Abstract
Keloids are benign skin tumors and are the effect of a dysregulated wound-healing process in genetically predisposed patients. They are characterized by formation of excess scar tissue beyond the boundaries of the wound. Keloids are often confused with hypertrophic scars because of an apparent lack of morphologic differences. The molecular distinction between scars and keloid is still controversial and, until today, there is no appropriate treatment yet for keloid disease. In this study, we have found, for the first time, p53 mutations in both hypertrophic scar and keloids fibroblasts from cultured cells to various extents. Since p53 plays a central role in the DNA damage response by inducing cell cycle arrest and/or apoptotic cell death, we also set up time course experiments making cell cultures at different times to investigate the phenomenon of apoptosis and its involvement in the process of pathological scarring in both hypertrophic scars and keloids. The extent of apoptosis in this study was investigated by DNA fragmentation and MTT assays, propidium iodide staining, p53 expression, and subcellular distribution. Moreover, the correlation of apoptosis and ROS levels in keloid and hypertrophic scars fibroblasts was assessed. Understanding the molecular mechanisms that determine the regulation of apoptosis during wound healing might allow us to therapeutically modulate these pathways so that apoptotic cell death is reactivated in dysregulated and hypertrophic cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Keloids are benign skin tumors occurring during wound healing in genetically predisposed patients. A hypertrophic scar looks similar to a keloid. It is estimated that keloids occur in about 10% of people and darkly pigmented people seem to be more prone to forming keloids. Hypertrophic scars are more common; they do not get as big as keloids, and may fade with time and occur in all racial groups [1–4].
Keloids and hypertrophic scars are dermal proliferative disorders, which occur following trauma, inflammation, surgery, burns, and sometimes spontaneously. Contrary to the fine-line scar characteristics of normal wound repair, the exuberant scarring of keloid and hypertrophic scars results typically in disfigurement, contractures, pruritis, and pain. Keloids enlarge and extend beyond the margins of the original wound, rarely regress, and possess more fibroblasts compared to normal skin.
Hypertrophic scars are raised, erythematous, pruritic, fibrous lesions, which typically remain within the confines of the original wound, usually undergo at least partial spontaneous resolution over widely varying time courses, and are often associated with contractures of healing tissues.
The clinical course and physical appearance define keloids and hypertrophic scars as separate entities; nevertheless, they are often confused because of an apparent lack of morphological differences. However, converting these differences into morphological or biochemical distinctions have led to a great extent of disagreement in the literature [5].
Keloid properties appear to be a multifaceted issue controlled by many gene products. In a previous study [6] we isolated ΔNp63 such as a keloid specific gene in specimens from keloids. Previous studies support the concept of a neoplastic and proliferative potential of aberrant Δp63 expression even in nonmalignant tumors [7–10].
In fibroblast, cell cultures from keloid ΔNp63 expression increased as compared to hypertrophic scar fibroblasts, and the signal of the transcript was quite weak in normal skin fibroblasts [11]. Moreover, the finding of ΔNp63 only in keloids (localized at the nuclei) as opposed to those from hypertrophic scar fibroblasts let this protein to appear as a molecular marker of this benign tumor distinct from hypertrophic scars.
In keloid fibroblasts, mutations in p53 were identified [12, 13], which may increase cell proliferation and decrease cell death in the dysregulated growth patterns. However, nothing is reported about potential p53 mutations in hyperthrophic scars fibroblasts and differential apoptosis mechanisms in both keloid and hyperthrophic scars fibroblasts.
The p53 tumor suppressor is a nuclear transcription factor, which has an ability to transactivate various p53-target genes implicated in the regulation of G1 cell cycle arrest and/or apoptosis such as p21/WAF1, MDM2, Bax, and NOXA [14–16]. p53 mutations are detected in more than half of all the human tumors [17, 18]. The tumor-suppressive activity of p53 is dependent on its sequence-specific transactivation function. Indeed, the vast majority of p53 mutations involve exons 5 through 8, a highly conserved DNA-binding domain critical for p53 function [19].
Under normal conditions, p53 is a short-lived protein whose expression levels are kept extremely low. Moreover, wild-type p53 is abnormally sequestered in the cytoplasm in certain human tumors, including breast and colon cancers [20, 21], although the detailed molecular mechanisms of the cytoplasmic accumulation of wild-type p53 remain unclear.
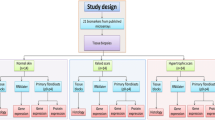
Since p53 has been shown to play an important role in controlling cell proliferation and apoptosis, first, both keloid and hyperthrophic scars fibroblasts were screened for p53 gene mutations in exons 5–8 sequence. Hence, we set up time course experiments making cell cultures at 24, 48, 72, and 96 h to underlying differential apoptosis markers of both keloid and hyperthrophic scars fibroblasts. The extent of apoptosis in this study was investigated by DNA fragmentation and MTT assays, propidium iodide staining, p53 expression, and subcellular distribution.
Growing evidences suggest that oxidative stress resulting from ROS (Reactive Oxygen Species) production is implicated in apoptosis [22, 23], and ROS have destructive actions on both DNA and proteins. Since ROS can play a role in the induction of apoptosis, especially in inflammatory cells [24], we also examined the increase of ROS generation in keloid and hypertrophic scar fibroblasts.
Materials and method
Cell cultures
Fresh tissues were obtained from skin lesions of 16 patients with keloids and hyperthrophic scars respectively, for establishing the primary cell cultures (Table 1). Normal skin was obtained from healthy patients.
Fibroblasts from keloids, hyperthrophic scars, and normal skin were isolated from samples following the same method established by Lim et al. [25]. All the experiments used cells between the second and third passage. No morphological and biochemical differences were found with the passage. Fibroblast cells were grown to confluence in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS), 10 μg/ml streptomycin, and 50 IU/ml penicillin in 5% carbon dioxide at 37°C. From the second passage time course experiments were set up for 48, 72, and 96 h without serum starvation.
p53 gene sequence analysis
Genomic DNA was isolated from cell cultures by proteinase K digestion and spin columns using QIAamp DNA mini kit (QIAGEN, Hilden, Germany). The quality of DNA was checked by running DNA on 1% agarose gel and the concentration of DNA was determined spectrophotometrically at 260 nm.
p53 exons 5–8 were amplified using Pwo Polymerase (Roche) analyzer by direct sequencing in the set of PCR primers of exons 5–8 of the p53 gene (Table 1). The sequencing was done by ABI PRISM 310 Genetic Analyzer using BigDye Terminator Cycle Sequencing Kit (PE Applied Biosystems, Foster City, CA). Experiment was performed according to the manufacturer’s instructions.
p53 gene analysis by DGGE assay
For control and confirmation of the p53 gene sequence analysis, a GC-clamped DGGE assay [26, 27] of the p53 exons 5–8 was done according to Beck [28].
DNA fragmentation assay
The cells were collected and washed twice with PBS. The cells were lysed with the digestion buffer (pH 7.4) containing 0.2% sodium dodecyl sulfate (SDS), 100 mM Tris–HCl, 0.5 mg/ml proteinase K, and 5 mM EDTA at 37°C, overnight. After extraction of cell lysates with phenol–chloroform (1:1) and chloroform, DNA was precipitated with 3 M sodium acetate (pH 5.2) and absolute ethanol, and was washed, dried, and resuspended in Tris-EDTA buffer containing 1 μl RNase A (0.5 mg/ml; Sigma–Aldrich) at 37°C for 1 h. Approximately 15 μg DNA was loaded on a 1.5% agarose gel. After electrophoresis, the gel was visualized with ethidium bromide staining under ultraviolet light.
Propidium iodide staining
Cell monolayers (5 × 105) were rinsed with serum-free medium and incubated with freshly prepared propidium iodide (Sigma P4170, 50 μg/ml in PBS) for 5 min. After rinsing with PBS, cells were fixed for 20 min in paraformaldeide (3% in PBS). Nuclear fluorescence indicative of chromatin condensation was revealed using confocal laser scanning microscopical analysis. For statistical evaluations at least 1,000 cells in randomly selected microscopic fields were counted. The obtained results were analyzed for significance by one-way ANOVA test and were presented as mean ± SD.
Cell viability analysis
To assess the cell metabolic activity, the 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide (MTT) assay was carried out using a modified method described by Li et al. [29]. In brief, 5 × 104 cells were seeded in each well containing 100 μl RPMI-1640 medium supplemented with 10% fetal bovine serum in a 96-well plate. After incubation for 48, 72, and 96 h, respectively, 50 μl MTT (Sigma–Aldrich, St. Louis, USA) (5 mg/ml stock solution) was added and incubated at 37°C for another 4 h. Then 0.2 ml of DMSO was added to stop the reactions. The absorbance of each well was determined spectrophotometrically at 490 nm by a microplate reader (Bio-Tek, Rockville, USA).
Measurement of ROS generation
The intracellular ROS generation was measured by incubating the cells in the presence of 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) (Calbiochem). DCFH-DA is a stable, non-fluorescence molecule able to cross the cell membranes and is hydrolyzed by intracellular esterases to the non-fluorescent probe DCFH, that in turn is rapidly oxidized in the presence of peroxides to the highly fluorescent 2′,7′-dichlorofluorescein (DCF). Cells (3.5 × 105/ml) were loaded with 1 μM DCFH-DA for 30 min at 37°C in standard medium, whose composition in mM was: NaCl 138, KCl 2.7, CaCl2 1.2, MgCl2 1.2, phosphate-buffered saline (PBS) 10, glucose, 10, pH 7.4. After the loading period, cells were washed twice with phosphate buffered medium before the experiment was performed. Acquisition data were obtained using a fluorescent sensitive camera Cool Snap (Media Cybernetics Inc., Silver Spring, MD, USA) and cells were evidenced by a complex software Image-Pro Plus 4.5 (Media Cybernetics Inc.) by which areas were examined.
RNA isolation
Total RNA of fibroblasts from keloids, hypertrophic scars, and normal skin were extracted with High Pure RNA Isolation Kit (Roche, Pleasanton, CA) according to the manufacturer’s instructions. A DNase I treatment was applied to remove traces of contaminating DNA. RNA quality and quantity was checked for all the samples before proceeding to all the analysis.
Semiquantitative RT-PCR
To improve the accuracy and sensitivity of the procedure, we performed a one-step quantitative RT-PCR, in which the reverse transcriptase enzyme and Pwo and Taq DNA polymerase were combined in the one tube, and a single, noninterrupted thermal cycling program was performed. Quantitation was achieved in a single reaction by using the housekeeping β-actin gene as internal standard. About 100 ng of RNA template was reverse transcribed in 50 μl reaction mix containing 200 mM deoxynucleotide triphosphates, 100 mM dithiothreitol, 0.25 μl RNase inhibitor, 1.5 mM magnesium chloride, and 1 μl enzyme mix (Titan One-tube RT-PCR Kit, Roche). The solution was incubated for 30 min at 50°C in an automated DNA thermal cycler (GeneAmp 2400, PerkinElmer, Waltham, MA).
To perform RT-PCR at optimal conditions, 40 cycles were chosen (30 s at 94°C, 30 s at 57°C, 60 s at 72°C) and were followed by a final extension for 7 min at 72°C. β-actin primers were designed to yield PCR products of 587 bp. The forward and reverse β-actin primers were 5′-CCAAGGCCAACCGCGAGAAGATGAC-3′ and 5′-AGGGTACATGGTGGTGCCGCCAGAC-3′; p53 gene primer pairs were designed to amplify a 199-bp human p53 product. The forward and reverse p53 primers were 5′-CCTCACCATCATCACACTGG-3′ and 5′-CTGGGGAGAGGAGCTGGTGTTGTT-3′, respectively. ΔNp63 gene primer pairs were designed to yield PCR products of 200 bp. The forward and reverse ΔNp63 primers were 5′CCCTTCAGTGCAACGGTCATTCAC-3′ and 5′-TGCCATTGGCCGTTTGTGTC-3′, respectively.
To rule out genomic DNA contamination, we performed a negative control, which contained RNA instead of cDNA. The signal intensities of PCR products were separated on a 1.2% agarose gel and were visualized by ethidium bromide staining. The products’ signal intensities were determined by computerized densitometric analysis using Fotoplot software. The expression of p53 and ΔNp63 mRNAs was normalized to β-actin mRNA levels.
DNA sequencing
To check the specificity of the amplified products, DNA bands were eluted from the gel and purified; sequence analysis was determined by the Big Dye Terminator Cycle Sequencing method (ABI-PRISM Sequencer 310, PerkinElmer).
Immunoblotting
Cells were washed twice in ice-cold PBS and lysed in lysis buffer (10 mm Tris–HCl, pH 7.5, 1 mm EDTA, 0.5% NP-40) containing a protease inhibitor mix (Sigma Chemical Co.) for 10 min at 4°C. Cell lysates were centrifuged at 10,000g for 10 min at 4°C to separate cytoplasmic fraction (supernatant). Insoluble materials were washed three times with the lysis buffer and nuclei were lysed in 1× SDS sample buffer. The nuclear lysates were sonicated, centrifuged, and the supernatant was collected. The protein concentrations were determined by the Bradford protein assay (Bio-Rad Laboratories). The nuclear and cytoplasmic fractions were subjected to immunoblot analysis. Fifty micrograms of protein in Laemmli sample buffer was boiled for 5 min, resolved by 10% SDS polyacrylamide gel electrophoresis (SDS-PAGE), and transferred onto polyvinylidene fluoride (PVDF) membranes. The membranes were then probed with polyclonal anti-p53 Ig (Santa Cruz Biotechnology), at 1:1000 dilution following the manufacturer’s instructions. To normalize for sample loading and protein transfer, the membranes were then stripped and reprobed with antibodies at 1:2000 dilution against monoclonal alpha-tubulin (Santa Cruz Biotechnology) and lamin-B (Santa Cruz Biotechnology), used for the cytoplasmic and nuclear markers, respectively.
Protein bands were revealed by enhanced chemiluminescence (Amersham Pharmacia Biotech, Milan, Italy) and quantified by densitometry.
Statistical analysis
All the results shown are mean ± SD of at least three separate experiments, measuring each parameter by triplicate (n = 3). Statistical significant differences were tested by one way analysis of variance (ANOVA), and, when the F value was significant, by Student-Newman-Keul’s test. P value <0.05 (*) was considered statistically significant. In addition, we computed Pearson correlation coefficient to describe associations between measures.
Results
While other authors screened for p53 mutations only in keloids, we have found in our study, for the first time, p53 mutations in both keloids and hypertrophic scar fibroblasts to various extents.
PCR/DGGE analysis of exons 5–8 of the p53 gene was performed on DNA extracted from cultured fibroblasts from keloids and hypertrophic scar specimens using primers showed in Table 2.
DNA sequence analysis was performed for all the samples that showed a positive PCR/DGGE mutational pattern. The results of the sequence analysis are listed in the Table 3. In cultured keloids fibroblasts, mutations in exons 5–8 of p53 gene have been studied (three cell-culture experiments for each skin lesion were performed, n = 3). Two mutations in exon 5 resulting in amino acid substitution were found in cultured keloid fibroblasts (codon 139, N → K, and codon 155, P → T). Frameshift mutations in exon 6 caused by the insertion of a G or deletion of a nucleotide A at different positions were found in keloid fibroblasts (codon 213, D → G). Mutations in exon 7 resulting in amino acid substitution were found in both keloid and hypertrophic scar fibroblasts (all in codon 261, S → R).
However, all these mutations were not present in the normal fibroblast cultures. We set up time course experiments performing cell culture experiments at 24, 48, 72, and 96 h; here, we obtained an increasing of apoptosis markers in hyperthrophic scars fibroblasts as more time elapsed compared to keloid and normal fibroblasts.
Cell nuclei were stained with propidium iodide (Fig. 1). Hyperthrophic scars fibroblasts resulted in a significant increase in a number of propidium iodide-positive cells at 72 (Fig. 1a) and 96 h (Fig. 1b), whereas fivefold decrease in a number of cells with apoptotic nuclei was observed in keloid fibroblasts at the same time intervals.
a–b Propidium iodide staining of normal, hypertrophic scar, and keloid fibroblasts cell cultures in time course experiments at 72 (panel A) and 96 h (panel B). Bar = 10 μm. The number of propidium iodide positive cells was counted, and expressed as a percentage of the total number of cells examined (histograms). * P < 0.05
Similar results were also obtained in MTT assays where cell viability in hypertrophic scars fibroblasts strongly decreased in a time-dependent manner compared to keloid fibroblasts (Fig. 2).
We compared the decrease of cell viability and the increased apoptosis in hypertrophic scars fibroblasts. A significant correlation between decreasing cell viability and increased apoptosis (as % of propidium iodide positive cells) was detected in hypertrophic scar fibroblasts (r = −0.94).
Because the presence of the genomic DNA ladder has been used extensively as a marker for apoptotic cell death, the next step was to examine DNA fragmentation using agarose gel electrophoresis. As shown in Fig. 3, the DNA ladders became apparent in hyperthrophic scars fibroblasts after 72 h and became stronger as more time elapsed. These results were consistent with the observation of morphological changes (Fig. 1).
As ROS can play a role in the induction of apoptosis, we examined the ROS generation in keloid and hypertrophic scar fibroblasts (Fig. 4).
ROS generation. Intracellular ROS generation was determined by measuring the fluorescent intensity of the oxidation-sensitive fluorescent DCFH-DA of normal, hypertrophic scar, and keloid fibroblasts in time course experiments at 48, 72 and 96 h. The histogram shows the mean values ± SD of ROS generation obtained by analysis of three separate experiments. * P < 0.05
As shown in Fig. 4 an increase in ROS generation at 72 and 96 h was detected in both keloid and hypertrophic scars fibroblasts. A statistically significant correlation was found between the increased ROS generation and the apoptosis data in hypertrophic scars fibroblasts (r = 0.8870). Pretreatment of the cell with a thiol-reducing antioxidant N-acetylcysteine (NAC) prevented ROS generation (data not shown).
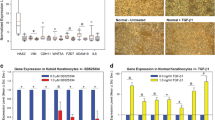
Since p53 plays a critical role in apoptosis, we examined p53 expression in hypertrophic scars and keloid fibroblasts. We found that in hypertrophic scars fibroblasts, p53 is more expressed compared to keloid and normal fibroblasts at 72 and 96 h (Fig. 5).
RT-PCR analysis of p53 expression of normal, keloid, and hypertrophic scar fibroblasts cell cultures in time course experiments at 48 (samples 1, 4, 7), 72 (samples 2, 5, 8), and 96 (samples 3, 6, 9) h. Panel A: RT-PCR assays; the histogram (panel B) shows the mean values ± SD of p53 expression obtained by densitometric analysis of three separate experiments. * P < 0.05
To examine the subcellular distribution of p53 in normal, keloid, and hypertrophic scar fibroblasts cells were biochemically fractionated into cytoplasmic and nuclear fractions, and subjected to Western blotting with the anti-p53 antibody. p53 was almost undetectable in each fraction of keloid fibroblasts, whereas p53 was expressed in nuclear fraction, though not in cytoplasm, of hypertrophic scar fibroblasts (Fig. 6). p53 immunoreactivity was detectable largely only in nucleus of hypertrophic scar fibroblasts, suggesting that p53 might lack its intact function due to mutation sequence, but not to an abnormal cellular localization. Furthermore, we assessed Δp63 mRNA expression in hypertrophic scar and keloid fibroblasts (Fig. 7). The results showed that Δp63 was highly expressed only in keloid at 72 and 96 h in a time-dependent manner, but not in hypertrophic scar and normal fibroblasts.
Nuclear localization of p53 in normal (1, 2), keloids (3, 4), and hypertrophic scars (5, 6) fibroblasts in time course experiments at 72 and 96 h. p53 is expressed in nuclei of hypertrophic scars and keloids fibroblasts. Normal, hypertrophic scars and keloids fibroblasts were biochemically fractionated into cytoplasmic (a) and nuclear (b) fractions as described under Experimental procedures. Equal amounts of cytoplasmic and nuclear extracts were subjected to Western blotting with the anti-p53. alpha-tubulin and lamin B were used for the cytoplasmic and nuclear markers, respectively
RT-PCR analysis of ΔNp63 expression of normal, keloid, and hypertrophic scar fibroblasts cell cultures in time course experiments at 48 (samples 1, 4, 7), 72 (samples 2, 5, 8), and 96 (samples 3, 6, 9) h. Panel A: RT-PCR assays; the histogram (panel B) shows the mean values ± SD of ΔNp63 expression obtained by densitometric analysis of three separate experiments. * P < 0.05
All together, these results showed that hypertrophic scars fibroblasts underwent apoptosis in a time-dependent manner though keloid fibroblasts maintained their progressive and hypertrophic nature.
Discussion
Resistance of tumor cells to apoptosis is an important mechanism for tumor progression. The objective of the current study has been the identification and characterization of the different genetic alterations involved in keloid and hypertrophic scar and their relations with apoptosis, which play a role in the process of pathological scarring [30, 31].
Direct evidence of genetic alterations in keloids is available in the literature [11–13]. The model of cellular transformation and progression of keloids has its origin in the accumulation of molecular alterations in genes like collagenase, TNF-alpha, Fas, and p53 [32–34]. More confused is the molecular distinction between hypertrophic scars and keloid [5, 35]. In our study, we have performed genetic analysis of p53 in fibroblast cell cultures from both cellular types. To our knowledge, this is the first study to completely characterize coding exons 5–8 of p53 in hypertrophic scars compared to keloids and differential apoptosis markers in time course experiments.
p53 acts as a tumor suppressor by inducing cell cycle arrest and/or apoptotic cell death in tumor cells. It has been well documented that p53 mutation is found in over 50% of all the human tumors, resulting in the loss of its pro-apoptotic function [17, 18]. The pro-apoptotic function of p53 can be also abrogated by non-mutational mechanisms [14].
We have found here, for the first time, mutations in exons with higher incidence of mutations (exons 5–8) of p53 in both keloids and hypertrophic scar fibroblasts using Polymerase chain reaction/DGGE analysis [28, 36].
The mutations detected in our research were unidentified by other authors [12, 37, 38, 39]. Three studies have examined mutations in fibroblasts from keloids using SSCP as a screening tool followed by DNA sequence analysis to confirm apparent mutations [12, 37, 38]. In the first study [12] mutations in exons 4, 5, 6 of p53 were detected.
Liu et al. [37] found points and frameshift mutations in the exon 4, 5, 6, 7 of p53 gene in nine out of the 12 keloid tissue samples and no p53 gene mutation was found in hypertrophic scar and normal skin samples. Duan et al. [38] reported p53 mutations in the exon 4 and exon 5 in fibroblasts derived from keloids and fibroblasts derived from the surrounding skin of keloids.
In this study, in hypertrophic scar fibroblasts, we found one mutation in exon 7 resulting in amino acid substitution that produced a still functional p53 protein.
A significant correlation between decreasing cell viability and increased apoptosis was detected in hypertrophic scar fibroblasts. This phenomenon correlates with the mutation in exon 7 of p53, since the mutation produced a still functional protein which induced a delayed apoptosis as more time elapsed. Indeed, in hypertrophic scar fibroblast, p53 was still expressed in nucleus (Figs. 5 and 6) leading to a time-dependent apoptosis that explained the better evolution of the scar.
On the other hand, keloid fibroblasts showed mutations in exons 5, 6, and 7 (located in codons 139, 155, 213, and 261) resulting in a unfunctional protein that was underexpressed in the time-course experiments.
Increased cell proliferation, accounting for the progressive and hypertrophic nature of keloids, correlates with the failure of apoptosis, which may play a role in the process of pathological scarring.
The mutations in exons 5, 6, and 7 in keloids fibroblasts can be considered of prognostic relevance. Indeed, some authors have suggested that mutations affecting the regions important for the function of p53 protein could present higher prognostic relevance [19]. They also have stated that different mutations in p53 are associated with different effects in patient evolution.
Our results strongly suggest that p53 mutation status is critical for p53 activation; the lack of p53 function is not due to an abnormal cytoplasmic localization as reported for certain human tumors [20, 21], since in keloid and hypertrophic scar fibroblasts, p53 localization is exclusively nuclear.
A number of conclusions can be reached based on these data. First, the results of prior studies suggested that there is an increased percentage of p53 mutations in keloids. It is likely that the differences come from the studied population in itself; the detected frequency of mutation will depend on the characteristics and the origin of the group of patients studied.
Second, identical mutations in exon 7 in both keloids and scars found in our research, supporting a precursor–product relationship between keloid and scars consistent with a clonal relationship between keloids and scars. Our results suggest that sequence alteration is relevant to p53 function. This indicates a possible mechanism of p53 underexpression that is related to mutations in the coding sequence.
p53 downregulation together with the increased ΔNp63 expression in keloids fibroblasts opposed to what we found in hypertrophic scar fibroblasts could explain the aberrant growth. The most well-known function of ΔNp63 is the inhibition of transcriptional activation on p53, TAp63, and p73; it is possible to hypothesize a mechanism where ΔNp63 works as a dominant-negative protein upon p53 preventing the latter from activating its target genes in apoptosis process [40]. Our data suggest that p53 underexpression, due to the sequence mutations, in concert with ΔNp63 activation is central in the mechanism involved in keloid proliferation. Suppression of apoptosis might contribute to keloid development by means of accumulation of continuously proliferating cells whereas the disruption or elimination of genetically altered cells might decrease tumor potential.
It is well known that p53 is a transcription factor which can be activated through DNA damage [41] and induce apoptosis through both extrinsic and intrinsic pathways [42–45].
The open question is how apoptosis can be triggered in hypertrophic scar and keloids. Apoptosis is a multifaceted process, in which different molecular mechanisms are involved. Results from Fig. 4 show that both cellular types indicate an intracellular oxidation that peaked at 72 h. There is growing evidence that reactive oxygen species (ROS) are important for the induction of apoptosis [24]. ROS can damage DNA and proteins, leading to the production of ROS-mediated modified products whereas catalase and high intracellular glutathione levels can also oppose the apoptosis triggered by ROS [46–49].
Since in hypertrophic scar fibroblasts we found a significant correlation between ROS generation and apoptosis data, we might hypothesize that ROS could be an upstream mediator for p53 activation in both keloids and hypertrophic scar fibroblasts, leading to p53 activation that failed in keloids due to p53 mutations together with Δp63 overexpression.
In conclusion, the results of our research point out to interesting different pathways in keloids and hypertrophic scars apoptosis that will be also useful in future researches. Further studies regarding the mechanisms concerning hypertrophic scars and keloids apoptosis in vitro and in vivo are therefore of interest. The understanding of the mechanisms by which keloids, in contrast to hypertrophic scars, escape from apoptosis would be beneficial in the design and the development of novel therapeutic strategies that warrant further investigations.
References
Slemp AE, Kirschner RE (2006) Keloids and scars: a review of keloids and scars, their pathogenesis, risk factors and management. Curr Opin Pediatr 18:396–402. doi:10.1097/01.mop.0000236389.41462.ef
Armour A, Scott PG, Tredget EE (2007) Cellular and molecular pathology of HTS: basis for treatment. Wound Repair Regen 15(Suppl 1):S6–S17. doi:10.1111/j.1524-475X.2007.00219.x
Berman B, Bieley HC (1995) Keloids. J Am Acad Dermatol 33:117–123. doi:10.1016/0190-9622(95)90035-7
Ehrlich HP, Desmouliere A, Diegelmann RF et al (1994) Morphological and immunochemical differences between keloid and hypertrophic scar. Am J Pathol 145:105–113
Atiyeh BS, Costagliola M, Hayek SN (2005) Keloid or hypertrophic scar: the controversy: review of the literature. Ann Plast Surg 54:676–680. doi:10.1097/01.sap.0000164538.72375.93
De Felice B, Wilson RR, Nacca M, Ciarmiello LF, Pinelli C (2004) p63 isoforms expression in human keloids. Mol Genet Genomics 272:28–34
Nishi H, Isaka K, Sagawa Y, Usuda S, Fujito A, Ito H, Senoo M, Kato H, Takayama M (1999) Mutation and transcription analyses of the p63 gene in cervical carcinoma. Int J Oncol 15:1149–1153
Park BJ, Lee SJ, Kim JI, Lee SJ, Lee CH, Chang SG, Park JH, Chi SG (2000) Frequent alteration of p63 expression in human primary bladder carcinomas. Cancer Res 60:3370–3374
Yamaguchi K, Wu L, Caballero OL, Hibi K, Trink B, Resto V, Cairns P, Okami K, Koch WM, Sidransky D, Jen J (2000) Frequent gain of the p40/p51/p63 gene locus in primary head and neck squamous cell carcinoma. Int J Cancer 86:684–689. doi:10.1002/(SICI)1097-0215(20000601)86:5<684::AID-IJC13>3.0.CO;2-M
Friedman DW, Boyd CD, Mackenzie JW, Norton P, Olson RM, Deak SB (1993) Regulation of collagen gene expression in keloids and hypertrophic scars. J Surg Res 55:214–222. doi:10.1006/jsre.1993.1132
De Felice B, Ciarmiello LF, Argenziano C, Nacca M, Mondola P, Damiano S, Garbi C (2007) Differential p63 and p53 expression in human keloid fibroblasts and hypertrophic scar fibroblasts. DNA Cell Biol 26:541–547. doi:10.1089/dna.2007.0591
Saed GM, Ladin D, Olson J, Han X, Hou Z, Fivenson D (1998) Analysis of p53 gene mutations in keloids using polymerase chain reaction-based single-strand conformational polymorphism and DNA sequencing. Arch Dermatol 134:963–967. doi:10.1001/archderm.134.8.963
Ladin DA, Hou Z, Patel D, McPhail M, Olson JC, Saed GM, Fivenson DP (1998) p53 and apoptosis alterations in keloids and keloid fibroblasts. Wound Repair Regen 6:28–37. doi:10.1046/j.1524-475X.1998.60106.x
Prives C, Hall PA (1999) The p53 pathway. J Pathol 187:112–126. doi:10.1002/(SICI)1096-9896(199901)187:1<112::AID-PATH250>3.0.CO;2-3
Sionov RV, Haupt Y (1999) The cellular response to p53: the decision between life and death. Oncogene 18:6145–6157. doi:10.1038/sj.onc.1203130
Vousden KH, Lu X (2002) Live or let die: the cell’s response to p53. Nat Rev Cancer 2:594–604. doi:10.1038/nrc864
Hollstein M, Sidransky D, Vogelstein B, Harris CC (1991) p53 mutations in human cancers. Science 253:49–53. doi:10.1126/science.1905840
Vogelstein B, Lane D, Levine AJ (2000) Surfing the p53 network. Nature 408:307–310. doi:10.1038/35042675
Cho Y, Gorina S, Jeffrey PD, Pavletich NP (1994) Crystal structure of a p53 tumor suppressor-DNA complex: understanding tumorigenic mutations. Science 265:346–355. doi:10.1126/science.8023157
Moll U, Riou G, Levine AJ (1992) Two distinct mechanisms alter p53 in breast cancer: mutation and nuclear exclusion. Proc Natl Acad Sci USA 89:7262–7266. doi:10.1073/pnas.89.15.7262
Bosari S, Viale G, Roncalli M, Graziani D, Borsani G, Lee AK, Coggi G (1995) p53 gene mutations, p53 protein accumulation and compartmentalization in colorectal adenocarcinoma. Am J Pathol 147:790–798
Greenlund LJ, Deckwerth TL, Johnson EM Jr (1995) Superoxide dismutase delays neuronal apoptosis: a role for reactive oxygen species in programmed neuronal death. Neuron 14:303–315. doi:10.1016/0896-6273(95)90287-2
Higuchi M, Honda, T, Proske RJ, Yeh ET (1998) Regulation of reactive oxygen species-induced apoptosis and necrosis by caspase 3-like proteases. Oncogene 17:2753–2760
Simon H-U, Haj-Yehia A, Levi-Schaffer F (2000) Role of reactive oxygen species (ROS) in apoptosis induction. Apoptosis 5:415–418. doi:10.1023/A:1009616228304
Lim IJ, Phan TT, Song C, Tan WT, Longaker MT (2001) Investigation of the influence of keloid-derived keratinocytes on fibroblast growth and proliferation in vitro. Plast Reconstr Surg 107:797–808. doi:10.1097/00006534-200103000-00022
Fischer SG, Lerman LS (1980) Separation of random fragments of DNA according to properties of their sequences. Proc Natl Acad Sci USA 77:4420–4424. doi:10.1073/pnas.77.8.4420
Sheffield VC, Cox DR, Lerman LS, Myers RM (1989) Attachment of a 40-base-pair G + C-rich sequence (GC-clamp) to genomic DNA fragments by the polymerase chain reaction results in improved detection of single-base changes. Proc Natl Acad Sci USA 86:232–236. doi:10.1073/pnas.86.1.232
Beck JS, Kwitek AE, Cogen PH, Metzger AK, Duyk GM, Sheffield VC (1993) A denaturing gradient gel electrophoresis assay for sensitive detection of p53 mutations. Hum Genet 91:25–30. doi:10.1007/BF00230217
Li SY, Ni JH, Xu DS, Jia HT (2003) Down-regulation of GluR2 is associated with Ca2+-dependent protease activities in kainate-induced apoptotic cell death in cultured rat hippocampal neurons. Neurosci Lett 352:105–108. doi:10.1016/j.neulet.2003.08.054
Derderian CA, Bastidas N, Lerman OZ, Bhatt KA, Lin SE, Voss J, Holmes JW, Levine JP, Gurtner GC (2005) Mechanical strain alters gene expression in an in vitro model of hypertrophic scarring. Ann Plast Surg 55:69–75. doi:10.1097/01.sap.0000168160.86221.e9
Moulin V, Larochelle S, Langlois C, Thibault I, Lopez-Vallé CA, Roy M (2004) Normal skin wound and hypertrophic scar myofibroblasts have differential responses to apoptotic inductors. J Cell Physiol 198:350–358. doi:10.1002/jcp.10415
Lu F, Gao J, Ogawa R, Hyakusoku H, Ou C (2007) Fas-mediated apoptotic signal transduction in keloid and hypertrophic scar. Plast Reconstr Surg 119:1714–1721. doi:10.1097/01.prs.0000258851.47193.06
Messadi DV, Le A, Berg S, Jewett A, Wen Z, Kelly P, Bertolami CN (1999) Expression of apoptosis-associated genes by human dermal scar fibroblasts. Wound Repair Regen 7:511–517. doi:10.1046/j.1524-475X.1999.00511.x
Tanaka A, Hatoko M, Tada H, Iioka H, Niitsuma K, Miyagawa S (2004) Expression of p53 family in scars. J Dermatol Sci 34:17–24. doi:10.1016/j.jdermsci.2003.09.005
Köse O, Waseem A (2008) Keloids and hypertrophic scars: are they two different sides of the same coin? Dermatol Surg 34:336–346. doi:10.1111/j.1524-4725.2007.34067.x
Borresen AL, Hovig E, Smith Sorensen B, Malkin D, Lystad S, Andersen TI, Nesland JM, Isselbacher KJ, Friend SH (1991) Constant denaturant gel electrophoresis as a rapid screening technique for p53 mutations. Proc Natl Acad Sci USA 88:8405–8409. doi:10.1073/pnas.88.19.8405
Liu W, Jiang Y-H, Li Y-L, Lin Z-H, Jiang H, Tan Q, Zhang J-L, Wang Z-M (2004) Experimental study on p53 gene mutation in keloid fibroblasts. Zhonghua Shao Shang Za Zhi = Chinese Journal of Burns 20:85–87
Duan HJ, Gao JH, Shen GY (2003) Cell cycle analysis and P53 gene mutation detection of fibroblasts derived from the surrounding skin of keloids. Zhonghua Zheng Xing Wai Ke Za Zhi 19:95–97
Liu YB, Gao JH, Duan HJ, Liu XJ (2003) Investigation of p53 gene mutations in keloids using PCR-SSCP. Zhonghua Zheng Xing Wai Ke Za Zhi 19:258–260
Yang A, McKeon F (2000) p63 and p73: p53 mimics, menaces, and more. Nat Rev Mol Cell Biol 1:199–207. doi:10.1038/35043127
Roos WP, Kaina B (2006) DNA damage-induced cell death by apoptosis. Trends Mol Med 12:440–450. doi:10.1016/j.molmed.2006.07.007
Bates S, Vousden KH (1999) Mechanisms of p53-mediated apoptosis. Cell Mol Life Sci 55:28–37. doi:10.1007/s000180050267
Hofseth LJ, Hussain SP, Harris CC (2004) p53: 25 years after its discovery. Trends Pharmacol Sci 25:177–181. doi:10.1016/j.tips.2004.02.009
Karen Vousden H (2006) Outcomes of p53 activation—spoilt for choice. J Cell Sci 119:5015–5020. doi:10.1242/jcs.03293
Helton ES, Chen X (2007) p53 modulation of the DNA damage response. J Cell Biochem 100:883–896. doi:10.1002/jcb.21091
Lloyd DR, Phillips DH, Carmichael PL (1997) Generation of putative intrastrand cross-links and strand breaks in DNA by transition metal ion-mediated oxygen radical attack. Chem Res Toxicol 10:393–400. doi:10.1021/tx960158q
Pierce GB, Parchment RE, Lewellyn AL (1991) Hydrogen peroxide as a mediator of programmed cell death in the blastocyst. Differentiation 46:181–186. doi:10.1111/j.1432-0436.1991.tb00880.x
Kasahara Y, Kazuyuki I, Yachie A et al (1997) Involvement of reactive oxygen intermediates in spontaneous and CD95 (Fas/APO-1)-mediated apoptosis of neutrophils. Blood 89:1748–1753
Watson RW, Rotstein OD, Jimenez M, Parodo J, Marshall JC (1997) Augmented intracellular glutathione inhibits Fas-triggered apoptosis of activated human neutrophils. Blood 89:4175–4181
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
De Felice, B., Garbi, C., Santoriello, M. et al. Differential apoptosis markers in human keloids and hypertrophic scars fibroblasts. Mol Cell Biochem 327, 191–201 (2009). https://doi.org/10.1007/s11010-009-0057-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-009-0057-x