Abstract
MicroRNAs (miRNAs) are endogenous non-coding small RNAs that inhibit gene expression post-transcriptionally. By regulating their target genes, miRNAs play important roles in tumor generation and development. Recently, the mir-200 family was revealed to inhibit the epithelial-mesenchymal transition, which is viewed as an essential step in early tumor metastasis. Here, we used luciferase assays to demonstrate that mir-200b interacts with predicted target sites in the 3′ untranslated region of RND3. In HeLa cells, mir-200b directly reduced the expression of RND3 at the mRNA and protein levels, which thereby promoted expression of the downstream protein cyclin D1 and increased S-phase entry. In conclusion, our study demonstrates a novel role for mir-200b in cell cycle progression and identifies RND3 as a novel mir-200b target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs or mirs) are 20–25-nt small non-coding RNAs, conserved across evolution, which regulate gene expression post-transcriptionally. miRNAs are usually transcribed by RNA polymerase II as long primary miRNA transcripts (pri-miRNAs) that are subsequently cleaved by drosha/pasha to form approximately 70-nt stem-loop pre-miRNAs in the nucleus [1, 2]. The pre-miRNAs are exported into the cytoplasm by exportin 5 and then processed by dicer to generate mature miRNAs [3, 4]. Finally, the mature miRNAs interact with argonaute proteins to form the RNA-induced silencing complex, which results in the decay of the target mRNAs or the inhibition of translation [5, 6].
By regulating target gene expression, miRNAs play important roles in many processes, such as cell proliferation, apoptosis, differentiation, invasion and metastasis [7–11]. Recently, the mir-200 family has been reported as a powerful marker and determining factor of the epithelial phenotype of cancer cells. By targeting the zinc finger E-box-binding homeobox (ZEB) proteins 1 and 2, the mir-200 family could regulate the epithelial to mesenchymal transition (which is known to promote tumor invasion and metastasis) and protect tumor cells from apoptosis [12, 13]. The mir-200 family can be divided into two clusters according to their chromosomal location: the mir-200a/mir-200b/mir-429 cluster on chromosome 1 and the mir-200c/mir-141 cluster on chromosome 12. They also can be grouped into two subfamilies according to their function: mir-200b, mir-200c and mir-429 have the same seed region while those of mir-200a and mir-141 are different.
In our previous study, we demonstrated that the mir-200 family is overexpressed in endometrial adenocarcinomas, and that mir-200b showed the most significant change [14]. Other groups have demonstrated that the mir-200 family is abnormally expressed in tumors of many other cancers, such as hepatocellular, ovarian and gastric cancers [15–17]. To investigate the function of mir-200b further, we transfected a mir-200b mimic into HeLa cells and the human hepatocellular liver carcinoma cell line HepG2. HeLa cells transfected with the mir-200b mimic showed a high percentage of S-phase entry. Using luciferase assays, quantitative (q) PCR and western blotting, we demonstrated that mir-200b could directly reduce the expression of RND3 in HeLa cells and promote expression of the downstream protein cyclin D1 (CCND1) and S-phase entry.
Materials and methods
Plasmid construction
The wild-type 3′ untranslated region (UTR) of the RND3 gene, containing predicted miRNA target sites, was amplified by PCR from HeLa cell genomic DNA, then cloned into a modified pGL3-control plasmid (Promega, USA) downstream of the firefly luciferase coding region between the PstI and EcoRI sites, as described [18]. Mutant constructs containing deletions of the predicted target sites were generated using a MutanBest Kit (Takara Bio, Japan).
Cell culture, miRNA and small interfering (si) RNA transfection
HeLa cells were grown in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, USA) containing 10% FBS and 100 μg/ml penicillin/streptomycin. The miRNA mimics, miRNA inhibitors, siRNA and negative control were synthesized by GenePharma (Shanghai, China). miRNA and siRNA transfections were performed using Lipofectamine 2000 (Invitrogen, USA). Cells were plated in 6-well plates at a density of to 2 × 105 cells per well. For each well, 5 μl siRNA or miRNA (100 pmol) was added to 250 μl DMEM, and 5 μl of Lipofectamine 2000 was added to 250 μl DMEM. The siRNA/miRNA and Lipofectamine dilutions were then mixed together and incubated for 20 min. The mixture was added to the cells and incubated for 4 h before replacing the medium with fresh DMEM. Total RNA and protein, for use in qRT-PCR or western blotting, respectively, were prepared 48 h after transfection.
The mimic and siRNA sequences were: mir-200b sense: 5′-UAAUACUGCCUGGUAAUGAUGA-3′; mir-200b anti-sense: 5′-AUCAUUACCAGGCAGUAUAAAU-3′; RND3 siRNA sense: 5′-GUAGAGCUCUCCAAUCACAdTdT-3′ and RND3 siRNA anti-sense: 5′-GUAGAGCUCUCCAAUCACAdTdT-3′.
Luciferase assays
HepG2 cells were transfected in 24-well plates using Lipofectamine 2000 (Invitrogen). The transfection mixtures contained 100 ng firefly luciferase reporter plasmid, 5 ng pRL-TK plasmid (Promega) for a normalization control and 1 μl (20 pmol) mir-200b mimic or negative control. Cells were harvested 48 h after transfection, and luciferase activity was measured using a dual-luciferase reporter assay system (Promega).
RNA extraction and qRT-PCR
Total RNA was extracted from the cultured cells using Trizol Reagent (Invitrogen) according to the manufacturer’s instructions. RNA was reverse transcribed using M-MLV reverse transcriptase (Promega), according to the manufacturer’s protocol. qPCR was performed according to the protocol of the SYBR Green I kit (Toyobo Life Science, Japan) with Mx3000p (Stratagene, USA). Beta-actin mRNA levels were used for normalization. The primer sequences were: β-actin forward: 5′-TGAAGTGTGACGTGGACATCCGC-3′; β-actin reverse: 5′-GCCAATCTCATCTTGTTTTCTGCGC-3′; RND3 forward: 5′-GTGCTTGCATTTTTGGGTTT-3′; RND3 reverse: 5′-ATCCCATGGGTCCTGATACA-3′; mir-200b forward: 5′-TAATACTGCCTGGTAATGATGA-3′; mir-200b reverse: 5′-GCGAGCACAGAATTAATACGAC-3′; U6 forward: 5′-CGCTTCGGCAGCACATATACTA-3′; U6 reverse: 5′-CGCTTCACGAATTTGCGTGTCA-3′.
Western blotting
Cells were washed with PBS, harvested in ice-cold PBS and centrifuged at 2,000 rpm at 4°C. Then, cells were lyzed for 30 min on ice in RIPA buffer in the presence of a cocktail proteinase inhibitor (Sigma-Aldrich, USA). The protein was harvested, subjected to PAGE and then transferred to Hybond™-ECL membrane (GE Healthcare, USA). Immunodetection was performed using standard techniques. Antibodies against β-tubulin (Santa Cruz Biotechnology, USA), RND3 (Proteintech Group, USA) and CCND1 (Bioworld Technology, USA) were used in western blot analysis. Signals were visualized with a SuperSignal® West Femto Trial Kit (Thermo Fisher scientific, USA) and exposure to film.
Flow cytometric analysis
Cells were transfected with mir-200b or siRNA against RND3 for 24 h, starved for 24 h in 0.5% FBS-containing medium and then stimulated for 8 h in medium containing 10% FBS. The cells were then trypsinized and collected by centrifugation, washed in PBS and fixed overnight at 4°C in 70% ethanol. After being washed twice with PBS, DNA was stained with propidium iodide (50 μg/ml) in the presence of 1 mg/ml RNase A for 30 min. Analysis was performed on a FACScalibur flow cytometer (Becton Dickinson, USA).
MicroRNA target prediction software
TargetScan Release 5.1: http://www.targetscan.org/
Results
miRNA-200b promotes S-phase entry in HeLa cells
The mir-200b mimic and negative control were transfected into HeLa cells for 24, 48 or 72 h, and we examined the level of mir-200b 24–72 h after transfected by qRT-PCR which was described in [19] (Fig. 1a). The cells transfected by mir-200b showed a higher percentage of S-phase entry compared with the negative controls, which suggested that mir-200b could promote S-phase entry in HeLa cells (Fig. 1b).
Target validation of miRNA-200b
The 3′ UTR of RND3 mRNA contains two elements complementary to mir-200b seed regions, and both are conserved among human, mouse, rat, dog and chicken (Fig. 2a, b). To investigate whether RND3 mRNA is regulated by mir-200b, the 3′ UTR of RND3 was cloned into a modified pGL-3 control vector, placing it downstream of the luciferase coding sequence. The construct was cotransfected into HepG2 cells with a mir-200b mimic or negative control siRNA. Luciferase assays revealed that overexpression of mir-200b could significantly reduce luciferase activity from the reporter vector containing the 3′ UTR of RND3.
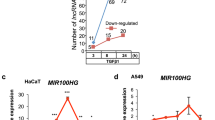
mir-200b downregulates RND3 mRNA and protein. a, b Human RND3 3′ UTR and target sites predicted by TargetScan. c Luciferase reporter assays indicated that mir-200b regulates RND3 expression by targeting both putative target sites. The transfection mixtures contained 100 ng of firefly luciferase reporter construct, 5 ng pRL-TK plasmid for a normalization control and 1 μl (20 pmol) mir-200b mimic or negative control (NC). Cells were harvested 48 h after transfection. d, e Overexpression of mir-200b downregulated RND3 mRNA (d) and protein (e) in vivo. f mir-200b mimics, mir-200b inhibitors and negative control were transfected into HeLa cells and the RND3 mRNA was up-regulated in the presence of mir-200b inhibitors
To identify which putative target site is regulated by mir-200b, two deletion constructs were generated in the modified pGL-3 control vector: the first contained putative target site 2 and the second contained no putative target site. Luciferase assays indicated that overexpression of mir-200b reduced the luciferase activity from the reporter vector containing the wild-type RND3 3′ UTR, but it had less effect on activity from the first deletion construct and no effect on activity from the second. Our results suggest that mir-200b regulates RND3 expression by targeting both putative target sites 1 and 2 (Fig. 2c).
miRNA-200b directly regulates the expression of endogenous RND3
Having confirmed the interaction of mir-200b and the RND3 3′ UTR by luciferase assays, we next analyzed the capability of mir-200b to regulate endogenous RND3 expression. The mir-200b mimic, siRNA against RND3 or the negative control RNA duplex was transfected into HeLa cells, and qRT-PCR and western blotting were used to detect the expression levels of RND3 mRNA and protein, respectively. The qRT-PCR assay showed that the mRNA level of RND3 was reduced in cells transfected by the mir-200b mimic and RND3 siRNA compared with the negative control, and western blotting gave the same results. We conclude that mir-200b can directly regulate the expression of RND3 in vivo (Fig. 2d, e). We also measured the RND3 mRNA in presence or absence of mir-200b mimics, mir-200b inhibitors and corresponding negative control, and found the reversion of RND3 mRNA in the presence of mir-200b inhibitors (Fig. 2f).
miRNA-200b down-regulates RND3 and activates the expression of CCND1
Because RND3 has been reported to regulate CCND1 expression at the post-transcriptional level, we used western blotting to investigate whether mir-200b can regulate CCND1 by targeting RND3, hence promoting cell cycle progression. In HeLa cells transfected with mir-200b or siRNA against RND3, CCND1 expression levels were higher than those in cells transfected with the negative control, corresponding to the lower levels of RND3 expression (Fig. 3a). Cell cycle analysis confirmed that mir-200b and siRNA against RND3 led to less G0/G1-cell accumulation and promoted cell cycle progression. Our results suggest that mir-200b regulates CCND1 expression and promotes cell cycle progression by targeting RND3 (Fig. 3b).
mir-200b promotes cell cycle progression by regulating cyclin D1 (CCND1). a Western blotting demonstrated that both mir-200b and siRNA against RND3 downregulated RND3, thereby upregulating CCND1. b Cell cycle analysis confirmed that mir-200b and siRNA against RND3 led to less G0/G1-cell accumulation and promoted S-phase entry
Discussion
Since miRNA was first identified in 1993 [20], over 700 miRNAs have been discovered in humans, and many of them are involved in tumor generation and development [21]. Recently, the mir-200 family has received much attention as it has been reported to inhibit the epithelial-mesenchymal transition [12, 13]. Members of the mir-200 family are abnormally expressed in hepatocellular tumors, ovarian cancer, gastric cancer and endometrial adenocarcinomas [15–17], and numerous studies have demonstrated that the mir-200 family inhibits migration and invasion by targeting ZEB1, ZEB2 and WAS protein family, member 3 (WASF3) in many kinds of cancer cells [22]. In embryonic stem cells, the mir-200 family is regulated by Myc and attenuates differentiation [23]. In bladder cancer cells, stable expression of mir-200 increases sensitivity to epidermal growth factor receptor (EGFR)-blocking agents by targeting ERBB receptor feedback inhibitor 1 (ERRFI1) [24]. The latest study has revealed that mir-200 could activate phosphoinositide-3-kinase by inhibiting feminization of germline 2 (FOG2) which suppresses cell growth; mir-200 and its target FOG2 are conserved components of the insulin pathway [25].
In this study, we investigated a novel role for mir-200b in cell cycle progression and identified a novel mir-200b target, RND3. We overexpressed mir-200b in HeLa cells, and found that the transfected cells showed a higher percentage of S-phase entry compared with the negative control cells. Luciferase assays, qPCR and western blotting suggested that mir-200b could directly regulate the expression of RND3 at the mRNA and protein levels. By comparing the data with that generated using an siRNA against RND3, we demonstrated that mir-200b could regulate the downstream protein CCND1 and promote S-phase entry by targeting RND3. CCND1 plays important role in cell cycle progression; overexpression of CCND1 may be connected with tumorigenesis. It is reported that RND3 could block CCND1 expression at the posttranscriptional level and induced cell cycle arrest [26]; as the downstream protein of RND3, CCND1 maybe up-regulated by mir-200b and promotes S-phase entry in HeLa cells.
Members of the Rho GTPase family are crucial regulators of cell shape and motility. RND3 is a member of the RND subfamily that is present in fish, birds and mammals, but not in invertebrates [27]. RND3 has two functions: one is to regulate the organization of the actin cytoskeleton [28] and the other is to inhibit cell cycle progression [26]. RND3 is regulated at multiple levels, from transcription to phosphorylation, protein stability and localization [29]. Here, we have demonstrated a novel miRNA-based mechanism for RND3 regulation.
In conclusion, this study demonstrated that mir-200b could promote CCND1 expression and S-phase entry by targeting RND3 in HeLa cells. Further research will explore the function of mir-200b in tumor generation and development.
References
Lee Y, Kim M, Han J et al (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060
Lee Y, Ahn C, Han J et al (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419
Lund E, Guttinger S, Calado A et al (2004) Nuclear export of microRNA precursors. Science 303:95–98
Yi R, Qin Y, Macara IG et al (2003) Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev 17:3011–3016
Gregory R, Chendrimada T, Cooch N et al (2005) Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123:631–640
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:287–297
Hwang HW, Mendel JT (2006) MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 94:776–780
Cheng AM, Byrom MW, Shelton J et al (2005) Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res 33:1290–1297
Kawasaki H, Taira K (2003) Hes1 is a target of microRNA-23 during retinoic-acid-induced neuronal differentiation of NT2 cells. Nature 423:838–842
Ma L, Teruya-Feldstein J, Weinberg RA (2007) Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449:682–688
Tavazoie SF, Alarco C, Oskarsson T et al (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451:147–152
Gregory PA, Bert AG, Paterson EL et al (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10:593–601
Park SM, Gaur AB, Lengyel E et al (2008) The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22:894–907
Liu Xia, Xia Wei, Dai Yinmei et al (2009) Expression of microRNAs in endometrioid adenocarcinoma. Natl Med J China 89:1365–1367
Ladeiro Y, Couchy G, Balabaud C et al (2008) MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology 47:1955–1963
Nam EJ, Yoon H, Kim SW et al (2008) MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14:2690–2695
Du Y, Xu Y, Ding L et al (2009) Down-regulation of miR-141 in gastric cancer and its involvement in cell growth. J Gastroenterol 44:556–561
Cui J, Fu H, Feng J et al (2007) The construction of miRNA expression library for human. Prog Biochem Biophys 34:389–394
Fu HJ, Zhu J, Yang M et al (2006) A novel method to monitor the expression of microRNAs. Mol Biotechnol 32:197–204
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Iorio MV, Croce CM (2009) MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol 27:5848–5856
Sossey-Alaoui K, Bialkowska K, Plow EF (2009) The miR200 family of microRNAs regulates WAVE3-dependent cancer cell invasion. J Biol Chem 284:33019–33029
Lin CH, Jackson AL, Guo J et al (2009) Myc-regulated microRNAs attenuate embryonic stem cell differentiation. EMBO J 28:3157–3170
Adam L, Zhong M, Choi W et al (2009) miR-200 expression regulates epithelial-to-mesenchymal transition in bladder cancer cells and reverses resistance to epidermal growth factor receptor therapy. Clin Cancer Res 15:5060–5072
Hyun S, Lee JH, Jin H et al (2009) Conserved MicroRNA miR-8/miR-200 and its target USH/FOG2 control growth by regulating PI3 K. Cell 139:1096–1108
Villalonga P, Guasch RM, Riento K et al (2004) RND3 inhibits cell cycle progression and Ras-induced transformation. Mol Cell Biol 24:7829–7840
Riento K, Villalonga P, Garg R et al (2005) Function and regulation of RND3. Biochem Soc Trans 33:649–651
Guasch RM, Scambler P, Jones GE et al (1998) RND3 regulates actin cytoskeleton organization and cell migration. Mol Cell Biol 18:4761–4771
Chardin P (2006) Function and regulation of Rnd proteins. Nat Rev Mol Cell Biol 7:54–62
Acknowledgments
This work was supported by the Chinese National Key Program on Basic Research (2005CB724600), Chinese National High-Tech Research and Development (2007AA021004) and the Chinese National Science Foundation (30600110, 30971630). We thank all the members of our laboratory for their help.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xia, W., Li, J., Chen, L. et al. MicroRNA-200b regulates cyclin D1 expression and promotes S-phase entry by targeting RND3 in HeLa cells. Mol Cell Biochem 344, 261–266 (2010). https://doi.org/10.1007/s11010-010-0550-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-010-0550-2






