Abstract
Context
The scale of environmental relationships is often inferred through the use of species distribution models. Yet such models are frequently developed at two distinct scales. Coarse-scale models typically use information-poor (e.g., presence-only) data to predict relative distributions across geographic ranges, whereas fine-scale models often use richer information (e.g., presence–absence data) to predict distributions at local to landscape scales.
Objectives
We unite presence–absence and presence-only data to predict occurrence of species, what we refer to as integrated distribution models. We determine if integrated models improve predictions of species distributions and identification of characteristic spatial scales of environmental relationships relative to presence–absence modeling and ensemble modeling that averages predictions from separate presence-only and presence–absence models.
Methods
We apply recent advances in integrated distribution models to predict Sherman’s fox squirrel (Sciurus niger shermani) distribution in north-central Florida. Presence-only data were collected through a citizen-science program across its geographic range, while presence–absence data were collected using camera trapping surveys across 40 landscapes.
Results
Integrated models estimated environmental relationships with greater precision and identified larger characteristic scales for environmental relationships than using presence–absence data alone. In addition, integrated models tended to have greater predictive performance, which was more robust to the amount of presence–absence and presence-only data used in modeling, than presence–absence and ensemble models.
Conclusions
Integrated distribution models hold much potential for improving our understanding of environmental relationships, the scales at which environmental relationships operate, and providing more accurate predictions of species distributions. Many avenues exist for further advancement of these modeling approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Understanding species distributions is essential to ecology, evolution, and conservation biology (Guisan and Thuiller 2005; Elith and Leathwick 2009). Species distributions models (SDMs) are often used to address ecological and conservation issues, such as quantifying habitat/environmental relationships, evaluating potential management actions, predicting the effects of land-use and climate change, and identifying priority areas for conservation planning (Loiselle et al. 2003; Evans et al. 2010; Lawler et al. 2010). In addition, these models have been frequently used for understanding the characteristic scales of environmental relationships (i.e., ‘scales of effects’; Urban et al. 1987; Smith et al. 2011; Jackson and Fahrig 2015) and the multi-level effects of landscapes on organisms (Fletcher and Hutto 2008; Thornton et al. 2011). However, the usefulness of such models is limited by the data used in model building, which often contain relatively limited information and sample bias (Norris 2004; Phillips et al. 2009; McCarthy et al. 2012).
Models of species distributions vary widely in spatial scale. Franklin (2009) categorizes species distribution models based on two contrasting spatial extents and how species data tend to vary with extent (cf. Wiens 1989). While there are exceptions to this categorization of distribution models (e.g., Albright et al. 2010), this tendency for data type to covary with spatial extent is common (e.g., Jones et al. 2010). One general category of models occurs at large extents, typically at the geographic range of species. In these situations, presence-only data are frequently used (Brotons et al. 2004; Elith et al. 2006), which often come from museum specimens or through citizen science programs (Dickinson et al. 2010). Such data are useful because of the large spatial extent, but they often suffer from sample selection bias (Phillips et al. 2009; Fourcade et al. 2014), such as presence records being more likely in areas that are easily accessible. This bias can limit the ability of using such data to identify environmental relationships (Kadmon et al. 2004) and make accurate predictions to new areas or times (Brotons et al. 2004). In addition, because absences are not available, only measures of relative suitability/probability can typically be modeled (but see Dorazio 2012; Royle et al. 2012).
In contrast, a second general category of models come from planned surveys typically collected at extents smaller than the geographic range, where interest lies in understanding local to landscape-scale distributions (Franklin 2009). In this category of models, presence–absence data (or more appropriately ‘detection–nondetection’ data; presence–absence hereafter) are frequently used (Rota et al. 2011; Lawson et al. 2014). These data are useful in that they typically have minimal sample selection bias regarding occurrence because at any survey location, presence or absence can be quantified (Dorazio 2014). This category of models contains richer information regarding the prevalence of species and may potentially allow less biased quantification of environmental relationships (Fithian et al. 2015); however, this category of models frequently suffers from small amounts of data and geographic extent. Consequently, such models may be limited in identifying the characteristic spatial scale of environmental relationships in situations where samples do not cover enough large-scale variation in landscape conditions (Oneill et al. 1996).
Given the strengths and limitations of both presence-only and presence–absence data, models that leverage both types of information may be valuable for quantifying environmental relationships and the scale of such relationships, as well as predicting species distributions across regions. Recent advances in species distribution modeling have focused on how to integrate these different sources of data to make more reliable predictions (Dorazio 2014; Keil et al. 2014; Fithian et al. 2015). We refer to this general group of models as integrated distribution models, borrowing this term from recent developments in population ecology where integrated population models link multiple data sources to estimate population dynamics (Schaub et al. 2007; Abadi et al. 2010). These approaches simultaneously use different sources of data to develop statistical models of species distribution. These advances are in contrast to increasingly used ensemble modeling consensus techniques (Araujo and New 2007; Marmion et al. 2009), which focus on summarizing predictions (e.g., average predictions) from separate models.
Advances in integrated distribution models have focused on two issues. First, models have integrated coarse-grain data, such as atlas data, with fine-grain data to downscale predictions of models (Keil et al. 2014). This approach is akin to a multi-level model (sensu Cushman and McGarigal 2002), in the sense that predictions at fine-grains are conditioned on predictions at coarse grains. One potential benefit of such models is that multi-level processes could be captured, yet these models could be limited when mis-matches occur in data (e.g., where atlas data suggest absence in a region where fine-scale data provides presence information). In addition, most distribution modeling data derive from point locations rather than coarse-grain grids, such that the spatial grain of different data sets (presence-only, presence–absence) are comparable (i.e., response data are single-level, rather than multi-level) while the spatial extents might vary considerably. A more general approach to integrate presence-only and presence–absence data has recently been independently described by Dorazio (2014) and Fithian et al. (2015). This approach is driven by viewing species distributions as spatial point processes (see below), and integrates presence-only data with planned surveys that include either presence–absence or detection–nondetection data (i.e., data that include imperfect detection). Here we focus on this latter approach. These models emphasize accounting for sample selection bias and leveraging planned surveys to improve presence-only modeling. While these models have been developed, our understanding of benefits and limitations of these models in real-world applications is still limited, with only one empirical example on Eucalpytus in Australia (Fithian et al. 2015). Furthermore, it is unclear if such models will provide insights to the characteristic scaling of environmental relationships (Thompson and McGarigal 2002; Holland et al. 2004; Jackson and Fahrig 2015). Yet given that such data integration may offer the potential to capture greater large-scale variability of the environment, integrated distribution models could provide better insight to the spatial scales of environmental relationships.
Here, we apply and compare recent statistical advances in integrated modeling of species distribution data to test if, and the extent to which, the inclusion of broad-scale presence-only data can improve modeling efforts, in terms of model predictions, estimation of environmental relationships, and the identification of the characteristic scales of environmental relationships. We first review recent modeling frameworks regarding unified models of species distribution data and how such frameworks can be applied to understand environmental relationships. We then apply this framework for interpreting the distribution of Sherman’s fox squirrels (Sciurus niger shermani) in north-central Florida. We contrast inferences from integrated distribution models with more conventional methods for species distribution, including conventional presence–absence models and ensemble models (Araujo and New 2007; Marmion et al. 2009), using a multi-scale analysis (Holland et al. 2004). We conclude by discussing the current limitations of integrated distribution models and potential extensions to improve such modeling efforts.
Methods
Uniting presence-only and presence–absence data
To integrate presence-only and presence–absence data, it is useful to view species distributions as being derived from inhomogeneous point processes (Warton and Shepherd 2010; Renner et al. 2015). Indeed, many species distribution modeling algorithms, such as MAXENT (Phillips et al. 2006), can be derived as spatial point process models (Renner and Warton 2013; Renner et al. 2015). Given that the application of such models to species distributions is recent, we briefly review and describe the relationship of inhomogeneous point processes to species distributions and its relevance for integrated distribution modeling based on recent model developments (Renner and Warton 2013; Dorazio 2014; Fithian et al. 2015; Renner et al. 2015). See Dorazio (2014) and Fithian et al. (2015) for derivation of these types of models and more details.
Point processes and thinned point processes
In spatial statistics, there has been a long history of viewing locations of species occurrence as point processes (Thompson 1955; Getis and Franklin 1987). In this context the focus is on species location, s, within a specified study region D. Point process models (PPM) focus on understanding the intensity (~density) of species, λ, in a bounded area or region across D. A point process is ‘inhomogeneous’ when intensity varies across D. This variation can be captured by spatially-explicit covariates by modeling intensity based on a log-linear relationship:
Consequently, PPMs are similar to generalized linear models (GLMs), but the focus is on spatial locations of point occurrences rather than focus being on the point occurrences themselves (Renner et al. 2015).
Thinned point process models are an extension of PPMs that acknowledge that observed point locations are a sample of all locations, and possibly a highly biased sample. For example, presence-only sampling is often biased near roads and such bias can influence predictions of models (Phillips et al. 2009). Fithian et al. (2015) extend the PPM to account for sample selection bias by adjusting the regression model to include covariates that help explain sampling bias:
where z is a bias covariate, such as distance from road (see also Warton et al. 2013). Note that such bias adjustment may only be identifiable when z(s) is not highly correlated with x(s). The log-likelihood of the inhomogeneous point process model that accounts for sample bias is then (Fithian et al. 2015):
where each observation i of a species is associated with a location s i and the covariates at that location (x i = x(s i ) and z i = z(s i )). The first component of this likelihood focuses on the information from the presence locations whereas the second component focuses on the covariate values across the region. The integral in this latter component cannot be directly estimated and is therefore approximated, typically from selecting background data points (sometimes called pseudo-absences). Background points are frequently used in presence-only modeling (Elith et al. 2006). See Warton and Shepherd (2010) and Renner et al. (2015) for interesting discussions on how PPMs help clarify the role of background points and the number background points that should be included in analyses of presence-only data. In general, Renner et al. (2015) emphasize that more background points should be used to estimate point processes than what is typically done in species distribution modeling (see below).
Joint likelihoods for integrating distribution data
Given the above point process model, a natural extension is to unite a presence-only point process model with a similar point process model based on presence–absence survey data. Joint analysis is commonly used in statistics and is increasingly used in population ecology, where likelihoods are developed for two (or more) independent data sets and a joint likelihood is formulated as the product of the two likelihoods (Schaub et al. 2007; Abadi et al. 2010). Because data from presence–absence and presence-only data are collected independently, the data can be analyzed together based on this joint likelihood. Joint analysis is thought to decrease bias and increase precision of estimates than modeling each data set in isolation (Dorazio 2014).
To appropriately link a presence-only PPM with planned survey data based on presence–absence information, Fithian et al. (2015) suggest that a model formulation for presence–absence data by relating counts at sample locations, A i (where |A i | is the area of A i ) to a point process:
The probability of occurrence can be derived based on using a complementary log–log link function:
See Royle and Dorazio (2008, pp. 150–151) for further discussion regarding the use of the complementary log–log link function for relating abundance to occupancy. Taken together, the log-likelihood of the presence–absence data can be described as (Fithian et al. 2015):
The joint log-likelihood is then the sum of these two log-likelihoods (Eqs. 3, 6). This modeling approach can be implemented with the multispeciesPP package in R (Fithian et al. 2015). To demonstrate the utility of this approach, we used presence-only and presence–absence (i.e., detection–nondetection; see below) data collected on Sherman’s fox squirrels.
Study species and study area
Fox squirrels (S. niger) are a common, widely-distributed tree squirrel native to North America and found east of the Rocky Mountains (Koprowski 1994). Of the 10 subspecies of fox squirrel, four subspecies occur in Florida. Our analysis focuses on Sherman’s fox squirrel, which primarily occurs in central and north Florida, but also ranges into southern Georgia. Sherman’s fox squirrels are sparsely distributed and in low densities and are listed as a State Species of Special Concern by Florida’s Endangered and Threatened Species Rule (Loeb and Moncrief 1993). Sherman’s fox squirrels and are known to use most major vegetative communities throughout their range; however, they are most commonly associated with inland pine-dominated vegetation found throughout most of peninsular Florida (Moore 1956, 1957).
We conducted our surveys within the core range of Sherman’s fox squirrels in central and northern Florida on public and private lands. The vegetative communities at our sites were highly variable, and included open grasslands, pine-dominated forests, hardwood-dominated forests, mixed pine–hardwoods, bottomland forest, and clear cuts. The canopy trees varied between sites, but the dominant pine trees included longleaf (Pinus palustris), slash (P. elliottii) and loblolly (P. taeda) pines, and the dominant oaks were turkey (Quercus laevis), live (Q. virginiana), laurel (Q. laurifolia), and water (Q. nigra) oaks. The sites varied in their vegetation management practices, which included cattle grazing, mowing, prescribed burning, and no active management. Nearly all sites where pines outnumbered hardwoods were managed for timber.
Presence-only data
We developed and promoted a publically available web-based tool (webpage) to generate presence-only locations of Sherman’s fox squirrel throughout their geographic range in Florida. The web-based tool recorded georeferenced locations (latitude and longitude in decimal degrees) using a Google map application to record sightings in the database. For this analysis, we extracted data from the website from August 20 2011 until May 1 2012 to coincide with the peak activity in fox squirrel activity typically related to mast availability (Weigl et al. 1989). We also subset these data further for the time period January–May 2012 to reflect only the time period of camera-trapping (see below). Results from this subset of data were similar, in terms of environmental relationships and metrics of predictive performance (Unpublished results).
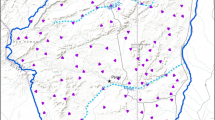
We reviewed each presence-only data point from the online survey. We quarantined location points which appeared to erroneous due to user error of mapping software (i.e. locations in the middle of a water body). To increase the validity of these suspicious occurrence data, we attempted to verify the locations and receive more specific location information from the participant that submitted the data. We removed all erroneous points which could not be verified. This resulted in 2785 presence-only points used for modeling (871 points from January to May 2012; Fig. 1).
Study area in Florida, USA, where presence–absence and presence-only data collection occurred for Sherman’s fox squirrels. Presence-only (citizen science) locations shown as small, grey dots. Landscapes for presence–absence sampling (camera trapping) shown in red. Major roads included as grey lines. (Color figure online)
Presence–absence data
We conducted intensive field surveys for Sherman’s fox squirrels using passive camera traps. We sampled across the study region using a hierarchal approach. We selected 40, 7.65 km2 landscapes across the study region using a stratified random design to capture major vegetative communities used by fox squirrels (Fig. 1). We sampled 10 landscapes in upland pine habitats, 10 in mesic flatwoods habitats and the remaining 20 without regard to a vegetative community. Within each landscape we random placed five 5.3 ha survey grids within each landscape. Each grid consisted of 9 sampling points in a 3 × 3-grid arrangement with 115 m spacing. For this analysis, we randomly selected a subset of points (n = 252 across the 40 landscapes) to use in models to reduce potential impacts of spatial autocorrelation influencing results (see “Discussion” section). Between January and June 2012 (a period of high activity for fox squirrels in Florida; Moore 1957), each sampling point was surveyed for 7 consecutive nights with a passive digital camera (Bushnell Trophy Cam model 119436c, Bushnell Outdoor Products, Overland Park, KS) baited with pecans (Carya illinoinensis) and cracked corn (Zea mays) placed at the base of a tree 1.5 m from the camera. When no photo of a fox squirrel was taken during the 7-night period, it was considered an absence (see below).
Analysis
To illustrate the approach and potential utility of integrated models, we contrasted conventional GLM models for presence–absence (detection–nondetection) data with integrated models and ensemble models that averaged predictions from separate presence-only and presence–absence models to make consensus predictions. In this comparison, integrated models use more data than conventional models, though such data contain less information (and possibly include sample selection bias; see below), while ensemble models use the same data used in integrated models, yet models are developed separately for presence–absence and presence-only data.
We considered three covariates for modeling the distribution of fox squirrels: canopy cover, distance from edge, and distance from roads. For canopy cover, we used circular buffers of different size radii (100 m, 500 m, 1 km, 2 km, 3 km, 4 km, and 5 km) to determine the characteristic scale of the environmental relationship of fox squirrel occurrence (Thornton and Fletcher 2014), and considered non-linear relationships with canopy cover via inclusion of quadratic terms. The scales we consider reflect area less than the average home range of fox squirrels (~25 ha on average) and up to approximately twice the average dispersal distance of fox squirrels (Kantola and Humphrey 1990). Canopy cover (percent tree cover) was taken from the 2011 National Land Cover Database and was re-scaled using a moving-window analysis. We did not adjust for overlapping landscapes being used in these buffers (sensu Holland et al. 2004) because we found no significant evidence for spatial autocorrelation in the residuals of models (Fig. S1; Zuckerberg et al. 2012). We did not consider buffers of different sizes in the same model to reduce multi-collinearity in model fitting. Because fox squirrels can concentrate their activities near habitat edges (Koprowski 1994), we considered the natural log of distance to edge of different vegetative communities using land-cover classes defined by the Florida Natural Areas Inventory (2010). We also considered distance to roads as a covariate (z(s) in Eq. 3 above) in integrated models to account for potential presence-only sample bias (Kadmon et al. 2004; McCarthy et al. 2012). While other factors may also influence fox squirrel distributions, these covariates were selected to illustrate the different ways in which covariates can be relevant for integrated models and the identification of characteristic scales of species–environment relationships.
The presence–absence GLM was based on a complementary log–log link function (see Eq. 6); note that a conventional logistic regression using a logit link function provided identical results. For the presence-only component of the integrated model, we selected background points based on a regular 2 × 2 km grid across the study area (Renner et al. 2015), which resulted in 13,713 points. Presence-only models were also used to create a consensus ensemble (Marmion et al. 2009), which were the same as the integrated model but with the presence–absence data removed (Eq. 3). Ensemble predictions were derived using a weighted average of model predictions (based on the area under the curve (AUC) statistic for each model) from the separate presence-only and presence–absence models (Marmion et al. 2009). Note that presence–absence and integrated models used here ignore the problem of imperfect detection. Currently, imperfect detection for occupancy has yet to be accounted for in integrated distribution models (see “Discussion” section). Nonetheless, based on an occupancy model using the camera-trapping data (MacKenzie et al. 2002), the estimated probability of detecting fox squirrels at a sample point at least one night of the seven night sample, given that the species occurred at the point, was 0.82.
To determine the characteristic scales of environmental relationships, we used a model selection approach to identify the most parsimonious scale for interpreting the relationship of fox squirrel distribution with canopy cover separately for the presence–absence GLM and the integrated GLM. In this assessment, we contrasted models using Akaike’s information criterion (AIC) that varied in the spatial scale of canopy cover (allowing for non-linearity in canopy cover relationships) while forcing distance from edge into the model and distance to roads in the integrated GLM to account for sample selection bias. We then took the selected model and further attempted to reduce model complexity by contrasting the model with a model that only included distance from edge and an intercept-only model. To determine effects on environmental relationships, we contrasted estimated β values, associated SEs, and predicted partial relationships.
To determine predictive performance, we used block validation (Wenger and Olden 2012). To do so, we split our presence–absence data into fourfolds at the scale of landscapes (7.65 km2) rather than points. Presence-only data were only used in model training (building) and were not used for validation. Block validation is helpful because creating folds at the sample unit (point) level results in validation data that can be more correlated with training data due to closer geographic proximity. We then assessed model predictions using two threshold independent measures—the AUC and a cross-validated log-likelihood (LL; Lawson et al. 2014). AUC is based on rank differences and is not impacted by predicted prevalence, whereas cross-validated log-likelihoods are measures of fit to the new data and incorporate predicted prevalence into assessments. We also used two threshold-dependent measures—the true skill statistic (TSS) and the kappa statistic (Fielding and Bell 1997; Liu et al. 2011). For TSS and kappa, we set a threshold cutoff based on maximizing the sum of the specificity and sensitivity (Liu et al. 2013).
Finally, we determined how the amount of presence–absence and presence-only data could alter the potential benefits of integrated models. To do so, we performed a similar block validation scheme as that described above; however, we altered model training in two ways. First, we sequentially reduced the amount of presence–absence data by reducing the number of landscapes used in model training from 30 (three of four folds above) down to 5 landscapes, using 10 landscapes for validation. For each sequential reduction, we calculated AUC, LL, TSS, and kappa based on integrated distribution models versus GLM on presence–absence data. We expected that GLMs would decline in predictive performance as the amount of training data declined, whereas integrated models to retain similar predictive performance. Second, we reduced the amount of presence-only data used in model training while keeping the amount of presence–absence data constant. In this scenario, we randomly removed 5–95 % of the original presence-only data. We expected that integrated models would decline in performance as the amount of presence-only data declined.
Results
The most parsimonious GLM model describing presence–absence data included a non-linear relationship of canopy cover at the 1-km scale and no significant relationship with distance from edge based on model selection criteria and parameter estimates (βedge = 0.99 ± 0.85 SE, P = 0.247; Table S1; Figs. 2, 3). For integrated models, the most parsimonious model identified a non-linear relationship of canopy cover at the 4-km scale and a positive, significant relationship with distance from edge (βedge = 0.19 ± 0.02, P < 0.0001; Table S2; Figs. 2, 3). For the integrated model, there was strong evidence for road-based sampling bias in presence-only locations identified through this model (δ road = −1.15 ± 0.04; z = −23.10, P < 0.0001). As expected, data for the integrated model spanned a larger environmental gradient than for the presence–absence GLM (Fig. S2). For example, the range of percent canopy cover at the 4 km scale for the presence–absence data was 0–82 %, while for the integrated data it was 0–100 %. In general, estimates from the integrated model were weaker (in terms of smaller β parameters) but also had greater precision (smaller SEs), such that the coefficient of variation of parameter estimates from integrated models was smaller and there was less uncertainty in partial relationships (Fig. 3).
Model selection indicates a larger characteristic scaling relationship of fox squirrel distribution with canopy cover using an integrated distribution model. Shown are changes in Akaike’s information criterion (AIC) as a function of the scale (radius) at which canopy cover is quantified for conventional (presence–absence) GLMs and integrated GLMs. In both sets of models, non-linear relationships with canopy cover were considered (via a quadratic term) and distance from edge was included
Predicted environmental relationships of Sherman fox squirrels with a percent canopy cover at the 1-km scale, b percent canopy cover at a 4-km scale; and c distance to edge based on a generalized linear model (GLM) with presence–absence data only (red/dashed line) and integrated distribution model (blue). (Color figure online)
Based on block validation, the most parsimonious integrated model tended to predict fox squirrel distribution better than the GLM model that used presence–absence data alone or an ensemble model that averaged predictions from separate presence–absence and presence-only models, based on measures of AUC (presence–absence = 0.74; ensemble = 0.76, integrated = 0.79), TSS (presence–absence = 0.51; ensemble = 0.53, integrated = 0.57), and kappa (presence–absence = 0.31; ensemble = 0.32, integrated = 0.41). For cross-validated log-likelihoods, performance was similar for presence–absence and integrated models and slightly less for ensemble models (presence–absence = −25.2; ensemble = −26.6, integrated = −25.3). In addition, predictions from the integrated models were less sensitive to the amount of presence–absence data used in model building, whereas predictive accuracy from the GLM and the ensemble model decreased as the amount of presence–absence data decreased in model building, particularly when less than 11 landscapes were used in model training (Fig. 4a–d). The integrated model also showed little sensitivity to the amount of presence-only data used in model building (Fig. 4e–h).
Effects of (a–d) the amount of presence–absence data (number of landscapes) and (e–h) amount of presence-only data used in distribution modeling on measures of predictive performance for presence–absence models, ensemble models, and integrated distribution models. Shown are results based on (a, e) AUC and the True Skill Statistic; other metrics (r and kappa) showed similar patterns
Discussion
Accurately predicting species distributions is essential for many ecological and conservation problems, yet the data used for building predicted models are often limited (Pearce and Boyce 2006). We found that by uniting both broad-scale presence-only data with finer-scale presence–absence data, we identified a larger characteristic scale for the relationship of fox squirrel distribution with canopy cover. In addition, metrics regarding predictive performance were generally greater. While each metric showed modest improvements (Fig. 4), this pattern is notable for two reasons: (1) the presence–absence data were used for model evaluation, such that presence–absence models could be expected to perform better due to similar sampling methods, and (2) benefits were more robust to sample size. Importantly, these benefits tended to be greater than from using common post hoc approaches of ensemble techniques (Araujo and New 2007; Marmion et al. 2009).
The value of uniting data in distribution models
The integrated distribution modeling framework we used was recently developed independently by Dorazio (2014) and Fithian et al. (2015). In both situations, the rationale for integrating data was to improve presence-only modeling. Presence-only modeling is commonly used, in a large part because of widely available data and because such data are frequently available across broad scales. However, presence-only data have limited information content (e.g., prevalence of the species is unknown) and such data often suffer from sample selection bias, where presence locations are frequently collected near roads or other easily accessible areas (Kadmon et al. 2004; Loiselle et al. 2008; Phillips et al. 2009). By integrating presence–absence information, this bias potentially can be ameliorated, even with relatively little amounts of presence–absence data. In addition, integrating presence–absence data provides information on species prevalence that is not available with presence-only data (Hastie and Fithian 2013), which can be helpful when the goal is to model the probability of occurrence.
While presence–absence data can improve presence-only modeling efforts, we argue the corollary that integration of presence-only data can improve presence–absence modeling efforts. There are at least three reasons why integration of presence-only data may prove helpful for presence–absence modeling efforts. First, presence–absence data are often more limited in amount, which can lead to greater uncertainty in environmental relationships. For fox squirrels, there was greater uncertainty in predicted environmental relationships when using presence–absence data than when using an integrated distribution model and significant relationships with distance from edge were only revealed with the integrated model (Fig. 3). Given the widespread influence of edge effects in shaping distributions across landscapes (Ries et al. 2004), this pattern illustrates how important landscape effects can potentially be better revealed through data integration. Second, presence-only data are often available across broader extents than presence–absence data. When there is interest in extrapolating predictions of presence–absence models into new areas (Miller et al. 2004), integrating presence-only information that may be available across a broader region may improve modeling efforts. Third, integrating presence-only data may provide more information regarding wider environmental gradients than those sampled with presence–absence information, particularly for large-scale gradients and conditions not captured in planned surveys (Smith et al. 2011). Such integration could improve the ability of presence–absence modeling to capture potential non-linearities of environmental relationships and ultimately environmental spaces of niches (Austin 2002; Soberon 2007).
Furthermore, this example illustrates how characteristic scales identified in species-environment relationships can be sensitive to the data used and highlights that by increasing the spatial extent considered, larger characteristic scales may emerge. Such issues may help explain variability observed in characteristic scales identified in species in different regions (Jackson and Fahrig 2015). Given the long-standing and widespread interest in understanding the importance of spatial scale in ecology and evolution (Wiens 1989; Horne and Schneider 1995; Fletcher et al. 2013; Jackson and Fahrig 2014), models that leverage different sources of data across scales may prove useful in many situations.
Current limitations and future extensions
Integrating multiple data sources into models of species distribution is a new perspective for distribution modeling and many advancements should occur. While the integrated distribution model improved predictions, there were some limitations of this modeling approach that could be improved.
First, the model we used only included simple non-linearities in environmental relationships that were accounted for by adding polynomial terms. Many other SDMs can capture highly non-linear relationships (e.g., Elith and Graham 2009). Fithian et al. (2015) argue that Eq. 3 could be extended to include non-linear basis functions and models could be extended to include splines, akin to generalized additive models. In addition, Renner et al. (2015) show how point process models can be implemented with MAXENT, an algorithm that allows for highly non-linear relationships to be estimated. Yet further developments are needed for formally integrating MAXENT with planned survey data.
Second, this approach ignored the problem of imperfect detection due to observation errors. Imperfect detection is common (Kellner and Swihart 2014), particularly in animal populations, and heterogeneity in imperfect detection can influence the predictive performance of models (Rota et al. 2011; Lahoz-Monfort et al. 2014). Dorazio (2014) formulated an integrated distribution model that linked presence-only data with abundance data using an N-mixture model formulation (Royle 2004; Kery et al. 2005), but analogous formulations that focus on occupancy, rather than abundance, have yet to be developed.
Third, this modeling approach does not formally account for spatial autocorrelation. While we found no significant autocorrelation in the residuals of models, there was a non-significant tendency for autocorrelation in the integrated model (Fig. S1). Renner et al. (2015) review some point process models that acknowledge the potential for spatial autocorrelation, which could be extended into an integrated distribution modeling framework. In addition, it may be feasible to extend this approach to other common regression-based approaches for dealing with spatial dependence (Beale et al. 2010; de Knegt et al. 2010).
Fourth, these frameworks could be extended to better accommodate some aspects of multi-scale environmental relationships. While the approach by Keil et al. (2014) naturally accounts for multi-level effects (e.g., different effects at the patch or landscape scale), the approach we used here did not accommodate covariates influencing distributions at different grains or levels for the independent data sets (Fletcher and Hutto 2008). The framework we used assumed that environmental covariates relevant to presence-only locations are the same as for presence–absence and occur at the same scales for the two data sources. Yet it would be possible to link differential covariate effects operating across regions (e.g., watersheds) that could influence distribution based on both presence–absence and presence-only data. Nonetheless, integrated models easily accommodate data arising from different spatial extents, which can help improve extrapolation and transferring model predictions across space (Miller et al. 2004; Wenger and Olden 2012).
Conclusions
Integrating multiple data sources into models of species distribution is a new frontier for distribution modeling and provides a means to reliably tackle some current limitations in our understanding of characteristic scales (Smith et al. 2011; Jackson and Fahrig 2015; Miguet et al. in press). Here we provide an example of integrating presence–absence and presence-only data, but other types of formal data integration could occur (e.g., information on breeding, dispersal, etc.). By integrating multiple sources of information into the modeling process, greater insights into environmental relationships and more accurate predictions can occur. Consequently, we expect such models will provide a powerful approach for addressing problems of species distributions and ongoing landscape change.
References
Abadi F, Gimenez O, Arlettaz R, Schaub M (2010) An assessment of integrated population models: bias, accuracy, and violation of the assumption of independence. Ecology 91:7–14
Albright TP, Pidgeon AM, Rittenhouse CD, Clayton MK, Flather CH, Culbert PD, Wardlow BD, Radeloff VC (2010) Effects of drought on avian community structure. Glob Change Biol 16:2158–2170
Araujo MB, New M (2007) Ensemble forecasting of species distributions. Trends Ecol Evol 22:42–47
Austin MP (2002) Spatial prediction of species distribution: an interface between ecological theory and statistical modelling. Ecol Model 157:101–118
Beale CM, Lennon JJ, Yearsley JM, Brewer MJ, Elston DA (2010) Regression analysis of spatial data. Ecol Lett 13:246–264
Brotons L, Thuiller W, Araujo MB, Hirzel AH (2004) Presence–absence versus presence-only modelling methods for predicting bird habitat suitability. Ecography 27:437–448
Cushman SA, McGarigal K (2002) Hierarchical, multi-scale decomposition of species-environment relationships. Landscape Ecol 17:637–646
de Knegt HJ, van Langevelde F, Coughenour MB, Skidmore AK, de Boer WF, Heitkonig IMA, Knox NM, Slotow R, van der Waal C, Prins HHT (2010) Spatial autocorrelation and the scaling of species–environment relationships. Ecology 91:2455–2465
Dickinson JL, Zuckerberg B, Bonter DN (2010) Citizen science as an ecological research tool: challenges and benefits. In: Futuyma DJ, Shafer HB, Simberloff D (eds) Annual review of ecology, evolution, and systematics, vol 41, pp 149–172
Dorazio RM (2012) Predicting the geographic distribution of a species from presence-only data subject to detection errors. Biometrics 68:1303–1312
Dorazio RM (2014) Accounting for imperfect detection and survey bias in statistical analysis of presence-only data. Glob Ecol Biogeogr 23:1472–1484
Elith J, Graham CH (2009) Do they? How do they? WHY do they differ? On finding reasons for differing performances of species distribution models. Ecography 32:66–77
Elith J, Leathwick JR (2009) Species distribution models: ecological explanation and prediction across space and time. Annual review of ecology evolution and systematics, pp 677–697
Elith J, Graham CH, Anderson RP, Dudik M, Ferrier S, Guisan A, Hijmans RJ, Huettmann F, Leathwick JR, Lehmann A, Li J, Lohmann LG, Loiselle BA, Manion G, Moritz C, Nakamura M, Nakazawa Y, Overton JM, Peterson AT, Phillips SJ, Richardson K, Scachetti-Pereira R, Schapire RE, Soberon J, Williams S, Wisz MS, Zimmermann NE (2006) Novel methods improve prediction of species’ distributions from occurrence data. Ecography 29:129–151
Evans JM, Fletcher RJ Jr, Alavalapati J (2010) Using species distribution models to identify suitable areas for biofuel feedstock production. Glob Chang Biol Bioenergy 2:63–78
Fielding AH, Bell JF (1997) A review of methods for the assessment of prediction errors in conservation presence/absence models. Environ Conserv 24:38–49
Fithian W, Elith J, Hastie T, Keith DA (2015) Bias correction in species distribution models: pooling survey and collection data for multiple species. Methods Ecol Evol 6:424–438
Fletcher RJ Jr, Hutto RL (2008) Partitioning the multi-scale effects of human activity on the occurrence of riparian forest birds. Landscape Ecol 23:727–739
Fletcher RJ Jr, Revell A, Reichert BE, Kitchens WM, Dixon JD, Austin JD (2013) Network modularity reveals critical scales for connectivity in ecology and evolution. Nat Commun 4:2572
Fourcade Y, Engler JO, Roedder D, Secondi J (2014) Mapping species distributions with MAXENT using a geographically biased sample of presence data: a performance assessment of methods for correcting sampling bias. PLoS ONE 9:e97122
Franklin J (2009) Mapping species distributions: spatial inference and prediction. Cambridge University Press, Cambridge
Getis A, Franklin J (1987) Second-order neighborhood analysis of mapped point patterns. Ecology 68:473–477
Guisan A, Thuiller W (2005) Predicting species distribution: offering more than simple habitat models. Ecol Lett 8:993–1009
Hastie T, Fithian W (2013) Inference from presence-only data; the ongoing controversy. Ecography 36:864–867
Holland JD, Bert DG, Fahrig L (2004) Determining the spatial scale of species’ response to habitat. Bioscience 54:227–233
Horne JK, Schneider DC (1995) Spatial variance in ecology. Oikos 74:18–26
Inventory FNA (2010) Guide to the natural communities of Florida: 2010. Tallahassee
Jackson HB, Fahrig L (2015) Are ecologists conducting research at the optimal scale? Glob Ecol Biogeogr 24:52–63
Jackson ND, Fahrig L (2014) Landscape context affects genetic diversity at a much larger spatial extent than population abundance. Ecology 95:871–881
Jones CC, Acker SA, Halpern CB (2010) Combining local- and large-scale models to predict the distributions of invasive plant species. Ecol Appl 20:311–326
Kadmon R, Farber O, Danin A (2004) Effect of roadside bias on the accuracy of predictive maps produced by bioclimatic models. Ecol Appl 14:401–413
Kantola AT, Humphrey SR (1990) Habitat use by Sherman’s fox squirrel (Sciurus niger shermani) in Florida. J Mammal 71:411–419
Keil P, Wilson AM, Jetz W (2014) Uncertainty, priors, autocorrelation and disparate data in downscaling of species distributions. Divers Distrib 20:797–812
Kellner KF, Swihart RK (2014) Accounting for imperfect detection in ecology: a quantitative review. PLoS ONE 9:e111436
Kery M, Royle JA, Schmid H (2005) Modeling avian abundance from replicated counts using binomial mixture models. Ecol Appl 15:1450–1461
Koprowski JL (1994) Sciurus niger. Mamm Species 479:1–9
Lahoz-Monfort JJ, Guillera-Arroita G, Wintle BA (2014) Imperfect detection impacts the performance of species distribution models. Glob Ecol Biogeogr 23:504–515
Lawler JJ, Shafer SL, Blaustein AR (2010) Projected climate impacts for the amphibians of the western hemisphere. Conserv Biol 24:38–50
Lawson CR, Hodgson JA, Wilson RJ, Richards SA (2014) Prevalence, thresholds and the performance of presence–absence models. Methods Ecol Evol 5:54–64
Liu C, White M, Newell G (2011) Measuring and comparing the accuracy of species distribution models with presence–absence data. Ecography 34:232–243
Liu C, White M, Newell G (2013) Selecting thresholds for the prediction of species occurrence with presence-only data. J Biogeogr 40:778–789
Loeb SC, Moncrief ND (1993) The biology of fox squirrels in the Southeast: a review. In: Moncrief ND, Edwards JW, Tappe PA (eds) Fox squirrels, Sciurus niger, Proceedings of the 2nd symposium of Southeast, pp 1–20
Loiselle BA, Howell CA, Graham CH, Goerck JM, Brooks T, Smith KG, Williams PH (2003) Avoiding pitfalls of using species distribution models in conservation planning. Conserv Biol 17:1591–1600
Loiselle BA, Jorgensen PM, Consiglio T, Jimenez I, Blake JG, Lohmann LG, Montiel OM (2008) Predicting species distributions from herbarium collections: does climate bias in collection sampling influence model outcomes? J Biogeogr 35:105–116
MacKenzie DI, Nichols JD, Lachman GB, Droege S, Royle JA, Langtimm CA (2002) Estimating site occupancy rates when detection probabilities are less than one. Ecology 83:2248–2255
Marmion M, Parviainen M, Luoto M, Heikkinen RK, Thuiller W (2009) Evaluation of consensus methods in predictive species distribution modelling. Divers Distrib 15:59–69
McCarthy KP, Fletcher RJ, Rota CT, Hutto RL (2012) Predicting species distributions from samples collected along roadsides. Conserv Biol 26:68–77
Miguet P, Jackson HB, Jackson ND, Martin AE, Fahrig L. What determines the spatial extent of landscape effects on species? Landscape Ecol (in press)
Miller JR, Turner MG, Smithwick EAH, Dent CL, Stanley EH (2004) Spatial extrapolation: the science of predicting ecological patterns and processes. Bioscience 54:310–320
Moore JC (1956) Variation in the fox squirrel in Florida. Amer Midland Nat 55:41–65
Moore JC (1957) The natural history of the fox squirrel, Sciurus niger shermani. Bull Am Mus Nat Hist 113:1–72
Norris K (2004) Managing threatened species: the ecological toolbox, evolutionary theory and declining-population paradigm. J Appl Ecol 41:413–426
Oneill RV, Hunsaker CT, Timmins SP, Jackson BL, Jones KB, Riitters KH, Wickham JD (1996) Scale problems in reporting landscape pattern at the regional scale. Landscape Ecol 11:169–180
Pearce JL, Boyce MS (2006) Modelling distribution and abundance with presence-only data. J Appl Ecol 43:405–412
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Model 190:231–259
Phillips SJ, Dudik M, Elith J, Graham CH, Lehmann A, Leathwick J, Ferrier S (2009) Sample selection bias and presence-only distribution models: implications for background and pseudo-absence data. Ecol Appl 19:181–197
Renner IW, Elith J, Baddeley A, Fithian W, Hastie T, Phillips SJ, Popovic G, Warton DI (2015) Point process models for presence-only analysis. Methods Ecol Evol 6:366–379
Renner IW, Warton DI (2013) Equivalence of MAXENT and poisson point process models for species distribution modeling in ecology. Biometrics 69:274–281
Ries L, Fletcher RJ, Battin J, Sisk TD (2004) Ecological responses to habitat edges: mechanisms, models, and variability explained. Annu Rev Ecol Evol Syst 35:491–522
Rota CT, Fletcher RJ Jr, Evans JM, Hutto RL (2011) Does accounting for detectability improve species distribution models? Ecography 34:659–670
Royle JA (2004) N-mixture models for estimating population size from spatially replicated counts. Biometrics 60:108–115
Royle JA, Chandler RB, Yackulic C, Nichols JD (2012) Likelihood analysis of species occurrence probability from presence-only data for modelling species distributions. Methods Ecol Evol 3:545–554
Royle JA, Dorazio RM (2008) Hierarchical modeling and inference in ecology: the analysis of data from populations, metapopulations, and communities. Academic Press, New York
Schaub M, Gimenez O, Sierro A, Arlettaz R (2007) Use of integrated modeling to enhance estimates of population dynamics obtained from limited data. Conserv Biol 21:945–955
Smith AC, Fahrig L, Francis CM (2011) Landscape size affects the relative importance of habitat amount, habitat fragmentation, and matrix quality on forest birds. Ecography 34:103–113
Soberon J (2007) Grinnellian and Eltonian niches and geographic distributions of species. Ecol Lett 10:1115–1123
Thompson CM, McGarigal K (2002) The influence of research scale on bald eagle habitat selection along the lower Hudson River, New York (USA). Landscape Ecol 17:569–586
Thompson HR (1955) Spatial point processes, with applications to ecology. Biometrika 42:102–115
Thornton DH, Branch LC, Sunquist ME (2011) The influence of landscape, patch, and within-patch factors on species presence and abundance: a review of focal patch studies. Landscape Ecol 26:7–18
Thornton DH, Fletcher RJ Jr (2014) Body size and spatial scales in avian response to landscapes: a meta-analysis. Ecography 37:454–463
Urban DL, Oneill RV, Shugart HH (1987) Landscape ecology. Bioscience 37:119–127
Warton DI, Renner IW, Ramp D (2013) Model-based control of observer bias for the analysis of presence-only data in ecology. PLoS ONE 8:e79168
Warton DI, Shepherd LC (2010) Poisson point process models solve the “pseudo-absence problem” for presence-only data in ecology. Ann Appl Stat 4:1383–1402
Weigl PD, Steele MA, Sherman LJ, Ha JC, Sharpe TL (1989) The ecology of the fox squirrel Sciurus niger in North Carolina: implications for survival in the Southeast. Bull Tall Timbers Res Stn 24(I-XII):1–93
Wenger SJ, Olden JD (2012) Assessing transferability of ecological models: an underappreciated aspect of statistical validation. Methods Ecol Evol 3:260–267
Wiens JA (1989) Spatial scaling in ecology. Funct Ecol 3:385–397
Zuckerberg B, Desrochers A, Hochachka WM, Fink D, Koenig WD, Dickinson JL (2012) Overlapping landscapes: a persistent, but misdirected concern when collecting and analyzing ecological data. J Wildl Manag 76:1072–1080
Acknowledgments
B. Reichert and three anonymous reviewers provided thoughtful reviews on previous versions of this manuscript, which greatly clarified the ideas presented here. R. Dorazio provided useful insight. We thank the University of Florida, the Florida Fish and Wildlife Conservation Commission, the private landowners, National Forest Service, Florida Forest Service, Florida Park Service, The Florida National Guard’s Camp Blanding Joint Training Center, and the University of Florida’s Ordway-Swisher Biological Station, Plant Science Research & Education Unit, and Austin Cary Forest for providing site access and logistical support. We also thank the U.S. Department of Agriculture, USDA-NIFA Initiative Grant No. 2012-67009-20090 for support. Finally, we thank the technicians and many volunteers for assistance in the field, and the voluntary participants for submitting their sightings to our online survey.
Author information
Authors and Affiliations
Corresponding author
Additional information
Special issue: Multi-scale habitat modeling.
Guest Editors: K. McGarigal and S. A. Cushman.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fletcher, R.J., McCleery, R.A., Greene, D.U. et al. Integrated models that unite local and regional data reveal larger-scale environmental relationships and improve predictions of species distributions. Landscape Ecol 31, 1369–1382 (2016). https://doi.org/10.1007/s10980-015-0327-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-015-0327-9