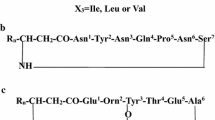Abstract
Microbes are known to interact and communicate with their neighbouring cells by releasing diverse types of low molecular weight diffusible metabolites. This paper describes the identification of iturins, fengycins and surfactins secreted by Bacillus tequilensis P15 isolated from sea coast of Jakhao, Kutch, India, using liquid chromatography coupled electron spray ionization tandem mass spectrometry. In methanol soluble fraction of acid precipitate harvested from cell free supernatant of B. tequilensis P15, 5 variants of iturins, 6 of fengycins and 39 surfactins could be identified. In particular, new surfactins with Ile/Leu at position 5 and Asp at position 6 in the peptide chain were discovered, which have not been previously reported. A novel class of novel surfactin consisting of Glu/methyl ester of Asp at position 5 in peptide chain was also identified. In addition, several linear forms of surfactins were also identified in the methanol soluble extracellular fraction of B. tequilensis P15. This is the first report on co-production of all the three classes of cyclic lipopeptides by a marine isolate B. tequilensis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The microbial metabolites have always remained fascinating subject of study owing to their ability to perform diversified physiological functions. The antibiotics are one such group of microbial metabolites, which have been demonstrated to play significant role in quorum sensing, cell adhesion, etc. in addition to their antagonistic activity on other organisms (Keller and Surette 2006). The members of genus Bacillus are well known for their ability to produce plethora of antibiotics such as surfactins, iturins, bacillomycins, mycosubtillins, plipastatins and fengycins (Koumoutsi et al. 2004; Madonna et al. 2003; Nihorimbere et al. 2012; Nishikiori et al. 1986 ; Pathak et al. 2012; Pecci et al. 2010; Peypoux et al. 1999). Surfactins are cyclic hepta-depsipeptides with β-hydroxy fatty (β-OH FA) acid while fengycin and plipastatins are cyclic deca-depsipeptides with a lactone ring consisting of C-terminal eight amino-acids and amino acid residue at N-terminal linked to β-OH FA of variable length (Kim et al. 2007; Nishikiori et al. 1986; Vanittanakom et al. 1986). The members of surfactin family exhibit variations of Ile/Leu/Val at position 2, 4 and 7 in cyclic depsipeptide ring as well as in length of β-OH FA chains varying from C13 to C15 (Kowall et al. 1998; Peypoux et al. 1991; Vater et al. 2002). The iturin family comprises of iturins, bacillomycins and mycosubtilins, which are cyclic octapeptides containing β-amino acid (βAA) side chains (Bonmatin et al. 2003; Isogai et al. 1982; Park et al. 1995; Peypoux et al. 1984; 1986). Iturins are classified into iturin A, C, D and E based on the variation in their amino acid sequence (Bonmatin et al. 2003; Park et al. 1995), while on the basis of variation in the βAA chain length, they are further classified into several sub-classes (Isogai et al. 1982). The fengycins are also cyclic lipodepsipeptides with octadepsipeptide ring and side chain composed of a dipeptide followed by β-hydroxy fatty acid chain varying from C12 to C19 (Vanittnakom et al. 1986; Volpon et al. 2000; Pathak et al. 2012). Although, Bacilli have been extensively explored for their secondary metabolites, many new antibiotics have been recently characterized from them employing highly sensitive mass spectrometric methods (Pathak et al. 2012; Watrous et al. 2012). This suggests the need for detailed re-investigation of the extra cellular bioactive extracts of potent microbial strains exhibiting antagonism using advanced technologies in order to identify novel antibiotics. The tandem mass spectrometry coupled with liquid chromatography (LC–ESI–MS/MS) analysis enables fingerprinting of very low concentration of metabolites from crude extracts prepared from natural sources with advantage of minimizing replication of compounds, saving time, energy and money spent on screening and identification of novel bioactive metabolites.
In this paper, we report the identification of lipopeptides secreted by Bacillus tequilensis P15 (isolated from sea coast, Jakhau, Kutch, India in our laboratory) employing liquid chromatography coupled with electron spray ionization tandem mass spectrometry (LC–ESI–MS/MS).
Methods
Cultivation of B. tequilensis
The pure culture of B. tequilensis P15 was inoculated in sterile 50 mL Luria–Bertani (LB) broth in 250 mL Erlenmeyer flask and incubated on orbital shaker (150 rpm) at 30 °C for 10–12 h. The liquid culture thus obtained was used to seed 500 mL LB broth in 1 L Erlenmeyer flasks to an initial O.D.600nm ≅ 0.05 and the flasks were incubated on orbital shaker (150 rpm) at 30 °C for 72 h for production of lipopeptides.
Extraction of Lipopeptides
The extraction of lipopeptide was carried out following the methods described by Pathak et al., (2012). A 72 h old culture broth of B. tequilensis P15 was centrifuged at 10,000×g, 4 °C for 15 min and the culture supernatant thus obtained was acidified to pH 2 using 6 N HCl. The precipitate was pelleted from acidified culture supernatant by centrifugation at 10,000×g, 4 °C for 25 min. The supernatant was discarded and the precipitated lipopeptides were solubilised in anhydrous methanol (MeOH). The methanol soluble fraction thus obtained was analyzed by mass spectrometry.
Mass Spectrometry
The lipopeptides in the methanolic extract obtained upon precipitation from culture supernatant of B. tequilensis P15 were analyzed by LC–ESI–MS by using HCT-Ultra ETD II ion trap mass spectrometer (Bruker Daltonics, Germany) coupled to an Agilent 1100 HPLC system as described by Pathak et al. (2012) with minor modifications. Briefly, 15 μL methanolic extract (1 mg/mL) was loaded onto a Zorbax 300SB-C18 reverse phase HPLC column (Santa Clara, CA, USA) (4.6 × 150 mm, 5 μm particle size) and separation of lipopeptides was achieved using a MeOH/H2O/0.1 % formic acid as mobile phase. The flow rate was maintained at 0.2 mL/min with a gradient of 50 min (50–95 %, vol/vol MeOH for 30 min; 95 %, vol/vol MeOH for 10 min and 95–50 %, vol/vol MeOH for 10 min). The eluted lipopeptides were directed to the ion trap mass spectrometer using ESI sprayer (Bruket Daltonics, Bremen, Germany). The separation was monitored by measuring both total ion chromatogram and UV at 226 nm. MS spectra were recorded on positive ion mode within the mass range of 50–2,000 m/z with a scan speed of 26,000 m/z s−1. The MS spectra were averaged over five scans. The molecular ions detected were further characterized by LC–ESI–MS/MS using the same instrument. The MS/MS experiment was performed in an auto mode. For MS/MS experiment, three most abundant ions within the 1.5 m/z isolation width and above the 5 % relative intensity threshold were selected for fragmentation. The fragmentation spectra were averaged over 3 scans with scan speed of 26,000 m/z sec−1. The precursor ion mass range for fragmentation was adjusted to 400–2,000 m/z in order to restrict analysis to only singly and doubly charged peptide precursor ions. The ion charge control target was kept at 200,000 with maximum acquisition time 200 ms. The helium was used as a collision gas and fragmentation amplitude (Vp-p) was kept at 1. The LC–ESI–MS/MS data were deconvoluted using Data analysis 4.0 (Bruker Daltonics, Bremen Germany).
Results and Discussion
Mass Spectrometry Based Characterization of B. tequilensis P15 Lipopeptides
In LC–ESI–MS profile of methanolic extract from B. tequilensis P15, molecular ions ranging from 700 to 1,800 m/z were considered for detailed investigation (Fig. 1a). The ion peaks in the range 730–764 m/z were found to be doubly charged molecules with their deconvoluted masses in the range 1,460–1,528 Da. The separated molecular ions were selected and further subjected to fragment analysis for their identification.
HPLC profile of methanolic antifungal extract of B. tequilensis P15. a The UV chromatogram at 226 nm and total ion chromatogram shows the separation profile of metabolites on Zorbax 300SB-column (4.6 × 150 mm, 5 μm particle size) at flow rate of 0.2 mL/min using a 50 min gradient. Each m/z value on RT peak top represents protonated mass ion belong to principal molecule eluted from the RT peak. b Primary structures of iturin, fengycin, surfactin isoforms
Characterizations of Iturins
In LC–ESI–MS/MS spectrum of [M+H]+ at 1,085.7 m/z (retention time (RT) 9.5 min), the product ions could be assigned as b− and y−ions, which enabled us to deduced the sequence: cyclo-[P–N–S–β-amino fatty acid (βAA)–N–Y–N–Q] (Fig. 2a). The molecular mass ion at 1,071.7 m/z with a 14 Da mass difference from ion at 1,085.7 m/z was found to be a βAA chain variant of 1,085.7 m/z with same peptide sequence on the basis of fragment ion profile obtained by LC–ESI–MS/MS. These lipopeptide precursor ions at 1,071.7 and 1,085.7 m/z were assigned as iturin A variants differing in βAA chain length. According to the nomenclature described by the Isogai et al. (1982), molecular ion at 1,071.7 m/z was assigned as iturin A6/A7 and 1,085.7 m/z as iturin A8. The ions at 1,093.7 and 1,107.7 m/z were assigned as sodium adducts of iturin A isoforms of ions at 1,071.7 and 1,085.7 m/z, respectively (Table 1).
On the basis of b− and y− ions observed in LC–ESI–MS/MS spectrum of [M+H]+ at 1,100.7 m/z (10.2 min RT), its sequence could be deduced as cyclo [P–N–S–βAA–D–Y–N–Q] (Fig. 2b). Further, on the basis of fragmentation profile in the MS/MS spectra, the molecular ions at 1,086.7, 1,100.7 and 1,114.6 m/z could be assigned as homologues (differing in mass by multiples of 14 Da) varying in chain length of βAA, but possessing identical peptide moiety. Subsequently, the ions at 1,086.7 m/z (with a mass difference of 1 Da from m/z 1,085.7) could be assigned as amino acid variant of 1,085.7 m/z with replacement of N-terminal of Asn to Asp in the peptide sequence. Further on the basis of literature, the molecular ion at 1,086.7 m/z was identified as iturin C with C17 β-amino fatty acid (Park et al. 1995). Thus, two new iturin C variants (1,100.7 and 1,114.6 m/z) with mass difference of 14 Da were identified in this study and were assigned as C18 and C19 βAA iturin C variants (Table 1).
Characterization of Fengycins
The molecular mass ions ranging from 1,445 to 1,555 m/z which eluted during 13.7–20.1 min RT were putatively assigned as fengycins (Figs. 3a, 4a) (Pathak et al. 2012). The [M+H]+ ions at 1,463.9, 1,477.8, 1,491.9, 1,505.9, 1,519.8 m/z varying from each other by multiple of 14 Da represent β-OH fatty acid chain length homologues of fengycin with identical peptide sequence, while the ions at 1,485.8, 1,499.9, 1,513.9 and 1,527.8 m/z were assigned as sodium adducts of corresponding protonated fengycin ions. Each of these putative fengycin ions was subjected to MS/MS analysis in order to confirm their assignments. In the LC–ESI–MS/MS spectrum of [M+H]+ ion at 1,477.8 m/z (Fig. 3b), the presence of two major intense paired product ions: −y 8 (966.4 m/z) and −y 9 (1,080.5 m/z) facilitated the assignment of 1,477.8 m/z as C15 fengycin A1 with Ala6 and Ile10 where C15 corresponds to the number of carbons in its β-hydroxy fatty acid (β-OH FA) moiety (Madonna et al. 2003; Pathak et al. 2012). The product ions at 980.4 (−y 8 ) and 1,094.5 (−y 9 ) m/z in MS/MS spectrum of parent ion [M+H]+ at 1,491.9 m/z (Fig. 3c) suggested it to be C16 fengycin B2 with Val at both positions 6 and 10 (Pathak et al. 2012). In MS/MS spectrum of parent ion [M+H]+ at 1,505.9 m/z (Fig. 3d), the presence of product ions at 994.5 (−y 8 ) and 1,108.6 (−y 9 ) m/z enabled it to be assigned as mono protonated C16 fengycin B1 with Val6 and Ile10 (Madonna et al. 2003; Pathak et al. 2012). In LC–MS spectrum, along with singly charged fengycin ions (1,445–1,555 m/z) (Fig. 3a, 4a), doubly charged fengycins ions with m/z varying from 730 to 764 could also be assigned. The Fig. 4b shows MS/MS spectrum of [M+2H]2+ ion at 753.4 m/z in which series of –b and –y as well as −b’ and −y’ type fragment ions could be assigned corresponding to C16 fengycin B1 and C17 fengycin B2, respectively. In the MS/MS spectrum of doubly charged fengycin ions, a complete series of −b and −y ions resulting from fragmentation across the depsipeptide ring along with attached Orn could be assigned (Fig. 4b), whereas only two paired intense product ions (−y 8 and −y 9 ) were observed in the MS/MS spectrum of singly charged fengycin species (Fig. 3b–d). The poor fragmentation of singly charged fengcins in gas phase collision induced dissociation may be attributed to the stability conferred by Orn residue (present outside the ring) to the octadepsipeptide ring (Geetha et al. 2010). In case of doubly charged fengycins, the second proton probably owing to its mobility leads to generation of fragment ions from ornithine to Ile. The fengycins sequences identified in this study are summarized in Table 2.
Characterization of Cyclic as Well as Linear Surfactin Variants
In LC–ESI–MS spectrum, protonated as well as sodium adducts of putative surfactins varying from each other multiple of 14 Da were observed, which eluted over a RT from 20.5 to 47.1 min (Fig. 5a) (Pecci et al. 2010; Vater et al. 2002). These protonated molecular species of putatively assigned surfactin ions were further subjected to MS/MS analysis. In LC–ESI–MS/MS of [M+H]+ at 1036.7 m/z (25.6 min RT) the fragment ions could be assigned as b 2 − b 7 , y 3 − y 6 and b’ 2 − b’ 7 , y’ 3 − y’ 6 ions, corresponding to two isobaric species of surfactin (Fig. 5b) with sequences: C15 β-OH FA-Cyclo[E–I/L–I/L–V–I/L–D–I/L] and C14 β-OH FA-Cyclo[E–I/L–I/L–I/L–I/L–D–I/L], respectively. These lipopeptide sequences represent novel positional variants of C15 surfactin (I/L2,7, V4) and C14 surfactin (I/L2,4,7) with Ile/Leu and Asp at position 5 and 6, respectively (Fig. 5b) (Grengemard et al. 1997; Stein 2005). In the same MS/MS spectrum along with −b/−b’ and −y/−y’ ions, ions at 454.5, 572.5, 685.5 m/z and 473.4, 586.4, 699.5 m/z represent the C-terminal fragment ions of two corresponding surfactins with gain of 18 Da in their mass. The C-terminal fragment ions of new C15 surfactin (I/L2,5,7,V4) at 454.5, 572.5 and 685.5 m/z represented V–I/L–D–I/L+OH2, I/L–V–I/L–D–I/L+OH2 and I/L–I/L–V–I/L–D–I/L+OH2, while the C-terminal fragment ions of new C14 surfactin (I/L2,4,5,7) at 473.4, 586.4 and 699.5 m/z corresponded to I/L–I/L–D–I/L+OH2, I/L–I/L–I/L–D–I/L+OH2 and I/L–I/L–I/L–I/L–D–I/L+OH2 peptide sequences, respectively (Fig. 5b) (Hue et al. 2001). These additional ions seem to have been generated upon the double hydrogen transfer (DHT) to the C-terminal amino acid residue followed by cleavage coupled with addition of OH2 (Yang et al. 2006). The DHT mechanism provides additional information: (1) it confirms the cyclization of peptide, (2) distinguish between cyclic depsipeptide or peptide cyclised by amide bond and (3) the generation of C-terminal peptide fragments which aid in the determination of cyclic peptide sequence with higher confidence (Yang et al. 2006). The Fig. 5c shows the b 2 − b 7 and y 3 − y 6 fragment ions assigned in the LC–ESI–MS/MS spectrum of [M+H]+ ion at 1022.7 m/z (26.0 min RT). The ions at 346.4 m/z (I/L–D–V+OH2), 445.5 m/z (V–I/L–D–V+OH2), 558.4 m/z (I/L–V–I/L–D–V+OH2) and 671.5 m/z (I/L–I/L–V–I/L–D–V+OH2) were identified as C-terminal fragment ions of C15 surfactin (I/L2,V4,7) generated upon DHT (Kowall et al. 1998). The Fig. 5d shows −b, −b’, −y and −y’ fragment ion assignments in LC–ESI–MS/MS spectrum of [M+H]+ ion at 1078.8 m/z (37.2 min RT). On the basis of −b, −y type ions new β-OH FA variant of surfactin [I/L2,4,7] with C17 chain length was identified and on the basis of −b’, −y’ ion assignments, a new cyclic peptide variant of C16 surfactin [I/L2,4,7] with substitution of Asp5 by either Glu or methyl ester of Asp (OmetD) is proposed. The additional fragment ions formed upon DHT were also identified for these two surfactin variants (Fig. 5d). The C15 surfactin (I/L2,7,Val4) with mono-methylation at Asp5 or Glu1 in the extract of B. licheniformis HSN 221 has been characterized previously using tandem mass spectrometry by Li et al. (2010). On the basis of earlier reports on occurrence of O-methyl Asp in surfactin, the mono protonated ion at 1,078.8 m/z is proposed to be a new C16 surfactin [I/L2,4,7] with O-methyl Asp5. Among the fragment ions generated upon DHT, intense bands corresponding to the ions at 671, 685, 699 and 713 m/z (with a general sequence, I/L2–I/L3–I/L/V4–D/OmetD/I/L5–I/L/D6–I/L/V7+OH2) (Fig. 5b–d) may be useful markers for quick identification of the proposed new surfactin variants. The identity of other protonated putative surfactin species at 994.7, 1,008.7, 1,022.7, 1,036.7, 1,064.7, 1,078.7 and 1,092.7 m/z was confirmed by auto LC–ESI–MS/MS analysis. These ions were found to be peptide as well as β-OH FA variants of surfactin (Table 3).
Figure 6a shows the LC–ESI–MS spectrum of protonated as well as sodium adduct mass ions varying from each other by multiples of 14 Da eluting within the time interval of 11.4–17.6 min. These ions, with mass difference of 18 Da from the surfactins characterized earlier, were thought to be the linear forms of corresponding surfactin ions with gain of H20. Moreover, these ions eluted faster than cyclic surfactins from C18 reverse phase column, suggesting their more polar nature in comparison to the cyclic counterparts. The presence of linear surfactins in the 4 day old B. globigii culture was reported by Madonna et al. (2003). The linear fengycins (lipopeptides) have also been reported in the 12 h old cell suspension of B. subtilis K1 (Pathak et al. 2012).The presence of these linear lipopeptides in the methanolic extract of B. tequilensis P15 may have resulted either during extraction from cell free fermentation broth or they represent uncyclized intermediates of surfactin biosynthetic pathway. The lipopeptides of surfactin, iturin and fengycins are synthesized by non ribosomal peptide synthetase (NRPS) machinery operated by multiple modules and their domains. Each of these domains acts as an enzyme and catalyzes the selection and activation of amino acid, peptide bond formation, elongation of peptide chain and termination of growing peptide chain. The termination is the last step where the linear lipopeptide undergoes cyclization by thioesterase domain (Te) of last module of NRPS machinery (Finking and Marahiel 2004). The surfactins and fengycins are the cyclic depsipeptides, the lactone ring in these lipopeptide is susceptible to mild alkali treatment or prolong exposure of acid. The extraction of lipopeptides from B. tequilensis was carried out by acid precipitation during which, the acidified broth (pH 2) was kept at 4 °C for 2 h to minimize the degradation of lipopeptides, thus it is less likely that linearization occurred during extraction. Moreover, the methanolic extract of B. tequilensis P15 consisted both surfactins as well as fengycins and while the linear forms of surfactins were observed, no linear fengycin could be detected. Thus, it is more likely that, the linear surfactins detected in the methanolic extract in this study represent the uncyclized intermediates. Each of these putative linear surfactins was further fragmented in order to confirm their structure and identity. Figure 6b shows LC–ESI–MS/MS of [M+H]+ ion at 1,068.8 m/z eluting at 13.4 min retention time. On the basis of −b and −y ion assignments, this molecule could be identified as linear C16 β-OH FA surfactin [I/L2,7 and V4]. Other linear surfactins identified in this study are summarized in Table 3. From the LC–MS/MS analysis, 39 surfactin species have been identified from the methanolic extract of B. tequilensis P15, out of which, 30 surfactins are found to be new and not reported previously (Table 3).
Various Bacillus species such as, B. subtilis, B. licheniformis, B. amyloliquefaciens, B. cereus, B. megaterium have been reported for their ability to produce cyclic lipopeptides (Nihorimbere et al. 2012; Nishikiori et al. 1986; Pathak et al. 2012; Pecci et al. 2010; Pueyo et al. 2009). Amongst them, cyclic lipopeptides from B. subtilis and B. amyloliquefaciens are studied in detail and well documented. B. subtilis is known to produce a concoction of more than 23 antibiotics. The genes involved in the biosynthesis of antibiotics occupy around 4–5 % of total B. subtilis genome (Stein 2005), whereas in B. amyloliquefaciens 7.5 % of total genome is devoted to antibiotic biosynthesis genes (Koumoutsi et al. 2004). Recently a molecular evidence for ability of B. tequilensis was demonstrated by identification and characterization of surfactin (sfp) gene from a marine isolate B. tequilensis NIOS11 by Porob et al. (2013). The present study shows the biosynthetic capability of this organism to produce antifungal compounds belonging to iturin, fengycin and surfactin cyclic lipopeptide families. Iturin and fengycin are potent fungicides while surfactin apart from its antifungal activity, exhibits synergism in fungal antagonism with iturins as well as fengycins (Gong et al. 2006; Ongena et al. 2007; Thimon et al. 1992; Vanittanakom et al. 1986). The antifungal cocktail of surfactins, iturins and fengycins produced by B. tequilensis P15 may have potential role in suppressing fungal diseases in agriculture crops. In addition, these lipopeptides can efficiently reduce the interfacial tension at oil water interphase. The iturin and fengycin are lower strength biosurfactants whereas surfactin is one of the most powerful biosurfactant with great surface tension lowering capacity (Peypoux et al. 1999; Deleu et al. 1999). Surfactin is also known to exhibit antiviral, antifungal, mosquitocidal, haemolytic, anticancer activities (Geetha et al. 2010; Kim et al. 2010; Kracht et al. 1999; Vater et al. 2002). The heterogeneity in population of surfactins in the extract of B. tequilensis P15 strain was found to be highest than any other Bacillus species investigated so far.
In this study, B. tequilensis P15 was used for the mass spectral characterization of lipopeptides secreted by it. The LC–MS/MS method used in this study enabled the characterization of iturins, fengycins and surfactins lipopeptides from the antifungal methanolic extract of B. tequilensis P15. The isolate P15 was found to produce highest number of surfactins with 30 novel species identified.
Abbreviations
- MS:
-
Mass spectrum
- MS/MS:
-
Tandem mass spectrum
- LC–ESI–MS:
-
Liquid chromatography electro spray ionization mass spectrometry
- LC–ESI–MS/MS:
-
Liquid chromatography electro spray ionization tandem mass spectrometry
- CID:
-
Collision induced dissociation
- βAA:
-
β-Amino acid
- βAA:
-
β-Hydroxy fatty acid
- RT:
-
Retention time
- HPLC:
-
High performance liquid chromatography
- TIC:
-
Total ion chromatogram
- SPS:
-
Smart parameter setting
- ICC:
-
Ion charge control
- Orn:
-
Ornithine
- Asn/N:
-
Aspargine
- Asp/D:
-
Aspartic acid
- OmetAsp/OmetD:
-
O-methyl aspartic acid
- Gln/Q:
-
Glutamine
- Glu/E:
-
Glutamic acid
- Tyr/Y:
-
Tyrosine
- Pro/P:
-
Proline
- Leu/L:
-
Leucine
- Ile/I:
-
Isoleucine
- Val/V:
-
Valine
- Thr/T:
-
Threonine
- Ser/S:
-
Serine
References
Bonmatin JM, Lapr′evote O, Peypoux F (2003) Diversity among microbial cyclic lipopeptides: iturins and surfactins. Activity structure relationships to design new bioactive agents. Comb Chem High Throughput Screen 6:541–556
Deleu M, Razafindralambo H, Popineau Y, Jacques P, Thonart P, Paquot M (1999) Interfacial and emulsifying properties of lipopeptides from Bacillus subtilis. Colloids Surf A Physicochem Eng Aspects 152:3–10
Finking R, Marahiel MA (2004) Biosynthesis of nonribosomal peptides1. Annu Rev Microbiol 58:453–488
Geeta L, Manonmani AM, Paily KP (2010) Identification and characterization of a mosquito pupicidal metabolite of Bacillus subtilis sub sp. subtilis strain. Appl Microbiol Biotechnol 86:1737–1744
Gong M, Wang JD, Zhang J, Yang H, Lu XF, Cheng JQ, Pei Y (2006) Study of the antifungal ability of Bacillus subtilis strain PY-1 in vitro and identification of its antifungal substance (Iturin A). Acta Biochim Biophys Sin (Shanghai) 38:233–240
Grengemard I, Peypoux F, Wallach J, Das BC, Labbe H, Caille A, Genest M, Maget-Dana R, Ptak M, Bonmatin JM (1997) Lipopeptides with improved properties: structure by NMR, purification by HPLC and structure-activity relationships of new isoleucyl-rich surfactins. J Pept Sci 2:145–154
Hue N, Serani L, Laprevote O (2001) Structural investigation of cyclic peptiolipids from Bacillus subtilis by high-energy tandem mass spectrometry. Rapid Commun Mass Spectrom 15:203–209
Isogai A, Takayama S, Murakoshi S, Suzuki A (1982) Structure of β-amino acids in antibiotics Iturin A. Tetrahedron Lett 23:3065–3068
Keller L, Surette MG (2006) Communication in bacteria: an ecological and evolutionary perspective. Nat Rev Microbiol 4:249–258
Kim SY, Kim JM, Kim SH, Bae HJ, Yi H, Yoon SH (2007) Surfactin from Bacillus subtilis displays anti-proliferative effect via apoptosis induction, cell cycle arrest and survival signalling suppression. FEBS Lett 581:865–871
Koumoutsi A, Chen XH, Henne A, Liesegang H, Hitzeroth G, Frank P, Vater J, Borriss R (2004) Structural and functional characterization of gene clusters directing nonribosomal synthesis of bioactive cyclic lipopeptides in Bacillus amyloliquefaciens strain FZB42. J Bacteriol 186:1084–1096
Kowall M, Vater J, Kluge B, Stein T, Franke P, Ziessow D (1998) Separation and characterization of surfactin isoforms produced by Bacillus subtilis OKB 105. J Colloid Interface Sci 204:1–8
Kracht M, Rokos H, Ozel M, Kowall M, Pauli G, Vater J (1999) Antiviral and haemolytic activities of surfactin isoforms and their methyl ester derivatives. J Antibiot 52:613–619
Li Y, Yang S, Mu B (2010) Structural Characterization of lipopeptide methyl esters produced by Bacillus licheniformis HSN 221. Chem Biodivers 8:2065–2075
Madonna AJ, Voorhees KJ, Taranenko NI, Laiko VV, Doroshenko VM (2003) Detection of cyclic lipopeptide biomarkers from Bacillus species using atmospheric pressure matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem 75:1628–1637
Nihorimbere V, Cawoy H, Sayer A, Brunelle A, Thonart P, Ongena M (2012) Impact of rhizosphere factors on cyclic lipopeptide signature from the plant beneficial strain Bacillus amyloliquefaciens S499. FEMS Microbiol Ecol 79:176–191
Nishikiori T, Naganawa H, Muraoka Y, Aoyagi T, Umezawa H (1986) Plipastatins: new inhibitors of phospholipase A2, produced by Bacillus cereus BMG302-fF67. II. structure of fatty acid residue and amino acid sequence. J Antibiot 39:745–754
Ongena M, Jourdan E, Adam A, Paquot M, Brans A, Joris B, Arpigny JL, Thonart P (2007) Surfactin and fengycin lipopeptides of Bacillus subtilis as elicitors of induced systemic resistance in plants. Environ Microbiol 4:1084–1090
Park JK, Hasumi K, Endo A (1995) Inhibition of the binding of oxidized low density lipoprotein to the macrophages by iturin C-related compounds. J Antibiot 48:226–232
Pathak KV, Keharia H, Gupta K, Thakur SS, Balaram P (2012) Lipopeptides from banyan endophyte, Bacillus subtilis K1: mass spectrometric characterization of a library of fengycins. J Am Soc Mass Spectrom 10:1716–1728
Pecci Y, Rivardo F, Martinotti MG, Allegrone G (2010) LC/ESI-MS/MS characterization of lipopeptide biosurfactants produced by Bacillus licheniformis V9T14 strain. J Mass Spectrom 45:772–778
Peypoux F, Pommier MT, Das BC, Besson F, Delcambe L, Michel G (1984) Structures of Bacillomycin D and Bacillomycin L peptidolipid antibiotics from Bacillus subtilis. J Antibiot 37:1600–1604
Peypoux F, Pommier MT, Marion D, Ptak M, Das BC, Michel G (1986) Revised structure of mycosubtilin, a peptidolipid antibiotic from Bacillus subtilis. J Antibiot 39:636–641
Peypoux F, Bonmatin JM, Labbe H, Das BC, Ptak M, Michel G (1991) Isolation and characterization of a new variant of surfactin, the [Val7]surfactin. Eur J Biochem 202:101–106
Peypoux F, Bonmatin JM, Wallach J (1999) Recent trends in the biochemistry of surfactin. Appl Microbiol Biotechnol 51:553–563
Porob S, Nayak S, Fernandes A, Padmanabhan P, Patil BA, Meena RM, Ramaiah N (2013) PCR screening of surfactin (sfp) gene in marine Bacillus strains and its molecular characterization from Bacillus tequilensis NIOS11. Turk J Biol 37:212–221
Pueyo MT, JrC Bloch, Carmona-Ribeiro AM, di Mascio P (2009) Lipopeptides produced by a soil Bacillus megaterium strain. Microb Ecol 57:367–378
Stein T (2005) Bacillus subtilis antibiotics: structures, synthesis and specific functions. Mol Microbiol 56:845–857
Thimon L, Peypoux F, Maget-Dana R, Roux B, Michel G (1992) Interactions of bioactive lipopeptides, iturin A and surfactin from Bacillus subtilis. Biotechnol Appl Biochem 16:144–151
Vanittanakom N, Loeffler W, Koch U, Jung G (1986) Fengycin—a novel antifungal lipopeptide antibiotic produced by Bacillus subtilis F-29-3. J Antibiot 39:888–901
Vater J, Kablitz B, Wilde C, Franke P, Mehta N, Cameotra SS (2002) Matrix-assisted laser desorption ionization-time of flight mass spectrometry of lipopeptide biosurfactants in whole cells and culture filtrates of Bacillus subtilis C-1 isolated from petroleum sludge. Appl Environ Microbiol 68:6210–6219
Volpon L, Besson F, Lancelin JM (2000) NMR structure of antibiotics plipastatins A and B from Bacillus subtilis inhibitors of phospholipase A2. FEBS Lett 485:76–80
Watrous J, Roach P, Alexandrov T, Heath BS et al (2012) Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci USA 109:E1743–E1752
Yang SZ, Wei DZ, Mu BZ (2006) Determination of the amino acid sequence in a cyclic lipopeptide using MS with DHT mechanism. J Biochem Biophys Methods 168:69–74
Acknowledgments
The authors are thankful to Prof. P. Balaram, Molecular Biophysics Unit, Indian Institute of Science, Bangalore, for providing mass spectrometry facility.
Conflict of interest
Khyati Pathak, Anjali Bose and Hareshkumar Keharia declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pathak, K.V., Bose, A. & Keharia, H. Characterization of Novel Lipopeptides Produced by Bacillus tequilensis P15 Using Liquid Chromatography Coupled Electron Spray Ionization Tandem Mass Spectrometry (LC–ESI–MS/MS). Int J Pept Res Ther 20, 133–143 (2014). https://doi.org/10.1007/s10989-013-9375-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-013-9375-7