Abstract
Eukaryotic protein kinases are fundamental factors for cellular regulation and therefore subject of strict control mechanisms. For full activity a kinase molecule must be penetrated by two stacks of hydrophobic residues, the regulatory and the catalytic spine that are normally well conserved among active protein kinases. We apply this novel spine concept here on CK2α, the catalytic subunit of protein kinase CK2. Homo sapiens disposes of two paralog isoforms of CK2α (hsCK2α and hsCK2α′). We describe two new structures of hsCK2α constructs one of which in complex with the ATP-analog adenylyl imidodiphosphate and the other with the ATP-competitive inhibitor 3-(4,5,6,7-tetrabromo-1H-benzotriazol-1-yl)propan-1-ol. The former is the first hsCK2α structure with a well defined cosubstrate/magnesium complex and the second with an open β4/β5-loop. Comparisons of these structures with existing CK2α/CK2α′ and cAMP-dependent protein kinase (PKA) structures reveal: in hsCK2α′ an open conformation of the interdomain hinge/helix αD region that is critical for ATP-binding is found corresponding to an incomplete catalytic spine. In contrast hsCK2α often adopts the canonical, PKA-like version of the catalytic spine which correlates with a closed conformation of the hinge region. HsCK2α can switch to the incomplete, non-canonical, hsCK2α′-like state of the catalytic spine, but this transition apparently depends on binding of either ATP or of the regulatory subunit CK2β. Thus, ATP looks like an activator of hsCK2α rather than a pure cosubstrate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eukaryotic protein kinases (EPKs) are central regulators of cellular key processes and require for this function a strict control of their own catalytic activity. Consequently the elucidation of regulatory mechanisms of EPKs and their structural bases are fundamental subjects of EPK research. The growing knowledge about these mechanisms and their common structural principles are reflected in a series of comprehensive reviews [1–4].
The latest approach to gain a thorough understanding of the structural constraints that govern the active state of EPKs was introduced by Kornev et al. [5] on the basis of “Local spatial pattern alignment”, a novel graph theoretical method for structural comparison [6]. According to this concept a fully active EPK requires two stacks of hydrophobic residues—called regulatory R-spine and catalytic C-spine—that start from the hydrophobic helix αF in the centre of the C-terminal EPK domain, penetrate the catalytic core and thereby cross the interface between the two main domains (Fig. 1). A continuous R-spine is the fundamental structural condition for catalytic activity of an EPK whereas the C-spine can be slightly disrupted by domain motions in the course of the catalytic cycle and is fully established only after integration of the adenine base of the cosubstrate ATP.
Catalytic spine (C-spine; yellow) and regulatory spine (R-spine; red) of cAMP-dependent protein kinase (PKA). The “gatekeeper” is a methionine residue in PKA (Met120), but Phe113 in hsCK2α. Reprinted from Biochim Biophys Acta, Vol. 1804, Kornev AP and Taylor SS, “Defining the conserved internal architecture of a protein kinase”, pp 440–444, 2010, with kind permission from Elsevier. (Color figure online)
In the context of protein kinase CK2—an ubiquitous, essential and pleiotropic Ser/Thr kinase composed of two catalytic subunits (CK2α) attached to a dimer of non-catalytic subunits (CK2β)—the spine concept was applied for the first time by Prudent et al. [7]. These authors hypothesized that the effect of polyoxometalates—a group of highly effective non-ATP-competitive CK2 inhibitors—might consist in a disruption of the R-spine of CK2α which is normally well conserved (Fig. 2c). We ourselves used the R-spine concept recently in the context of a structural study on human CK2α′ [8]. There we interpreted a hydrophobic pocket at the surface of the N-terminal domain that can be filled with a conserved tryptophan residue (Trp34 in CK2α′) and that is equivalent to the PIF-pocket of PKA and other AGC family kinases [9] as an extension of the R-spine.
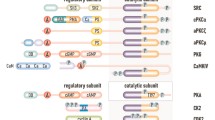
Multiple alignments of residues forming the catalytic spine and the regulatory spine. Illustration of the R-spine (a) and the C-spine (b) as published by Taylor and Kornev [4]. The residue numbers refer to cAMP-dependent protein kinase (PKA). This part of the figure was reprinted from Trends Biochem Sci, Vol. 36, Taylor SS and Kornev AP, “Protein kinases: evolution of dynamic regulatory proteins”, pp 65–77, 2011, with kind permission from Elsevier. R-spine (c) and C-spine (d) in CK2α. Only in the closed hinge/helix αD conformation Phe121 overlaps with Met128 of PKA. In part (d) the CK2β interaction “hot spots” Leu41 and Phe54 [36] were drawn in addition to the C-spine residues (grey background). The colour code of the superimposed structures is given in (e). The PKA structure 1ATP [20] was added as a reference (red side chains and labels). The CK2α residues were numbered according to the hsCK2α sequence. (Color figure online)
Here we will present a closer look at the C-spine of the catalytic CK2 subunit. CK2 has been called “a challenge to the canons” [10], a qualification that is also valid with respect to the C-spine. This is indirectly apparent from Fig. 2a/b which stems from a recent review by Taylor and Kornev [4]: in Fig. 2a the conservation of the R-spine is demonstrated by an overlay of 23 active EPK structures (plus 2 structures of prokaryotic EPK-like kinases) among them CK2. In Fig. 2b, however, in which Taylor and Kornev [4] illustrated the canonical C-spine, CK2 was left out. Obviously CK2 does not fit to the C-spine canon, and we will show here why.
Materials and methods
Crystal structures
For the C-spine analysis we used several crystal structures mainly of human CK2α (hsCK2α), but also of human CK2α′ (hsCK2α′), maize CK2α, human CK2 holoenzyme and cAMP-dependent protein kinase (PKA). HsCK2α and hsCK2α′ are the two paralogous isoforms of CK2α encoded in the human genome [11]. Their amino acid sequences are to 90.7% identical up to position 330 whereas the C-terminal segments are completely unrelated (Fig. 3).
Sequence alignment of C-terminal regions of CK2α subunits. The C-terminal segments of human CK2α (hsCK2α) and CK2α′ (hsCK2α′) differ completely in length and sequence. Both paralogs are significantly longer than maize CK2α for which the majority of CK2α crystal structures exist [32]. The bordered boxes indicate the sequence of the hsCK2α_α′ construct the structure of which was described here (Table 1). The highlighted SP- and TP-motifs in the hsCK2α sequence were reported to be phosphorylated during mitosis [37]
The identities of the compared structures can be seen in Fig. 2e. Their Protein Data Bank codes and references are as follows: 3NSZ [12], 2PVR [13], 3OFM [8], 1LP4 [14], 1JWH [15], 3JUH [16], 3BQC [17], 3H30 [18], 3FWQ [19] and 1ATP [20]. Two further structures included in the comparison had not been published previously; the details of their determination are described below.
The first of these structure, a co-crystal structure of hsCK2α1−335 with the ATP-competitive inhibitor 3-(4,5,6,7-tetrabromo-1H-benzotriazol-1-yl)propan-1-ol (abbreviated here as “MB002”) [21] that was recently crystallized together with the hsCK2α′ mutant hsCK2α′Cys336Ser [8], was determined to allow a structural comparison of both CK2α paralogs with the same ligand at the ATP site. The second novel structure contains a chimeric construct comprising residues 1–325 of hsCK2α and residues 327–350 of hsCK2α′ (see residues in bordered boxes of Fig. 3) and is called hsCK2α_α′ from here on. We explained the rationale behind hsCK2α_α′ elsewhere [8]; briefly, the motivation was to determine the structure of the C-terminal region of hsCK2α′ in which it is unrelated to hsCK2α (Fig. 3).
Mutagenesis
The construction of the C-terminally truncated mutant hsCK2α1−335 was described by Ermakova et al. [22]. For the construction of hsCK2α_α′ the DNA sequence coding for the kinase core of hsCK2α (amino acids 1–325) was amplified with oligonucleotides introducing a NdeI restriction site (forward primer: 5′-GGAATTCCATATGTCGGGACCCGTGCCAAG-3′) and SnaBI plus HindIII restriction sites (reverse primer: 5′-CCCAAGCTTTACGTAGAAATAGGGGTGCTCCATTGCC-3′). The fragment was ligated into the NdeI and HindIII cut pT7-7 expression vector.
To obtain the C-terminus of hsCK2α′, complementary oligonucleotides with a 3′-end overlap suitable for ligation with a HindIII-cut restriction site were used (P-5′-CCTGTGGTGAAGGAGCAGTCCCAGCCTTGTGCAGACAATGCTGTGCTTTCCAGTGGTCTCACGGCAGCACGATG-3′ and P-5′-AGCTTTCATCGTGCTGCCGTGAGACCACTGGAAAGCACAGCATTGTCTGCACAAGGCTGGGACTGCTCCTTCACCACA-3′). The annealing reaction mixture contained both oligonucleotides and 100 mM potassium acetate, 30 mM HEPES pH 7.4 and 2 mM magnesium acetate and was incubated at 95°C for 4 min followed by an incubation at 70°C for 10 min. After that the reaction mixture was slowly cooled down to room temperature to ensure annealing of the complementary oligonucleotides. The annealed fragment was then ligated into the SnaBI/HindIII cut pT7-7-hsCK2α1−325-plasmid adjacent to the hsCK2α1−325-sequence.
Enzyme production and crystallization
The construct hsCK2α_α′ and the established mutant hsCK2α1−335 [22] were prepared according to a protocol described previously [13]. For hsCK2α_α′ an additional gel-filtration run was necessary. The purified proteins were concentrated and rebuffered in 500 mM NaCl and 25 mM Tris/HCl, pH 8.5, by ultrafiltration using AMICON Ultra-15 tubes.
Crystallization experiments were performed at 20°C with the sitting drop variant of the vapour diffusion technique (sitting drop). For hsCK2α_α′ initial crystallization experiments with “Crystal Screen Cryo” from Hampton Research led to first crystals. The crystallization conditions were optimized by varying the concentrations of the reservoir components and using “Detergent Screen 2” from Hampton Research. The composition of the optimized crystallization drop was as follows: 0.8 μl hsCK2α_α′ (12.6 mg/ml), 1.5 μl 5 mM AMPPNP, 1.5 μl 10 mM MgCl2, 1.5 μl 1 mM CK2 substrate peptide (sequence RRRADDSDDDDD), 0.5 μl 10% (v/v) Anapoe®305, 0.8 μl reservoir solution (0.17 M ammonium sulfate, 0.1 M sodium cacodylate, pH 6.2, 15% PEG 8000 and 15% glycerol). Cryo conditions were obtained by reequilibration after changing the reservoir to 0.17 M ammonium sulfate, 0.1 M sodium cacodylate, pH 6.5, 25% PEG 8000 and 25% glycerol.
HsCK2α1−335 was co-crystallized with 3-(4,5,6,7-tetrabromo-1H-benzotriazol-1-yl)propan-1-ol (MB002) [21], i.e., with the same ATP-competitive inhibitor as used before in a co-crystal structure determination with hsCK2α′Cys336Ser [8]. Prior to crystallization 99 volume parts of a protein stock solution that contained 6 mg/ml hsCK2α1−335 in 0.5 M NaCl, 25 mM Tris/HCl, pH 8.5 was mixed with 1 volume part 10 mM MB002 in dimethyl sulfoxide. After 30 min incubation at room temperature the hsCK2α1−335/MB002 mixture was used for vapour diffusion crystallization (sitting drop). In the most successful setup the reservoir contained 4 M NaCl, 0.1 M sodium citrate buffer, pH 5.0. In the drop 0.5 μl of this reservoir solution was mixed with 0.5 μl hsCK2α1−335/MB002 mixture. HsCK2α1−335/MB002 crystals grown under these conditions could be directly frozen in liquid nitrogen and mounted for X-ray diffractometry.
X-ray diffraction data collection and structure determination
In both cases X-ray diffraction data were collected at a wavelength of 0.91841 Å and a temperature of 100 K at beamline BL-1 at the BESSY synchrotron in Berlin (Germany). The diffraction data were processed with XDS [23]. The structures were solved by molecular replacement using PHASER [24] and refined with REFMAC [25] from the CCP4 program suite [26]. A parameter file for the MB002 ligand [21] was calculated with PRODRG [27]. Manual model building was performed with COOT [28].
Results and discussion
Quality and overview of the two novel structures
The two novel co-crystal structures (hsCK2α1−335 plus MB002 and hsCK2α_α′ plus AMPPNP) were refined to reasonable R-factors and stereochemical geometries (Table 1). In spite of the low resolution the hsCK2α_α′ structure fits well to its final electron density; however, the original aim—to see an ordered structure in the C-terminal region originating from hsCK2α′—was not reached due to disorder after Glu330. Hence, the C-terminal segments of both hsCK2α and hsCK2α′ remain structurally uncharacterized similar to all previous studies in which full-length constructs were used [8, 15, 29]. Probably it requires specific interaction partners to induce ordered conformations in these regions.
Nevertheless the hsCK2α_α′ structure (basically thus a C-terminally truncated hsCK2α structure) turned out to be valuable for this analysis since in two of the four subunits within its asymmetric unit it contains—as the first human CK2α structure—a complete and well ordered AMPPNP molecule plus two magnesium ions (Fig. 4a). The whole arrangement is fully functional and similar to the ternary complex of PKA, an inhibitor peptide and ATP (black bonds in Fig. 4a) [20], meaning the γ-phospho group is correctly placed for the transfer reaction. In the two other subunits no ligands are visible so that they can be regarded as apo-forms.
Stereo pictures to illustrate conformations and ligands at and around the ATP-binding site of hsCK2α and hsCK2α′. In all panels of the figure the illustrated part of the respective main structure (yellow carbon atoms) was embedded in the final electron density with cutoff level 1 σ. The figure was prepared with BOBSCRIPT [38] in combination with Raster3D [39]. (a) Adenylyl imidodiphosphate (AMPPNP) plus Mg2+ ions bound to the ATP site of the chimeric construct hsCK2α_α′. From the enzyme the catalytic loop and three C-spine residues (Met163, Val66, Val53) were drawn. For comparison ATP plus Mn2+ ions from the PKA ternary complex structure 1ATP [20] were added after structural superimposition of the protein matrices. (b) The ATP-competitive inhibitor 3-(4,5,6,7-tetrabromo-1H-benzotriazol-1-yl)propan-1-ol (MB002) [21] bound either to hsCK2α1−335 (yellow carbon atoms; thick bonds) or to hsCK2α′Cys336Ser (PDB file 3OFM; black carbon atoms; thin bonds) [8]. (c) Structural ambiguity of Phe121 and Tyr125 in hsCK2α. The “closed hinge/canonical C-spine”-state was drawn with black carbon atoms and is represented by the hsCK2α1–335/MB002 structure (Table 1; Fig. 4b). The alternative “open hinge/non-canonical spine”-conformation (yellow carbon atoms) was extracted from the hsCK2α_α′/AMPPNP structure (Table 1; Fig. 4a). (Color figure online)
In all four chains of the hsCK2α_α′ structure the β4/β5-loop as a central element of the CK2β interface adopts an open and stretched conformation which is present also in the CK2 holoenzyme [15]. Since hitherto, in all structures of unbound (CK2β-free) hsCK2α the β4/β5-loop was found with a closed conformation, incompatible with CK2β binding, this is a remarkable detail. It is, however, not really surprising since in the recently published near-atomic resolution structure of hsCK2α2−335 in complex with an ADP-analog the β4/β5-loop was also in the open state [12].
In the case of the hsCK2α1−335/MB002 complex structure only the ATP site of chain A is filled by the inhibitor while chain B is the second subunit within the asymmetric unit shows an uncomplexed apo-state. In this structure the β4/β5-loop conformation differs between the two chains, i.e., it is open in one subunit and closed in the other which shows that the crystalline environment is a major determinant of the loop conformation.
Differences between hsCK2α and hsCK2α′ in MB002 binding
MB002 is so far the only ATP-competitive CK2 inhibitor which has been crystallized with both paralogous forms of human CK2α. A superimposition of these structures (Fig. 4b) reveals that the binding mode of MB002 differs in both cases: in complex with hsCK2α′Cys336Ser [8] (black carbon atoms in Fig. 4b) the propanol tail of the inhibitor points to the inner region of the ATP site and forms a hydrogen bond to the conserved water molecule pointed out by Battistutta et al. [30]; in contrast when MB002 is bound to hsCK2α1−335 (yellow carbon atoms in Fig. 4b) the propanol moiety approaches the glycine-rich loop (ATP-binding loop; not shown in Fig. 4b). Here, the conserved lysine Lys68 bends significantly in the direction of the inhibitor and forms a halogen bond to one of the bromine substituents.
A further halogen bond to the MB002 inhibitor not found in the hsCK2α′Cys336Ser/MB002 complex is established by the carbonyl oxygen of Asn117 (Fig. 4b). The region around Asn117 is part of the interdomain hinge/helix αD region for which two principle conformations—an open and a closed one—are known [18]. “Open” means that relatively much space is left in the ribose binding region, a necessary condition for the typical “dual cosubstrate specificity” of CK2 [14]. The open conformation was found both in the CK2 holoenzyme [15] and in two maize CK2α structures with ATP- and GTP-analogs [14] showing a similarly well defined state of the nucleotides and adjacent magnesium ions as seen in Fig. 4a. These findings suggest a correlation between the open hinge/helix αD region conformation and a fully functional way of ATP or GTP binding whereas the closed conformation seems to be associated with a non-productive ATP-binding mode [16, 31]. In Fig. 4b the open state is represented by the hsCK2α′Cys336Ser/MB002 complex [8] (black carbon atoms in Fig. 4b) and the closed one by the novel hsCK2α1−335/MB002 complex (Table 1).
The structural ambiguity in the protein environment of the MB002 inhibitor shows that the latter is not able to enforce a particular protein conformation. In fact, as we pointed out recently [8] hsCK2α′ just like maize CK2α prefers the open hinge conformation irrespective of the ligand, the crystallization condition and the crystalline environment. In contrast hsCK2α is so far the only CK2α protein in which either of the two conformations have been found [32]. Significantly the two novel structures presented here (Table 1) reflect this conformational plasticity: one of them, hsCK2α1−335/MB002 (Fig. 4b), belongs to the cluster of closed hinge structures, but the other one, hsCK2α_ α′/AMPPNP (Fig. 4a), to the open ones.
Phe121: a switchable member of the catalytic spine
The most conspicuous conformational difference in the protein matrices surrounding the two MB002 ligands refers to Phe121 (Phe122 in hsCK2α′) (Fig. 4b). The ambivalent character of this residue in hsCK2α, its ability to switch from a buried position in the closed state to a more solvent exposed one in the open conformation and the correlation of this switch to the productivity of ATP-binding have been noted and discussed before [18, 31, 32]. Now, the novel spine paradigm [4] provokes an interpretation in the light of this concept since on a sequence level [33] Phe121 is equivalent to Met128 of PKA which is a member of the catalytic spine (Figs. 1, 2b–d).
As illustrated by Fig. 1 PKA-Met128 is located at the beginning of the small surface helix αD. Its side chain forms the centre of a hydrophobic cluster whose further components are the other C-spine members Leu227, Met231, Leu172 and Ile174 (Fig. 1, 2b). Figure 2b emphasizes the structural conservation of this hydrophobic ensemble.
The more surprising is the fact that Kornev and Taylor [4] left CK2α out from Fig. 2b although including it in Fig. 2a. The reason is indicated in Fig. 2d and more precisely in Fig. 4c: CK2α fits only with the closed hinge/helix αD conformation which does not correspond to the functional arrangement in Fig. 4a to the C-spine canon; for full functionality, however, it requires the open hinge/helix αD state in which the C-spine is incomplete in the sense that the central (Met128-equivalent) position of the hydrophobic cluster remains more or less unoccupied. “More or less” means that in this case Tyr125 partially fills the relevant space (Fig. 4c). That is why Prudent et al. [8] incorporated Tyr125 in a picture illustrating the two spines of CK2α. In fact, the “local spatial pattern alignment” procedure [6] underlying the spine concept does not take into account sequence or structural homology so that it is principally justified to include a non-equivalent residue. However, we prefer to classify the C-spine corresponding to the open hinge conformation as “incomplete” since Tyr125 is less hydrophobic than Phe121 and since it occupies only to a minor extend the PKA-Met128 equivalent space at the centre of the C-spine hydrophobic cluster and even this only with structural strain, i.e., with an unfavourable side chain conformation.
Correlations and conclusions
According to the spine concept ATP is not a pure cosubstrate of the kinase reaction. Rather it is, as recently shown for PKA [34], additionally an activating factor since a correctly positioned adenine moiety is required to connect the N-lobal and the C-lobal parts of the catalytic spine (Fig. 1). Against this background it is remarkable that the incomplete and non-canonical C-spine corresponding to an open hinge/helix αD conformation was found in hsCK2α whenever co-crystallized with an ATP- or ADP analog (Fig. 2d, e). In particular the most recent hsCK2α structures, the above-mentioned high-resolution structure 3NSZ [12] and the hsCK2α_α′-structure of this work, suggest that ATP, ADP or its analogs stabilize the incomplete C-spine state or at least select it for binding. The only apparent counter-example is the complex structure of the mutant hsCK2α1−335,Val66Ala,Met163Leu with AMPPNP [16]. Here, specific mutations render the ATP site and the C-spine “PKA-like” which supports the combination of canonical C-spine (Fig. 2d, e) and closed hinge conformation. This shift of a conformational preference correlates with a partial breakdown of the dual-cosubstrate specificity, i.e. ATP is now clearly favoured as a cosubstrate compared to GTP [16].
Taken together, in CK2α molecules the R-spine is absolutely conserved (Fig. 2c), but the C-spine can be ambiguous (Fig. 2d). For full functionality CK2α requires an incomplete and non-canonical catalytic spine which is necessarily connected with an open conformation of the hinge/helix αD region. This special structural arrangement seems to be an inherent feature of the maize CK2α molecule, since in more than 40 crystal structures it was never observed with the alternative, i.e., with a closed hinge region combined to a canonical C-spine. The same may be true for hsCK2α′ but given the limited set of two structures [8, 35] it is too early for a generalization.
In the case of its paralog hsCK2α, however, specific interaction partners are required to stabilize the non-canonical C-spine state. It was mentioned already [31] that CK2β is probably such a factor, and it is noteworthy that the two recently identified CK2β interaction “hot spot” residues Leu41 and Phe54 [36] are located in direct prolongation of the C-spine axis across the plane of the N-lobal β-sheet (Fig. 2b). The analysis presented here suggests that ATP after CK2β might be the second stabilizing and activating interaction partner of hsCK2α.
Protein data bank accession codes
The two structures of Table 1 are available from the Protein Data Base (www.rcsb.org) under the accession codes 3RP0 (hsCK2α_α′/AMPPNP) and 3RPS (hsCK2α1−335/MB002).
References
Johnson LN, Noble ME, Owen DJ (1996) Active and inactive protein kinases: structural basis for regulation. Cell 85:149–158
Huse M, Kuriyan J (2002) The conformational plasticity of protein kinases. Cell 109:275–282
Nolen B, Taylor S, Ghosh G (2004) Regulation of protein kinases: controlling activity through activation segment conformation. Mol Cell 15:661–675
Taylor SS, Kornev AP (2011) Protein kinases: evolution of dynamic regulatory proteins. Trends Biochem Sci 36:65–77
Kornev AP, Taylor SS, Ten Eyck LF (2008) A helix scaffold for the assembly of active protein kinases. Proc Natl Acad Sci USA 105:14377–14382
Kornev AP, Taylor SS, Ten Eyck LF (2008) A generalized allosteric mechanism for cis-regulated cyclic nucleotide binding domains. PLoS Comput Biol 4:e1000056
Prudent R, Sautel CF, Cochet C (2010) Structure-based discovery of small molecules targeting different surfaces of protein-kinase CK2. Biochim Biophys Acta 1804:493–498
Bischoff N, Olsen B, Raaf J, Bretner M, Issinger OG, Niefind K (2011) Structure of the human protein kinase CK2 catalytic subunit CK2α′ and interaction thermodynamics with the regulatory subunit CK2β. J Mol Biol 407:1–12
Biondi RM, Cheung PC, Casamayor A, Deak M, Currie RA, Alessi DR (2000) Identification of a pocket in the PDK1 kinase domain that interacts with PIF and the C-terminal residues of PKA. EMBO J 19:979–988
Pinna LA (2002) Protein kinase CK2: a challenge to canons. J Cell Sci 115:3873–3878
Pyerin W, Ackermann K (2003) The genes encoding human protein kinase CK2 and their functional links. Prog Nucleic Acid Res Mol Biol 74:239–273
Ferguson AD, Sheth PR, Basso AD, Paliwal S, Gray K, Fischmann TO, Le HV (2011) Structural basis of CX-4945 binding to human protein kinase CK2. FEBS Lett 585:104–110
Niefind K, Yde CW, Ermakova I, Issinger OG (2007) Evolved to be active: sulfate ions define substrate recognition sites of CK2α and emphasise its exceptional role within the CMGC family of eukaryotic protein kinases. J Mol Biol 370:427–438
Niefind K, Pütter M, Guerra B, Issinger O-G, Schomburg D (1999) GTP plus water mimic ATP in the active site of protein kinase CK2. Nat Struct Biol 6:1100–1103
Niefind K, Guerra B, Ermakova I, Issinger O-G (2001) Crystal structure of human protein kinase CK2: insights into basic properties of the CK2 holoenzyme. EMBO J 20:5320–5331
Yde CW, Ermakova I, Issinger O-G, Niefind K (2005) Inclining the purine base binding plane in protein kinase CK2 by exchanging the flanking side-chains generates a preference for ATP as a cosubstrate. J Mol Biol 347:399–414
Raaf J, Klopffleisch K, Issinger O-G, Niefind K (2008) The catalytic subunit of human protein kinase CK2 structurally deviates from its maize homologue in complex with the nucleotide competitive inhibitor emodin. J Mol Biol 377:1–8
Raaf J, Brunstein E, Issinger OG, Niefind K (2008) The CK2α/CK2β interface of human protein kinase CK2 harbors a binding pocket for small molecules. Chem. Biol. 15:111–117
Raaf J, Issinger O-G, Niefind K (2009) First inactive conformation of CK2α, the catalytic subunit of protein kinase CK2. J Mol Biol 386:1212–1221
Zheng J, Trafny EA, Knighton DR, Xuong NH, Taylor SS, Ten Eyck LF, Sowadski JM (1993) 2.2 Å refined crystal structure of the catalytic subunit of cAMP-dependent protein kinase complexed with MnATP and a peptide inhibitor. Acta Crystallogr D 49:362–365
Bretner M, Najda-Bernatowicz A, Łebska M, Muszyńska G, Kilanowicz A, Sapota A (2008) New inhibitors of protein kinase CK2, analogues of benzimidazole and benzotriazole. Mol Cell Biochem 316:87–89
Ermakova I, Boldyreff B, Issinger OG, Niefind K (2003) Crystal structure of a C-terminal deletion mutant of human protein kinase CK2 catalytic subunit. J Mol Biol 330:925–934
Kabsch W (2010) XDS. Acta Crystallogr D66:125–132
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) Phaser crystallographic software. J Appl Crystallogr 40:658–674
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D53:240–255
Collaborative Computational Project Number 4 (1994) The CCP4 suite: programs for protein crystallo-graphy. Acta Crystallogr. D50:760–763
Schuettelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D60:1355–1363
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D66:486–501
Pechkova E, Zanotti G, Nicolini C (2003) Three-dimensional atomic structure of a catalytic subunit mutant of human protein kinase CK2. Acta Crystallogr D59:2133–2139
Battistutta R, Mazzorana M, Cendron L, Bortolato A, Sarno S, Kazimierczuk Z, Zanotti G, Moro S, Pinna LA (2007) The ATP-binding site of protein kinase CK2 holds a positive electrostatic area and conserved water molecules. Chembiochem 8:1804–1809
Niefind K, Issinger OG (2010) Conformational plasticity of the catalytic subunit of protein kinase CK2 and its consequences for regulation and drug design. Biochim Biophys Acta 1804:484–492
Niefind K, Raaf J, Issinger O-G (2009) Protein kinase CK2 in health and disease: protein kinase CK2: from structures to insights. Cell Mol Life Sci 66:1800–1816
Hanks SK, Hunter T (1995) Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J 9:576–596
Masterson LR, Cheng C, Yu T, Tonelli M, Kornev A, Taylor SS, Veglia G (2010) Dynamics connect substrate recognition to catalysis in protein kinase A. Nat Chem Biol 6:821–828
Nakaniwa T, Kinoshita T, Sekiguchi Y, Tada T, Nakanishi I, Kitaura K, Suzuki Y, Ohno H, Hirasawa A, Tsujimoto G (2009) Structure of human protein kinase CK2α2 with a potent indazole-derivative inhibitor. Acta Crystallogr F65:75–79
Raaf J, Bischoff N, Klopffleisch K, Brunstein E, Olsen BB, Vilk G, Litchfield DW, Issinger OG, Niefind K (2011) Interaction between CK2α and CK2β, the subunits of protein kinase CK2: thermodynamic contributions of key residues on the CK2α surface. Biochemistry 50:512–522
St-Denis NA, Derksen DR, Litchfield DW (2009) Evidence for regulation of mitotic progression through temporal phosphorylation and dephosphorylation of CK2α. Mol Cell Biol 29:2068–2081
Esnouf RM (1997) An extensively modified version of MolScript that include greatly enhanced coloring capabilities. J Mol Graph 15:132–134
Merrit EA, Bacon DJ (1997) Raster3D: photorealistic molecular graphics. Methods Enzymol 277:505–524
Acknowledgments
We are grateful to the staff of the BESSY synchrotron in Berlin, Germany, for assistance with X-ray diffraction experiments. The work was funded by the “Deutsche Forschungsgemeinschaft” (Grant NI 643/4-1) and by the “Forskningsråd for Natur og Univers” (Grant 272-07-0257).
Author information
Authors and Affiliations
Corresponding author
Additional information
Nils Bischoff and Jennifer Raaf—Common first authors.
Rights and permissions
About this article
Cite this article
Bischoff, N., Raaf, J., Olsen, B. et al. Enzymatic activity with an incomplete catalytic spine: insights from a comparative structural analysis of human CK2α and its paralogous isoform CK2α′. Mol Cell Biochem 356, 57–65 (2011). https://doi.org/10.1007/s11010-011-0948-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-011-0948-5