Abstract.
Previous work has shown that mutations in muscle LIM protein (MLP) can cause hypertrophic cardiomyopathy (HCM). In order to gain an insight into the molecular basis of the disease phenotype, we analysed the binding characteristics of wild-type MLP and of the (C58G) mutant MLP that causes hypertrophic cardiomyopathy. We show that MLP can form a ternary complex with two of its previously documented myofibrillar ligand proteins, N-RAP and α-actinin, which indicates the presence of distinct, non-overlapping binding sites. Our data also show that, in comparison to wild-type MLP, the capacity of the mutated MLP protein to bind both N-RAP and α-actinin is significantly decreased. In addition, this single point mutation prevents zinc coordination and proper folding of the second zinc-finger in the first LIM domain, which consequently renders the protein less stable and more susceptible to proteolysis. The molecular basis for HCM-causing mutations in the MLP gene might therefore be an alteration in the equilibrium of interactions of the ternary complex MLP–N-RAP–α-actinin. This assumption is supported by the previous observation that in the pathological situation accompanied by MLP down regulation, cardiomyocytes try to compensate for the decreased stability of MLP protein by increasing the expression of its ligand N-RAP, which might finally result in the development of myocyte disarray that is characteristic of this disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The precise propagation of signalling cascades and the establishment of supramolecular assemblies like cell contacts depend to a large degree on the activity of adapter proteins. The function of these versatile proteins is based on the seemingly unlimited combinatorics of a great variety of protein-binding domains. One such domain type is the so-called LIM domain, named after the three transcriptional factors, lin11, isl-1 and mec-3, in which it was first identified (Freyd et al. 1990; Karlsson et al. 1990; Way and Chalfie 1988). LIM domains are characterized by the cysteine-rich consensus sequence CX2CX16–23HX2CX2CX2CX16–21CX2(CHD) (Freyd et al. 1990; Sadler et al. 1992), of which the conserved cysteine, histidine and aspartic acid residues coordinate two zinc ions (Archer et al. 1994). Consequently, each LIM domain folds into two zinc fingers. LIM domains that do not function as transcription factors are classified into two subgroups: (a) so-called “LIM only proteins”, in which two or more of these protein-binding interfaces are arranged in tandem (Dawid et al. 1998); (b) alternatively, LIM domains occur in combination with other functional domains which in turn may also act as protein-binding motifs (Dawid et al. 1998).
Cross-striated muscle cells contain several LIM domain proteins, for instance, MLP (also called cysteine-rich protein 3, CRP3; Arber et al. 1994), N-RAP (Luo et al. 1997), FHL (also called SLIM; Morgan and Madgwick 1996) and ALP (Xia et al. 1997). Probably the most versatile member of this protein family is MLP: it was detected at the cell membrane (Flick and Koniecny 2000), in Z-discs (Arber et al. 1994, 1997) as well as in the nucleus (Arber et al. 1994; Kong et al. 1997). In concert with these multiple distinct localizations, several binding partners have been identified to date: α-actinin (Louis et al. 1997), β-spectrin (Flick and Koniecny 2000), MyoD (Kong et al. 1997), N-RAP (Ehler et al. 2001) and telethonin (Knöll et al. 2002). Ablation of the MLP gene in mice leads either to a myocardial hypertrophy reaction with increased heart weight and subsequent rapid progressive congestive heart failure or to a dilated cardiomyopathy phenotype (Arber et al. 1997). Thus, the phenotype of MLP−/− mice presents characteristics of both hypertrophic (HCM) and dilated (DCM) cardiomyopathy in humans. Recently a mutation in the human MLP gene was reported to be associated with DCM (Knöll et al. 2002), while three further MLP mutations were associated with HCM (Geier et al. 2003).
HCM is among the most frequent inherited cardiac diseases and it is the most commonly identified cause of sudden death in adolescents and young adults (Maron et al. 1995). The predominant clinical feature is myocardial hypertrophy—most frequently at the interventricular septum—without preceding pressure overload. Structurally, this hypertrophy is typically accompanied by fibrosis and a disorganized arrangement of myocytes, giving rise to myocardial disarray (McKenna et al. 1997). Consequently, electrical conduction in the heart may be disturbed, leading to arrhythmias. HCM is an autosomal dominant disease caused by mutations in several genes, most of them encoding myofibrillar proteins. Depending on the precise nature of the mutation and on mainly unidentified modifiers, the clinical manifestation of HCM may vary considerably (Marian and Roberts 2001; Seidman and Seidman 2001).
Dilated cardiomyopathy is one of the leading causes of heart failure and a primary cause of heart transplantation in the young. About 25–30% of all DCM cases are of familial etiology. DCM can be transmitted as autosomal, X-linked or mitochondrial traits, with autosomal dominant inheritance being the most frequently observed. Several disease genes encoding cytoskeletal, sarcomeric or nuclear membrane proteins have been identified so far (Seidman and Seidman 2001).
To better understand the molecular pathway underlying the HCM or DCM phenotype, we investigated whether a mutation in MLP alters its binding capabilities to two of its ligands, α-actinin and N-RAP. The latter appeared particularly interesting, because its expression was shown to be upregulated in two mouse models for DCM, the MLP knock-out mouse (Ehler et al. 2001) and tropomodulin-overexpressing transgenic (TOT) mice (Ehler et al. 2001; Sussman et al. 1998).
The results of our binding studies provide direct evidence for an alteration of the α-actinin- and N-RAP-binding characteristics of a mutant MLP protein that may help to understand the molecular pathway that leads to the dramatic pathologic changes in HCM and DCM hearts.
Materials and methods
RT-PCR for N-RAP cDNA
Total RNA was extracted from non-differentiated and differentiated cultured human skeletal muscle cells using an RNA isolation kit (Stratagene, Heidelberg, Germany) as previously described (Schröder et al. 2000). Specific fragments of the N-RAP cDNAs were obtained by reverse transcriptase-polymerase chain reaction (RT-PCR) using this total RNA purified from differentiated cultured human skeletal muscle cells as a template using the Expand reverse transcriptase and the “Expand long template PCR system” according to the manufacturer’s instructions (Roche Diagnostics, Mannheim, Germany). The position of the amplified fragments is shown in Fig. 1and primer sequences are given in Table 1. Some portions of the N-RAP cDNA were also amplified by PCR (Saiki et al. 1985) using a human skeletal muscle cDNA library (BD Biosciences Clontech, Heidelberg, Germany) as a template.
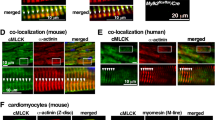
Expression constructs used for protein-protein binding assays. a Schematic representation of the expression constructs used in this study. Full-length N-RAP is also given for comparison, witharrows indicating alternatively spliced exons. Amino-terminal N-RAP constructs as well as full-length α-actinin 2 were cloned into pET23aW2 (constructs therefore carry an amino-terminal T7 tag), whereas wild-type and mutant MLP were cloned into the vector pET23aW1 (recombinant proteins therefore carry a carboxy-terminal EEF tag). Theasterisk indicates the site of the mutation.b Immunodetection of the recombinant proteins described above after SDS-PAGE and Western blotting. Lanes were loaded with the following samples: 1 N-RAP LIr02, 2 N-RAP LI, 3α-actinin 2, 4 MLP WT, 5 MLP C58G mutant. Lanes 1–3 detection by T7 tag; lanes 4and 5 detection via EEF tag. Numbers display the molecular masses (×1,000) of marker proteins at the respective positions.Asterisk indicates proteolysis products derived from the mutant protein C58G MLP
Design of protein expression constructs
The complete human α-actinin 2 and MLP cDNA sequences were amplified by PCR using a human skeletal muscle cDNA library (BD Biosciences Clontech) as a template. Mutagenesis to obtain the C58G mutation in MLP has been described previously (Geier et al. 2003).
These three constructs and the two amino-terminal fragments N-RAP LI and N-RAP LIr02 (see Fig. 1) were cloned either into the prokaryotic expression vector pET23a-EEF, resulting in fusion proteins carrying a carboxy-terminal His6-tag and an EEF-immunotag (Obermann et al. 1996), or into pET23a-T7 (amino-terminal T7-tag and carboxy-terminal His6-tag; Obermann et al. 1998). Plasmids were transformed into E. coli JM109 (Stratagene) and purified using standard protocols (Ausubel et al. 1987). Plasmid integrity was verified by DNA sequencing (Sequence Laboratories, Göttingen, Germany, or Agowa, Berlin, Germany).
Protein expression and purification
Protein expression in E. coli BL21(DE3)pLysS (Novagene, Heidelberg, Germany) was performed as previously described (Obermann et al. 1996, 1997, 1998). Cells were grown in SOC medium supplemented with 100 mg/l carbenicillin and 34 mg/l chloramphenicol, and protein expression was induced at OD600=1 by addition of 0.5 mM IPTG. ZnCl2 was added to 0.2 mM to ensure proper folding of LIM domains. After 3 h at 30°C, cells were harvested and stored at −20°C until further purification. For the expression of α-actininE. coli BL21(DE3)Codon Plus cells (Strategene) were used, following the same protocol, except that ZnCl2 was omitted from the culture medium.
Purification of His-tagged protein was performed using the QiaExpressionist kit (Qiagen, Hilden, Germany) as described by the manufacturer. Briefly, bacterial cells were lysed by the addition of lysis buffer (50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 10 mM imidazole, 1 mg/ml lysozyme, 5 µM E64, 1 µg/ml leupeptin, 10 µg/ml trypsin inhibitor, 0.25 µg/ml pepstatin A), DNA was fragmented by sonification and cell debris sedimented by centrifugation (18,000×g, 30 min, 4°C). The supernatant containing the soluble protein fraction was incubated with Ni2+-NTA agarose beads under constant agitation at 4°C for 1 h. After filling the beads into a column, they were washed twice with washing buffer (50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 20 mM imidazole) and bound protein was eluted with elution buffer (50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 250 mM imidazole, 5 µM E64, 1 µg/ml leupeptin, 10 µg/ml trypsin inhibitor, 0.25 µg/ml pepstatin A). Protein solutions were subsequently stored on ice.
Protein concentrations were determined using absorption measurements in 6 M guanidine hydrochloride as previously described (Pace et al. 1995) using the following molar absorption coefficients: NRAP LI, ε=17,510 l/mol/cm; NRAP LIr02, ε=35,430 l/mol/cm; MLP wild type (WT), ε=23,650 l/mol/cm; MLP C58G, ε=23,525 l/mol/cm and α-actinin 2, ε=125,620 l/mol/cm.
Antibodies
Since recombinant protein constructs were tagged either amino-terminally with a peptide of the T7 major capsid protein (“T7 tag”) or carboxy-terminally with the amino acids EEF, respective fusion proteins could be detected using either a T7 tag murine monoclonal antibody (mAb; Novagen, Heidelberg, Germany) or the rat mAb YL1/2 (a kind gift from Dr. J. Wehland, Braunschweig, Germany). Alternatively, α-actinin was detected with mAb EA53 (Sigma, Taufkirchen, Germany).
For immunoprecipitations the rabbit polyclonal antiserum 653 against α-actinin (Van der Ven et al. 2000) was used.
Zinc-binding experiments
Defined amounts of purified recombinant protein were lyophilized and digested in 0.25 ml 65% (w/v) nitric acid (Suprapur, Merck, Darmstadt, Germany) at room temperature overnight. After digestion the volume was adjusted to 10 ml with water, and zinc concentrations were determined by inductively coupled plasma optical emission spectroscopy using an IRIS Advantage Duo ER/S (Thermo Jarrell Ash, Franklin, MA, USA).
Dot blot protein-binding assays
Recombinant proteins were spotted on nitrocellulose membranes (BA-85, Schleicher and Schüll, Dassel, Germany). After air drying, the strips were blocked with 4% (w/v) low fat milk powder in TBST (TRIS-buffered saline with Tween-20). Individual strips were overlaid with the respective protein in blocking solution for 90 min. After three washes with TBST, bound protein was immunodetected with antibodies specific for the respective immunotag (see below).
Immunoprecipitation
For co-immunoprecipitation experiments, a mixture of 0.2 nmol of each recombinant protein was diluted in IP buffer (phosphate-buffered saline, PBS, with 0.05% Triton X-100, 1% BSA, 5 µM E64, 1 µg/ml leupeptin, 10 µg/ml trypsin inhibitor, 0.25 µg/ml pepstatin A). After the addition of 5 µl polyclonal α-actinin antibody 653, the mixture was incubated at 4°C for 60 min with gentle shaking. Subsequently, 40 µl Dynabeads protein G (Dynal Biotech, Hamburg, Germany) was added, and the mixture was further incubated for 30 min with gentle shaking. The beads were washed three times with IP buffer without BSA. Subsequently, beads were boiled in sodium dodecyl sulphate (SDS) sample buffer, and bead-associated proteins were separated by SDS-polyacrylamide gel electrophoresis (PAGE), blotted onto nitrocellulose membranes and immunodetected (see below).
Competition experiments were performed under identical conditions, except that varying amounts of wild-type MLP protein (40 pmol to 1 nmol) were applied.
SDS-PAGE and Western blotting
SDS-PAGE was performed as previously described (Laemmli 1970) using 10, 12 or 14% acrylamide gels.
Separated proteins were blotted onto nitrocellulose membranes (Roth, Karlsruhe, Germany) overnight using a tank blot buffer system (25 mM TRIS base, 18% (v/v) methanol, 192 mM glycine, 0.01% SDS). Transfer of the protein was confirmed by staining the membranes with Ponceau red dye.
Immunodetection
Nitrocellulose membranes were blocked with 4% (w/v) low fat milk powder in TBST. Primary antibodies were diluted in blocking solution, secondary antibodies in TBST and used in the following combinations: anti-T7 antibody and anti-α-actinin EA53 antibody followed by peroxidase-conjugated goat anti-mouse secondary antibody (Jackson Immuno Research Laboratories, West Grove, PA, USA); anti-EEF antibody YL1/2 and peroxidase-conjugated goat anti-rat secondary antibody (Jackson Immuno Research Laboratories). Conjugated enzymes were detected by enhanced chemiluminescence using “SuperSignal West Pico Chemiluminescent Substrate” (Pierce, Rockford, IL, USA) and Kodak XAR-351 film.
Results
Characterization of the human N-RAP gene structure and splice variants
Because of the involvement of MLP in human cardiac disease, and the observation that a ligand of MLP, the intercalated disc component N-RAP, is abnormally expressed in myopathic hearts, we aimed to characterize the interaction of human N-RAP with a recently identified HCM-causing mutant MLP (Geier et al. 2003). Our attempts to amplify and clone specific portions of N-RAP were initially hampered by discrepancies in database entries that existed at that time. This necessitated establishing the human N-RAP cDNA sequence independently. As a result, we noticed the molecular mass to be 197 kDa instead of the previously reported 133 kDa (Luo et al. 1997). In addition, we identified two alternatively spliced exons/repeats: (1) exon 12/repeat 8, and (2) exon 39/repeat 41 (see arrows in Fig. 1). During revision of this work the complete human N-RAP cDNA sequence and gene structure were described (Mohiddin et al. 2003). While the first of our splice variants was also reported in this paper, the second one is novel.
Protein chemical characterization of wild-type and mutant MLP
All proteins and defined subfragments used in this study were expressed as recombinant proteins containing His6and T7 or EEF tags (Fig. 1). Interestingly, the C58G mutant of MLP was consistently more prone to proteolytic degradation already in expression cultures, which implies that this mutation results in a less stable ternary structure of the recombinant protein. Since the cysteine residue in question (Cys58) is one of the four zinc-coordinating amino acids in the second zinc finger of the first LIM domain of N-RAP, it was logical to assume that MLP C58G might bind one zinc ion less efficiently. This was tested by comparing the amount of Zn2+ bound to wild-type MLP versus MLP C58G using atomic emission spectroscopy. Interestingly, the mutant protein was only capable of binding 75% of the amount of Zn2+ bound to the wild-type protein (see Fig. 2), confirming our hypothesis that the mutation abolished one of the four Zn2+-binding sites. The observations described above are strong indications for an abnormal and less stable folding of the second zinc finger of the first LIM domain.
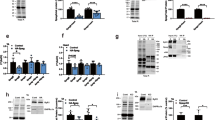
Comparison of zinc content of wild-type and mutant (C58G) MLP proteins. Protein samples were digested by nitric acid and zinc concentrations were determined by atomic emission spectroscopy as described in “Materials and methods”. The number of coordinated zinc atoms per protein molecule is expressed in arbitrary units, and the wild-type content was set to 100% (*p<0.01). Note that mutant MLP (C58G) shows a significant reduction of its zinc-binding ability compared to the wild-type MLP protein (WT)
Protein-protein interaction studies with MLP
At the heart of our study was an investigation of the binding properties of MLP to its protein ligands. MLP has been reported to bind to both N-RAP and α-actinin (Louis et al. 1997; Ehler et al. 2001). Since recent studies have shown mutations in the human MLP gene are associated with cardiomyopathies (Knöll et al. 2002; Geier et al. 2003), we speculated that differences in the binding capacity of the mutant MLP protein to its ligands might be involved in the pathogenesis of these diseases. We therefore expressed and purified MLP (both wild-type and the C58G mutant that is associated with HCM), α-actinin and two amino-terminal subfragments of N-RAP: first, the LIM domain plus the subsequent linker sequence (N-RAP LI) and, secondly, a larger polypeptide containing in addition the first two nebulin repeats (N-RAP LIr02; see Fig. 1). Subsequently their capacity for mutual association was analysed using dot blot overlay assays. These results confirmed the previously described strong and specific binding of wild-type MLP to both N-RAP and α-actinin (Fig. 3). Thus, both N-RAP fragments and α-actinin 2 bound to spotted MLP. Likewise, α-actinin 2 bound to both spotted N-RAP constructs (Fig. 3). Interestingly, the N-RAP construct comprising solely the non-modular region (N-RAP LI) bound considerably more weakly to both MLP and α-actinin than N-RAP LIr02, the construct that additionally contained the first two nebulin repeats. Since nebulin repeats were shown to bind to F-actin (Luo et al. 1997), we assume that these differences in binding strength are due to more stable folding of the construct N-RAP LIr02.
Binding properties of purified recombinant proteins in dot blot overlay experiments. aBinding of N-RAP LI, N-RAP LIr02 and α-actinin 2 to immobilized wild-type MLP (MLP WT, 5 pmol) as detected via the T7 immunotag of the bound protein. In addition, the binding of α-actinin to both N-RAP LI and N-RAP LIr02 (5 pmol each) is shown. Bound α-actinin 2 was in this case detected by mAb EA53. b Binding of amino-terminal N-RAP constructs to wild-type and mutant MLP. Either wild-type MLP (MLP WT), mutant MLP (MLP C58G) or BSA were immobilized and bound N-RAP LI or N-RAP LIr02 were detected via their T7 tag. Ponceau red dye staining visualizes spotted protein. Note that the MLP mutant C58G shows considerably weaker binding to N-RAP
Distinct binding properties of wild-type and mutant MLP
Subsequently, we compared the binding properties of wild-type MLP with the cardiomyopathy-associated C58G MLP mutant to N-RAP. Increasing amounts of wild-type and mutant MLP were spotted onto nitrocellulose and overlaid with either N-RAP LI or N-RAP LIr02. Under these conditions N-RAP LI exhibited an obvious, concentration-dependent binding to wild-type MLP, while binding of the construct N-RAP LIr02 was considerably stronger and already saturated at the lowest MLP concentration applied (Fig. 3). In contrast, the MLP mutant C58G behaved quite differently under the same conditions: only minimal binding to N-RAP LI was observed. The binding to N-RAP LIr02 was also highly reduced. While 10 pmol of mutant MLP resulted in a strong signal, 3 pmol was hardly detected (Fig. 3). This indicates that MLP C58G has a considerably reduced binding capacity to N-RAP and α-actinin.
These weaker interactions offer the first experimentally supported potential explanation for the pathomechanism of HCM as a result of MLP mutations at the molecular level.
Identification of a ternary complex between MLP–N-RAP–α-actinin
The work of other laboratories and our own data (see Fig. 3) have shown single binding activities: MLP plus α-actinin (Louis et al. 1997), MLP plus N-RAP (Ehler et al. 2002) and N-RAP plus α-actinin (Zhang et al. 2001; Lu et al. 2003). We have therefore addressed the question of whether these bindings might influence one another, be it in a negative, competitive manner, or by the formation of a ternary complex. To do so, a series of co-immunoprecipitation experiments were performed, the results of which are documented in Fig. 4. Recombinant α-actinin was mixed with wild-type or mutant MLP and/or N-RAP, and the formed protein complexes were precipitated by the α-actinin polyclonal antiserum. Co-precipitated, added MLP and N-RAP fragments were detected using tag-specific antibodies. Thus, both wild-type MLP and N-RAP LIr02 alone bound in large quantities, indicating strong and specific binding of both proteins to α-actinin (Fig. 4, lanes 2, 6). The mutant MLP C58G was revealed in significantly lower amounts, indicating, in accordance with the dot blot data (Fig. 3), a reduced binding strength to α-actinin (Fig. 4, lane 3). In mixtures of all three proteins, both MLP and N-RAP were readily co-precipitated with α-actinin (Fig. 4, lanes 4, 5). Since this indicated a cooperative effect, we performed a series of coIPs in which constant amounts of α-actinin and N-RAPLIr02 were mixed with increasing quantities of MLP. In these experiments, the amount of bound N-RAP fragment was independent of the amount of MLP added over a wide range of concentrations even beyond saturation levels (Fig. 4, lanes 7–11). Both results indicated the formation of a true ternary complex rather than competitive binding.
Co-immunoprecipitation of MLP and N-RAP with α-actinin. a Wild-type (WT) as well as mutant (C58G) MLP co-precipitated with α-actinin either in the presence or in the absence of N-RAP LIr02. In addition, this N-RAP construct also co-immunoprecipitated with α-actinin in the absence of wild-type or mutant MLP protein. b Competition experiments. Increasing amounts of wild-type MLP (40 pmol, 0.1 nmol, 0.2 nmol, 0.4 nmol, 1 nmol) did not interfere with the co-immunoprecipitation of N-RAP LIr02 (N-RAP) along with α-actinin
Discussion
LIM domain-containing proteins have variously been described to act as adaptors in signal transduction processes. In cross-striated muscle two such proteins have been studied in greater detail: MLP and N-RAP. MLP was suggested to be involved in myofibril development and, initially based on the phenotype of the MLP knockout mouse, in cardiac disease (Arber et al. 1997). Recently, ten patients with dilated cardiomyopathy carrying a single missense mutation in the MLP gene were identified (Knöll et al. 2002). Our own recent data showed co-segregation of three different MLP mutations in three families with hypertrophic cardiomyopathy (Geier et al. 2003). In addition, MLP was found to be downregulated in chronic human heart failure (Zolk et al. 2000). Taken together, these studies suggest that MLP may mainly function as a key component of signalling pathways involved in both cardiomyocyte growth and stress response. Biophysical experiments also support the hypothesis that MLP could be part of the cardiac mechanosensor machinery (Knöll et al. 2002).
Where and how precisely MLP exerts its activities has, however, remained largely enigmatic. Since this protein was detected both in the cytoplasm and in the nucleus, MLP may be considered a typical dual compartment protein, but the regulatory events controlling its localization at a specific time point are still unknown. In the nucleus of skeletal muscle cells MLP binds to and controls the activity of the myogenic transcription factor MyoD (Kong et al. 1997). In the cytoplasm of skeletal and cardiac muscle cells molecular interactions were demonstrated with α-actinin (Louis et al. 1997), β-spectrin (Flick and Koniecny 2000), N-RAP (Ehler et al. 2001) and telethonin (Knöll et al. 2002), and the protein was localized either to Z-discs (Arber et al. 1994; Kong et al. 1997) or to subsarcolemmal regions (Flick and Koniecny 2000). It is conceivable that this discrepancy reflects adaptive, functional distinctions in particular signalling processes rather than real discrepancies in experimental implementations.
In this study we have aimed to elucidate the interactions of both wild-type and mutant MLP with N-RAP and α-actinin. These interactions are of immediate significance for the myofibrillar apparatus. However, during our efforts to design defined subfragments of N-RAP for protein expression and binding studies, we noticed conflicting sequence database entries which necessitated establishing the N-RAP sequence independently. The resulting sequence turned out to be identical to a sequence published shortly after the initial submission of this work (Mohiddin et al. 2003). Thus, the carboxy-terminal 95% of N-RAP is strictly organized into 46 nebulin-like repeats (Labeit and Kolmerer 1995), which in part form a superrepeat structure. In addition, we have identified a further, novel variant on top of the one described previously (see Fig. 1 and Mohiddin et al. 2003). We may therefore anticipate that even further N-RAP isoforms may be identified in the future.
In order to perform the intended protein-protein interaction studies, we used purified bacterially expressed purified proteins. Already during the purification steps we noticed the MLP mutant C58G behaving distinctly from the wild-type protein: The mutant protein was more prone to proteolysis (see Fig. 1) and was less stable in solution. Most likely this is due to the fact that the mutated cysteine is one of the four residues involved in complexing a zinc ion in the second zinc finger of the first LIM domain of MLP.
A similar situation was described in an earlier study, in which one of the zinc-coordinating cysteine residues in the second zinc finger of a zyxin LIM domain was mutated to alanine by site-directed mutagenesis (Schmeichel and Beckerle 1997). The resulting loss of ability to bind zinc within the affected second zinc finger led to the destabilization of both zinc fingers of this LIM domain, and not only the one carrying the mutation. We therefore found it important to compare the zinc-binding properties of wild-type and mutant MLP proteins. These experiments revealed that MLP C58G bound precisely one zinc ion less than wild-type MLP, i.e. three instead of four ions (see Fig. 2). This suggests that the mutant protein has greater problems folding properly into its native ternary structure at least in its first LIM domain, making it more susceptible to proteolysis.
The amino-terminal LIM domain of N-RAP has been shown to be dispensable for the association with MLP (Ehler et al. 2001). Based on a comparison of the construct applied in that study with the constructs used in our experiments, we conclude that the MLP-binding site in the N-RAP polypeptide must be confined to the ~60-amino-acid portion located between the LIM domain and the nebulin repeat region, because this is the only region that both constructs have in common.
Our binding data provide the first proof for the existence of a ternary complex between MLP, N-RAP and α-actinin. Since varying concentrations of MLP did not affect the binding of N-RAP to α-actinin, distinct binding sites are the only logical explanation. Therefore, HCM-causing mutations in the MLP gene may result in a disturbance of the homeostasis of the interactions of MLP with N-RAP and α-actinin. In this respect it is interesting to note that in MLP knock-out mice the expression of N-RAP was shown to be significantly increased (Ehler et al. 2001), while α-actinin is downregulated in samples of DCM patients (our unpublished results). This indicates that in the pathological situation (cardiac disease caused by MLP mutations) cardiac myocytes might try to counterbalance the instability of the mutant MLP protein and/or the MLP–N-RAP–α-actinin ternary complex by increasing the expression of the functionally important MLP ligand N-RAP. This unbalanced expression may be involved in triggering the development of myocyte disarray and hypertrophy. This scenario will need to be investigated in further studies including patient material.
References
Arber S, Halder G, Caroni P (1994) Muscle LIM protein, a novel essential regulator of myogenesis, promotes myogenic differentiation. Cell 79:221–231
Arber S, Hunter JJ, Ross Jr J, Hongo M, Sansig G, Borg J, Perriard JC, Chien KR, Caroni P (1997) MLP-deficient mice exhibit a disruption of cardiac cytoarchitectural organization, dilated cardiomyopathy, and heart failure. Cell 88:393–403
Archer VE, Breton J, Sanchez-Garcia I, Osada H, Forster A, Thomson AJ, Rabbitts TH (1994) Cysteine-rich LIM domains of LIM-homeodomain and LIM-only proteins contain zinc but not iron. Proc Natl Acad Sci U S A 91:316–320
Ausubel FM, Brent R, Kingston RE et al. (eds) (1987) Current protocols in molecular biology. John Wiley & Sons, New York
Dawid IB, Breen JJ, Toyama R (1998) LIM domains: multiple roles as adapters and functional modifiers in protein interactions. Trends Genet 14:156–162
Ehler E, Horowits R, Zuppinger C, Price RL, Perriard E, Leu M, Caroni P, Sussmann M, Eppenberger HM, Perriard JC (2001) Alterations at the intercalated disc associated with the absence of muscle LIM protein. J Cell Biol 153:763–772
Flick MJ, Koniecny SF (2000) The muscle regulatory and structural protein MLP is a cytoskeletal binding partner of beta I-spectrin. J Cell Sci 113:1553–1564
Freyd G, Kim SK, Horvitz HR (1990) Novel cysteine-rich motif and homeodomain in the product of the Caenorhabditis elegans cell lineage gene lin-11. Nature 344:876–879
Geier C, Perrot A, Özcelik C, Binner P, Counsell D, Hoffmann K, Pilz B, Martiniak Y, Gehmlich K, van der Ven PMF, Fürst DO, Vornwald A, von Hodenberg E, Nürnberg P, Scheffold T, Dietz R, Osterziel KJ (2003) Mutations in the human muscle LIM protein gene in families with hypertrophic cardiomyopathy. Circulation 107:1390–1395
Karlsson O, Thor S, Olsson H, Edlund T (1990) Insulin gene enhancer binding protein Isl-1 is a member of a novel class of proteins containing both a homeo- and a Cys-His domain. Nature 344:879–882
Knöll R, Hoshijima M, Hoffmann HM, Person V, Lorenzen-Schmidt I, Bang ML, Hayashi T, Shiga N, Yasukawa H, Schaper W, McKenna W, Yokoyama M, Schork NJ, Omens JH, McCulloch AD, Kimura A, Gregorio CC, Poller W, Schaper J, Schultheiss HP, Chien KR (2002) The cardiac mechanical stretch sensor machinery involves a Z disc complex that is defective in a subset of human dilated cardiomyopathy. Cell 111:943–955
Kong Y, Flick MJ, Kudla AJ, Konieczny SF (1997) Muscle LIM promotes myogenesis by enhancing the activity of MyoD. Mol Cell Biol 17:4750–4760
Labeit S, Kolmerer B (1995) The complete primary structure of human nebulin and its correlation to muscle structure. J Mol Biol 28:308–315
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Louis HA, Pino JD, Schmeichel KL, Pomies P, Beckerle MC (1997) Comparison of three members of the cysteine-rich protein family reveals functional conservation and divergent patterns of gene expression. J Biol Chem 272:27484–27491
Lu S, Carroll SL, Herrera AH, Ozanne B, Horowits R (2003) New N-RAP-binding partners alpha-actinin, filamin and Krp1 detected by yeast two-hybrid screening: implications for myofibril assembly. J Cell Sci 116:2169–2178
Luo G, Zhang JQ, Nguyen TP, Herrera AH, Patterson B, Horowits R (1997) Complete cDNA sequence and tissue localization of N-RAP, a novel nebulin-related protein of striated muscle. Cell Motil Cytoskel 38:75–90
Marian AJ, Roberts J (2001) The molecular genetic basis for hypertrophic cardiomyopathy. J Mol Cell Cardiol 33:655–670
Maron BJ, Gardin JM, Flack JM, Gidding SS, Kurosaki TT, Bild DE (1995) Prevalence of hypertrophic cardiomyopathy in a general population of young adults. Echocardiographic analysis of 4111 subjects in the CARDIA Study. Coronary Artery Risk Development in (Young) Adults. Circulation 92:785–789
McKenna WJ, Spirito P, Desnos M, Dubourg O, Komjada M (1997) Experience from clinical genetics in hypertrophic cardiomyopathy: proposal for new diagnostic criteria in adult members of affected families. Heart 77:130–132
Mohiddin SA, Lu S, Cardoso JP, Carroll S, Jha S, Horowits R, Fananapazir L (2003) Genomic organization, alternative splicing, and expression of human and mouse N-RAP, a nebulin-related LIM protein of striated muscle. Cell Motil Cytoskeleton 55:200–12
Morgan MJ, Madgwick AJ (1996) Slim defines a novel family of LIM-proteins expressed in skeletal muscle. Biochem Biophys Res Comm 14:632–638
Obermann WMJ, Gautel M, Steiner F, van der Ven PFM, Weber K, Fürst DO (1996) Molecular structure of the sarcomeric M band: localization of defined domains of myomesin, M-protein and the 250 kDa carboxyterminal region of titin. J Cell Biol 134:1441–1453
Obermann WMJ, Gautel M, Weber K, Fürst DO (1997) Molecular structure of the sarcomeric M band: identification of a novel PKA phosphorylation site and mapping of titin- and myosin-binding domains in myomesin. EMBO J 16:211–220
Obermann WMJ, van der Ven PFM, Steiner F, Weber K, Fürst DO (1998) Mapping of a myosin binding domain and a regulatory phosphorylation site in M-protein, a structural protein of the sarcomeric M band. Mol Biol Cell 9:829–840
Pace CN, Vajdos F, Fee L, Grimsley G, Gray T (1995) How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4:2411–2423
Sadler I, Crawford AW, Michelsen JW, Beckerle MC (1992) Zyxin and cCRP: two interactive LIM domain proteins associated with the cytoskeleton. J Cell Biol 119:1573–1587
Saiki RK, Scharf SJ, Faloona F, Mullis GT, Ehrlich HA (1985) Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230:1350–1354
Schmeichel KL, Beckerle MC (1997) Molecular dissection of a LIM domain. Mol Biol Cell 8:219–230
Schröder R, Fürst DO, Klasen C, Reimann J, Herrmann H, van der Ven PFM (2000) The association of plectin with Z-discs is a prerequisite for the formation of the intermyofibrillar desmin cytoskeleton. Lab Invest 80:455–464
Seidman JG, Seidman C (2001) The genetic basis for cardiomyopathy: from mutation identification to mechanistic paradigms. Cell 104:557–567
Sussman MA, Welch S, Cambon N, Klevitsky R, Hewett TE, Price R, Witt SA, Kimball TR (1998) Myofibril degeneration caused by tropomodulin overexpression leads to dilated cardiomyopathy in juvenile mice. J Clin Invest 101:51–61
Van der Ven PFM, Obermann WMJ, Lemke B, Gautel M, Weber K, Fürst DO (2000) Characterization of muscle filamin isoforms suggests a possible role of γ-filamin/ABP-L in sarcomeric Z-disc formation. Cell Motil Cytoskeleton 45:149–162
Way M, Chalfie M (1988) Mec-3, a homeobox-containing gene that specifies differentiation of the touch receptor neurons in C. elegans. Cell 54:5–16
Xia H, Winokur ST, Kuo WL, Altherr MR, Bredt DS (1997) Actinin-associated LIM proteins: identification of a domain interaction between PDZ and spectrin-like repeat motifs. J Cell Biol 139:507–515
Zhang JQ, Elzey B, Williams G, Lu S, Law DJ, Horowits R (2001) Ultrastructural and biochemical localization of N-RAP at the interface between myofibrils and intercalated disks in the mouse heart. Biochemistry 40:14898–14906
Zolk O, Caroni P, Böhm M (2000) Decreased expression of the cardiac LIM domain protein MLP in chronic human heart failure. Circulation 101:2672–2673
Acknowledgements.
We gratefully acknowledge the technical assistance of A. Guhlan and J. Kühnisch. The authors thank Dr. Jürgen Wehland (Braunschweig, Germany) for the donation of mAb YL1/2. We would also like to thank Ute Krämer and Astrid Schröder (Potsdam-Golm, Germany) for help with the zinc measurements.
Author information
Authors and Affiliations
Corresponding author
Additional information
This study was supported by a grant from the Deutsche Forschungsgemeinschaft to D.O.F.
Rights and permissions
About this article
Cite this article
Gehmlich, K., Geier, C., Osterziel, K.J. et al. Decreased interactions of mutant muscle LIM protein (MLP) with N-RAP and α-actinin and their implication for hypertrophic cardiomyopathy. Cell Tissue Res 317, 129–136 (2004). https://doi.org/10.1007/s00441-004-0873-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-004-0873-y