Abstract
Tuberculosis (TB) remains a major threat to human health. This due to the fact that current drug treatments are less than optimal and the increasing occurrence of multi drug-resistant strains of etiological agent, Mycobacterium tuberculosis (Mt). Given the wide-spread significance of this disease, we have undertaken a design and evaluation program to discover new anti-TB drug leads. Here, we focused on ketol-acid reductoisomerase (KARI), the second enzyme in the branched-chain amino acid biosynthesis pathway. Importantly, this enzyme is present in bacteria but not in humans, making it an attractive proposition for drug discovery. In the present work, we used molecular docking to identify seventeen potential inhibitors of KARI using an in-house database. Compounds were selected based on docking scores, which were assigned as the result of favourable interactions between the compound and the active site of KARI. The inhibitory constant values for two leads, compounds 14 and 16 are 3.71 and 3.06 µM respectively. To assess the mode of binding, 100 ns molecular dynamics simulations for these two compounds in association with Mt KARI were performed and showed that the complex was stable with an average root mean square deviation of less than 3.5 Å for all atoms. Furthermore, compound 16 showed a minimum inhibitory concentration of 2.06 ± 0.91 µM and a 1.9 fold logarithmic reduction in the growth of Mt in an infected macrophage model. The two compounds exhibited low toxicity against RAW 264.7 cell lines. Thus, both compounds are promising candidates for development as an anti-TB drug leads.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tuberculosis (TB) is a contagious and lethal malady caused by Mycobacterium tuberculosis (Mt). The 2016 World Health Organization tuberculosis report shows that there were an estimated 10.4 million new TB cases and 1.4 million TB deaths in 2015. Although the number of TB deaths reduced by 22% between 2000 and 2015, TB continued as one of the top ten causes of fatalities worldwide in 2015 [1].
Of great concern is that one-third of the world’s population is infected with latent TB and therefore at risk of developing the active form of this disease. Current best-practice therapy for TB is 6-months treatment with rifampicin and isoniazid, the two front-line drugs, supplemented with pyrazinamide and ethambutol [2]. However, the emergence of extensively drug-resistant Mt (XDR-TB) [3], multidrug-resistant Mt (MDR-TB) [4] and totally drug resistant TB have become increasing threats to the control and eradication of TB. As a result, there is a necessity for new anti-TB agents to be discovered with new mechanisms of action.
An attractive approach to antimicrobial drug discovery is to target enzymes that are essential for the survival of the pathogen and where the target is not present in humans. Thus, inhibitors that are designed for such targets will have a low probability of producing toxic side-effects for the host. Enzymes of the branched-chain amino acid biosynthesis pathway (BCAA) fits this criterion; being present in bacteria, fungi, and plants but not animals. Furthermore, transposon mutagenesis studies by Griffin and colleagues have demonstrated that all enzymes of this pathway are essential to the growth and survival of M. tuberculosis in culture [5]. In support of this concept, the first enzyme of BCAA pathway acetohydroxyacid synthase (AHAS) has been explored as a target for anti-TB drug discovery, with AHAS inhibitors having MIC values of 2.5–80 mg L−1 in cell-based assays against M. tuberculosis H37Rv strain [6]. Furthermore, high throughput screening of small molecule databases has also revealed that inhibitors which target M. tuberculosis AHAS also have anti-TB activity [7].
The second enzyme of the BCAA pathway, ketol-acid reductoisomerase (KARI), converts 2-acetolactate to (2R)-2,3-dihydroxy-3-isovalerate, or 2-aceto-2-hydroxybutyrate to (2R,3R)-2,3-dihydroxy-3-methylvalerate in two steps. The first step is semi-ordered in that the cofactors Mg2+ and NADPH bind independently before 2-acetolactate or 2-keto-2-hydroxybutyrate [8]. Next, the movement of the alkyl group occurs. This step has a strict requirement for Mg2+. However, it is unclear as to whether the alkyl migration and NADPH-dependent reduction occur simultaneously [9] or consecutively [8]. Either way, several divalent metal ions (Mg2+, Mn2+, Co2+ or Ni2+) can be used in the reduction step (Fig. 1) [8].
A predicted transition state analog for KARI, N-hydroxy-N-isopropyloxamate (IpOHA), has been shown to inhibit plant and E. coli KARI and it also has anti-TB activity with an IC50 value of 62 µM against the M. tuberculosis T13704 strain [10].
The crystal structure of M. tuberculosis KARI (Mt KARI) in complex with Mg2+ has been determined to 1.0 Å resolution [11]. Based on this structure, the active site of Mt KARI is made from two different polypeptides. Hence the requirement for the enzyme to be dimeric (Fig. 2). In this complex there are two magnesium ions located deep in the active site pocket and these are separated by 4.7 Å. One of the magnesium ions is coordinated by the side chains of D188 and E192, and four water molecules and the second magnesium ion is coordinated by the side-chains of D188, E224 and E228. Overall, the active site environs are expansive with an outer entrance of ~ 20 Å in length and 7–12 Å wide and 7 Å in depth. Below this there is a small pocket that houses the magnesium ions. Thus, the active site is exposed to the solvent, allowing ready access to the substrates, the NADPH or a newly designed inhibitor. Inhibition of this form of the enzyme (i.e. in the absence of NADPH) is attractive since there is likely to be no direct competition with the naturally occurring substrate and the expansive NADPH binding site is available for inhibitor binding. In the present study, we have used molecular docking, free energy calculations and molecular dynamics simulations to identify hits from an in-house compound library that could bind to this form of the enzyme. The lead compounds have then been subjected to biological assays to determine their activity against M. tuberculosis grown in culture.
Materials and methods
Computational details
In silico analysis was performed using a Dell Precision T7610 workstation (8 processors; 8 GB RAM; NVIDIA 3 GB graphics; Maestro 9.8, Schrödinger, New York, U.S.A) workstation running on RHEL 6.1 Linux environment.
Protein preparation and binding site analysis
The coordinates for Mt KARI (PDB code: 4YPO) [11] were obtained from the protein data bank (PDB). Hydrogen atoms, bond orders, and formal charges were added using the protein preparation wizard of the Maestro software package (Maestro, Schrödinger, LLC, New York, NY, 2017). Optimized protein structures were minimized using optimized potential for liquid simulations (OPLS-2005) force field, by converging heavy atoms to an RMSD of 0.3 Å.
The complete dimer was used for all calculations since the active site is at the interface of two subunits. The receptor grid was generated such that the ligand poses bind within the predicted active site during docking. In Glide, grids were generated keeping the default parameters of van der Waals scaling factor to 1.00 and charge cutoff of 0.25 subject to an OPLS-2005 force field.
A “metal co-ordination” constraint is a necessity that a ligand atom fall within an ideal distance of an optimal coordination site for a metal atom. The ligands feature for a metal constraint is “charged acceptors” so in the docking process the ligand satisfies the constraint parameter. Ligands will dock to the metal present in the receptor grid with Glide, using the metal constraints function within the program. Interactions with metals were considered in the optimization of the Glide score. The protonation states of the ionizable residues were predicted by the PROPKA tool in the Protein Preparation Wizard. The following residues within the active site were protonated H105, K128. The negatively charged residues D188, E192, E224, and E228 were deprotonated to allow binding to the metals (Mg2+). The binding site was defined by a rectangular 20 Å box [12] surrounding the two Mg2+ ions.
Ligand database preparation
Virtual screening was performed using the three-dimensional structures of the ~ 3000 compounds from our BITS (Birla Institute of Technology and Science) in-house database [13]. This library contains compounds that only have properties commonly observed in drug or drug-like compounds (ADME properties). The in-house ligand set was further preprocessed by adding explicit hydrogen atoms, and ionizable compounds that were converted to their most probable charged forms at pH 7.0 ± 2.0 using the LigPrep module of Schrödinger software. A maximum of eight tautomeric forms were generated for each ligand. Chiralities were retained if specified and if undefined, at most 32 stereoisomers were generated per ligand and used for the screening (LigPrep, Schrödinger, LLC, New York, NY, 2017). While performing the ligand preparation we have considered that they have to dock with the receptor active-site and its metals. The metal charges and coordination spheres were taken into account by Glide when docking the ligands.
Docking study
The compounds in the prepared in-house database were docked into the binding site of Mt KARI using the using the Glide SP (standard precision). Post-docking minimization was then implemented to optimize the ligand geometries. The highest scoring compounds from the SP docking calculation were then subjected to docking in Glide XP (extra precision) to reduce the possibility of false positives. This was achieved by extensive sampling and progressive scoring, resulting in advanced enrichment [14].
In-vitro Enzyme inhibition study
Enzyme expression and purification
The Mt KARI gene has a hexa-histidine tag (–LEHHHHHH–) attached at the C-terminal as described previously [11]. The plasmid containing this gene was then transformed into E. coli BL21 cells for expression. The cells were then grown at 37 °C in LB media containing 100 µg·ml−1 ampicillin until an A600 of ~ 0.6 was achieved. Expression was induced for 7 h at room temperature by using 1 mM isopropyl β-d-1-thiogalactopyranoside (IPTG). An Ni-NTA-agarose resin was used to purify the enzyme. The binding buffer was 10 mM Tris–HCl, 250 mM MgCl2, 25 mM imidazole, 500 mM NaCl and 10% glycerol, pH 8.0. The buffer to elute the enzyme was the same as the binding buffer, except that the concentration of imidazole was increased from 25 to 500 mM [11]. Gel filtration was performed to further purify the enzyme. This was by using a Superdex 200 Hiload 26/60 column and buffer containing 10 mM Tris–HCl, 250 mM MgCl2, 500 mM NaCl and 10% glycerol [11]. The purity of the concentrated enzyme was assessed by SDS-PAGE. The sample was stored in small aliquots at − 80 °C until use.
Kinetic assay
IC50 and inhibitory constant (Ki) values were obtained by measuring the conversion of NADPH to NADP+ at 340 nm at 25 °C. For kinetic assays the buffer consisted of 0.1 M Tris–HCl at pH 8 and 50 mM MgCl2, 50 µM NADPH and 0.6 mM 2-acetolactate. The concentration of the enzyme was 179 nM. The enzyme was incubated with varying concentrations of inhibitors in assay buffer for 30 min followed by adding 0.6 mM 2-acetolactate. Ki values were calculated using
where Vi is rate at inhibitor concentration [I], Vu is uninhibited rate.
Molecular dynamics simulation
The Desmond module from Schrödinger was utilized for the MD simulations with periodic boundary conditions. The enzyme-inhibitor complexes were immersed in an orthorhombic simulation box, with the TIP4P explicit water model using the System Builder Panel with the minimum thickness of a solvent layer, 10 Å. In order to neutralize the system, counter ions were added. The system was energy minimized and then subjected to 100 ns MD simulation with the NPT ensemble at 300 K and 1.013 bar pressure with default settings. The temperature and pressure were maintained during simulation using a Nose–Hoover chain thermostat at 300 K and a Matrtyna–Tobias–Klein barostat at 1.01325, respectively. A time step of 1.2 fs was used. Energies and the structures enumerated were saved every 9.6 ps during the simulation. To analyze the trajectory simulation, the simulation event analysis tool was used [15,16,17].
Binding affinity calculation
Binding free energies for the selected compounds after dynamics were calculated using Prime MM-GBSA methods [18]. The prime MM-GBSA module (version 3.0, Schrödinger) calculates the free energy change upon formation of the complex by comparison to total individual energy based on a change in the solvent accessible surface area [19]. Initially equally spaced snapshots (every 9.6 ps) for each complex were generated from MD trajectories. Later binding free energy (ΔGbind) values were calculated at different time scales (0–10 to 0–100 ns) after removing all water molecules and counter ions. The equations for estimating binding free energy are as follows
Whereas ΔEMM is the difference in energy between the KARI and ligand complex and the sum of the energies of the KARI and ligand alone, ΔGsolv is the difference in GBSA solvation energy of the KARI-ligand complex and the sum of the solvation energies for the ligand and KARI, ΔGSA is the difference in the surface area energy for the KARI ligand complex and the sum of the surface area energies for the ligand and KARI.
In-vitro anti-tubercular screening
Minimum inhibitory concentrations (MIC) for the most potent inhibitors identified from the enzyme assays were determined using the microplate Alamar blue assay method (MABA). Mt H37Rv cultures in Middlebrook 7H9 broth with 10% OADC (0.1% casitone, 0.5% glycerol, supplemented oleic acid, albumin, dextrose, and catalase) (Himedia) having OD590 1.0 were diluted 1:20, of which 100 µl culture was used as inoculum for screening. Stock solutions for each test compound were fourfold diluted in Middlebrook 7H9 broth. Thus, the highest concentration to be tested in any assay was 100 µM. Compounds were serially diluted in sterile 96-well microtiter plates using 100 µl Middlebrook 7H9 broth. All the sampling was performed in duplicate. Assay readings were compared against the TB drugs, rifampicin (Rif), ethambutol and isoniazid (INH). Humidity was retained during the incubation period by adding sterile water into all the boundary wells of each plate. The plates were incubated at 37 °C for a week. After the incubation period, 30 µl of Alamar blue solution was added to each well, and the plate was re-incubated for 12 h. Growth was indicated by a color change from blue to pink with the lowermost concentration of compound that did not change color recorded as its MIC value [20].
Nutrient starvation assay
A culture of M. tuberculosis H37Rv (O.D. of 0.8–1.0) grown in Middlebrook 7H9 medium supplemented with OADC was pelleted and washed twice with PBS. The pellet was suspended in PBS in sealed bottles and incubated at 37 °C for 6 weeks. Aliquots of these cultures were then treated with the lead compounds and standard drugs, INH, Rif and moxifloxacin for 7 days. The lead compounds were tested at a concentration of 30 µM for compound 14 and 26.5 µM for compound 16 (i.e. ~ 10 µg ml−1). To do this, cell suspensions were diluted 10-fold using Middlebrook 7H9 broth with 10% OADC. 100 µl of each dilution was plated in sterile 48 well plates containing 450 µl of Middlebrook 7H9 broth. The plates were then incubated at 37 °C for 4 weeks. Wells with illusory bacterial gathering were counted as positive, and most probable number (MPN) values were calculated using standard statistical methods [21, 22]. These experiments were performed in triplicate.
Biofilm assay
4.5 millilitre of Sauton’s media (HiMedia) was added to each well in a 12 well sterile plate with the Mt culture which had an A590 between 0.7 and 1.0. Biofilm was developed within 5 weeks of incubation at 37 °C. To the matured biofilm, test compounds were added at 30 µM (compound 14) and 26.5 µM (compound 16), swirled (n = 4) and sealed. After 7 days incubation, Tween-80 (0.1% v/v) was added by swirling at room temperature for about 15 min. The contents of each well were centrifuged and the pellet was suspended in 5 ml of wash buffer (PBS with 10% glycerol and 0.05% Tween-80) and washed three times. The pellet was then re-suspended in 5 ml of wash buffer and rocked overnight at room temperature. Substances were homogenized through syringe passage five times. The persistence of bacteria in the biofilm population was determined by comparing antibiotic-treated plates with positive control plates by MPN [23, 24].
Cytotoxicity determination
Cell toxicity was tested using an inhibition assay with mouse macrophage cell lines (RAW 264.7). Various concentrations of compounds were added to sterile 96 well microtiter plate 5 × 103 cells and incubated for 48 h at 37 °C. After the incubation period, 10 µl of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT reagent) (5 mg ml−1) was added and then incubated for 3 h. Next, media was removed and 100 µl of DMSO was added to each well. DMSO dissolves the formazan crystals formed in wells. The absorbance was measured at 560 nm using Perkin Elmer Victor X3 microplate reader against the blank. The assay was performed in triplicates for each concentration. The cytotoxicity is represented as % inhibition at each concentration [25].
Mt infected macrophage assay
The most active compound from the in-vitro anti-TB assays was selected and its activity evaluated in the Mt infected macrophage model. The mouse macrophage primary cell line was cultured in RPMI-1640 media supplemented with 10% FBS. Mt H37Rv cultured in 7H9 broth was added at a multiplicity of infection of 10:1 for macrophages, and then incubated for 4 h. The drug at 10 µM final concentration was added to the cells and incubated for 48 h. Macrophage cells were lysed using 1 ml 1× PBS containing 0.1% Triton X 100. Bacteria were then suspended in 7H9 broth. Serial dilutions were then performed. Plates were incubated at 37 °C and growth was observed after 3 weeks [26].
Results and discussion
Mt has a complex lifecycle involving four stages when infecting humans (i.e. transmission, infection, latency and reactivation). It almost exclusively resides in the lungs, where it enters phagocytic cells including macrophages, neutrophils and dendritic cells. When Mt is ingested by macrophages, most of the bacteria are found in fused phagolysosomes, but many escape. The majority of cells are extruded into tightly apposed membrane vesicles, whereas some cells may appear free in the cytoplasm. Knowing the efficiency of the lead molecules in this model can help us to understand how the drug is inhibiting Mt in the macrophages by total count of bacteria in macrophage lysates, which should exceeded the number of CFU by several orders of magnitude.
Significantly, in the microenvironment of granulomas Mt faces challenges such as hypoxia, nutrient starvation, and oxidative stress characterized by changes in phenotype, genotype and metabolism. Thus, it is important to perform biological assays that closely mimic the conditions where this bacterium resides. Mt also has the ability to form biofilms, a distinct physiological state that harbours drug-tolerant persisters. Here, the Mt bacilli are embedded in a lipid-rich extracellular matrix containing free methoxy mycolic acids. New drugs that could specifically inhibit biofilm formation are highly desired because application of these could dramatically reduce the length of time required for the current treatments of tuberculosis that are based on the use of well-established antibiotics. The safety profile of compounds was evaluated by checking their in vitro cytotoxicity against murine RAW 264.7 cell line by using MTT assay.
In the present study an in-house database of compounds was virtually screened to identify potential inhibitors of Mt KARI. The compounds with the highest docking scores were then be assayed as enzyme inhibitors and then as anti-TB drug leads.
Docking study
A Glide grid was generated for the active site of Mt KARI centered on the two bound Mg2+ ions. All 3000 compounds from the in-house database were assessed by Glide SP docking. The 236 survivors of these steps were then assessed by the Glide XP docking program. From the final docked compounds 17 compounds were short listed for further study (Supplement Fig. 1). All of these compounds had docking scores better than that reported for IpOHA (− 5.214 kcal mol−1) (Supplement Table 1) a known inhibitor of Mt KARI with a Ki value of 97.4 ± 15.8 nM [11]. Since the in-house library was developed to contain only compounds with good ADME and pharmacological properties these leads represent good starting points for further drug development (Supplement Table 1).
Enzyme inhibition studies
The inhibition rates of the 17 compounds at a final concentration of 100 µM were compared with a control assay containing only 1% DMSO (Supplement Fig. 2). At this concentration, compounds 14 and 16 inhibited Mt KARI by at least 90%, suggesting that they are good inhibitors of the enzyme. The Ki values of these two compounds for Mt KARI were then determined (Fig. 3). For compound 14, the Ki value was measured to be 3.71 ± 0.39 µM and for compound 16, the Ki value was measured to be 3.06 ± 0.25 µM.
Molecular dynamics studies
Compounds 14 and 16, in the active site of Mt KARI were subjected to a 100 ns simulation using Desmond. The root mean square deviation (RMSD) analysis plots for the protein–ligand complexes were calculated so as to measure the movement of atoms during simulation. The RMSD for the Cᾳ atoms of Mt KARI and 14 were within an average distance of 3.61 and 1.52 Å, and for 16 the Cᾳ atoms and ligand atoms were within an average distance of 3.61 and 1.517 Å (Fig. 4), respectively.
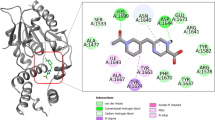
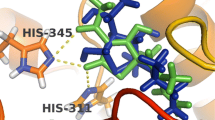
Compound 14 interacts with Mg2+ (I) through its hydroxyl group, which is deprotonated. After 100 ns molecular dynamics the orientation of the compound changed to show additional hydrogen bonding with S249, though the binding to the metal binding was maintained throughout. In a similar way, compound 16 makes direct interactions with the magnesium ions, through its nitrogen and oxygen atoms. After the 100 ns simulation its orientation had also changed to show new interactions in the form of hydrogen bonding between the nitro group of ligand and Q83, N108 residues. Thus, both ligands were stabilized whilst still retaining interactions with the metal ions. The interactions between the enzyme and the inhibitors after docking are shown in Fig. 5 and the interactions after MD simulation are provided in Figs. 6 and 7.
Enzyme inhibitor interactions for compounds 14 and 16. a, b Pymol images for compound 14 and 16, respectively. Left images: the interactions of the ligand with the metal ions (yellow dashed lines). Water molecules (red spheres) were added to fill the coordination space around the metal centres. Right images: the Connolly surface of the enzyme showing that the compounds occupy a deep cleft within the active site
Binding mode for compound 14 to Mt KARI.Mg2+. a Comparison of the binding mode of 14 after docking (white carbons) and after 100 ns of molecular dynamics simulation (blue). b The binding of 14 with Mt KARI after 100 ns of molecular dynamics simulation with a transparent surface for Mt KARI in the background. c Interactions of 14 with Mt KARI after 100 ns of molecular dynamics simulation. Amino acid side-chains are shown as thin sticks. Mg2+ ions are shown as green spheres. Ligand bonds to metals are shown as cyan dashed lines
Binding mode for compound 16 to Mt KARI.Mg2+. a Comparison of the binding mode of 16 after docking (white carbons) and after 100 ns of molecular dynamics simulation (blue). b The binding of 16 with Mt KARI after 100 ns of molecular dynamics simulation with a transparent surface for Mt KARI in the background. c Interactions of 16 with Mt KARI after 100 ns of molecular dynamics simulation. Amino acid side-chains are shown as thin sticks. Mg2+ ions are shown as green spheres. Ligand bonds to metals are shown as cyan dashed lines and hydrogen bonds are shown as salmon coloured dashed lines
MM-GBSA binding free energy calculations
The binding free energies of the two compounds in the active site of Mt KARI were determined using the snapshots collected from the trajectories obtained from MD simulations by MM-GBSA method. A total of 10,416 snapshots for each complex (every 9.6 ps) were generated. The ΔGbind values were then calculated for every 10 ns and shown in Supplement Table 2. Binding energies were also calculated from snapshots extracted from trajectories after 75 ns with the time interval of 2.5 ns such as 75–77.5 to 75–100 ns, as the system appeared more stable after 75 ns (Supplement Table 3). Therefore in the case of these ligands the 100 ns simulation time appears to be sufficient for binding energies calculation. The binding energies thus estimated around 100 ns simulation time in complex with compound 14 and 16 can provide a scope for predicting the biological activity. These leads were then assessed for their ability to inhibit M. tuberculosis growth in several different Mt models.
In-vitro anti-tubercular screening
The MIC values of 14 and 16 against M. tuberculosis grown in cell culture were determined using the MABA method. The MIC values for isoniazid, rifampicin, and ethambutol were also measured as controls (Table 1). The results show that the MIC of 16 compares favourably with the three well established drugs to treat human TB, especially with respect to ethambutol where its MIC value is 7.63 µM, or 3.7-fold higher than for compound 16.
Nutrient starvation model
Whilst the first line drugs such as isoniazid and rifampicin are successful in clearing TB in individuals, they cannot control the spread of the bacteria. This may, in part be due to the fact that these drugs are not as effective in killing persistent or inactive TB, which can survive in supplement and oxygen denied conditions. Supplement starvation triggers an inactive condition in the bacilli that changes it to a non-replicating state. To simulate this stage of the life-cycle, M. tuberculosis culture was supplement starved in phosphate buffer saline (PBS) for 6 weeks. After that, the culture was treated with compounds 14 and 16 at 30 µM and 26.5 µM concentrations, showing 2.0 and 1.8 log reductions in growth, respectively. This is a similar activity to that observed for isoniazid with a 1.5 log reduction, rifampicin with a 1.8 log reduction and moxifloxacin with a 2.0 log reduction (Fig. 8).
Biofilm model
Mycobacterium tuberculosis has developed an innate ability to create organized cellular structures in the form of biofilms. These provide additional protection from the environment and in so doing reduce the effectiveness of therapeutic drugs such as isoniazid and rifampicin. Thus, there is a need to develop new drugs that can act under these conditions. Both compounds were tested for their activity on a biofilm of Mt cells. At a concentration of 30 µM of compound 14 and 26.5 µM of compound 16 show a log reduction of cell growth of 1.0 and 0.7, respectively (Fig. 9). By comparison, INH, Rif and Moxifloxacin (Moxi) shown a log reduction of 1, 0.9 and 1.8, respectively compared to the control.
Cytotoxicity determination
To determine the toxicity of the compounds against normal host cells (macrophages), we used the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay. Both 14 and 16 were tested at 10 and 25 µM. Compound 16 inhibits the growth of these cells by 17.3% at 25 µM with no inhibition observed at 10 µM. Compound 14 is nontoxic, even at a concentration of 25 µM. Hence, even at this early stage of development a therapeutic window is apparent for this class of compound.
Mt infected macrophage model
Compound 16, with an MIC value of 2.06 ± 0.91 µM was evaluated in an Mt infected macrophage model. At 10 µM this compound has a log reduction of 1.9 fold. By comparison, INH has a 2.5 fold reduction compared to control (Fig. 10).
Conclusion
Based on transposon mutagenesis studies, KARI appears to be an excellent target for the development of new anti-TB chemotherapeutics. Here we have made use of the X-ray crystal structure of Mt KARI and virtual screening to identify new inhibitors of this enzyme. In combination with molecular docking, MD simulations and MM-GBSA free energy calculations we identified 17 compounds that have the potential to form tight binding interactions with this enzyme. Inhibition studies showed two compounds, 16 and 14, with Ki values of ~ 3 µM. In cell based assays and against virulent M. tuberculosis they have MIC values of 2.06 ± 0.91 and 76.6 µM, respectively. In addition, these compounds are also active in nutrient starvation and biofilm model assays. Furthermore, both compounds have low toxicity in RAW 264.7 cell lines and have shown 16 is effective in an Mt infected macrophage assay. All the results taken together suggest that these compounds can be further optimized for further anti-TB drug development.
References
WHO (2017) WHO global tuberculosis report 2017. World Health Organization, Geneva
Zumla A, Raviglione M, Hafner R, Von Reyn CF (2013) Tuberculosis. N Engl J Med 368:745–755
Jain A, Mondal R (2008) Extensively drug-resistant tuberculosis: current challenges and threats. FEMS Immunol Med Microbiol 53:145–150
Rawat DS (2013) Antituberculosis drug research: a critical overview. Med Res Rev 33:693–764
Griffin JE, Gawronski JD, DeJesus MA, Ioerger TR, Akerley BJ, Sassetti CM (2011) High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1002251
Wang D, Zhu X, Cui C (2013) Discovery of novel acetohydroxyacid synthase inhibitors as active agents against Mycobacterium tuberculosis by virtual screening and bioassay. J Chem Inf Model 53:343–353
Choi KJ, Yu YG, Hahn HG, Choi JD, Yoon MY (2005) Characterization of acetohydroxyacid synthase from Mycobacterium tuberculosis and the identification of its new inhibitor from the screening of a chemical library. FEBS Lett 579:4903–4910
Chunduru SK, Mrachko GT, Calvo KC (1989) Mechanism of ketol acid reductoisomerase—steady-state analysis and metal ion requirement. Biochemistry 28:486–493
Tadrowski S, Pedroso MM, Sieber V, Larrabee JA, Guddat LW, Schenk G (2016) Metal ions play an essential catalytic role in the mechanism of ketol-acid reductoisomerase. Chem-A Eur J 22:7427–7436
Grandoni JA, Marta PT, Schloss JV (1998) Inhibitors of branched-chain amino acid biosynthesis as potential antituberculosis agents. J Antimicrob Chemother 42:475–482
Lv Y, Kandale A, Wun SJ et al (2016) Crystal structure of Mycobacterium tuberculosis ketol-acid reductoisomerase at 1.0 Å resolution—a potential target for anti-tuberculosis drug discovery. FEBS J 283:1184–1196
Friesner RA, Banks JL, Murphy RB et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47:1739–1749
Beesetti H, Tyagi P, Medapi B et al (2018) A quinoline compound inhibits the replication of dengue virus serotypes 1–4 in Vero cells. Antivir Ther. https://doi.org/10.3851/IMP3231
Friesner RA, Murphy RB, Repasky MP et al (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49:6177–6196
Tripathi SK, Muttineni R, Singh SK (2013) Extra precision docking, free energy calculation and molecular dynamics simulation studies of CDK2 inhibitors. J Theor Biol 334:87–100
Katari SK, Natarajan P, Swargam S, Kanipakam H, Pasala C, Umamaheswari A (2016) Inhibitor design against JNK1 through e-pharmacophore modeling docking and molecular dynamics simulations. J Recept Signal Transduct 36:558–571
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101:4177–4189
Mobley DL, Dill KA (2009) Binding of small-molecule ligands to proteins: ‘what you see’ is not always ‘what you get’. Structure 17:489–498
Lyne PD, Lamb ML, Saeh JC (2006) Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring. J Med Chem 49:4805–4808
Collins LA, Franzblau SG (1997) Microplate Alamar blue assay versus BACTEC 460 system for high-throughput screening of compounds against Mycobacterium tuberculosis and Mycobacterium avium. Antimicrob Agents Chemother 41:1004–1009
Betts JC, Lukey PT, Robb LC, McAdam RA, Duncan K (2002) Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol Microbiol 43:717–731
Malapati P, Krishna VS, Nallangi R, Srilakshmi RR, Sriram D (2018) Identification and development of benzoxazole derivatives as novel bacterial glutamate racemase inhibitors. Eur J Med Chem 145:23–34
Kulka K, Hatfull G, Ojha AK (2012) Growth of Mycobacterium tuberculosis biofilms. J Vis Exp. https://doi.org/10.3791/3820
Malapati P, Siva Krishna V, Nallangi R, Meda N, Srilakshmi RR, Sriram D (2018) Lead identification and optimization of bacterial glutamate racemase inhibitors. Bioorg Med Chem 26:177–190
Reck F, Alm RA, Brassil P et al (2012) Novel N-linked aminopiperidine inhibitors of bacterial topoisomerase type II with reduced pKa: antibacterial agents with an improved safety profile. J Med Chem 55:6916–6933
Biketov S, Mukamolova GV, Potapov V et al (2000) Culturability of Mycobacterium tuberculosis cells isolated from murine macrophages: a bacterial growth factor promotes recovery. FEMS Immunol Med Microbiol 29:233–240
Acknowledgements
VSK and EMR are thankful to Department of Science and Technology, Government of India for the INSPIRE fellowship. DS is thankful to Department of Biotechnology, Government of India for the Tata innovation fellowship (BT/HRD/35/01/04/2015). LWG is supported by an NHMRC Project Grant 1147297.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Krishna, V.S., Zheng, S., Rekha, E.M. et al. Discovery and evaluation of novel Mycobacterium tuberculosis ketol-acid reductoisomerase inhibitors as therapeutic drug leads. J Comput Aided Mol Des 33, 357–366 (2019). https://doi.org/10.1007/s10822-019-00184-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-019-00184-1