Abstract
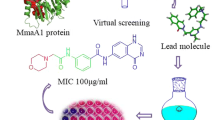
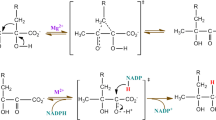
The worldwide TB structural genomics initiative has identified several new drug targets for Mycobacterium tuberculosis (M. tb). Dihydrofolate reductase (DHFR) catalyzes the NADPH-dependent reduction of dihydrofolate to tetrahydrofolate that is essential for DNA synthesis. Inhibition of its activity leads to arrest of DNA synthesis and hence cell death. Thus, M. tb DHFR (mtDHFR) is an attractive novel drug target for developing anti-TB drugs. Structural comparison of mtDHFR and human DHFR (hDHFR) reveals key differences in the active sites. These differences can be exploited for the design of selective inhibitors for mtDHFR. Based on the recently determined high resolution crystal structure of mtDHFR complexed with known inhibitor methotrexate (MTX) and cofactor NADPH, a tri-peptide inhibitor has been identified using a structure-based drug design approach. Docking studies indicate that the designed tripeptide inhibitor has a high potency (K d = 1.78 nM) and is a selective (approximately 120 fold over hDHFR) inhibitor for mtDHFR. Hence, the tripeptide is a suitable lead compound for the development of novel anti-TB drugs.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
References
Tuberculosis (as of 17 March 2009) -> Estimated TB -> Estimated TB deaths (MDG indicator 23). WHO Global Tuberculosis Database
Chan ED, Iseman MD (2002) Current medical treatment for tuberculosis. Br Med J 325: 1282–1286
Corbett EL, Watt CJ, Walker N, Maher D, Williams BG, Raviglione MC, Dye C (2003) The growing burden of tuberculosis: global trends and interactions with the HIV epidemic. Arch Intern Med 163: 1009–1021
Nachega JB, Chaisson RE (2003) Tuberculosis drug resistance: a global threat. Clin Infect Dis 36(Suppl 1): S24–S30
Raviglione M, Smith I (2007) XDR tuberculosis—implications for global public health. N Engl J Med 356((7): 656–659
Schweitzer BI, Dicker AP, Bertino JR (1990) Dihydrofolate reductase as a therapeutic target. FASEB J 4: 2441–2452
White EL, Ross LJ, Cunningham A, Escuyer V (2004) Cloning, expression and characterization of Mycobacterium tuberculosis dihydrofolate reductase. FEMS Microbiol Lett 232: 101–105
Sardarian A, Douglas KT, Read M, Sims PFG, Hyde JE, Chitnumsub P, Sirawaraporn R, Sirawaraporn W (2003) Pyrimethamine analogues as strong inhibitors of double and quadruple mutants of dihydrofolate reductase in human malaria parasites. Org Biomol Chem 1: 960–964
Hekmat-Najad M, Rathod PK (1997) Plasmodium falciparum: kinetic interactions of WR99210 with pyrimethamine-sensitive and pyrimethamine-resistant dihydrofolate reductase. Exp Parasitol 87: 222–228
Wiktor SZ, Sassan-Morokro M, Grant AD, Abouya L, Karon JM, Marice C, Djomand G, Ackah A, Domoua K, Kadio A, Yapi A, Combe P, Tossou O, Roels TH et al (1999) Efficacy of trimethoprim-sulfamethoxazole prophylaxis to decrease morbidity and mortality in HIV-1 infected patients with tuberculosis in Abijian, Cote d’Ivoire: a randomized control trial. Lancet 353: 1469–1475
Argyrou A, Vetting MW, Aladegbamil B, Blanchard JS (2006) Mycobacterium tuberculosis dihydrofolate reductase is a target for isoniazid. Nat Struct Mol Biol 13(5): 408–413
Li R, Sirawaraporn R, Chitnumsub P, Sirawaraporn W, Wooden J, Athappilly F, Turley S, Hol WG (2000) Three-dimensional structure of M. tuberculosis dihydrofolate reductase reveals opprtunities for the design of novel tuberculosis drugs. J Mol Biol 295: 307–323
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28: 235–242
Cody V, Galitsky N, Luft JR, Pangborn W, Rosowsky A, Blakley RL (1997) Comparison of two independent crystal structures of human dihydrofolate reductase ternary complexes reduced with nicotinamide adenine dinucleotide phosphate and the very tight binding inhibitor PT523. Biochemistry 36: 4399–4411
Johnson JM, Meiering EM, Wright JE, Pardo J, Rosowsky A, Wagner G (1997) NMR solution structure of the antitumor compound PT523 and NADPH in the ternary complex with human dihydrofolate reductase. Biochemistry 36: 13897–13903
da Cunha EFF, Ramalho TC, Reynolds RC (2008) Binding mode analysis of 2,4-diamino-5-methyl-5-deaza-6-substituted pteridines with Mycobacterium tuberculosis and human dihydrofolate reductase. J Biomol Struct Dyn 25(4): 377–385
Otvos L (2008) Peptide-based drug design: here and now. Pept-Based Drug Des 494: 1–8
Accelrys Software Inc (2003) Cerius2 modeling environment, release 4.7. Accelrys Software Inc, San Diego
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4: 187–217
Momany FA, Rone R (1992) Validation of the general purpose QUANTA ®3.2/CHARMm® force field. J Comput Chem 13: 888–900
Wu G, Robertson DH, Brooks CL, Vieth M (2003) Detailed analysis of grid-based molecular docking: a case study of CDOCKER—a CHARMm-based MD docking algorithm. J Comput Chem 24: 1549–1562
Hamdouchi C, Zhong B, Mendoza J, Collins E, Jaramillo C, Diego JE, Robertson D, Spencer CD, Anderson BD, Watkins SA, Zhang F, Brooks HB (2005) Structure-based design of a new class of highly selective aminoimidazo[1,2-a]pyridine-based inhibitors of cyclin dependent kinases. Bioorg Med Chem Lett 15: 1943–1947
Böhm HJ (1994) The development of a simple empirical scoring function to estimate the binding constant for a protein-ligand complex of known three-dimensional structure. J Comput Aided Mol Des 8: 243–256
Böhm HJ (1994) On the use of LUDI to search the fine chemicals directory for ligands of proteins of known three-dimensional structure. J Comput Aided Mol Des 8: 623–632
Böhm HJ (1998) Prediction of binding constants of protein ligands: a fast method for the prioritization of hits obtained from de novo design or 3D database search programs. J Comput Aided Mol Des 12: 309–323
Wang R, Lu Y, Wang S (2003) Comparative evaluation of 11 scoring functions for molecular docking. J Med Chem 46: 2287–2303
Berendsen HJC, Postma JPM, DiNola A, van Gunsteren WF, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81: 3684–3690
Subba Rao G, Vijayakrishnan R, Kumar M (2008) Structure-based design of a novel class of potent inhibitors of InhA, the Enoyl acyl carrier protein reductase from Mycobacterium tuberculosis: a computer modelling approach. Chem Biol Drug Des 72: 444–449
DeLano WL (2002) The PyMOL molecular graphics system. http://www.pymol.org
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kumar, M., Vijayakrishnan, R. & Subba Rao, G. In silico structure-based design of a novel class of potent and selective small peptide inhibitor of Mycobacterium tuberculosis Dihydrofolate reductase, a potential target for anti-TB drug discovery. Mol Divers 14, 595–604 (2010). https://doi.org/10.1007/s11030-009-9172-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-009-9172-6