Abstract
SHP2 is a protein tyrosine phosphatase (PTP) that can regulate the tyrosine phosphorylation level. Overexpression of SHP2 will promote the development of cancer diseases, so SHP2 has become one of the popular targets for the treatment of cancer. Studies have reported that both SHP099 and SHP844 are inhibitors of SHP2 and bind to different allosteric sites 1 and 2, respectively. Studies have shown that combining SHP099 with SHP844 will enhance pharmacological pathway inhibition in cells. This study uses molecular dynamic simulations to explore the dual allosteric targeted inhibition mechanism. The result shows that the residues THR108-TRP112 (allosteric site 1) move to LEU236-GLN245 (αB-αC link loop in PTP domain) , the residues of GLN79-GLN87 (allosteric site 2) get close to LEU262-GLN269 (αA-αB link loop in PTP domain) and HIS458-ARG465 (P-loop) come near to ARG501-THR507 (Q-loop) in SHP2-SHP099-SHP844 system, which makes the “inactive conformation” more stable and prevents the substrate from entering the catalytic site. Meanwhile, residue GLU110 (allosteric site 1), ARG265 (allosteric site 2), and ARG501 (Q-loop) are speculated to be the key residues that causing the SHP2 protein in auto-inhibition conformation. It is hoped that this study will provide clues for the development of the dual allosteric targeted inhibition of SHP2.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cancer, medically known as malignant neoplasm, has become a major problem that plagues the world. It has the characteristics of rapid increase in morbidity and mortality, which seriously threatens the health and safety of people [1]. Malignant tumors often have biological characteristics, such as abnormal cell differentiation and proliferation, uncontrolled growth, invasion and metastasis [2]. The occurrence is a complex process with multiple factors and multiple steps. At present, surgery, chemotherapy, and radiotherapy are the main methods of treating malignant tumors, but the curative ratio for patients is extremely low [3]. In search of a better treatment, targeted drug therapy has gradually become the focus of tumor research.
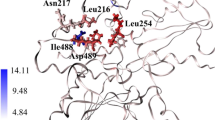
Some studies have reported that protein tyrosine phosphatase non-receptor type 11 (PTPN11; encoding protein SHP2) is related to cancer types, such as lung cancer, breast cancer, leukemia [4]. SHP2 is a protein tyrosine phosphatase (PTP) widely expressed in the human body. It can co-regulate the phosphorylation of tyrosine residues in the body with protein tyrosine kinases (PTK) [5]. Reversible phosphorylation of tyrosine residues in proteins is not only the key to regulate the activation of cell signal transduction pathways, but also to regulate cell growth, differentiation, metabolism, gene transcription and immune response [6,7,8]. SHP2 includes a trade N-SH2 domain (residues 3-104), an N-SH2 domain (residues 112-216), and a PTP catalytic area (residues 221-524). PTP catalytic area is composed by pTyr-loop (residues 277-284) which confers specificity to pTyr, WPD-loop (residues 421-431) which contains the general acid-base catalyst D425, P-loop (residues 458-465) which harbors the active site nucleophile C459 and R465 for recognition of the phosphoryl moiety in the substrate, and Q-loop (residues 501-507) which contains the conserved Q506 required to position and activated a water molecule for hydrolysis of the phosphoenzyme intermediate. Substrate could be dephosphorylated in PTP catalytic area [9], as shown in Fig. 1a. Substrate could be dephosphorylated in PTP catalytic area. SHP2 has two states: “close state”=“auto-inhibited conformation” and “open state”=“active conformation”. Inactive state (Fig. 1a), N-SH2 of SHP2 is inserted into PTP domain, which can prevent the substrate from entering the catalytic site. SHP2 plays an important role in multiple cell signal transduction pathways (RAS-ERK, JAK-STAT, PI3K-AKT, PD-1/PD-L1)[10,11,12]. SHP2 is the upstream positive regulator of Ras-Raf-MEK-ERK signaling pathway, and ERK can promote the development of cancer diseases. Inhibiting SHP2 to block the cancer-promoting function of Ras-Raf-MEK-ERK pathway is particularly important in the treatment of cancer diseases [13]. In addition, the expression of SHP2 is positively correlated with PD1 in T cells of patients with neck squamous cell carcinoma. When PD-1 is combined with PD-L1/L2, it can recruit SHP2 and release its auto-inhibition state, thereby blocking the activation of T cells. Therefore, it is very important to block the binding of PD-1 to SHP2 and releases the tumor-specific T-cell killing ability that is suppressed [14]. In short, SHP2 has become an important potential target for the treatment of cancer diseases.
a (PDB code: 6BMY) displaying the structure component of SHP2. b The allosteric site location of SHP2. The pink area is N-SH2, the yellow area is C-SH2, and the cyan area is PTP domain. A: The violet loop is pTyr–loop, the black loop is β5-β6 loop, and the green loop is WPD loop, the blue loop is P–loop, and the red loop is Q–loop. B: The SHP099 and SHP844 are color in blue and green, respectively
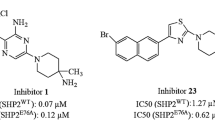
Since the first discovery of SHP2 as a potential anti-cancer target, the identification of pharmacologically relevant inhibitors has aroused widespread interest in the scientific community [15]. At present, SHP2 protein allosteric inhibitors could stabilize the combination of N-SH2 and PTP domain, which can make SHP2 protein keeping in auto-inhibition state [16]. However, an engineered, double mutations in the allosteric pocket (SHP2T253M/Q257L) disrupt the binding of SHP099 [17] and rescue pathway inhibition in cancer cells. After testing single- and double-mutant alleles, SHP2T253M/Q257L is found to retain the catalytic activity and auto-inhibited basal state of SHP2, but is 1000-fold less sensitive to SHP099 inhibition as compared to wild-type SHP2 in vitro [17]. These mutations facilitate release of allosteric ligands. Dual allosteric targeted protein inhibition can stabilize the auto-inhibition state of SHP2 protein through binding to different allosteric sites at the same time, which may improve the inhibition on SHP2 protein. Combining two distinct but compatible SHP2 inhibitors may offer advantages in enhancing SHP2 inhibition and overcoming resistance. Michelle Fodor and his team members tested the pharmacological activity of the SHP2 protein using a dual allosteric targeted protein inhibition method to inhibit the SHP2 protein. Combining SHP099 (allosteric site 1) with SHP844 [18] (allosteric site 2) interact with different allosteric sites in SHP2 protein shown in Fig. 1b. The structure of SHP099 and SHP844 can be seen in Fig. S1. SHP099 (IC50 value: 0.07 μM) is an effective, selective, highly soluble, SHP2 inhibitor with oral bioactivity [18]. It can inhibit the corresponding signaling pathways and has antitumor activity in a concentration-dependent manner in animal models of transplanted tumors. SHP099 simultaneously binds the interface of N-terminal SH2, C-terminal SH2 and protein tyrosine phosphatase domain, inhibits the activity of SHP2 through an allosteric mechanism, and stabilizes SHP2 in an auto-inhibiting conformation [19]. SHP844 is a weak SHP2 inhibitor with IC50 of 18.9 μM [18]. SHP844 enhances the inhibition of SHP099 in biochemical phosphatase experiments, and SHP844 enhances down regulation of the MAPK pharmacodynamic marker DUSP6 by SHP099 in cellular experiments [18]. Although SHP844 has less inhibitory activity on SHP2 than SHP099, the binding site of SHP844 is a new site. The crystal structure shows that SHP844 binds at the gap formed at the interface between the N-SH2 and the PTP domain, stabilizing the SHP2 concept of the auto-inhibited state, and thus plays a certain inhibitory role. Research showed that when the inhibitors (SHP099 and SHP244) acted simultaneously with the allosteric sites 1 and site 2 showed a certain synergistic effect on the pharmacological pathway. Therefore, the double target inhibition of SHP2 is feasible. However, the mechanism of the combining SHP099 and SHP844 inhibiting SHP2 at the molecular level is unclear. In order to promote the development of dual allosteric SHP2 inhibitor, a series of studies are performed to explore the changes of residues at allosteric sites and catalytic area.
The impact of molecular dynamics (MD) simulations on drug discovery is increasing [20,21,22]. MD methods simulate dynamic biological and chemical events at a molecular level, which can pave the way for drug development [23, 24]. For instance, Ali Rahimi's research group used MD simulation to simulate all atoms of poly (L-lysine) (PLL) dendrimer of different generations [25]. Magnus Lundborg's group used MD simulations to predict the permeability of drugs through the skin [26]. MD simulation can provide information about protein molecular levels on a reasonable time scale. Therefore, MD simulation and post-analysis (RMSD, RMSF, PCA, DCCM, ANM and RIN analyses) were performed to investigate the conformational changes of the SHP2 protein at the molecular level.
Materials and methods
The crystal structure of the complex of SHP2 protein and ligand (SHP099 and SHP244) was applied in this study (PDB code: 6BMY) [18]. The topology parameters files of the compounds SHP099 and SHP244 are obtained from the ACPYPE Portal website [27], which is designed to generate topology parameters files for unusual organic chemical compounds. SHP2 protein and ligand preparation are performed in Discovery Studio software. The methods in detail are in the supporting information. The MD simulations of SHP2, SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 system are performed by the AMBER99SB force field in GROMACS 4.5.5 [28,29,30].
MD simulations
The MD simulation process is carried out in GROMACS 4.5.5 [31]. MD simulation is applied to analyze the effect of dual allosteric inhibitors on the conformation of SHP2 protein [32]. First, the coordinate file (gro) and topology file of the SHP2 protein were generated using pdb2gmx program of the GROMACS package by AMBER99SB force field [33, 34]. In addition, it is necessary to merge the topology file and gro file of protein and ligand [35]. The coordinate file of the ligand is obtained by conversed from the Acpype Server-Bio2Byte website. Second, a regular cube box containing proteins is established through the edit-conf program and the distance from the edge of the protein to the box is less than 1 nm. Third, SPC water molecules are added into the cube box through the genbox program. The “amber99sb.ff/ions.itp” forcefield is used for Na+ and Cl–. Fourth, minimization is done. The energy is minimized using the steepest descent algorithm. There are two very important factors to evaluate whether EM (energy minimization) is successful. One factor is that the benchmark for convergence is either the maximum force of the system (<10 kJ/mol), another is no drastic energy changes during minimization steps. The system is then heated to 310 K during a 200 ps NVT (constant number of particles, volume, and temperature) simulation with 1 fs time step. The pressure is then equilibrated to 1 atm during a 500 ps NPT (constant number of particles, pressure, and temperature) simulation with 2 fs time step. The time constant for the temperature and pressure coupling is kept at 0.1 and 2 ps, respectively. Both temperature and pressure are regulated by V‐rescale, a modified Berendsen thermostat, and the pressure is regulated by the Parrinello-Rahman method [36]. The short-range electrostatic cutoff is 1.3 nm and the short-range van der Waals cutoff was 1.3 nm. The long-range electrostatic interactions are calculated using particle-mesh-Ewald (PME) summation method with fourth-order cubic interpolation and 0.16 Å grid spacing. All bonds are constrained using the parallel LINCS method [37]. Finally, the 100 ns MD simulation can be officially carried out.
Binding free energy
The calculation of free energy between ligand and protein has always been an important component in computer-aided drug design, and has been widely used in drug design [38, 39]. In order to understand the binding between protein and ligand, Alchemistry and MM-PBSA methods are applied in this study to investigate the binding strength of protein to ligand. Alchemistry first calculates the binding free energy of the intermediate state, and finally obtains the free energy between two physical end states. The thermodynamic cycle of alchemy is shown in Fig. 2. The calculation formula binding free energy of MM-PBSA is as follows:
Diagram of the thermodynamic cycle of protein-ligand complexes. The fully interacting ligand in the solution (a) is transformed into a non-interacting solute (b) in a series of equilibrium simulations, where the electrostatic and van der Waals interaction ratio is zero, providing ΔGsolvelec+vdw item. The ligand is restrained while still not interacting with the environment (c), calculate ΔGsolvrestr. This state is equivalent to restrain non-interacting ligands in the protein cavity (d). The electrostatic and vdw interactions with the restrained non-interacting ligands in the protein complex are reopened (e), thereby making ΔG protelec+vdw. Then remove the restriction between the ligand and the protein (ΔGprot restr) to end this cycle, and the final state is an unrestricted ligand complex that completely interacts with the protein (f)
ΔGgas represents the energy of the gas phase. ΔEint, ΔEele, and ΔEvdw represent internal energy, electrostatic and van der Waals contributions, respectively. The solvation free energy (ΔGsol) is composed of nonpolar (ΔGGB) and polar energy (ΔGSA). ΔGSA is estimated from the solvent–accessible surface area (SASA) obtained from a water probe radius of 1.4 Å. T and S corresponds to temperature and total solute entropy, respectively.
Principal component analysis
PCA (Principal component analysis),method, is one of the most widely accepted data dimensionality reduction algorithms [40]. In different fields, it is usually necessary to observe multiple data variables [41]. Therefore, PCA is used to analyze the conformational change information of SHP2 protein in MD simulation. The calculation formula as follows:
The Cartesian coordinates of the ith and jth Cα atoms are represented by xa and xb, respectively. Mean value during molecular dynamics simulation is represented by ⟨⟩.
Domain cross-correlation map analysis
DCCM can calculate the correlation between each amino acid Cα atom and other amino acid Cα atoms, which can effectively provide information on the movement of proteins in molecular dynamics simulations [42, 43]. Therefore, DCCM is utilized to analyze the movement of SHP2. The calculation formula is as follows:
Cij represents the strength of the correlation between atoms in the protein chain, ranging from - 1 to 1. Δri and Δrj represent the vector displacements of atoms i and j, respectively. In the research, the trajectory files of SHP2 are processed by Bio3D library and R-studio software, and the DCCM diagrams are finally obtained, which can show the correlation between atoms.
Residue interaction network
RIN is a commonly adopted method to analyze various interactions between residues and residues (VDW, H-bond, etc.) [44]. At present, RINs have been widely applied to analyze internal mutations, folding, and catalytic activities of proteins [45, 46]. In order to better analyze the interaction between protein and ligand, RINs analysis is performed on the files generated by MD simulation in this study.
Results and discussion
The reliability of the initial SHP2 structures
Ramachandran plot is often be accustomed to explain the dihedral angles (psi) of the bond between the Cα atom and the carbonyl C atom, the dihedral angles (phi) of the bond between the Cα atom and the N atom in the protein or peptide chain [47, 48]. At the same time, it can also reflect whether the conformation of the protein is reasonable. Therefore, ramachandran plot analysis of SHP2 protein could select out a reasonable protein conformation for MD simulation. In the ramachandran plot, it could be divided into three types of regions: the best region (blue region), the allowed region (purple region), and the not–allowed region. In addition, if the amino acid conformation of the best region and the allowable region accounts for more than 90% of the total amino acids, it illustrates that the protein model obeys the rules of stereochemistry. The simulation conformers extracted at intervals of 20-picoseconds were obtained from performing 10-nanosecond MD simulation. The Ramachandran plot of the 133th simulated conformation of the SHP2 protein is shown in Fig. S2, with 505 amino acids distributed in the allowed region and 18 amino acids distributed in the not–allowed region where amino acids distributed in the allowed region account for 96.6% of the total amino acids. Therefore, the 133th conformation of the protein was applied to the initial structure to run the MD simulation.
Stability analysis
In this study, the RMSD was an evaluative criterion to estimate the convergence of the simulation along the MD trajectories for SHP2,SHP2-SHP099, SHP2-SHP844 and SHP2-SHP099-SHP844 system, respectively. The “general properties” (total pressure, pressure tensor, density and energy) were also calculated to analyze the equilibration systems by g_energy and g_traj program. Based on these analyses, it is found that the RMSD calculated with respect to the first frame could better characterize the convergence. The RMSD analysis could be used as a method to assess the overall fluctuations of the main chain Cα atoms. The high the RMSD value was, the more flexible the main chain Cα atom was. In Fig. 3a, the results showed that the systems tend to be in equilibrium within 8–100 ns, and the average value was less than 0.4 nm. Therefore, MD files after 8 ns were utilized for post-MD analysis.
a: The RMSD of SHP2, SHP2-SHP099, SHP2-SHP844 and SHP2-SHP099-SHP844 system. The black line represents the outcome for the SHP2 system, the red line represents the outcome for the SHP2-SHP099 system, the blue line represents the outcome for the SHP2-SHP844 system, and the cyan line represents the outcome for the SHP2-SHP099-SHP844 system. b The RMSF of the side-chain atoms for SHP2 system and SHP2-SHP099-SHP844system. The yellow ellipse represents the areas not in the allosteric site or in the active that RMSF had large fluctuation. The black box represents the areas in the allosteric site or in the active that RMSF had large fluctuation
The effect of dual allosteric targeted protein inhibitors on the side–chain atoms of SHP2 protein were studied by RMSF. Figs. 3 and S3 were the RMSF figures that were generated by the SHP2,SHP2-SHP099, SHP2-SHP844 and SHP2-SHP099-SHP844 system at 8–100 ns, respectively. The smaller the RMSF value was, the more stable the side chain atoms were. The areas with large fluctuations in the two systems were marked with yellow ellipse and black box, respectively. In addition, there were the six residue regions (GLN79-GLN87, THR108-TRP112, LEU236-GLN245, LEU262-GLN269, HIS458-ARG465, and ARG501-THR507) marked by the black box and three regions (THR153-SER165, ASN200-GLN211, and HIS293-VAL301) marked by the yellow ellipse, respectively. The area marked by the yellow ellipse was not in the allosteric site area or in the active area, so it was not within the scope of this study. Residues THR108-TRP112 and LEU236-GLN245 belong to allosteric site 1, residues GLN79-GLN87, LEU262-GLN269 belong to allosteric site 2. Residues HIS458-ARG465 and ARG501-THR507 belong to P-loop and Q-loop, respectively. The average RMSF of residue GLN79-GLN87, THR108-TRP112, LEU236-GLN245, LEU262-GLN269, HIS458-ARG465, and ARG501-THR507 was 0.168, 0.128, 0.206, 0.142, 0.113 and 0.099 nm in SHP2 system, respectively, were 0.145, 0.093, 0.174, 0.159, 0.105, and 0.93 nm in SHP2-SHP099 system, respectively, were 0.108, 0.099, 0.196, 0.095, 0.125 and 0.087 nm in SHP2-SHP844, respectively. While in the SHP2-SHP099-SHP844 system, the average RMSF of residues GLN79-GLN87, THR108-TRP112, LEU236-GLN245, LEU262-GLN269, HIS458-ARG465, and ARG501-THR507 was 0.101, 0.100, 0.150, 0.099, 0.067 and 0.076 nm, respectively. Compared with the SHP2 system, in SHP2-SHP099 system, the value of RMSF at residues THR108-TRP112, LEU236-GLN245, were significantly reduced, and at residue GLN79-GLN87, HIS458-ARG465, and ARG501-THR507 was also slightly reduced. In the SHP2-SHP844 system, the value of RMSF at residues GLN79-GLN87 and LEU262-GLN269 was significantly reduced, and at residue THR108-TRP112, LEU236-GLN245 was also slightly reduced. While the value of RMSF at residues GLN79-GLN87, THR108-TRP112, LEU236-GLN245, LEU262-GLN269, HIS458-ARG465, and ARG501-THR507 was obviously reduced in SHP2-SHP099- SHP844. From the above data, it is found that the fluctuation of allosteric site 1 (residues THR108-TRP112, LEU236-GLN245) changed to a more stable state in SHP2-SHP099 system and the fluctuation of allosteric site 2 (GLN79-GLN87, LEU262-GLN269) also changed to a stable state in SHP2-SHP844. While, the residues fluctuation at allosteric site 1, allosteric site 2, and catalytic area had become more stable in SHP2-SHP099-SHP844 system. Therefore, RMSF analysis showed that combining SHP099 (an allosteric site 1 inhibitor) with SHP844 (an allosteric site 2 inhibitor) would enhance the stability of SHP2 residues fluctuations in allosteric and catalytic area, which would help SHP2 protein keep in inactive state.
Study on the binding stability of ligand to SHP2 in MD simulation
In order to further study the binding SHP099 and SHP844 to the internal residues of the SHP2 protein, the binding free energies of the SHP2-SHP099, SHP2-SHP844, SHP2-SHP099-SHP844 system were calculated by Gromacs.4.5.5 [49], respectively. Alchemical free energy calculations describe the change from state A to state B by defining the parameter λ. In this study, a 4ns molecular dynamics simulation was performed on 20 intermediate states. Table 1 shows the results obtained by the six calculation methods. The sum of the absolute values of the binding free energy of the SHP2-SHP099 and SHP2-SHP844 systems is less than that of the SHP2-SHP099-SHP844 system, which indicates that the combination of SHP099 and SHP844 in the SHP2-SHP099-SHP844 system is more stable than single system (SHP2-SHP099, SHP2-SHP844).
The calculated binding free energies were decomposed into the individual residue contributions using the MM/PBSA method to identify the key residues and their contributions for elucidating the SHP2 interactions with the SHP099 and SHP844. The stronger the interactions (H-bond, hydrophobic, conjugation interactions) between protein and ligand were, the more negative the calculated binding free energy value became. The binding energies of SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 system generated by the ligand and the protein mainly included (Table.S1): van der Waal energy (−98.593 +/−19.905 kJ/mol, −164.863 +/−14.521, and −492.983 +/−12.207), Electrostatic energy (−51.241 +/−8.047, −714.916 +/−12.834, and −661.303 +/−13.953 kJ/mol), polar solvation energy (107.251 +/−13.209, 441.402 +/−24.702, and 520.524 +/−34.560 kJ/mol ) and SASA energy(−7.566 +/−0.531, −17.675 +/−2.284, and −1.324 +/−1.092 kJ/mol) . The overall binding energy value was −50.149 +/−6.430, −456.052 +/−23.087, and −675.086 +/−21.846 kJ/mol, respectively. The sum of the absolute values of the binding free energy of the SHP2-SHP099 and SHP2-SHP844 systems is also less than that of the SHP2-SHP099-SHP844 system. Compared with alchemical method, the results obtained by MM-PBSA method are consistent, but alchemical method is more accurate and the error estimates are smaller.
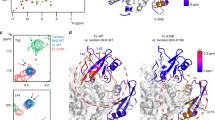
The binding energies between residues of allosteric sites (THR108, GLU110, ARG111, PHE113, GLU250, GLU252, THR253, PRO491, and LYS492) and SHP099 in SHP2-SHP099 system and SHP2-SHP099-SHP844 system are shown in Fig. 4a, respectively. The value of binding energies was 0.6178 +/−0.0360, 3.9424 +/−0.5699, −3.981 +/−0.3185, 0.3428 +/−0.0772, −2.0284 +/−0.1986, 0.5132 +/−0.0733, −4.4403 +/−0.3178, −1.2303 +/−0.3118 kJ/mol, and −2.2209 +/−0.2132 kJ/mol in SHP2-SHP099 system, respectively, whereas the value of binding energies were −7.2500 +/−0.3538, −24.2746 +/−0.9576, −10.6974 +/−0.4625, −5.6764 +/−0.3178, −24.6568 +/−0.1780, −25.5066 +/−1.3382, −8.0207 +/−0.9083, −7.2894 +/−0.2746 kJ/mol, and −23.0974 +/−1.9596 in the SHP2-SHP099-SHP844 system, respectively. By comparing the binding energy, is found that SHP099 bound more stably to the residues at the allosteric site 1 in the SHP2-SHP844-SHP099 system. Meanwhile, SHP844 bound to the residues GLN79, TYR80, GLN87, TYR263, ARG264, ARG265, GLN269, LYS280, and LYS266 (Fig. 4b), and the binding energy was 1.3294 +/−0.2365, −4.4566 +/−0.2741, −0.9685 +/−0.3327, 0.5631 +/−0.1346, −1.1838 +/−0.1226, −8.4755 +/−0.8240, −5.9755 +/−0.2975, −25.1962 +/−2.1586, and −20.6195 +/−0.9402 kJ/mol in SHP2-SHP844 system, respectively, while the binding energy were −2.8856 +/−0.6263, −27.9870 +/−2.5437, −3.1199 +/−0.0876, −58.7989 +/−3.2648, −2.1882 +/−0.1068, −5.6728 +/−0.7917, −8.1144 +/−1.2917, −36.6236 +/−3.0240, and −79.7206 +/−6.7512 kJ/mol in the SHP2-SHP099-SHP844 system, respectively. The binding ability of SHP844 at allosteric site 2 is significantly improved in SHP2-SHP099-SHP844 system. Therefore, according to the performance of the binding energy of the residues at the allosteric sites 1 and 2 with the ligands, it showed that SHP099 and SHP844 bound stronger in SHP2-SHP099-SHP844 system than that in single systems.
a: The binding free energy of SHP2-SHP099 system is represented by black stripe. The binding free energy of SHP2-SHP099-SHP844 system is represented by red stripe. b: The binding free energy of SHP2-SHP844 system is represented by black stripe. The binding free energy of SHP2-SHP099-SHP844 system is represented by red stripe
The H-bond interaction was monitored in 8–100 ns MD simulation of SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 systems, respectively, and the top five H-bond occupancy rates are shown in Fig. 4. In SHP2-SHP099 system, the H-bond occupancy rates of GLU250(O)–SHP099(H8), PHE113(O)–SHP099(H10), ASN217(D22)–SHP099(N22), ARG111(O)–SHP099(H10), and HIS114–SHP099(H10) were 63.0, 53.6, 34.0, 31.3, and 20.8%, respectively. In SHP2-SHP844 system, the H-bond occupancy rates of ASN218(D22)–SHP844(O5), LYS280(HZ3)–SHP844(O5), LYS274(HZ3)–SHP844(O4), LYS274(HZ3)–SHP844(O3), and LYS280(HZ3)–SHP844(O4) were 59.3, 56.9, 49.6, 46.9, and 42.8%, respectively. In SHP2-SHP099-SHP844 system, the H-bond occupancy rates of GLU250(O)–SHP099(H15), GLN79(E22)–SHP844(O2), PHE113(O)–SHP099(H21), TYR80(HH)–SHP844(O4), and LYS274(HZ3)–SHP844(N8) were 78.2, 76.3,56.3, 56.2, and 55.3%, respectively. Obviously, the H-bong occupancy rates had significant increase in SHP2-SHP099-SHP844 system, indicating that ligands SHP099 and SHP844 together formed stable H-bond interactions with SHP2 protein than either inhibitor alone. The analysis result was consisted with RMSF analysis.
Conformation transitions of the SHP2 and SHP2-SHP099-SHP844 system
The biological function of a protein is determined by its specific structure, and the intervention of the ligand will cause the change of conformations of the SHP2 protein and affect its function. PCA was utilized to obtain the information on the conformation status of SHP2, SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 system within 8–100 ns, respectively. Based on the calculation and diagonalization of the covariance matrix, principal component analysis was to project the protein dynamics information into the form of eigenvectors and eigenvalues. As shown in Figs. 5and S5, the top 20 PC values account for 77.5, 77.8, 80.1, and 79.3% of the total variation in the SHP2, SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 system, respectively. In the SHP2 system, the first two PCs accounted for 26.4 and 15.5% respectively, while the highest value of other PCs was only 8.7%. In the SHP2-SHP099 system, the first two PCs accounted for 36.6 and 10.2% respectively, while the value of other PCs was less than 5.5%. In the SHP2-SHP844 system, the first two PCs accounted for 25.9 and 22.7% respectively, while the value of other PCs was less than 8.4%. At the same time, the first two PCs in the SHP2-SHP099-SHP844 system accounted for 34.8 and 14.2% of the total variance, respectively, while the maximum value of the other PCs was not higher than 5.1%. Because the first two feature vectors (PC1, PC2) account for a large proportion of the overall protein conformation, the conformational transitions could be reflected by projecting the PC1 and PC2 into a two-dimensional (2D) map, respectively. The two conformational states marked as red dots and blue dots and the intermediate state are shown as white dots in the Figs. 5 and S5 . Compared with the SHP2 system, the red and blue dots in the SHP2-SHP099 system were concentrated in the middle position, the red and blue dots were distributed diagonally in SHP2-SHP844 system, the red dots and blue dots were distributed on both sides of the midline in the SHP2-SHP099-SHP844 system. According to the distribution of the conformation in the scatter diagram, the conjoined distributions of the PCs in the two systems revealed that the conformations of the SHP2 system were scattered and energetically less stable than that of the SHP2-SHP099-SHP844 system.
The fluctuations of the protein backbone of four system were displayed in the Anisotropic Network Model (ANM), which was done by PyoDy [50]. The off distance of ANM was 15, and the “ANM. calcModes ()” method was used to generate ANM modes. ANM could analyze how the SHP099 and SHP844 together affected the motions of allosteric sites 1 and 2 and the catalytic sites of SHP2, respectively [51]. The direction of the motion was indicated by the orientation of the arrow and the magnitude of the motion was expressed by the length of the arrow. As shown in Figs. 6 and S6, compared with the SHP2 system, the arrow length of residue THR108-TRP112 (yellow, allosteric site (1), LEU236-GLN245 (green, allosteric site 1), GLN79-GLN87 (orange, allosteric site (2), LEU262-GLN269 (pink, allosteric site 2), HIS458-ARG465 (red, catalytic area) and ARG501-THR507 (cyan, catalytic area) is significantly reduced in SHP2-SHP099-SHP844 system, which shows that the magnitude of the motion at above residue region is reduced. Meanwhile, the residues GLN79-GLN87 (orange) tends to be close to LEU262-GLN269 (pink) in SHP2-SHP099-SHP844 system, which indicates that the ligand (SHP099, SHP844) may lead to more stable amino acid binding at allosteric site 1. Moreover, after ligands (SHP099, SHP844) are bound to SHP2 protein, the overall fluctuations of the residues of the SHP2 protein are significantly reduced.
ANM analysis of the SHP2 system (a) and SHP2-SHP099-SHP844 system (b) calculated using ProDy and VMD programs. The regions of residues GLN79-GLN87, THR108-TRP114, LEU236-GLN245, and LEU262-GLN269, HIS458-ARG465 and ARG501-THR507 were colored in orange, yellow, green, pink, red, and blue, respectively
The correlation motion of SHP2 and SHP2-SHP099-SHP844 system
To further study the effects of SHP099 and SHP844 on the interactions between residues and residues of SHP2, DCCM analysis was performed on the SHP2, SHP2-SHP099, SHP2-844 and SHP2-SHP099-SHP844 system, respectively. In Figs. 7 and S7, the cyan and pink regions represent the positive and negative correlation of movement of specific residue, respectively. The darker the color was, the stronger the correlation was, while the white areas (0.25 to 0.25) represent no significant correlation. It can be seen from Figs. 7 and S7, that the cyan and pink regions were darker in the SHP2-SHP099-SHP844 system than that in SHP2 system, which pointed that the combination of SHP099 and SHP844 in the allosteric site enhanced the correlation movement of SHP2 residues. The area with obvious differences in Fig. 7 between the SHP2 and SHP2-SHP099-SHP844 system were marked by the black boxes. In Fig. 7, the areas where residue GLN79-GLN87 (N-SH2) forming correlation motion with LEU262-GLN269 (αB-αC link loop), HIS458-ARG465 (P-loop), and ARG501-THR507 (Q-loop), were marked as “a”, “b”, “c”, respectively. The areas where residue THR108-TRP112 (C-SH2) forming correlation motion with LEU236-GLN245 (αA-αB link loop) were marked as “d”. The residues ARG421-ASP431 (WPD-loop) forming correlation motion with HIS458-ARG465 (Q-loop) and ARG501-THR507 (P-loop) were marked as “e” and “f”, respectively. There was strong positive correlation motion at the area “a”, “b”, “c” “d” “e” and “f” in SHP2-SHP099-SHP844 system. However, these correlations were negative or weak positive in the SHP2, SHP2-SHP099, and SHP2-SHP844 system, revealing the interactions between residues GLN79-GLN87 and LEU262-GLN269, residues GLN79-GLN87 and HIS458-ARG465, residues GLN79-GLN87 and ARG501- THR507, residues THR108-TRP112 and LEU236-GLN245, residues ARG421-ASP431 and HIS458-ARG465, residues ARG421-ASP431 and ARG501-THR507 were stronger in SHP2-SHP099-SHP844 system than that in the other three systems. In addition, the correlation between the residues in the SHP2-SHP099-SHP844 system is overall enhanced, and the difference at the “d” region is the most obvious, changing from a negative correlation to a positive correlation. The positive correlation of the above-mentioned residue regions suggested that N-SH2 (allosteric site 1) got close to αB-αC link loop, P-loop, and Q-loop; C-SH2 (allosteric site 2) moved to αA-αB link loop; WPD-loop (catalytic area) came near to Q-loop, P-loop. It can be seen from the DCCM analysis results that the correlations at catalytic area, allosteric site (1, 2) in the SHP2-SHP099-SHP844 system are enhanced, which may be the reason for the decreased activity of SHP2.
DCCM for the Cα atoms fluctuations and their average positions of SHP2 (a) and SHP2-SHP099-SHP844 (b) systems during 8–100 ns of MD trajectories. Blue represents positive correlations, whereas red represents negative correlations. The areas marked represent that the correlations of residual have obviously transformation. The a, b, c, d, e, and f regions represent the correlation motion between residues GLN79-GLN87 and LEU262-GLN269, residues GLN79-GLN87 and HIS458-ARG465, residues GLN79-GLN87 and ARG501-THR507, residues THR108-TRP112 and LEU236-GLN245, residues ARG421-ASP431 and HIS458-ARG465, residues ARG421-ASP431 and ARG501-THR507
The ligand SHP-099-844 influencing the interactions of the residues in SHP2
By analyzing the RIN of SHP2 system and protein-ligand system, the effect of ligands on the structure and function of protein can be further explained. RIN was generated based on the average trajectory file, and could display various interactions between amino acids (for example: H-bond, VDW, electrostatic). There were two important parameters (shortest path betweenness and closeness centrality) in RIN analysis. Residues with high shortest path betweenness value could stabilize the structure of the protein, and residues with high closeness centrality value could affect the function of the protein. [52,53,54] The values of the shortest path betweenness and closeness centrality for the SHP2, SHP2-SHP099, SHP2-SHP844, and SHP2-SHP099-SHP844 system are shown in Table.S2, respectively. The notable residues were the residues with the value of shortest path betweenness greater than 0.1. The notable residues included: ARG5, PHE7, PHE41,TYR63, PHE71, and CYS104 (N-SH2), PHE135, THR191, and VAL194 (C-SH2), ILE282, ASN306, GLN506, and PHE513 (PTP domain) in SHP2 system; PHE7, PHE41, ASN58, and ASP61 (N-SH2), GLU232, PHE247, PHE251, GLU252, ILE463, GLN506, and TYR511 (PTP domain) in the SHP2-SHP099 system; ASP61 (N-SH2), GLU250, PHE251, TRP342, PHE469, ILE472, GLN506, TYR511, and ILE514 (PTP-domain) in SHP2-SHP844 system; N-SH2 (GLU110), PHE113, VAL137 (C-SH2), GLU250, GLU252, THR253, GLN255, GLU258, ARG265, ASN306, ARG498, and ARG501 (PTP domain) in the SHP2-SHP099-SHP844 system. In comparison, the notable residue of the SHP2-SHP99-SHP844 system included more residues around allosteric sites 1, 2, indicating that the connection between the residues was observed to be tightened. Moreover, the shortest path betweenness value of notable residue GLU110 (allosteric site 1), ARG501 (Q-loop), and ARG265 (allosteric site 2) were 0.196, 0.144, and 0.101 in SHP2-SHP099-SHP844 system, respectively, while the value were 0.019, 0.038, and 0.019 in SHP2-SHP099 system, respectively, were 0.083, 0.056, and 0.028 in SHP2-SHP844 system, were 0.004, 0.047, and 0.035 in SHP2 system, respectively. Compared to the other three systems, the Shortest Path Betweenness value of notable residues GLU110, ARG501, and ARG265 had obviously increased. Therefore, GLU110, ARG501, and ARG265 were key residues to keep SHP2 in the inactive state when SHP099 and SHP844 bound to allosteric sites 1 and 2 of SHP2.
The results of the interactions between residues and residues at the allosteric site 1, allosteric site 2 and the binding site of the SHP2, SHP2-SHP099, SHP2-SHP844 and the SHP2-SHP099-SHP844 system are shown in Figs. 8 and S8. In RINs, the network edge line styles corresponded to no covalent and covalent residue interactions that were preserved in both two systems (black solid lines), presented only in the SHP2 system (green dotted lines) [55], or only in the SHP2-ligand system (red dashed lines). It can be seen from the Fig. 8 that the interactions between the residues at allosteric site 1 were significantly increased in the SHP2-SHP099-SHP844 system than that in SHP2-SHP099 system, and the interactions between the residues at allosteric site 2 were also obviously increased in the SHP2-SHP099-SHP844 system than that in SHP2-SHP844 system. By comparing RINs of four systems, the interactions between the key regions in the SHP2-SHP099-SHP844 systems were increased significantly due to the existence of dual ligands. The interactions of key residues GLU110 (allosteric site 1), ARG501 (Q-loop), and ARG265 (allosteric site 2) made it easy to maintain the inactive state of SHP2, and the catalytic area are difficult to expose the active site, when SHP099 and SHP844 bound to allosteric sites 1 and 2 of SHP2. RIN analysis was consistent with RMSD, RMSF, DCCM, and ANM analysis results.
The interactions of the key residues for allosteric site 1, allosteric site 2, and catalytic area in the comparison network of protein systems. The edge styles correspond to mutual interactions that are preserved in both complexes (black solid lines), only in SHP2 system (green dashed lines) or only in SHP2-SHP099-SHP844 (red dotted lines)
Conclusion
The purpose of this study was to investigate the effects of dual allosteric inhibitors on SHP2 protein and to provide new ideas for the development of SHP2 protein inhibitors. First of all, 100–ns molecular dynamics simulations were performed for the SHP2, SHP2-099, SHP2-SHP844, SHP2-SHP099-SHP844 system, respectively. Then, post-MD simulation analyses (RMSD, RMSF, PCA, ANM, DCCM, and RIN) were applied to explore the mechanism of dual allosteric SHP2 inhibitors on SHP2 protein. According to the results of RMSD, RMSF, it showed that SHP099 bound to residues THR108, GLU110, ARG111, PHE113, GLU250, GLU252, THR253, PRO491, LYS492 and SHP844 bound to residues GLN79, TYR80, GLN87, LEU262, TYR263, ARG265, GLN269, causing the decrease in fluctuations of residues THR108-TRP112 (N-SH2 and C-SH2 link loop), LEU236-GLN245 (C-SH2 and PTP link loop), GLN79-GLN87 (N-SH2), LEU262-GLN269 (αB and αC link loop of PTP domain), HIS458-ARG465 (P-loop), ARG501-THR507 (Q-loop). Furthermore, the PCA, ANM, DCCM analysis suggested that the conformations of SHP2 were more stable and the interactions between residue areas (allosteric site 1, allosteric site 2, P-loop, and Q-loop) had visibly enhanced due to the combining SHP099 with SHP844, causing the active site was hardly to be exposed. Finally, RIN result speculated that the residues GLU110, ARG265, and ARG501 were the key residues, which made the residues of allosteric site 1, allosteric site 2, and catalytic area to be tightened. Above all, those results provided the comprehensive structural information to understand the effects of dual allosteric inhibitors on SHP2 protein.
Abbreviations
- ANM:
-
Anisotropic network model
- DCCM:
-
Domain cross-correlation map
- DS:
-
Discovery studio
- EM:
-
Energy minimization
- MD:
-
Molecular dynamics
- NPT:
-
Constant number of particles, pressure, and temperature
- NVT:
-
Constant number of particles, volume, and temperature
- PCA:
-
Principal component analysis
- PTK:
-
Protein tyrosine kinases
- PLL:
-
Poly (L-lysine)
- PTP:
-
Protein tyrosine phosphatase
- PTPN11:
-
Protein tyrosine phosphatase non-receptor type 11
- RIN:
-
Residue interaction network
- RMSD:
-
Root mean square deviation
- RMSF:
-
Root mean square fluctuation
References
Li M, Maso LD, Vaccarella S (2020) Global trends in thyroid cancer incidence and the impact of overdiagnosis. The Lancet Diabetes Endocrinol 8(6):468–470. https://doi.org/10.1016/S2213-8587(20)30115-7
Khazaei S, Rezaeian S, Khazaei S, Mansori K, Sanjari Moghaddam A, Ayubi E (2016) Effects of human development index and its components on colorectal cancer incidence and mortality: a global ecological study. Asian Pacific J Cancer Prevention : APJCP 17(S3):253–256. https://doi.org/10.7314/apjcp.2016.17.s3.253
Nolan GS, Wormald JCR, Kiely AL, Totty JP, Jain A (2020) Global incidence of incomplete surgical excision in adult patients with non-melanoma skin cancer: study protocol for a systematic review and meta-analysis of observational studies. Syst Rev 9(1):83. https://doi.org/10.1186/s13643-020-01350-5
Idrees M, Xu L, Song SH, Joo MD, Lee KL, Muhammad T, El Sheikh M, Sidrat T, Kong IK (2019) PTPN11 (SHP2) Is Indispensable for Growth Factors and Cytokine Signal Transduction During Bovine Oocyte Maturation and Blastocyst Development. Cells 8 (10). doi:https://doi.org/10.3390/cells8101272
Grossmann KS, Rosario M, Birchmeier C, Birchmeier W (2010) The tyrosine phosphatase Shp2 in development and cancer. Adv Cancer Res 106:53–89. https://doi.org/10.1016/S0065-230X(10)06002-1
Griger J, Schneider R, Lahmann I, Schowel V, Keller C, Spuler S, Nazare M, Birchmeier C (2017) Loss of Ptpn11 (Shp2) drives satellite cells into quiescence. Elife. https://doi.org/10.7554/eLife.21552
Spalinger MR, McCole DF, Rogler G, Scharl M (2016) Protein tyrosine phosphatase non-receptor type 2 and inflammatory bowel disease. World J Gastroenterol 22(3):1034–1044. https://doi.org/10.3748/wjg.v22.i3.1034
Tiganis T, Bennett AM (2007) Protein tyrosine phosphatase function: the substrate perspective. The Biochem J 402(1):1–15. https://doi.org/10.1042/BJ20061548
Zhang X, He YT, Liu SJ, Yu ZH, Jiang ZX, Yang ZY, Dong YS, Nabinger SC, Wu L, Gunawan AM, Wang LN, Chan RJ, Zhang ZY (2010) Salicylic acid based small molecule inhibitor for the oncogenic src homology-2 domain containing protein tyrosine phosphatase-2 (SHP2). J Med Chem 53(6):2482–2493. https://doi.org/10.1021/jm901645u
Bunda S, Burrell K, Heir P, Zeng L, Alamsahebpour A, Kano Y, Raught B, Zhang ZY, Zadeh G, Ohh M (2015) Inhibition of SHP2-mediated dephosphorylation of Ras suppresses oncogenesis. Nat Commun 6:8859. https://doi.org/10.1038/ncomms9859
Ehrman LA, Nardini D, Ehrman S, Rizvi TA, Gulick J, Krenz M, Dasgupta B, Robbins J, Ratner N, Nakafuku M, Waclaw RR (2014) The protein tyrosine phosphatase Shp2 is required for the generation of oligodendrocyte progenitor cells and myelination in the mouse telencephalon. J Neurosci 34(10):3767–3778. https://doi.org/10.1523/JNEUROSCI.3515-13.2014
Agazie YM, Hayman MJ (2003) Molecular mechanism for a role of SHP2 in epidermal growth factor receptor signaling. Mol Cell Biol 23(21):7875–7886. https://doi.org/10.1128/mcb.23.21.7875-7886.2003
Easton JB, Royer AR, Middlemas DS (2006) The protein tyrosine phosphatase, Shp2, is required for the complete activation of the RAS/MAPK pathway by brain-derived neurotrophic factor. J Neurochem 97(3):834–845. https://doi.org/10.1111/j.1471-4159.2006.03789.x
Yuan X, Bu H, Zhou J, Yang CY, Zhang H (2020) Recent Advances of SHP2 inhibitors in cancer therapy: current development and clinical application. J Medicin Chem. https://doi.org/10.1021/acs.jmedchem.0c00249
LaRochelle JR, Fodor M, Xu X, Durzynska I, Fan L, Stams T, Chan HM, LaMarche MJ, Chopra R, Wang P, Fortin PD, Acker MG, Blacklow SC (2016) Structural and functional consequences of three cancer-associated mutations of the oncogenic phosphatase SHP2. Biochemistry 55(15):2269–2277. https://doi.org/10.1021/acs.biochem.5b01287
Garcia Fortanet J, Chen CH, Chen YN, Chen Z, Deng Z, Firestone B, Fekkes P, Fodor M, Fortin PD, Fridrich C, Grunenfelder D, Ho S, Kang ZB, Karki R, Kato M, Keen N, LaBonte LR, Larrow J, Lenoir F, Liu G, Liu S, Lombardo F, Majumdar D, Meyer MJ, Palermo M, Perez L, Pu M, Ramsey T, Sellers WR, Shultz MD, Stams T, Towler C, Wang P, Williams SL, Zhang JH, LaMarche MJ (2016) Allosteric Inhibition of SHP2: identification of a potent, selective, and orally efficacious phosphatase inhibitor. J Med Chem 59(17):7773–7782. https://doi.org/10.1021/acs.jmedchem.6b00680
Chen YN, LaMarche MJ, Chan HM, Fekkes P, Garcia-Fortanet J, Acker MG, Antonakos B, Chen CH, Chen Z, Cooke VG, Dobson JR, Deng Z, Fei F, Firestone B, Fodor M, Fridrich C, Gao H, Grunenfelder D, Hao HX, Jacob J, Ho S, Hsiao K, Kang ZB, Karki R, Kato M, Larrow J, La Bonte LR, Lenoir F, Liu G, Liu S, Majumdar D, Meyer MJ, Palermo M, Perez L, Pu M, Price E, Quinn C, Shakya S, Shultz MD, Slisz J, Venkatesan K, Wang P, Warmuth M, Williams S, Yang G, Yuan J, Zhang JH, Zhu P, Ramsey T, Keen NJ, Sellers WR, Stams T, Fortin PD (2016) Allosteric inhibition of SHP2 phosphatase inhibits cancers driven by receptor tyrosine kinases. Nature 535(7610):148–152. https://doi.org/10.1038/nature18621
Fodor M, Price E, Wang P, Lu H, Argintaru A, Chen Z, Glick M, Hao HX, Kato M, Koenig R, LaRochelle JR, Liu G, McNeill E, Majumdar D, Nishiguchi GA, Perez LB, Paris G, Quinn CM, Ramsey T, Sendzik M, Shultz MD, Williams SL, Stams T, Blacklow SC, Acker MG, LaMarche MJ (2018) Dual allosteric inhibition of SHP2 phosphatase. ACS Chem Biol 13(3):647–656. https://doi.org/10.1021/acschembio.7b00980
Sun XJ, Ren Y, Gunawan S, Teng P, Chen ZM, Lawrence HR, Cai JF, Lawrence NJ, Wu J (2018) Mutation selectivity of the allosteric SHP2 inhibitor SHP099. Cancer Res 78 (13). doi:https://doi.org/10.1158/1538-7445.AM2018-4879
Ravindran A, Anishetty S, Pennathur G (2020) Molecular dynamics of the membrane interaction and localisation of prodigiosin. J Mol Graph Model 98:107614
Durrant JD, McCammon JA (2011) Molecular dynamics simulations and drug discovery. Bmc Biol 9(1):1–9
Salsbury FR (2010) Molecular dynamics simulations of protein dynamics and their relevance to drug discovery. Curr Opin Pharmacol 10(6):738–744. https://doi.org/10.1016/j.coph.2010.09.016
Borhani DW, Shaw DE (2012) The future of molecular dynamics simulations in drug discovery. J Comput-Aided Mol Design 26(1):15–26. https://doi.org/10.1007/s10822-011-9517-y
Perilla JR, Hadden JA, Goh BC, Mayne CG, Schulten K (2016) All-atom molecular dynamics of virus capsids as drug targets. J Phy Chem Lett 7(10):1836–1844. https://doi.org/10.1021/acs.jpclett.6b00517
Rahimi A, Amjad-Iranagh S, Modarress H (2016) Molecular dynamics simulation of coarse-grained poly(L-lysine) dendrimers. J Mol Model 22 (3). doi:ARTN 5910.1007/s00894-016-2925-0
Lundborg M, Wennberg CL, Narangifard A, Lindahl E, Norlen L (2018) Predicting drug permeability through skin using molecular dynamics simulation. J Control Release 283:269–279. https://doi.org/10.1016/j.jconrel.2018.05.026
Sousa da Silva AW, Vranken WF (2012) ACPYPE - Antechamber python parser interface. BMC Res Notes 5:367. https://doi.org/10.1186/1756-0500-5-367
Hess B (2009) GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. Abstr Pap Am Chem S 237
Pronk S, Pall S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29 (7):845-854. doi:https://doi.org/10.1093/bioinformatics/btt055
Rampogu S, Son M, Baek A, Park C, Rana RM, Zeb A, Parameswaran S, Lee KW (2018) Targeting natural compounds against HER2 kinase domain as potential anticancer drugs applying pharmacophore based molecular modelling approaches. Comput Biol Chem 74:327–338. https://doi.org/10.1016/j.compbiolchem.2018.04.002
Makarewicz T, Kazmierkiewicz R (2013) Molecular Dynamics simulation by GROMACS using GUI plugin for PyMOL. J Chem Inf Model 53(5):1229–1234. https://doi.org/10.1021/ci400071x
Gruebele M (2014) Protein dynamics in simulation and experiment. J Am Chem Soc 136(48):16695–16697. https://doi.org/10.1021/ja510614s
Nava M (2018) Implementing dimer metadynamics using gromacs. J Comput Chem 39(25):2126–2132. https://doi.org/10.1002/jcc.25386
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA, Wei S, Buckner J, Jeong JC, Qi Y, Jo S, Pande VS, Case DA, Brooks CL 3rd, MacKerell AD Jr, Klauda JB, Im W (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405–413. https://doi.org/10.1021/acs.jctc.5b00935
Rakhshani H, Dehghanian E, Rahati A (2019) Enhanced GROMACS: toward a better numerical simulation framework. J Mol Model 25 (12). doi:Artn 35510.1007/S00894-019-4232-Z
Bealing C, Fugallo G, Martonak R, Molteni C (2010) Constant pressure molecular dynamics simulations for ellipsoidal, cylindrical and cuboidal nano-objects based on inertia tensor information. Phy Chem Chem Phy 12(30):8542–8550. https://doi.org/10.1039/c004053c
Hess B (2008) P-LINCS: a parallel linear constraint solver for molecular simulation. J Chem Theory Comput 4(1):116–122. https://doi.org/10.1021/ct700200b
Wang L, Chambers J, Abel R (2019) Protein-ligand binding free energy calculations with FEP. Methods in molecular biology 2022:201–232. https://doi.org/10.1007/978-1-4939-9608-7_9
Irwin BWJ, Huggins DJ (2018) Estimating atomic contributions to hydration and binding using free energy perturbation. J Chem Theory Comput 14(6):3218–3227. https://doi.org/10.1021/acs.jctc.8b00027
Giuliani A (2017) The application of principal component analysis to drug discovery and biomedical data. Drug Discov Today 22(7):1069–1076. https://doi.org/10.1016/j.drudis.2017.01.005
Latry P, Martin-Latry K, Labat A, Peter C, Molimard M (2010) Use of principal component analysis in the evaluation of adherence to statin treatment: a method to determine a target population for public health intervention. Fund Clin Pharmacol 24:70–70
Arnold GE, Ornstein RL (1997) Molecular dynamics study of time-correlated protein domain motions and molecular flexibility: cytochrome P450BM-3. Biophys J 73(3):1147–1159. https://doi.org/10.1016/S0006-3495(97)78147-5
Upadhyay SK (2014) Dynamics of Gal80p in the Gal80p-Gal3p complex differ significantly from the dynamics in the Gal80p-Gal1p complex: implications for the higher specificity of Gal3p. Mol Biosyst 10(12):3120–3129. https://doi.org/10.1039/c4mb00371c
Liu R, Hu J (2011) Computational prediction of heme-binding residues by exploiting residue interaction network. Plos One 6(10):e25560. https://doi.org/10.1371/journal.pone.0025560
Hu G, Yan WY, Zhou JH, Shen BR (2014) Residue interaction network analysis of Dronpa and a DNA clamp. J Theor Biol 348:55–64. https://doi.org/10.1016/j.jtbi.2014.01.023
Xia Q, Ding YR (2019) Thermostability of lipase A and dynamic communication based on residue interaction network. Protein and Peptide Lett 26(9):702–716. https://doi.org/10.2174/0929866526666190617091812
Mannige RV, Kundu J, Whitelam S (2016) The Ramachandran Number: An Order Parameter for Protein Geometry. Plos One 11 (8). doi:ARTN e016002310.1371/journal.pone.0160023
Ho BK, Brasseur R (2005) The Ramachandran plots of glycine and pre-proline. Bmc Struct Biol 5:14. https://doi.org/10.1186/1472-6807-5-14
Klimovich PV, Mobley DL (2015) A Python tool to set up relative free energy calculations in GROMACS. J Comput-Aided Mol Design 29(11):1007–1014. https://doi.org/10.1007/s10822-015-9873-0
Eyal E, Lum G, Bahar I (2015) The anisotropic network model web server at 2015 (ANM 2.0). Bioinformatics 31 (9):1487-1489. doi:https://doi.org/10.1093/bioinformatics/btu847
Sarkar R (2018) Stiffening of flexible SUMO1 protein upon peptide-binding: analysis with anisotropic network model. Math Biosci 295:67–72. https://doi.org/10.1016/j.mbs.2017.11.008
Singh SS, Khundrakpam B, Reid AT, Lewis JD, Evans AC, Ishrat R, Sharma BI, Singh RK (2016) Scaling in topological properties of brain networks. Sci Rep 6:24926. https://doi.org/10.1038/srep24926
Jiao X, Chang S, Li CH, Chen WZ, Wang CX (2007) Construction and application of the weighted amino acid network based on energy. Physical Review E Stat Nonlinear Soft Matter Physics 75(5 Pt 1):051903. https://doi.org/10.1103/PhysRevE.75.051903
Xue W, Jiao P, Liu H, Yao X (2014) Molecular modeling and residue interaction network studies on the mechanism of binding and resistance of the HCV NS5B polymerase mutants to VX-222 and ANA598. Antiviral Res 104:40–51. https://doi.org/10.1016/j.antiviral.2014.01.006
Jiao X, Ranganathan S (2017) Prediction of interface residue based on the features of residue interaction network. J Theor Biol 432:49–54. https://doi.org/10.1016/j.jtbi.2017.08.014
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant No. 81773569), the Natural Science Foundation of Tianjin (Grant No. 18JCQNJC13700), the Science & Technology Development Fund of Tianjin Education Commission for Higher Education (Grant No. 2017KJ229).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors report no conflicts of interest in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yangchun, M., WenYu, Y., Liang, Z. et al. Exploring the cause of the dual allosteric targeted inhibition attaching to allosteric sites enhancing SHP2 inhibition. Mol Divers 26, 1567–1580 (2022). https://doi.org/10.1007/s11030-021-10286-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-021-10286-4