Abstract
Free energy calculations based on molecular dynamics (MD) simulations have seen a tremendous growth in the last decade. However, it is still difficult and tedious to set them up in an automated manner, as the majority of the present-day MD simulation packages lack that functionality. Relative free energy calculations are a particular challenge for several reasons, including the problem of finding a common substructure and mapping the transformation to be applied. Here we present a tool, alchemical-setup.py, that automatically generates all the input files needed to perform relative solvation and binding free energy calculations with the MD package GROMACS. When combined with Lead Optimization Mapper (LOMAP; Liu et al. in J Comput Aided Mol Des 27(9):755–770, 2013), recently developed in our group, alchemical-setup.py allows fully automated setup of relative free energy calculations in GROMACS. Taking a graph of the planned calculations and a mapping, both computed by LOMAP, our tool generates the topology and coordinate files needed to perform relative free energy calculations for a given set of molecules, and provides a set of simulation input parameters. The tool was validated by performing relative hydration free energy calculations for a handful of molecules from the SAMPL4 challenge (Mobley et al. in J Comput Aided Mol Des 28(4):135–150, 2014). Good agreement with previously published results and the straightforward way in which free energy calculations can be conducted make alchemical-setup.py a promising tool for automated setup of relative solvation and binding free energy calculations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alchemical free energy calculations [5, 9, 10] have increased in popularity since their appearance three decades ago, when the first such calculation of practical use was performed [11]. Currently, these calculations find their major application in biomolecular modeling [8], and computer-aided drug design in particular [6]. In the early stage of computer-aided drug design, a library of compounds is screened to find a molecule that could be suitable to become a drug. Traditionally, this has often been performed by docking [17], in part because of the low computational cost of this method. Alchemical free energy calculations, however, have not attained the same level of success as docking. While part of the reason for this may be the relatively higher computational cost, a larger factor is likely the lack of high-level automation, such as seen for docking techniques. During lead optimization in a drug discovery project, one might want to rank tens to hundreds or perhaps even thousands of possible compounds by their predicted affinity to a target or targets. This requires setting up myriads of relative free energy calculations and correctly analyzing all of the large amount of data generated. Recently, we developed a tool which standardizes analysis, helping with this task [13]. But far more automation is still needed to set up free energy calculations.
The free energy calculation workflow can be broken down into three distinct steps: planning, i.e. figuring out between which molecules the calculations are to be performed; preparing input files for the simulations; and analyzing the output produced by the MD simulation package. The whole workflow is schematically shown in Fig. 1. In our lab, some progress has been reached in developing the terminal stages of this workflow, planning [14] and analysis [13]. In this paper we present the missing link of the workflow, a Python tool, alchemical-setup.py, for automated relative free energy setup, freely accessible on GitHub at https://github.com/MobleyLab/alchemical-setup.
alchemical-setup.py provides the missing link between planning and analysis of relative free energy calculations for GROMACS. Previous work in the group has focused on automated planning and on analysis of these calculations; alchemical-setup.py automates the setup of the calculations which can then be run to arrive at the analysis stage
Theory and methodology
There are two main types of relative free energy calculations
Here, our focus is on relative free energy calculations, which compute the free energy change associated with transforming or perturbing one chemical entity into another, or replacing one with another. These two different approaches—transformation versus replacement—are typically called “single topology” and “dual topology” relative free energy calculations, respectively [20]. To understand these types of transformations, consider a transformation between two molecules, A and B, sharing a common substructure (CSS). Single topology calculations rely on this substructure as their foundation. There are two types of single-topology free energy calculations [13], “single topology, implicit intermediate” and “single topology, explicit intermediate”. In the single topology, implicit intermediate approach, we perturb A to B directly, using the CSS to determine which atoms are appeared or disappeared and which will simply have modified non-bonded interactions (possibly including modifications to atom type) as a result of the transformation. Missing atoms of either molecule are represented as “dummy” atoms in the CSS, atoms without non-bonded interactions. An example of the single topology, implicit intermediate approach is shown in Fig. 2b. In the alternative (single topology, explicit intermediate) approach depicted in Fig. 2a, an intermediate state corresponding to the CSS is introduced so that the transformation of end state A into end state B is split into two steps (hence the name): A → CSS and CSS → B. Dummy atoms are again used to replace missing atoms. In contrast, dual topology free energy calculations involve replacement. The molecule A is turned entirely into dummy atoms, while the molecule B, initially present entirely as dummy atoms, has its interactions with the remainder of the system turned on.
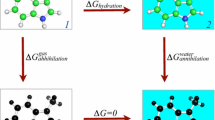
Two main types of relative free energy calculations. a Single topology, explicit intermediate. The transformation of molecule A into molecule B is split into two steps: each molecule is transformed to an intermediate whose atoms that need to be replaced are turned into dummy atoms. At this point, the two intermediates are equivalent (essentially corresponding to the CSS) except that their dummy atoms differ. However, these differences cancel when computing the total free energy change (it is not that they actually are equivalent, nor that their free energies are equal—rather, it is that the free energy difference between the two is equal in the different environments (water and gas) and so it cancels). b Single topology, implicit intermediate. Molecule A (left) is transformed into molecule B (right). Here, the endpoints of the transformation are identical in that the number of the atoms the molecules are comprised of is intact. As described in text, we are dealing with a single entity here, comprised of the CSS region (highlighted in blue) and peripheral atoms that are subject to being appeared and disappeared
We need to introduce several terms
Here, we specifically focus on single topology relative free energy calculations [10, 15]—calculations that compute free energy changes via transformation rather than replacement.
To describe our perturbation protocol, we need to define several terms. We will use common substructure Footnote 1 (CSS) to refer to an assembly of atoms that remain during the perturbation, though their chemical identities (“atom types”, more formally) may be perturbed. Again, typically it is the CSS which is of interest here. Single topology free energy calculations transform molecule A into molecule B, rather than replacing A with B. Particularly, we perform simulations of a single set of atoms, and alchemical free energy calculations (as employed here) involve changing the interactions of these atoms with one another and their environment. All other atoms of the molecule aside from the common substructure we call peripheral. It is these atoms that either appear or disappearFootnote 2 during the course of the perturbation and are referred to as dummy atoms when their nonbonded interactions with the rest of the system are removed.
The relative free energy calculations are planned with LOMAP
Our first step when considering a pair of molecules is to identify the common substructure. For our purposes, we ignore the coordinates of the atoms and consider the topology. Each molecule can be thought of as a graph whose vertices are atoms and edges bonds. For example, the phenol benzene ring and the cyclohexane subgraph of methylcyclohexane correspond to the same common substructure because they are isomorphic. With two isomorphic subgraphs comprised of the same number of atoms it is possible to find the one-to-one atom correspondence, or, bijection. Finding bijection for a pair of molecules normally does not seem burdensome—it requires inspecting the graph or overlay of the two molecules—but it may become an onerous task as the number of molecules to be screened grows. Therefore, we need to automatically find bijection, a task handled by LOMAP [14].
Given a set of molecules with binding affinities to be compared, one cannot proceed directly to relative free energy calculations. Instead, a planning step is required first. Calculations must span between molecules in order to achieve at least a minimum spanning tree across the set of molecules, so specific molecules must be connected by calculations to achieve this. In general, however, more calculations are desirable, and some attention needs to be paid to transformation efficiency. These issues have been dealt with by the LOMAP tool developed previously in the group [14], so here we will assume that the desired relative free energy calculations have already been planned, and the task of finding common substructures and performing bijection is also complete. These are handled by LOMAP [14], but other tools can presumably perform a similar task.
Our approach is based on single topology calculations
Our approach is to construct a molecule comprised of the common substructure atoms plus all the peripheral atoms of both molecules. Then its end states can be described as follows. State A is composed of the substructure atoms plus the first molecule’s peripheral atoms that are to be disappeared plus the second molecule peripheral atoms that are to be appeared (these are represented as dummies). State B is composed of the substructure atoms plus the second molecule’s peripheral atoms plus the first molecule’s peripheral atoms represented as dummies. We refer to the resulting “molecule” as a chimeric molecule. The dummy atoms have no non-bonded interactions with the rest of the system. This is controlled by turning off their Coulomb and LJ interactions. In this approach the peripheral atoms from one end state are to be disappeared, while those from the other end state are appeared, as shown in Fig. 2b.
Our tool generates the topology and coordinate files of this chimeric molecule by parsing the topology and coordinate files of the molecules in question. The geometry of the chimeric molecule is found as a result of the optimal overlay of the two molecules realized through the Kabsch method [12] which is based on the singular value decomposition algorithm [7]. The moieties that are overlaid are the parts of the molecules that correspond to the common substructure. This approach is advantageous in that it does not require the construction of an extra GROMACS topology file as there is no need to introduce an intermediate dummy state, although linear scaling of electrostatic interactions becomes impossible due to the presence of dummy atoms in both end states and the use of the soft-core Coulomb potential [2] is required to avoid numerical instabilities in situations with little or no separation between countercharges [3, 18].
Constructing the final topology and coordinate files
To construct the final topology and coordinate files we take as input the topology and coordinate files of molecule A and molecule B (to be referred to as A and B subsequently) and the CSS atom indices. These atom indices include which atoms from both A and B are present in the CSS and the mapping onto their atom numbers in the CSS, as would be provided by the output of LOMAP [14], for example. The output will be a final topology and coordinate file for the full transformation. First, we determine the number of atoms in the chimeric molecule by counting the number of atoms in the CSS and adding the number of excess atoms in the molecules A and B. Then, we reassign the indices of the atoms in A and B to map them onto the final chimeric molecule. Figure 3 depicts the content of the [atoms] directive of the topology file of the mannitol-to-tetrahydropyran interconversion. Figure 4 shows the corresponding chimeric molecule. Then, we copy all of the A-state parameters (for molecule A) into the appropriate places in the topology file and all of the B-state parameters (for molecule B) similarly. Any atom which is an excess atom (not present in A or B) is set to be a dummy atom in the appropriate place. For any bond, angle, or torsional parameter involving a dummy atom, its parameters are copied from the state where it does not involve dummy atoms (either the A or B state). For the [pairs] section, all entries corresponding to retained atoms are copied to the new topology file; the atom indices are changed accordingly. At the same time, we construct the final coordinate file by first overlaying the two molecules, then retaining the coordinates of every atom that exists in the chimeric molecule. These coordinates plus the environment comprise our final coordinate file. The atom indices are changed to be consecutive, as appropriate (Fig. 4).
An excerpt from the topology file of the mannitol-to-tetrahydropyran interconversion displaying the content of the [atoms] directive. The entries corresponding to to-be-annihilated and to-be-appeared atoms are highlighted in green and purple, respectively, while those of the CSS are left plain. As discussed in the text, soft-core potentials must be used for both types of non-bonding interactions, as the dummy atoms are present in both end states. The chimeric molecule for this transformation is shown in Fig. 4, and the atom numbering and coloring schemes here correspond to those in Fig. 4 as well
The chimeric molecule for the mannitol-to-tetrahydropyran transformation. The atoms colored as follows: atoms to be appeared are red, atoms to be disappeared are green, and the common substructure is black (the blue oxygen atoms of the common substructure are those that will be converted in the hydrogen atoms, i.e. will change their atom type). The –CH2– fragment (index 2 in mannitol and index 7 in tetrahydropyran) was not included in the common substructure to avoid biasing the conformation of mannitol. A section of the GROMACS topology file describing this transformation is shown in Fig. 3, and uses the same coloring scheme and atom numbering
The major stages of how alchemical-setup.py fits into the relative free energy workflow are diagrammatically shown in Fig. 5. To provide more details on the algorithmic side, we achieve the above by performing the following steps:
-
Initialize two objects, one for each molecule, A and B (these objects store .top and .gro file names and a dictionary with the mapping of atom indices)
-
Initialize an object for chimeric molecule whose attributes are
-
a list tracking the types of dummy atoms,
-
a dictionary containing atom types of the atoms with new indices,
-
a nested dictionary with lists storing entries for the [bonds], [angles], and [dihedrals] directives,
-
a nested dictionary with dictionaries storing atom type sequences for the [bonds], [angles], and [dihedrals] directives
-
-
Identify the atom indices of each molecule in CSS by reading in a provided mapping (from the map.txt file)
-
Build the [atoms] directive
-
read in and store appropriately all the fields of the [atoms] directive for both .top files
-
to avoid duplicates, skip the CSS atoms of molecule B
-
assign new atom names
-
sort atoms by their new name
-
-
Build the [atomtypes] directive
-
Build the [pairs] directive
-
Build directives for the bonded interactions
-
Prepare the final .gro file
-
extract the coordinates of atoms of both molecules
-
find the coordinates of atoms of molecule B when its CSS overlaid on that of molecule A
-
append coordinates of the peripheral atoms of molecule B to the molecule A .gro file
-
renumber entries in the .gro file
-
-
Write out the final .top file
A diagram of how input for relative alchemical free energy calculations is constructed in GROMACS. Two molecules (A and B) along with corresponding coordinates and topologies are provided; a mapper is then used to determine a common substructure shared by the two molecules (as in Fig. 2a, for example); the resulting map or bijection is then used by the alchemical-setup.py algorithm to define a chimeric molecule which is a hybrid of the two (Fig. 4, for example) and generate output topology and coordinate files. Thus, alchemical-setup.py covers the final green box shown here. The main text provides a more detailed description of what is involved
Simulation details
Free energy calculations consisted of a minimization at each λ value, followed by an NVT equilibration phase and then an NPT equilibration phase. Then we collected production data at each lambda value in the NVT ensemble (to be consistent with our prior work [16], which provides reference data). Simulation protocols were as described previously [16] except as noted, though principal details will be highlighted here. Minimizations consisted of up to 1500 steps of steepest descents minimization, followed by dynamics simulations with the leap-frog stochastic dynamics integrator for temperature control. NVT equilibration was done for 5 ps, followed by NPT equilibration for 100 ps with the Berendsen barostat [1]. Production was 500 ps at each λ. To capture the dynamics of hydrogen atoms (some of which change their chemical identity) the time step is lessened from the standard value of 2 fs to 1 fs and we run our simulations without constraints on hydrogen bond lengths, since these need to change if a heavy atom is changing to a hydrogen or vise versa, or even sometimes if the type of a hydrogen atom is changing. Other standard run protocols (cutoffs, PME parameters, etc.) were as used previously [16], but free energy specific settings were different and need to be discussed in more detail. Soft core potentials were used for both Coulomb and van der Waals interactions [2], with sc-alpha set to 0.5 and sc-power set to 1, as standard. We used 12 λ values as follows: 0.0, 0.02, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9 and 1.0.
In previous relative free energy calculations we have done within GROMACS [4, 19] we separated electrostatic and van der Waals transformations, with Coulomb interactions of any disappearing atoms first being turned off using linear scaling of the atomic partial charges, then van der Waals interactions being modified (turned to zero for disappearing atoms) via soft core potentials. If atoms were simultaneously being appeared, we would then add a second Coulomb transformation to introduce charges on any appearing atoms. In contrast, here, we implement the entire transformation within a single GROMACS topology file which simultaneously changes both Coulomb and van der Waals interactions to move from molecule A to molecule B. In other words our previous work used a single-topology explicit intermediate approach, whereas here we use an implicit intermediate approach. This means that here we also use soft core potentials for Coulomb interactions [2] to avoid crashes due to numerical instabilities mentioned in the “Our approach is based on single topology calculations” section.
Results and discussion
To validate our tool we performed relative hydration free energy calculations for a handful of molecules from the SAMPL4 hydration free energy challenge with absolute hydration free energies which were computed recently in our group [16]. It is worth noting though that alchemical-setup.py has much broader application and can be employed to setup any type of relative transfer free energy calculations, be it solvation, binding, or even partitioning.
The choice of the molecules within the SAMPL4 set is somewhat arbitrary and is dictated mainly by the desire to cover the common scenarios one may encounter in relative free energy calculations. Thus, the transformations selected here include a transformation between structurally similar molecules (a planar-ring to planar ring and a heterocycle to heterocycle, as in transformations 2, 3, and 4 in Fig. 6), and a transformation between structurally distinct molecules (transformation 1 in Fig. 6).
A table of relative hydration free energies found for four pairs of molecules depicted in the panels above. The free energies were computed with a new scheme (left column) and obtained as a difference between the absolute hydration free energies reported earlier (right column). The units are kcal/mol
The table of Fig. 6 shows relative hydration free energies obtained by alchemical-setup.py and those found as a difference between the absolute hydration free energies computed earlier in our lab [16]. We find that for the majority of the transformations there are no statistically significant differences between the two sets of relative hydration free energies. A slight discrepancy in the free energy estimates for transformation 1 can be attributed due to alternate lambda schedules used in the two approaches.Footnote 3 For the purpose of validating our tool, a comparison with experiment is irrelevant since the approach needs to give correct results for the force field which may or may not agree well with experiment. Thus, this consistency between the two sets of results justifies the usage of our tool for setting up relative free energy calculations.
In addition to validating alchemical-setup.py on hydration free energies, we also validated the resulting topologies via visual inspection to ensure parameters in every GROMACS directive were as expected.
Conclusion
Relative transfer free energy calculations in many cases remain suitable only for experts for a variety of reasons. One reason is that their setup can be complicated and can require substantial expert knowledge and scripting. In an effort to ease the setup of these calculations, we here provide alchemical-setup.py, a Python tool which automatically constructs topology and coordinate input files for relative free energy calculations in GROMACS.
alchemical-setup.py implements the following features:
-
1.
the topology builder which produces the .top file with properly defined end states
-
2.
the coordinate file builder based on the geometry optimal overlay realized through the singular value decomposition algorithm
Results from free energy calculations set up by this tool exhibit no statistically significant deviations from those obtained from absolute free energy calculations; and the calculations themselves are substantially easier to set up than those with the explicit intermediate approach we used previously [14] and are now fully automated. Additionally, relative free energy calculations are typically expected to be more efficient than absolute free energy calculations (and offer some other advantages) in the context of binding free energy calculations, further highlighting the utility of this approach. We believe this tool will be helpful in allowing better automation of relative free energy calculations. Along with LOMAP and alchemical-analysis.py, alchemical-setup.py (available at https://github.com/MobleyLab/alchemical-setup) forms a powerful triad of tools to plan, set up, and analyze relative free energy calculations in an automated manner. Thus, these calculations can now be set up and conducted on a large scale with much less human intervention and time.
Notes
Here, we do not prepend it with the adjective “maximal” as this would imply the largest possible number of atoms the two molecules can have in common which is not what is always wanted. For example, in the mannitol-to-tetrahydropyran transformation, the inclusion of an extra –CH2– fragment in the common substructure would certainly make the substructure larger but only at the expense of favoring certain conformations of mannitol which, in general, should be avoided.
This somewhat loose term means that the “disappearing” atoms contribute less and less to the total potential energy of the system as the coupling parameter lambda grows. “Appearing” is the opposite process. Ultimately, a fully “disappeared” atom (a dummy atom) no longer retains any non-bonded interactions with the rest of the system, while a fully “appeared” atom has full non-bonded interactions with the rest of the system (and is a normal atom).
Because of the necessity to use soft-core potentials for both non-bonded interactions in this study it is impossible to employ exactly the same lambda schedule used previously [16].
References
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690
Beutler TC, Mark AE, van Schaik RC, Gerber PR, van Gunsteren WF (1994) Avoiding singularities and numerical instabilities in free energy calculations based on molecular simulations. Chem Phys Lett 222(6):529–539
Beveridge DL, DiCapua FM (1989) Free energy via molecular simulation: applications to chemical and biomolecular systems. Annu Rev Biophys Biophys Chem 18:431–492
Boyce SE, Mobley DL, Rocklin GJ, Graves AP, Dill KA, Shoichet BK (2009) Predicting ligand binding affinity with alchemical free energy methods in a polar model binding site. J Mol Biol 394(4):747–763
Chipot C (2014) Frontiers in free-energy calculations of biological systems. Wiley Interdiscip Rev Comput Mol Sci 4(1):71–89
Chodera JD, Mobley DL, Shirts MR, Dixon RW, Branson K, Pande VS (2011) Alchemical free energy methods for drug discovery: progress and challenges. Curr Opin Struct Biol 21(2):150–160
Demmel JW (1997) Applied numerical linear algebra. SIAM
Deng Y, Roux B (2009) Computations of standard binding free energies with molecular dynamics simulations. J Phys Chem B 113(8):2234–2246
Frenkel D, Smit B (2001) Understanding molecular simulation. From algorithms to applications. Academic Press, New York
Hansen N, van Gunsteren WF (2014) Practical aspects of free-energy calculations: a review. J Chem Theory Comput 10:2632–2647
Jorgensen WL, Ravimohan C (1985) Monte Carlo simulation of differences in free energies of hydration. J Chem Phys 83(6):3050–3054
Kabsch W (1976) A solution for the best rotation to relate two sets of vectors. Acta Cryst A 32(5):922–923
Klimovich PV, Shirts MR, Mobley DL (2015) Guidelines for the analysis of free energy calculations. J Comput Aided Mol Des 29(5):397–411
Liu S, Wu Y, Lin T, Abel R, Redmann JP, Summa CM, Jaber VR, Lim NM, Mobley DL (2013) Lead optimization mapper: automating free energy calculations for lead optimization. J Comput Aided Mol Des 27(9):755–770
Mobley DL, Klimovich PV (2012) Perspective: alchemical free energy calculations for drug discovery. J Chem Phys 137(23):230,901
Mobley DL, Wymer KL, Lim NM, Guthrie JP (2014) Blind prediction of solvation free energies from the SAMPL4 challenge. J Comput Aided Mol Des 28(4):135–150
Muddana HS, Fenley AT, Mobley DL, Gilson MK (2014) The SAMPL4 host-guest blind prediction challenge: an overview. J Comput Aided Mol Des 28(4):305–317
Pitera JW, van Gunsteren WF (2002) A comparison of non-bonded scaling approaches for free energy calculations. Mol Simul 28(1–2):45–65
Rocklin GJ, Boyce SE, Fischer M, Fish I, Mobley DL, Shoichet BK, Dill KA (2013) Blind prediction of charged ligand binding affinities in a model binding site. J Mol Biol 425(22):4569–4583
Shirts MR, Mobley DL (2013) An introduction to best practices in free energy calculations. Methods Mol Biol 924:271–311
Acknowledgments
We acknowledge the financial support of the National Institutes of Health (1R15GM096257-01A1, 1R01GM108889-01) and the National Science Foundation (CHE 1352608) and computing support from the UCI GreenPlanet cluster, supported in part by NSF Grant CHE-0840513. We thank Shuai Liu (UCI) for useful comments on the LOMAP functionality, and Michael Shirts (University of Virginia) for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10822_2015_9873_MOESM1_ESM.zip
In the Supporting Information, we provide a copy of the script, as well as the input files used to set up the relative free energy calculations for a set of molecules.
Rights and permissions
About this article
Cite this article
Klimovich, P.V., Mobley, D.L. A Python tool to set up relative free energy calculations in GROMACS. J Comput Aided Mol Des 29, 1007–1014 (2015). https://doi.org/10.1007/s10822-015-9873-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-015-9873-0