Abstract
Muscle LIM Protein (MLP) is small, just 198 amino acid long protein, which is specifically expressed in slow skeletal muscle and cardiac tissues. This article will focus on the cardiac functions of MLP: the current knowledge about localisation data, binding partners and animal models for the protein will be summarised, and the role of MLP in maintaining a healthy heart be discussed. This review will furthermore attempt to identify gaps in our knowledge—and hence future research potential—with a special focus on MLP’s role in cardiac mechano-signalling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Since Muscle LIM Protein (MLP) was first described in 1994 (Arber et al. 1994), almost a hundred articles have been published on the protein, many interaction partners identified, several localisations described and relation to diseases established in both mice and humans. However, we still do not fully understand the precise functions of the protein. In particular, more recent publications have highlighted that we need to critically re-asses our current knowledge about MLP’s role in the healthy and diseased heart.
This article will summarise what we know about this small, just 198 amino acid long protein and its role in maintaining a healthy heart.
MLP in cardiac disease
A knock-out of the CSRP3 gene coding for MLP was presented in 1997 by Arber and colleagues. The animals were viable, but had cardiac phenotypes: a subset of the MLP−/− mice died with in the first two weeks (“early phenotype”) due to congestive heart failure. The remaining animals survived until adulthood and developed a late phenotype resembling dilated cardiomyopathy in humans. Signs of cardiac hypertrophy as shown by an induction of a foetal gene expression programme were evident in both groups of animals. A more detailed examination of the adult MLP+/− mice revealed abnormalities at the intercalated disc (Ehler et al. 2001), accompanied by an up-regulation of adherens junction proteins, down-regulation of the gap junction protein connexin-43 and a more convoluted appearance of the intercalated disc. In contrast, heterozygous MLP+/− mice do not display an overt phenotype, but are characterised by attenuated cardiac remodelling after myocardial infarction (Heineke et al. 2005).
The first observation that MLP might be implicated in human cardiac disease was reported by Zolk et al. (2000), who showed a significant reduction in MLP protein levels in human failing heart. The effect was found in cardiac samples from both dilated and ischemic cardiomyopathy patients, suggesting that reduced MLP protein levels might be a general marker for heart failure. Unfortunately, the huge variability of MLP protein content in non-failing control hearts (Zolk et al. 2000, and Fig. 1) preclude using MLP levels for diagnostic purposes in the case of heart failure. When we investigated MLP expression in non-failing versus DCM heart samples, we only found slightly reduced levels of MLP in the failing hearts (Fig. 1). Interestingly, mechanical support of failing hearts with left ventricular assist devices (VAD) did not restore MLP levels but decreased them in the majority of patients (Fig. 1).
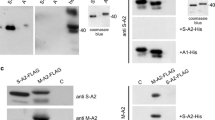
Expression of MLP in myocardial samples of DCM patients. A MLP and alpha-actinin were detected by Western blotting in eight samples of non-failing hearts (NF) and 19 samples of end-stage DCM hearts, ten without (DCM) and nine with left ventricular assist device support (VAD), respectively, for experimental details see Gehmlich et al. 2007; B Quantification of MLP in myocardial samples: both MLP and alpha-actinin levels were determined by densitometry (Biorad QuantityOne software) and expression levels of alpha-actinin were used to normalise MLP contents of the samples. The box plots visualise minimum, maximum, quartiles and mean left of the individual samples. MLP is significantly reduced in samples with left ventricular assist device support, compared to both non-failing and DCM hearts without mechanical support, (NF vs DCM: p = 0.19)
Subsequently, mutations in the human CSRP3 gene coding for MLP were associated with cardiomyopathies: A MLP W4R missense mutation was reported in some DCM patients (Knöll et al. 2002), and several missense mutations (MLP L44P, S46R, S54R/E55G, C58G) were described in patients suffering from HCM (Geier et al. 2003). A remarkable feature of HCM caused by these MLP mutations—and of HCM in general—is the mostly delayed onset of disease. This indicates that altered homeostasis of MLP levels or functions may precede the eventual development of structural changes seen in overt HCM (Gehmlich et al. 2004; Geier et al. 2008).
Linkage analysis in large German families has dispelled any doubt about the causative role of MLP in HCM (Geier et al. 2008), however the association of the MLP W4R mutation with human disease has remained less clear: the mutation was reported in both DCM and HCM patients (Bos et al. 2006). The low penetrance in affected families (Geier et al. 2008; Knöll et al. 2002), and the occurrence of the mutation in healthy controls (Bos et al. 2006; Geier et al. 2008) suggest that the MLP W4R mutation should rather be understood as a polymorphism, possibly with (negative) disease modifying potential.
MLP localisation and binding partners
To understand the mechanism how MLP mutations cause cardiac disease in humans, functional differences between wild-type protein and the mutated versions have to be established. A prerequisite for such comparative studies is to identify the functions of wild-type MLP in the healthy heart. Both, localisation data and information about binding partners may provide means to elucidate particular MLP functions.
MLP was demonstrated to bind to a variety of muscle-specific proteins: It interacts with the Z-disc proteins telethonin (also called T-Cap, Knöll et al. 2002) and alpha-actinin (Gehmlich et al. 2004). It binds to N-RAP at the intercalated disc (Ehler et al. 2001), to beta-spectrin at the costameres (Flick and Konieczny 2000), and to MyoD and other transcription factors in the nucleus (Kong et al. 1997). On top of that, numerous additional binding partners have been identified in in vitro or yeast two hybrid experiments: MLP can interact with itself (Zolk et al. 2000), with other LIM domain proteins such as zyxin (Louis et al. 1997), with kinases, e.g., integrin linked kinase (Postel et al. 2008) and with other enzymes such as histone deacetylases (Gupta et al. 2008). Whether all of these protein-protein interactions are functionally relevant in vivo remains to be proven: it is known that LIM domain proteins such as MLP may exhibit a certain degree of low affinity, unspecific binding in the test tube. Therefore, the proof of co-localisation of MLP and its ligands is the first evidence of the functional relevance of an interaction.
Unfortunately, the data about MLP localisation are not as straightforward as one would expect. MLP has been reported to be either a Z-disc protein (Knöll et al. 2002), or a doublet flanking the Z-disc (Arber et al. 1997). Beyond these sarcomeric localisations, both nuclear and costameric localisations were suggested (Boateng et al. 2007; Flick and Konieczny 2000). The use of polyclonal rabbit anti-sera in these studies may explain such contradicting findings, since these antibodies have the risk of additional unspecific reactivity, especially if they are not thoroughly characterised. A newly generated mouse monoclonal antibody 79D2 against MLP may have overcome this problem (Geier et al. 2008): This antibody is specific for MLP, and does not cross-react with closely related cysteine rich proteins CRP1 or CRP2. Careful characterisation has shown that this antibody recognises the human and rodent MLP in Western blot, immuno-precipitation and immuno-fluorescence applications, and antigen-blocking assays have proven the specificity of the observed signals. Surprisingly, this antibody indicates that MLP is a diffuse cytoplasmic protein with very little preference for sarcomeric structures in the myocardium. Its loose association with the sarcomere has been underlined by extraction experiments. MLP is easily extracted from cardiomyocytes by mild detergent treatment, i.e. under conditions which do not allow the extraction of sarcomeric proteins (Geier et al. 2008). Additional support for a mostly diffuse cytoplasmic localisation of MLP comes from transfection studies. When a fusion protein of MLP with a small immuno-tag, such as hemagglutinin (HA), is expressed in neonatal rat cardiomyoctes, it shows a diffuse cytoplasmic localisation (Fig. 2—top panel). Occasionally observed Z-disc localisation of MLP fused with green fluorescent protein (GFP) may thus have to be attributed to the GFP portion (Fig. 2—bottom panel).
Transfection of different MLP constructs into neonatal rat cardiomyocytes. Cells were transfected using HA-tagged MLP protein (top row, a kind gift of Stephan Lange) or MLP-GFP fusion protein (bottom row). 48 h post transfection cells were labelled for the HA-tagged construct (top row first column), or GFP (bottom row first column), a sarcomeric marker (second column: Z-disk alpha-actinin in top row, M-band myomesin in bottom row). Merged pictures: MLP green, sarcomeric marker red; scale bar represents 10 μm. Note that transfected HA-labelled MLP protein is not confined to sarcomeric structures, but rather diffusely distributed throughout cytoplasm, in this particular case also excluding the nucleus. In comparison, MLP-GFP fusion protein shows more preference for Z-disk localisation, however this has to be attributed to the GFP portion of the fusion construct (“GFP stickiness”)
Redefining the role of MLP in mechano-signalling
In 2002, the concept of a cardiac stretch sensor emerged (Knöll et al. 2002). Based on the observation that MLP−/− cardiomyocytes are defective in brain natriuretic peptide (BNP) induction upon mechanical stimulation, it was suggested that MLP is part of a cardiac stretch sensor together with titin and telethonin. This concept was further supported by several expression studies in striated muscle systems, where changes in mechanical load are often associated with changes of MLP expression.
However, the new data on the subcellular localisation of MLP (Geier et al. 2008) demand a critical revision of this concept: Whereas the giant titin protein spanning from Z-disk to M-band is an ideal candidate molecule to sense mechanical load, it is unclear how a cytoplasmic, not sarcomeric-anchored protein like MLP could possibly be involved in sensing mechanical signals. We suggest, therefore, that MLP should rather be understood as a downstream signal transducer in mechano-signalling cascades. Several lines of evidence indicate that the NFAT-calcineurin pathway is activated down-stream of mechanical stimulation (Heineke et al. 2005), and MLP’s interaction with PICOT might be involved in the control of this cascade (Jeong et al. 2008). However, how exactly MLP transduces signals in response to mechanical stress remains to be explored. One hypothesis is that post-translational modifications of MLP may serve as a molecular switch, among them protein phosphorylation or lysine acetylation (Gupta et al. 2008). These might alter binding affinities for ligands, and hence even lead to temporary chances in protein localisation, e.g., from cytoplasmic to nuclear predominance (Boateng et al. 2007).
Future research will have to address the molecular basis of how healthy cardiomyocytes sense and respond to mechanical stress. The analysis of disease situations, such as hypertrophic cardiomyopathy, may help us explore signalling pathways/molecules involved in this process. As access to human patient material is limited, animal models will be useful tools to dissect mechano-signalling pathways in the heart.
References
Arber S, Halder G, Caroni P (1994) Muscle LIM protein, a novel essential regulator of myogenesis, promotes myogenic differentiation. Cell 79:221–231. doi:10.1016/0092-8674(94)90192-9
Arber S, Hunter JJ, Ross J Jr, Hongo M, Sansig G, Borg J, Perriard JC, Chien KR, Caroni P (1997) MLP-deficient mice exhibit a disruption of cardiac cytoarchitectural organization, dilated cardiomyopathy, and heart failure. Cell 88:393–403. doi:10.1016/S0092-8674(00)81878-4
Boateng SY, Belin RJ, Geenen DL, Margulies KB, Martin JL, Hoshijima M, de Tombe PP, Russell B (2007) Cardiac dysfunction and heart failure are associated with abnormalities in the subcellular distribution and amounts of oligomeric muscle LIM protein. Am J Physiol Heart Circ Physiol 292:H259–H269. doi:10.1152/ajpheart.00766.2006
Bos JM, Poley RN, Ny M, Tester DJ, Xu X, Vatta M, Towbin JA, Gersh BJ, Ommen SR, Ackerman MJ (2006) Genotype-phenotype relationships involving hypertrophic cardiomyopathy-associated mutations in titin, muscle LIM protein, and telethonin. Mol Genet Metab 88:78–85. doi:10.1016/j.ymgme.2005.10.008
Ehler E, Horowits R, Zuppinger C, Price RL, Perriard E, Leu M, Caroni P, Sussman M, Eppenberger HM, Perriard JC (2001) Alterations at the intercalated disk associated with the absence of muscle LIM protein. J Cell Biol 153:763–772. doi:10.1083/jcb.153.4.763
Flick MJ, Konieczny SF (2000) The muscle regulatory and structural protein MLP is a cytoskeletal binding partner of betaI-spectrin. J Cell Sci 113:1553–1564
Gehmlich K, Geier C, Osterziel KJ, van der Ven PF, Fürst DO (2004) Decreased interactions of mutant muscle LIM protein (MLP) with N-RAP and alpha-actinin and their implication for hypertrophic cardiomyopathy. Cell Tissue Res 317:129–136. doi:10.1007/s00441-004-0873-y
Gehmlich K, Pinotsis N, Hayess K, van der Ven PF, Milting H, El BA, Korfer R, Wilmanns M, Ehler E, Fürst DO (2007) Paxillin and ponsin interact in nascent costameres of muscle cells. J Mol Biol 369:665–682. doi:10.1016/j.jmb.2007.03.050
Geier C, Perrot A, Ozcelik C, Binner P, Counsell D, Hoffmann K, Pilz B, Martiniak Y, Gehmlich K, van der Ven PF, Fürst DO, Vornwald A, Von HE, Nurnberg P, Scheffold T, Dietz T, Osterziel KJ (2003) Mutations in the human muscle LIM protein gene in families with hypertrophic cardiomyopathy. Circulation 107:1390–1395. doi:10.1161/01.CIR.0000056522.82563.5F
Geier C, Gehmlich K, Ehler E, Hassfeld S, Perrot A, Hayess K, Cardim N, Wenzel K, Erdmann B, Krackhardt F, Posch MG, Bublak A, Nagele H, Scheffold T, Dietz R, Chien KR, Spuler S, Fürst DO, Nurnberg P, Ozcelik C (2008) Beyond the sarcomere: CSRP3 mutations cause hypertrophic cardiomyopathy. Hum Mol Genet 17:2753–2765. doi:10.1093/hmg/ddn160
Gupta MP, Samant SA, Smith SH, Shroff SG (2008) HDAC4 and PCAF bind to cardiac sarcomeres and play a role in regulating myofilament contractile activity. J Biol Chem 283:10135–10146. doi:10.1074/jbc.M710277200
Heineke J, Ruetten H, Willenbockel C, Gross SC, Naguib M, Schaefer A, Kempf T, Hilfiker-Kleiner D, Caroni P, Kraft T, Kaiser RA, Molkentin JD, Drexler H, Wollert KC (2005) Attenuation of cardiac remodeling after myocardial infarction by muscle LIM protein-calcineurin signaling at the sarcomeric Z-disc. Proc Natl Acad Sci USA 102:1655–1660. doi:10.1073/pnas.0405488102
Jeong D, Kim JM, Cha H, Oh JG, Park J, Yun SH, Ju ES, Jeon ES, Hajjar RJ, Park WJ (2008) PICOT attenuates cardiac hypertrophy by disrupting calcineurin-NFAT signaling. Circ Res 102:711–719. doi:10.1161/CIRCRESAHA.107.165985
Knöll R, Hoshijima M, Hoffman HM, Person V, Lorenzen-Schmidt I, Bang ML, Hayashi T, Shiga N, Yasukawa H, Schaper W, McKenna W, Yokoyama M, Schork NJ, Omens JH, McCulloch AD, Kimura A, Gregorio CC, Poller W, Schaper J, Schultheiss HP, Chien KR (2002) The cardiac mechanical stretch sensor machinery involves a Z disc complex that is defective in a subset of human dilated cardiomyopathy. Cell 111:943–955. doi:10.1016/S0092-8674(02)01226-6
Kong Y, Flick MJ, Kudla AJ, Konieczny SF (1997) Muscle LIM protein promotes myogenesis by enhancing the activity of MyoD. Mol Cell Biol 17:4750–4760
Louis HA, Pino JD, Schmeichel KL, Pomies P, Beckerle MC (1997) Comparison of three members of the cysteine-rich protein family reveals functional conservation and divergent patterns of gene expression. J Biol Chem 272:27484–27491. doi:10.1074/jbc.272.43.27484
Postel R, Vakeel P, Topczewski J, Knöll R, Bakkers J (2008) Zebrafish integrin-linked kinase is required in skeletal muscles for strengthening the integrin-ECM adhesion complex. Dev Biol 318:92–101. doi:10.1016/j.ydbio.2008.03.024
Zolk O, Caroni P, Bohm M (2000) Decreased expression of the cardiac LIM domain protein MLP in chronic human heart failure. Circulation 101:2674–2677
Author information
Authors and Affiliations
Corresponding author
Additional information
Katja Gehmlich and Christian Geier contributed equally to this manuscript.
Rights and permissions
About this article
Cite this article
Gehmlich, K., Geier, C., Milting, H. et al. Back to square one: what do we know about the functions of Muscle LIM Protein in the heart?. J Muscle Res Cell Motil 29, 155–158 (2008). https://doi.org/10.1007/s10974-008-9159-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10974-008-9159-4