies raised against the COOH-terminus of OsNHX1, revealing a reactive protein band located at 40 kDa
Growth performance of seedlings in water after salt stress treatment
To examine whether the enhanced expression of the vacuolar Na+/H+ antiporter conferred salt resistance to the plants, seeds of three transgenic lines and WT were germinated and grown in 100 mM NaCl solution or water for 5 days, then were transferred to water for 15 days. In water, nearly all transgenic and control seeds germinated well. Nearly no difference was seen in their seed germination rate. However, in 100 mM NaCl both transgenic and control plants germinated slowly, with a delay of about 2 or 3 days for the emergence of shoots and roots. After germination, seedlings were transferred into water. Here, both 35S::OsNHX1 and control seedlings recovered and resumed normal growth. However the 35S::OsNHX1 plants grew faster than the WT plants did during this recovery period. The shoots of transgenic plants were significantly taller than controls after 15 days growth, especially lines 1 and 3, which grew more vigorously than line 5 and the WT. This result correlated positively with the expression levels of OsNHX1 mRNA and protein (Figs. 1C, D and 3; Table 1). No significant difference was observed between WT and 35S::OsNHX1 plants when the seeds were germinated and grown continuously in water.
Over-expression of OsNHX1 improves the salt tolerance of upland rice by enhancing recovery upon the removal of salt stress conditions. A Seeds of WT and transgenic upland rice plants (lines 1–16, lines 3–12, and lines 5–14) were germinated and grown in water (without NaCl) for 20 days. B Seeds of WT and transgenic plants (lines 1–16, lines 3–12, and lines 5–14) were germinated in 100 mM NaCl for 5 days then transferred to water for 15 days
Transgenic plants have improved salt tolerance in agar under salt-stress conditions
To further confirm the enhanced salt tolerance of transgenic plants, we tested 5-day-old seedlings germinated in MS medium from seeds of both T2 generation transgenic and WT plants for their response to salt stress. The growth of both sets of seedlings was inhibited in agar media containing salt, and inhibition was aggravated progressively with increasing NaCl concentration, regardless of whether they were grown in plates (four–five seedlings in each plate) or flasks (ten seedlings in each flask). In the MS plus 200 mM NaCl agar plates, old leaves of WT plants started necrosis on the 11th day, and 75% had died by the 18th day. However, old leaves of transgenic plants did not undergo necrosis until the 15th day and were still alive on the 18th day (Fig. 4). Interestingly, the death of WT plants was delayed, and the old, lower leaves of these plants became yellower than those of transgenic plants grown in flasks (photograph not shown). Triplicate experiments on plants from lines 1–16, lines 3–12, and lines 5–14 produced similar results.
Salt-stress assay of wild-type and transgenic upland rice plants in agar medium. After 5 days of germination in MS medium, the wild type and transgenic plants (lines 1–16) were transferred to MS medium plus NaCl (0 mM, 50 mM, 100 mM, 150 mM, or 200 mM) in plates. After 18 days, the plants were photographed. A, B Plants (not shown on MS plus 50 mM NaCl medium) in plates. A) WT, B transgenic plants (lines 1–16)
Growth performance of transgenic plants in soil under salt-stress conditions
In order to investigate whether tolerance for high salinity soil was improved in the transgenic plants, we planted WT and transgenic four-leaf-stage plants in soil in trays and allowed them to grow and recover for 2 weeks, then we irrigated them with different NaCl solutions (50 mM, 100 mM, and 200 mM). We found that severe salt stress (200 mM NaCl) significantly affected the growth and survival of both transgenic and control plants. After being watered with 200 mM NaCl in soil, the seedlings of the control plants gradually wilted within 4 days and died within 1 week. In contrast, the wilting of transgenic seedlings was delayed by 3–4 days and the transgenic plants continued to survive for 2 weeks (photo not shown). Under more moderate salt stress, irrigation with 50 mM or 100 mM NaCl in each tray, the transgenic plants continued to grow for more than 1 month, whereas the WT plants died within 2 weeks.
Determination of the Na+ and K+ content in WT and transgenic plants
We determined the Na+ and K+ content in leaves and roots from both WT and transgenic plants grown under control and differing concentrations of NaCl in MS agar media (Fig. 5). As anticipated, the Na+ content in both the leaves and roots of WT and transgenic plants increased as the NaCl concentration was raised (Fig. 5A, B). However, the Na+ content of the leaves and roots of the transgenic plants grown in MS medium containing 200 mM or 50 mM NaCl was markedly higher than in WT plants grown in the same medium. These results indicated that the transgenic plants accumulated more vacuolar Na+ than did the WT plants under the same conditions. Conversely, the K+ content in leaves progressively decreased in both WT and transgenic plants with increasing NaCl concentration, and no significant difference between the WT and transgenic plants was observed, except in the 200 mM NaCl condition (Fig. 5C, D). In addition, the root K+ content of both WT and transgenic plants grown in concentrations between 50 mM and 200 mM NaCl was significantly lower than under normal conditions, and no significant difference was seen among the different NaCl concentrations. There were slightly lower K+ levels in transgenic plant roots than in WT plants.
Na+ and K+ content of leaves and roots of wild-type and transgenic plants over-expressing OsNHX1 grown in medium with various salt concentrations for 18 days. A, C Leaves; B, D roots. Solid black bar wild type plants, open bar lines 1–16 transgenic plants. Values shown are the mean ± SD (n = 4). The letters in each graph (a–d in A, B, a–c in C, D) indicate significant differences (P < 0.05, analysis of variance (ANOVA) and Tukey’s test). DW dry weight
Lower osmotic potential in transgenic plants grown under severe salt stress
We measured the leaf osmotic potential of both WT and transgenic plants grown in soil with a series NaCl concentrations, revealing that there was no difference between WT and transgenic plants grown in the range of 0–100 mM NaCl (Fig. 6). Nevertheless, transgenic plants treated with 200 mM NaCl had lower osmotic potentials than the controls had, suggesting that transgenic plants had absorbed more Na+ in their vacuoles. Thus, osmotic potentials decreased and the capacity of osmotic adjustment increased to absorb more water in transgenic plants.
Discussion
Increased expression levels of OsNHX1 mRNA and protein, and salt tolerance in transgenic upland rice plants over-expressing OsNHX1
Several dicotyledonous and monocotyledonous species, including Arabidopsis (Apse et al. 1999), tomato (Zhang and Blumwald 2001), Brassica napus (Zhang et al. 2001), wheat (Xue et al. 2004), and maize (Yin et al. 2004) over-expressing AtNHX1, a vacuolar Na+/H+ antiporter gene from Arabidopsis, have exhibited enhanced salt tolerance. In addition, Ohta et al. (Wu et al. 2005a) also observed that transgenic rice plants over-expressing this type of gene from a halophytic plant could survive 300 mM NaCl for 3 days, while WT plants could not. Fukuda 2004b recently reported that over-expression of OsNHX1 improved the salt tolerance of transgenic rice cells and plants. We obtained similar results in upland rice, further verifying the importance of OsNHX1 in salt tolerance of rice.
By assessing the response to salt stress in solution, medium, and soil, we found that over-expression of the OsNHX1 in the upland rice variety, IRAT109, improved salt tolerance. Symptoms of major damage caused by salt stress, such as wilting, necrosis, yellowing of old leaves, and death of older leaves or whole plants were moderated or delayed in transgenic plants (Fig. 4). Furthermore, enhanced salt tolerance was also exhibited when the salt stress was removed, transgenic plants displayed improved recovery in comparison with that of WT plants (Fig. 3). The three transgenic lines that we tested exhibited greatly improved performance over WT plants during salt stress. The increased salt tolerance of T2 generation plants correlated positively with the expression level of OsNHX1 transcript and protein (Fig. 1). Plants from lines 1–16 and lines 3–12 showed better salt tolerance than the plants from lines 5–14 did, indicating that increased expression of the rice vacuolar Na+/H+ antiporter confers enhanced salt tolerance in upland rice. This phenotype was reminiscent of rice over-expressing barley HVA1, a stress protein (Xu et al. 1996). These results further suggest the potential use of transgenic plants that over-express OsNHX1 for agricultural practice in otherwise arable saline soil.
Over-expression of OsNHX1 in upland rice increased the Na+ content and decreased osmotic potential under salt stress
Over-expression of OsNHX1 in lowland rice improved the salt tolerance of transgenic rice suspension cells and whole plants and correlated with increased Na+ content of rice suspension cells. However, it did not correlate with increased Na+ content of young leaves when cells were grown in high NaCl medium or when plants were grown in hydroponic solutions containing 50 mM or 100 mM NaCl (Fukuda et al. 2004b). Ohta et al. 2002 observed a similar result of enhanced salt tolerance in transgenic plants, without a significant difference of leaf Na+ content between WT and transgenic plants. These results implied that compartmentalization of Na+ in mature leaves or tissues may improve salt tolerance in rice plants.
In this report, we also monitored the Na+ content of transgenic upland rice treated with NaCl (0–200 mM) and found that the Na+ content in leaves of transgenic plants was higher than in WT controls under severe salt conditions, but not in moderate saline conditions (Fig. 5). These results clearly demonstrate that transgenic plants over-expressing the Na+/H+ antiporter has enhanced ability to efficiently compartmentalize sodium into vacuoles of leaf cells when faced with severe salt stress, thus reducing the osmotic potential of leaf cells and increasing water uptake capacity and, consequently, maintenance of turgor and survival [corroborated by measurements of leaf osmotic potential (Fig. 6)]. When plants were grown under low or moderate NaCl conditions, the Na+ was primarily accumulated in mature leaves. This Na+ exclusion to mature leaves effectively protected younger leaves from damage caused by excess Na+, thus maintaining their function (Tester and Davenport 2003). Moreover, a comparative analysis of Na+ accumulation in the roots of upland rice was performed for the first time in this study. In roots, the transgenic plants had higher levels of Na+ than the controls had at 50 mM NaCl. This may be due to greater Na+ transport into the vacuoles of root hair or cortex cells (Shi and Zhu 2002), thus reducing cytoplasmic or cell wall Na+ and consequently limiting Na+ transport into shoots through the xylem stream.
Determination of the osmotic potential of leaves from wild-type and transgenic plants grown under 0–200 mM NaCl. The osmotic potential of leaves from four-leaf-stage seedlings of WT and transgenic plants that were allowed to recover in soil for 1 week and were then treated with NaCl concentrations from 0 mM to 200 mM for 24 h. Fully expanded leaves were collected as material for this assay. Each value is the mean ± SD of four independent experiments. Triangles WT, (circles) lines 3–12
Previous studies (Kinclova-Zimmermannova et al. 2004) have shown that OsNHX1 has the ability to compartmentalize K+ into vacuoles from cytoplasm. However, the K+ content of suspension cells derived from transgenic rice (Fukuda et al. 2004b) or of roots from upland rice was less than or similar to the K+ content of control cells and roots under nearly all conditions. When treated with 200 mM NaCl, the leaf Na+ content of transgenic upland rice plants was higher than that of WT leaves, whereas the K+ content of leaves from transgenic plants was lower than the K+ content of leaves from WT plants (Fig. 5). These results suggest that Na+ is likely to be transported into cells through K+ carriers (Blumwald 2000; Fukuda et al. 2004b). Previous research has conjectured that Na+ can enter cells via several high-affinity transporters (such as the HAK or HKT potassium transporters) or low-affinity K+ transporter [i.e., LCT (Amtmann et al. 2001) and non-selective cation channels (NSCCs), including cyclic-nucleotide-gated channels and glutamate-activated channels, although the relative roles of each component seem likely to vary among species (Maser et al. 2002; Tester and Davenport 2003; Flowers 2004]. For example, parallel experiments measuring K+ and Na+ uptake in yeast expressing the wheat or rice HKT1 transporters verified that they were very different; TaHKT1 transported K+ and Na+, OsHKT1 only Na+ (Garciadeblas et al. 2003), while OsHKT2 was found to function as a Na+/K+ symporter (Horie et al. 2001). We believe that the explanation for upland rice’s having higher Na+ levels and lower K+ levels in the leaves or roots, respectively, under high salt conditions, is likely because K+ transport can be affected by Na+ levels through competition for the K+ binding site of potassium transporters (Blumwald 2000; Xue et al. 2004).
Recently, Apse et al. (2003) have reported that nhx1 plants, harboring a T-DNA insertion mutant of AtNHX1 from Arabidopsis, had much lower Na+/H+ and K+/H+ exchange activity in leaves, had altered leaf development, and had reduced frequency of large epidermal cells and overall leaf area in comparison with WT plants. Those results suggest that the vacuolar Na+/H+ antiporter(s) contribution to ion homeostasis is important not only for salinity tolerance but also for development (Apse et al. 2003). The 35S::OsNHX1 upland rice plants showed a little accelerated growth during seedling stage (Fig. 3, Table 1), but, in the late vegetative stage, transgenic plants exhibited delayed flowering and slight retardant growth (Fig. 2), indicating that OsNHX1 may also play an important role in rice development. However, the mechanism behind this remains to be further studied.
In summary, the physiological traits, including increased Na+ content and decreased osmotic potential, of transgenic plants grown under severe saline stress illustrate well that these transgenic plants had enhanced salt tolerance. This, of course, implies that these plants, over-expressing the OsNHX1, could be cultivated in saline soil. Furthermore, these transgenic plants also had herbicide resistance, because the bar gene was used as a selection gene during transformation. It is known that upland fields nearly always have more weeds to compete for nutrients and light than lowland fields have. Therefore, the application of bialaphos may make possible simultaneous weed control in upland fields, reducing the labor cost of cleaning weeds. This transgenic salt- and herbicide-resistant upland rice could be widely used in upland fields, providing significant economic value.
References
Amtmann A, Fischer M, Marsh EL, Stefaovic A, Sander D, Schachtman DP (2001) The wheat cDNA LCT1 generates hypersensitivity to sodium in a salt-sensitive yeast strain. Plant Physiol 126:1061–1071
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Apse MP, Sottosanto JB, Blumwald E (2003) Vacuolar cation/H+ exchange, ion homeostasis, and leaf development are altered in a T-DNA insertional mutant of AtNHX1, the Arabidopsis vacuolar Na+/H+ antiporter. Plant J 36:229–239
Blumwald E (2000) Sodium transport and salt tolerance in plants. Curr Opin Cell Biol 12:431–434
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55:307–319
Fukuda A, Yazaki Y, Ishikawa T, Koike S, Tanaka Y (1998) Na+/H+ antiporter in tonoplast vesicles from rice roots. Plant Cell Physiol 39:196–201
Fukuda A, Nakamura A, Tanaka Y (1999) Molecular cloning and expression of the Na+/H+ exchanger gene in Oryza sativa. Biochim Biophys Acta 1446:149–155
Fukuda A, Chiba K, Maeda M, Nakamura A, Maeshima M, Tanaka Y (2004a) Effect of salt and osmotic stresses on the expression of genes for the vacuolar H+-pyrophosphatase, H+-ATPase subunit A, and Na+/H+ antiporter from barley. J Exp Bot 55:585–594
Fukuda A, Nakamura A, Tagiri A, Tanaka H, Miyao A, Hirochika H, Tanaka Y (2004b) Function, intracellular localization and the importance in salt tolerance of a vacuolar Na+/H+ antiporter from rice. Plant Cell Physiol 45:146–159
Garciadeblas B, Senn M, Banuelos M, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801
Hamada A, Shono M, Xia T, Ohta M, Hayashi Y, Tanaka A, Hayakawa T (2001) Isolation and characterization of a Na+/H+ antiporter gene from the halophyte Atriplex gmelini. Plant Mol Biol 46:35–42
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Horie T, Yoshida K, Nakayama H, Yamada K, Oiki S, Shinmyo A (2001) Two types of HKT transporters with different properties of Na+ and K+ transport in Oryza sativa. Plant J 27:129–138
Huang JQ, Wei ZM, An HL, Xu SP, Zhang B (2000) High efficiency of genetic transformation of rice using Agrobacterium mediated procedure. Acta Bot Sin 42:1172–1178
Kinclova-Zimmermannova O, Flegelova H, Sychrova H (2004) Rice Na+/H+-antiporter Nhx1 partially complements the alkali–metal–cation sensitivity of yeast strains lacking three sodium transporters. Folia Microbiol (Praha) 49:519–525
Maser P, Gierth M, Schroedr J (2002) Molecular mechanisms of potassium and sodium uptake in plants. Plant Soil 247:43–54
Ohta M, Hayashi Y, Nakashima A, Hamada A, Tanaka A, Nakamura T, Hayakawa T (2002) Introduction of a Na+/H+ antiporter gene from Atriplex gmelini confers salt tolerance to rice. FEBS Lett 532:279–282
Shi H, Zhu JK (2002) Regulation of expression of the vacuolar Na+/H+ antiporter gene AtNHX1 by salt stress and abscisic acid. Plant Mol Biol 50:543–550
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Tester N, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot 91:1–25
Wang HQ, Bouman BAM, Zhao DL, Wang CG, Maoya PF (2002) Aerobic rice in northern China: opportunities and challenges. In: Bouman BAM, Hengsdijk H, Hardy B, Binddraban PS, Tuong TP, Ladha JK (eds) Water-wise rice production. International Rice Research Institute, Los Banos, Philippines, pp 143–154
Wang J, Zuo K, Wu W, Song J, Sun X, Lin J, Li X, Tang K (2003) Molecular cloning and characterization of a new Na+/H+ antiporter gene from Brassica napus. DNA Seq 14:351–358
Wu CA, Yang GD, Meng QW, Zheng CC (2004) The cotton GhNHX1 gene encoding a novel putative tonoplast Na+/H+ antiporter plays an important role in salt stress. Plant Cell Physiol 45:600–607
Wu LQ, Fan ZM, Guo L, Li YQ, Chen ZL, Qu LJ (2005a) Over-expression of the bacterial nhaA gene in rice enhances salt and drought tolerance. Plant Sci 168:297–302
Wu YY, Chen QJ, Chen M, Wang XC (2005b) Salt-tolerant transgenic perennial Ryegrass (Lolium perenne L.) obtained by Agrobacterium tumefaciens-mediated transformation of vacuolar Na+/H+ antiporter gene. Plant Sci 169:65–73
Xia T, Apse MP, Aharon GS, Blumwald E (2002) Identification and characterization of a NaCl-inducible vacuolar Na+/H+ antiporter in Beta vulgaris. Physiol Plant 116:206–212
Xu D, Duan X, Wang B, Hong B, David Ho T, Wu R (1996) Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110:249–257
Xue ZY, Zhi DY, Xue GP, Zhang H, Zhao YX (2004) Enhanced salt tolerance of transgenic wheat (Tritivum aestivum L.) expressing a vacuolar Na+/H+ antiporter gene with improved grain yields in saline soils in the field and a reduced level of leaf Na+. Plant Sci 167:849–859
Yin XY, Yang AF, Zhang KW, Zhang JR (2004) Production and analysis of transgenic maize improved salt tolerance by the introduction of AtNHX1 gene. Acta Bot Sin 46:854–861
Zhang HX, Blumwald E (2001) Transgenic salt-tolerant tomato plants accumulate salt in foliage but not in fruit. Nat Biotechnol 19:765–768
Zhang HX, Hodson JN, Williams JP, Blumwald E (2001) Engineering salt-tolerant Brassica plants: characterization of yield and seed oil quality in transgenic plants with increased vacuolar sodium accumulation. Proc Natl Acad Sci U S A 98:12832–12836
Zhu ZQ, Wang JJ, Sun JS, Xu Z, Yin GC, Zhu ZY, Bi FY (1975) Establishment of an efficient medium for another culture of rice through comparative experiments on the nitrogen sources. Sci Sin 18:659–668
Acknowledgments
We thank Dr. Fukada (Department of Plant Physiology, National institute of Agrobiological Resources, Japan) for kindly providing the vector: pOsNHX1, containing the OsNHX1 cDNA, as well as for supplying the OsNHX1 antibodies. This work was supported by grants from the National Basic Research Program of China (grant nos. 2006CB100100 and 2003CB114307) and from the National Science Foundation of China (grant nos. 30370129 and 30421002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, H., An, R., Tang, JH. et al. Over-expression of a vacuolar Na+/H+ antiporter gene improves salt tolerance in an upland rice. Mol Breeding 19, 215–225 (2007). https://doi.org/10.1007/s11032-006-9048-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-006-9048-8