Abstract
The immune system is designed to protect the organism from infection and to repair damaged tissue. An effective response requires recognition of the threat, the appropriate effector mechanism to clear the pathogen and a return to homeostasis with minimal damage to self-tissues. T cells play a central role in orchestrating the immune response at all stages of the response and have been the subject of intense study by both experimental immunologists and modelers. This review examines some of the more critical questions in T cell biology and describes the latest attempts to address those questions using approaches that combine mathematical modeling and experiments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The main function of the immune system is to protect the organism from infection with pathogens. This is achieved by an elaborate network of cells and secreted molecules that together coordinate a successful immune response. Three main stages of an immune response can be characterized; (1) recognition i.e. is the perceived threat a danger to the organism?; (2) removal of the pathogen that is achieved via a variety of effector mechanisms and (3) return to homeostasis which involves contraction of expanded immune cell populations and repair of damaged tissues. Each of these stages has generated a great deal of interest from both experimental immunologists and modelers.
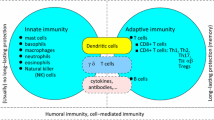
Recognition of a pathogen occurs at two main levels. Cells of the innate immune system, such as dendritic cells (DC), macrophages, natural killer (NK) cells possess receptors for pathogen associated molecular patterns (PAMPs) such as lipopolysaccharide (LPS), bacterial DNA, viral RNA and others. These pattern recognition receptors (PRR) include toll-like receptors (TLR) [1, 2], NOD-like receptors (NLR) [3], RIG-I-like receptors (RLR) and C-type lectin receptors (CLR) [4], which are specific for various pathogen-specific structures. An additional stimulus is required to fully activate the innate immune system in the form of damage associated molecular patterns (DAMPs) or alarmins, which are released from dead or dying cells and are indicative of an invasive pathogen. Many DAMPs, including HMGB1, uric acid, ATP, DNA, bind to the same PRR as a PAMP and may initiate the same inflammatory responses [5]. Thus, the innate immune system recognizes a pathogen invasion and detects tissue damage. The second level of recognition is mediated by cells of the adaptive immune system, namely T and B cells, which express clonally restricted receptors that have a high degree of specificity for their target antigens. Each B and T cell expresses a unique receptor that displays a high degree of specificity and sensitivity. As discussed in more detail below the recognition of antigen by T cells, mediated by the T cell receptor (TCR), has been a topic of intense study. DCs activated by pathogens in the peripheral tissues migrate to draining lymph nodes (LNs) where they present antigen to specific B and T cells. B cell receptors (BCR) recognize antigens in their natural conformation, whereas the TCR recognizes peptide fragments of the same antigen bound in the groove of major histocompatibility complex (MHC) molecules. Once BCR and TCR encounter their specific antigen, an adaptive immune response is initiated which results in the expansion of specific cells, differentiation into appropriate effectors, and the generation of B and T cell memory [6, 7].
The immune system has developed a wide array of effector mechanisms to deal with specific pathogens, since infection with extracellular bacteria, viruses, parasites such as Helminth worms, etc., pose very different challenges. Thus, T cells have evolved the ability to differentiate into several different subsets whose effector functions are best suited to deal with a specific pathogen threat. T helper (Th)1 cells secrete interferon (IFN)-γ, which is required for optimal macrophage activation, cytotoxic T cell maturation and for infections caused by intracellular organisms such as viruses and certain bacteria. Th17 cells secrete interleukin (IL)-17, which acts to attract neutrophils to the site of an infection with extracellular bacteria. Th2 cells secrete IL-5, IL-4 and IL-13, which activate eosinophils in infections with Helminth worms. How does the naïve T cell determine which type of pathogen is causing an infection? DCs migrating from the site of infection carry this information to the naïve T cell, because the interaction of pathogens with specific PRR on the DC triggers the production of unique sets of cytokines that drive the differentiation of specific Th subsets. For example, IL-12 induced following interaction with several TLRs drives Th1 differentiation, IL-23 produced following activation of other PRRs stimulates Th17 differentiation.
Successful elimination of a pathogen depends on the correct choice of effector response, and there are instances where the wrong choice is made and the infection is not cleared. An example of this is the lepromatous form of Mycobacterium leprae infection, which causes leprosy [8]. In these cases an inappropriate Th2 response is generated that fails to control the growth of this intracellular bacterium and there is widespread bacterial dissemination resulting in nerve damage. In contrast infected individuals who generate Th1 responses to this pathogen develop the tuberculous form of the disease, which is characterized by low bacterial burden and a granulomatous reaction that walls off the infection. Damage to nerves still occurs in this form of leprosy but it is caused by the immune response and the bacterial growth is controlled.
The case of tuberculous leprosy highlights the importance of the return to homeostasis. Even when a Th1 response is initiated to respond to Mycobacterium leprae infection macrophages are unable to completely clear the infection and a persistent and chronic infection ensues. It is important to prevent over-activation of effector cells and to turn these off when the pathogen has been cleared. This is achieved through immunosuppressive mechanisms, including the generation of both cytokines such as IL-10, IL-27 and TGF-β, and regulatory T (Treg) cells [9–11]. Defects in Treg and IL-10 may lead to complete clearance of a pathogen but, often, with severe immunopathological consequences [12, 13].
Thus, the regulation of the immune response at all of these stages is critical to ensure the elimination of invading pathogens while preventing excessive immune-mediated tissue damage [14]. When these regulatory mechanisms fail disease may result. For example a defect in the ability of immune system to distinguish between an invading and dangerous pathogen and self-tissues can result in autoimmune disease, such as type 1 diabetes or multiple sclerosis. Excessive immune recognition of commensal bacteria in the gut can lead to inflammatory bowel diseases, such as Crohn’s disease and ulcerative colitis. Early on we realized the value of mathematical and computational modeling in attempting to understand these complex interactions [15–18]. Experimentalists tend to examine the role of a particular protein or cell in the system by creating model systems in which the protein of interest is either removed from the system, by gene targeting, or over-expressed. This reductionist approach has yielded many important insights but also has limits. For example when the cytokine IL-2 [19], an important growth factor for T cells in vitro, was knocked out in a mouse model, no defects in T cell proliferation in vivo were observed [20]. Rather, the mice developed signs of autoimmunity and excessive T cell activation [21], which was subsequently attributed to the non-redundant role of IL-2 in the development and maintenance of Treg cells [22, 23]. There are many such examples, both in and out of the literature, in which targeted gene deletion in a mouse fails to show the expected phenotype. These findings reveal both a great deal of redundancy in the immune system, such that other factors can replace one that is missing, and also pleiotropy, as demonstrated by the IL-2 example, where previously unknown functions of a protein are revealed. Predicting the effect of such manipulations on phenotype is difficult because it involves the interplay of complex and competing mechanisms, such as feedback loops and competition that resolve in a context-dependent manner [24]. Computational modeling allows us to build representations of the system as a whole, which can be used to test hypotheses and provide predictions that can then be tested experimentally.
There are many other important and interesting questions that are covered in other contributions to this issue, and in this article we are choosing to focus on issues related to T cell recognition, activation and regulation. These are topics that are pivotal to the immune response, due to the central role that T cells play in orchestrating the immune response and these have motivated a large number of modeling studies. This review will also highlight how technological advances in the study of immune responses are providing new quantitative data that inform computational models and may lead to new insights into T cell dynamics and function.
Big questions in T cell immunology
T cell recognition of antigen has been a subject of intense study for many years and the discovery of MHC restriction [25] led to intense speculation concerning the nature and structure of antigen receptors on T cells. The fact that T cell recognition required the presence of self MHC molecules led to speculations that T cells express two receptors; one for MHC and one for antigen. The description of the elegant structure of MHC molecules [26] clarified the issue when it was revealed that MHC molecules contain a peptide-binding groove into which small peptides derived from self and foreign antigens could bind. In addition it was shown that the well known polymorphic regions of MHC genes were localized to the peptide binding groove [27], thus giving each MHC allele the ability to bind a unique set of peptides. Further structural analysis of the TCR [28, 29] revealed that this structure binds components of both MHC and peptide. Unlike the high affinities with which immunoglobulin binds antigen, TCR have very low measured affinities for the peptide (p)MHC complex [30, 31]. The low affinity of this interaction is in contrast to the high degree of specificity and sensitivity exhibited by the TCR. This raises important questions about how T cell acquire these receptors, what is necessary for T cell activation and how does the signal received from the TCR determine the T cell fate following activation.
How is the T cell repertoire generated and maintained?
T cells develop in the thymus where they acquire different TCRs, through random DNA rearrangements. Interaction with self MHC is required for a cell to survive during the process of positive selection, but cells expressing TCRs that interact too strongly with self pMHC complexes are deleted in negative selection. It is thought that positive and negative selection are governed by different activation thresholds in the developing thymocyte [32]. Thus, the TCR/pMHC affinity required for positive selection is lower than that needed for negative selection. Modeling in this area has focused on many aspects of T cell development [33] including the role of the thymic involution on TCR diversity [34, 35], how signals are integrated by developing thymocytes [36] and the population dynamics of developing T cells [37–39]. Early ordinary differential equation (ODE) models of population dynamics in the thymus used measurements of the proportions of different thymocyte populations to infer the dynamics of thymic selection [40]. These models concluded that most thymocytes fail to pass the positive selection step and predicted that positive selection signaled proliferation of selected thymocytes. More recently a multi-compartment ODE model, which tracks lymphocyte populations in distinct parts of the immune system (i.e., blood, spleen, and LNs), used data from studies in which dividing thymocytes were specifically depleted to calculate kinetic parameters for the flux and steady state levels of relevant thymic populations [41]. Sinclair et al. [39] have taken advantage of detailed experimental data using a transgenic system in which T cell development can be turned on at selected times and thymocyte differentiation can be followed over time. These data suggested that the observed asymmetric differentiation of CD8 versus CD4 T cells could be explained by increased death rate in the CD8 population [39]. Another more recent model [38] used novel experimental data from bim −/− NUR77GFP mice [42]; NUR77 is downstream of TCR signaling and levels of NUR77 expression are directly proportional to the strength of TCR signal whereas the proapoptotic molecule Bim is necessary for negative selection. These data provided estimates for the number of thymocytes undergoing positive and negative selection and thus allowed a more detailed kinetic study of thymocyte differentiation incorporating death rates, proliferation and selection rates [38]. These and other models [33] have developed quantitative estimates of the lifetimes of the individual thymocyte populations and the timing of the selection steps during T cell development. Most of these models assume that TCR engagement depends on the overall affinity of the pMHC/TCR interaction [33], i.e., the total number of TCR that are engaged with pMHC at a given time, but recent data suggest that the duration of the TCR engagement also plays an important role in thymic selection [43–45].
The result of thymic differentiation is the generation of a diverse repertoire of T cells that are restricted to self MHC, lack autoreactivity and maintain enough diversity to withstand pathogen challenge. Another feature of T cell recognition is that it has been estimated that individual TCR may crossreact on a wide array of pMHC complexes with shared structural features [33, 46, 47]. In addition, the influence of so-called coagonist peptides on T cell activation was shown in a recent study, which combined modeling and experiments, to depend on the affinity of the co-receptor CD8 for individual MHC class I molecules [48]. In this context a coagonist peptide is derived from endogenous proteins, is unable to stimulate T cells alone but enhances the recognition of agonist peptides [49]. Recently it has been observed that virus-specific T cells with a memory phenotype, indicative of prior activation, are found in individuals who had never previously been exposed to that virus [50, 51] suggesting a protective role for this cross-reactivity. These studies were possible because of new technology allowing the isolation of rare antigen-specific T cells in a polyclonal population using soluble pMHC tetramers [52]. As a result of these studies estimates for the frequencies of antigen-specific T cells have been made [53] and these range from 1 in a million to 90 in a million T cells. Recent technical advances in deep sequencing, flow cytometry, spectratyping and analysis of T cell precursor frequencies have allowed for more quantitative assessments of TCR diversity [54]. Spectratyping is a method by which the length of the CDR3 region of TCR α and β chains can be determined and this gives an indication of the diversity of the TCR sequences within a population; diverse T cell populations exhibit a normal distribution of CDR3 lengths whereas a population dominated by a small number of clones may only have one of two CDR3 lengths that dominate [55]. Several statistical approaches have been developed to analyze spectratyping data [56–58] and these have been used to estimate the size of the repertoire [59]. Advances in next generation sequencing allow for detailed sequence analysis of TCR gene diversity at various stages of development and new estimates of diversity are being generated using these data [47]. One challenge with the sequencing data is that each α and β chain is sequenced separately and it is not possible to know how they are paired unless the T cells are first cloned, although novel single cell techniques are now making it possible to pair individual TCR chains [ 60 ]. New analysis tools are being developed to handle the large amount of data that is generated from these sequencing platforms [61]. In addition, advances in single cell sequence analysis will allow more accurate assessments of diversity [60, 62], and combined with isolation of T cells with a given specificity [63] a deeper understanding of how the naïve T cell repertoire is generated and how it changes following antigen exposure will be obtained.
Treg cells arise in the thymus following interaction with self pMHC class II complexes and there has been a great deal of interest in analyzing the Treg repertoire. A recent study performed high throughput sequencing of TCR genes from defined human T cell populations [64] and this showed considerable overlap between sequences derived from T cell subsets with different functional properties. A more recent study demonstrated that increased TCR diversity in Treg was necessary for optimal suppressive function [65]. In a commentary to this study Wing and Sakaguchi proposed a simple mathematical model that explained these findings [66]. This model states that the degree of protection from autoimmunity is proportional to the number of Treg cells with specificity for the self-antigen in question. A prediction from this model is that mouse strains, or individuals, which develop autoimmunity have a reduced Treg repertoire diversity. This prediction has been validated in the non-obese diabetic mouse, which spontaneously develops type 1 diabetes [67].
Treg induction following peripheral stimulation of naïve T cells is also critical to determining that nature and strength of the immune response to antigen. Mathematical modeling has recently been used to explore the factors that lead to variability in the level of Treg induction in response to different types of stimulation. We have recently investigated the role of TCR signaling strength in the induction of FoxP3+ Treg via the interaction of multiple signaling pathways, as discussed in more detail below [Miskov-Zivanov, 2013]. Another study combined CFSE labeling and careful quantification of apoptotic cells to determine how differential regulation of both proliferation and cell death can create conditions favorable and unfavorable for Treg induction [68].
How do T cells get activated?
There are two major challenges faced by the immune system in mounting a specific T cell response to a given antigen: (1) the low precursor frequency of T cells specific for a given foreign pMHC; and (2) the small number of antigen-containing pMHC complexes presented by each DC in the draining LN. Understanding how the immune system overcomes each of these challenges has generated a great deal of interest from modelers and experimental immunologists.
It has been known for some time that naïve T cells are constantly circulating between LN and blood [69, 70] and that this circulation involves interactions between T cells and self pMHC necessary for survival [71–73]. Early attempts at quantitating the migratory behavior of lymphocytes used the adoptive transfer of radiolabeled cells followed by enumeration of their location at various time points after injection [74, 75]. Based on such results a partial differential equation (PDE) model was developed [76] that was able to approximate the experimental results. In this model, the blood, spleen, and lymphatic systems were modeled as separate compartments and in addition the spleen and lymphatic compartments were spatially resolved in one dimension to model T cell transport dynamics. Interestingly this model suggested that T cells do not simply enter and exit LN but they also encounter and interact with other cells within the LN [76].
Advances in intra-vital imaging have allowed the direct visualization of T cell DC interactions within a living LN and thus the actual interaction times can now be measured [77, 78]. A recent study determined the transit times of CD4 and CD8 T cells through lymph nodes using these techniques [79]. Labeled T cells were adoptively transferred, the drug FTY720 was administered to block sphingosine-1-phosphate receptors and sequester lymphocytes in LN and cells were counted in multiple LN [79]. These experiments revealed that CD4 T cells spend less time in LN compared with CD8 T cells (12 h vs 22 h). In addition these studies also showed that the duration of the contact between DC and CD4 T cells depended on the presence of MHC [79], and this has also been shown in an antigen specific system [80]. In this last study it was shown that the overall avidity of the TCR/pMHC interaction was more important than pMHC density in determining contact time [80].
These new and more quantitative data have been used to develop more detailed models of lymphocyte trafficking and activation [81–84]. An ODE-based model [82] was used to model the efficiency with which an antigen-specific T cell migrates through LN and finds the DCs that are presenting the cognate antigen. The results of this model suggest a trade-off between transit times and pMHC density such that detection of a low abundance pMHC is optimal when transit times are slow. Fast transit times favor a more rapid response when the relevant pMHC is in high abundance [82]. Another modeling study using a hybrid two-compartment model reached similar conclusions [81]. In these studies the LN is modeled using a 3D agent based model (ABM) [83] and the blood compartment is modeled using ODEs. The model considers T cell trafficking as well as the generation of primary and secondary immune responses in CD4 and CD8 T cells [81]. This model demonstrated that T cell fate was determined in part by the relative abundance of pMHC such that low abundance favored the development of memory CD4 T cells, whereas intermediate pMHC levels led to CD4 effector T cells. [81]. Interestingly, CD8 T cells required higher levels of pMHC to achieve these fates compared with CD4 T cells.
When a naïve T cell encounters a DC presenting the cognate pMHC complex, a series of signaling events are triggered by the interaction, resulting in activation, proliferation and differentiation of the T cell. T cell activation has fascinated immunologists and modelers for many years since it is unclear how T cells can have such a high degree of specificity and sensitivity when the interaction between TCR and pMHC has an inherently low affinity [85, 86]. Early models proposed that T cells engage in serial triggering [87] along with kinetic proofreading [88–90]. In these models TCR are internalized following engagement with pMHC thereby “counting” the number of interactions (serial triggering) and each interaction stimulates a series of conformational changes in the TCR as it encounters ligands. This provided an explanation for the observed differences in activation profiles between agonist and non-stimulatory ligands [91]. These models associated the degree of T cell activation with the duration of the interaction between T cell and DC and more recent imaging analyses of T cell/DC interactions have leant support to this notion [80, 92].
TCR activation leads to a complex series of early and late signaling events [93]. Many of the early signaling responses follow a digital pattern of response with the number of responding cells in a population increasing over time [86, 94]. This has led to several models based on feedback loops and leading to digital responses necessary for full activation based on ERK [94] or Ras activation [95]. Another important signaling mediator is calcium and a rise in intracellular calcium levels occurs within seconds following TCR stimulation. A recent multi-compartment quantitative model of calcium dynamics investigated the role of the calcium release activated channel (CRAC) in controlling calcium levels, concluding that CRAC played a predominant role in preventing calcium depletion in T cells [96].
These early signaling events are followed by the formation of an immunological synapse (IS), an ordered structure in which TCR and signaling molecules are clustered in the center and they are surrounded by a ring of adhesion molecules, such as LFA-1 [97, 98]. New imaging techniques have allowed for a better understanding of the formation and potential function of the IS [99], although its exact role in T cell activation is still being debated [98]. New advances in imaging and the development of specific tools are further defining the important role that the IS plays in communications between cells of the immune system [98].
Many of the signaling cascades stimulated in T cells are also found in other cells and one good example is the activation of the NFκB pathway, which plays an important role downstream of TCR signaling. This pathway has been extensively studied in multiple systems and has also been the subject of many models [100]. Recently, modeling of this pathway has been used in the development of approaches to identify signal-specific pharmacological targets [101]. In this study the topology of signaling hubs in the NFκB pathway along with dynamical modeling were used to identify pharmacological targets that would inhibit responses to one input signal without disrupting responses to other signals and also to reshape temporal responses to select desirable features and suppress undesirable ones. This approach proved successful in identifying some stimulus-specific inhibitors in the context of the NFκB response to LPS and TNF [101], and could also be applied to development of specific inhibition strategies in T cells. The fact that T cells also contain many of the signaling pathways that have been widely studied in other cell types suggests that it will be useful to integrate models that have been developed for other systems into modeling of T cell responses.
How the immune response is regulated by the strength of the TCR signal
Signaling via the TCR not only stimulates T cell activation and proliferation but it has become apparent that the differentiation of naïve T cells into specific Th subsets is influenced by the perceived strength of the TCR signal. It has been known for many years that the dose of a stimulating antigen or pathogen has important implications for the nature of the immune response [102, 103]. In these studies it was observed that the development of cell-mediated or humoral immunity was controlled by the dose of the immunizing antigen [102]. This was later found to depend on Th1 and Th2 responses respectively and it was possible to alter the nature of the response by altering the dose and thereby the TCR signal strength [104, 105]. This has important consequences in the context of infection as demonstrated by the Leishmania infection model. Certain mouse strains develop an inappropriate Th2 response to this infection, which proves fatal. It was shown that exposing these mice to a low dose of bacteria would induce a Th1 response that was then able to protect the mice from subsequent infection with the higher and potentially fatal dose [103]. The differentiation of Th cell subsets involves the expression of unique transcription factors that drive the production of the signature cytokines produced by each of the Th subsets. It is known that Th differentiation requires the presence of certain signature cytokines in the milieu: IFN-γ for Th1 cells, IL-4 for Th2 cells, TGF-β for Treg cells and IL-6 and TGF-β for Th17 cells. These cytokines act by inducing transcription factors that augment the production of relevant cytokines and induce expression of specific cytokine receptors that are required for full differentiation and maintenance of the phenotype [106]. Several models have been developed that address Th cell differentiation networks [107–110], although these models did not take TCR signal strength into account. Because of the large number of cytokines, receptors, and signaling components involved in these processes, these models have used more coarse-grained modeling approaches than the studies cited so far. In such models, some or all of the variables are restricted to a small number of possible values, reducing the number of states of the system and simplifying formulation of the model, as we will discuss in more detail below. One cost of such simplifications is that quantitative factors, such as TCR signal strength, are more difficult to incorporate into the model, but, as we also discuss below, it is possible to do so.
We have been interested in this phenomenon in the context of the differentiation of Treg from naïve T cells. We [111] and others [80, 112–114] have shown that Foxp3 expressing Treg are induced when naïve T cell are exposed to low doses of antigen. This has been correlated with signaling via the Akt/mTOR pathway [115, 116] such that the degree of Akt/mTOR signaling is inversely correlated with the induction of Treg [111]. Treg that are expanded following low dose antigen are able to prevent autoimmune diabetes in vivo [112, 117]. We developed a mathematical model, based on Boolean logic, to further understand how TCR signal strength could contribute to Treg expansion and induction [118]. The model was developed through an iterative process of reading the literature, tuning the components and interactions, and comparison with experimental data, which was particularly important for calibrating the time scales of various processes. The components of the signaling pathway are depicted as elements within the model, and each element has only a small number of possible states—either ON or OFF in the case of most variables, or OFF, LOW, and HIGH in the case of a few critical variables, such as TCR and PI3K. Figure 1a presents the overall structure of the model showing model components as nodes and positive and negative influences between these elements as arrows with pointed and flat arrowheads respectively. This diagram is translated into an executable model by constructing update rules for each element. At each point in time, the overall state of the network is represented by the values of each of the variables in the system. The value of each element at the next time step is determined from the current values by the rules, which for each element take the form of logical functions of the influencing elements involving the basic logic operators AND, OR and NOT. For example, Akt, the element circled in red in Fig. 1a, is updated according to the rule
which means that the next value of AKT is ON only if both PDK1 and MTORC2 are ON and otherwise the next value is OFF. If the rule used “or” instead of “and”, AKT would become ON if either MTORC2 or PDK1 were ON. A “trajectory” corresponding to a single cell containing this network evolving in time is simulated by selecting an initial set of values for the network elements representing the resting state of the cell plus the initial stimulation that is applied in the experiments, e.g., stimulating TCR and CD28. Different methods exist for updating the state of the network through application of the rules [119]. In our work we have chosen an asynchronous stochastic scheme that takes into account stochastic variation among cells [118]. We then compute a large number of individual trajectories and plot the percentage of cells exhibiting a given characteristic, e.g., expression of IL2 or FoxP3, as shown in Fig. 1b. These results can be directly compared with experimental results obtained from flow cytometry (Fig. 1c). A major advantage of this kind of modeling is that signaling pathways can be followed from the cell surface to the end result of gene transcription without having to determine the large number of kinetic constants needed in an ODE based model.
Computational modeling of T cell differentiation. a Signaling network governing differentiation. Nodes represent network elements, which include external ligands such as MHC and IL-2, receptors, such as TCR and IL-2R, signaling intermediates such as Ras, Akt, and JAK3, and transcription factors such as AP-1, NF-AT, and FoxP3. Edges represent interactions between these elements with pointed arrowheads representing activation, blunt arrowheads representing inhibition, and dashed lines representing translocation of molecules between cellular compartments. The state of a given element in the network, such as Akt (circled in red) is updated according to the states of its incoming influences—PDK1 and mTORC2 in the case of Akt. b Simulated time courses of the percentage of cells destined to become IL-2+ Th cells (blue line) or Foxp3+ Treg cells (red line). In this simulation the TCR signal was interrupted after 6 times steps and the appearance of Th and Treg cells were followed at each time step. Results represent the combined trajectories, from a total of 1,000 simulations, leading to Th (390/1,000) or Treg (419/1,000) outcomes. c Corresponding experimental results for percentages of purified CD4+ T cells stimulated on plates coated with anti-CD3 antibody at high (blue line) and low (blue line) dose in the presence of soluble anti-CD28 antibody. High dose stimulation results in rapid stabilization of the Th (Foxp3− CD25+) phenotype whereas Treg (Foxp3+ CD25+) cells take much longer to appear in low dose stimulation. Results shown are mean ± SEM of three similar experiments (Color figure online)
The model made several predictions that we were able to verify experimentally [118]. In particular the model predicted that the induction of Treg by adding TGF-β to a high TCR signal strength scenario would not result in reduced Akt/mTOR signaling, which was confirmed in experiments. More significantly, we found that the modeling of low TCR signal strength most accurately reproduced the experimental results when the duration of the TCR engagement was reduced [118]. Interrupting the TCR signal after varying periods of time revealed the plasticity of the activated T cells. Thus, if the TCR signal was allowed to persist throughout the course of the experiment only Th cells were generated, whereas if the signal was interrupted after a period of time a mixed population of cells was generated that included Th, Treg, and also non-activated cells [118]. Another important insight that we gained from the model was that Treg cells take longer to stabilize as a phenotype than Th cells (Fig. 1b). Subsequent experiments confirmed this prediction (Fig. 1c). The model also allowed us to examine the trajectory of cells destined to become Th or Treg and identified the importance of the lipid phosphatase PTEN in determining the ultimate cell fate. Several new avenues of experimental research are being pursued based on these predictions. These results suggest that a population of T cells has the capacity to differentiate down several paths. It seems likely that differences in the behavior of individual cells are related to the observed heterogeneity in the expression levels of critical proteins in the TCR signaling cascade [120].
Treg expansion and differentiation is optimized by low TCR signal strength but other factors also play an important role in Treg biology [121]. These include the cytokines IL-2, TGF-β, IL-10 and costimulatory pathways such as CD28, PD-1. IL-2 plays a critical role in the development and homeostasis of Treg; this is illustrated by the complete lack of Treg in mice deficient for the signaling chain of the IL-2R [122]. Treg constitutively express the high affinity IL-2R (CD25) whereas conventional Th cells only transiently express this receptor following activation. In addition, Treg do not make IL-2 whereas Th cells are the main source of IL-2 that is used by both Th and Treg to stimulate proliferation and expansion. Models of this interaction between the cells have illustrated how Treg can “steal” IL-2 from neighboring Th cells thereby not only ensuring their own survival but suppressing the proliferation of the Th cells [123]. Defects in IL-2 production have been associated with autoimmune diseases such as type 1 diabetes [124, 125] and this has been correlated with a decrease in Treg [124]. Due to its initial description as a T cell growth factor, IL-2 was also used clinically for the treatment of cancer with the aim of boosting T cell responses which were further enhanced by the addition of complexes of IL-2 with anti-IL-2 [126]. However, it was observed that depending on the type of complex used the result could be either expansion of CD8 T cells or Treg [126]. This phenomenon has been explored in models with the aim of developing the best therapeutic strategies for the use of IL-2 to boost Treg and therefore prevent autoimmunity [127, 128]. A recent model has also suggested that collective decisions by a population of cells are important in determining whether T cells become activated and that IL-2 availability may be one of the mechanisms by which this is achieved [129]. This is a situation where computational modeling can be of great benefit for the optimal design of an immunotherapy because intuition generally fails in systems that can exhibit such complex dynamics. Similarly, modeling has been used to optimize therapeutic intervention with anti-CD3 antibodies [130]; a therapy that involves a short-term course of low dose anti-CD3 antibodies in combination with islet antigens designed to boost antigen-specific Treg and preserve islet function [131, 132]. This model has identified several biomarkers that can be used to predict who may respond to this form of therapy [130].
Modeling approaches and communication between modelers and experimentalists
The field of modeling in immunology has advanced a great deal since the early days and this is in part due to an increasing awareness among immunologists of the need for modeling to increase insights into complex biological processes. In addition, more modelers are becoming familiar with the experimental techniques that are used and there is an increasing number of individuals who are trained in both disciplines [133, 134]. The improvements in technology have also made it possible for detailed quantitative measurements to be made. There is however still a need for closer collaboration and communication. For example modeling TCR activation has involved the use of deterministic ODE-based models that have focused on the earliest steps in T cell activation [94, 135, 136]. The development of rule-based approaches has simplified the development of complex models and has allowed the use of continuous as well as stochastic simulation methods [137]. Because these models require detailed quantitative data that may not be available in many cases, alternative approaches based on logical (aka Boolean) modeling have also been used to model T cell differentiation. Logical models do not require detailed quantitative measurements, but rather allow the development of complex qualitative networks and several have been developed to investigate T cell activation [118, 138, 139] and differentiation [108, 109, 118]. The models of Th cell differentiation focused on the interactions between cytokines, cytokine receptors, signaling molecules and transcription factors in defining a network and identifying scenarios for Th cell differentiation and plasticity [108, 109].
One concern with the present modeling efforts is that few published models are ever revisited or used by other investigators. Different tools and approaches are often used to address similar questions and it can be difficult to compare results from these kinds of models. In addition, the field is in need of more developed systems immunology approaches [140, 141]. These are models that incorporate the spatial and time scales of the whole immune response. These would include molecular and signaling events occurring within the cell, cellular interactions and then effects on the tissue or organism as a whole. Multi-compartment models of antigen presentation [142] and of lymph nodes [81] have been developed. A recent study described a novel methodology that can be used to capture the dynamics of complex biological systems using data from multiple levels of intra and inter-cellular behavior [143]. This method uses algorithmic information theory and the study was able to perform successful analysis of the response of Th and Treg cells to heat shock proteins [143]. Another issue is that experimentalists are often intimidated by the technical aspects of modeling and thus may fail to see the value of the exercise. Several modeling platforms have been developed that allow non-professional modelers to build models and test out ideas in an easy to use platform [144, 145]. These use graphic interfaces that are familiar to immunologists, although at this stage the models are mainly focused on the signaling aspects.
In conclusion, modeling of events in T cell differentiation, activation and function have proved to be extremely valuable in helping to elucidate the role of these important cells in the immune system. There are now numerous modeling approaches and platforms available and a challenge for the future will be the coordination of these diverse methodologies. Many new therapies are targeting T cell immune functions, such as IL-2 or immune checkpoint blockade in cancer [146], and modeling is likely to play a key role in optimizing these approaches and in identifying new ones
References
Akira S, Takeda K (2004) Toll-like receptor signalling. Nat Rev Immunol 4:499–511
Takeda K, Kaisho T, Akira S (2003) Toll-like receptors. Annu Rev Immunol 21:335–376. doi:10.1146/annurev.immunol.21.120601.141126
Davis BK, Wen H, Ting JP-Y (2011) The inflammasome NLRs in immunity, inflammation, and associated diseases. Annu Rev Immunol 29:707–735 10.1146/annurev-immunol-031210-101405
Sancho D, Reis e Sousa C (2012) Signaling by myeloid C-type lectin receptors in immunity and homeostasis. Annu Rev Immunol 30:491–529 10.1146/annurev-immunol-031210-101352
Zelenay S, Reis e Sousa C (2013) Adaptive immunity after cell death. Trends Immunol 34:329–335. doi:10.1016/j.it.2013.03.005
McHeyzer-Williams LJ, McHeyzer-Williams MG (2005) Antigen-specific memory B cell development. Annu Rev Immunol 23:487–513. doi:10.1146/annurev.immunol.23.021704.115732
Mueller SN, Gebhardt T, Carbone FR, Heath WR (2013) Memory T cell subsets, migration patterns, and tissue residence. Annu Rev Immunol 31:137–161. doi:10.1146/annurev-immunol-032712-095954
Nath I, Vemuri N, Reddi AL, Jain S, Brooks P, Colston MJ, Misra RS, Ramesh V (2000) The effect of antigen presenting cells on the cytokine profiles of stable and reactional lepromatous leprosy patients. Immunol Lett 75:69–76
Berg DJ, Davidson N, xFc, hn R, xFc, ller W, Menon S, Holland G, Thompson-Snipes L, Leach MW, Rennick D (1996) Enterocolitis and colon cancer in interleukin-10-deficient mice are associated with aberrant cytokine production and CD4(+) TH1-like responses. J Clin Invest 98:1010–1020. doi:10.1172/jci118861
Hunter Christopher A, Kastelein R (2012) Interleukin-27: balancing protective and pathological immunity. Immunity 37:960–969. doi:10.1016/j.immuni.2012.11.003
Rouse BT, Suvas S (2004) Regulatory cells and infectious agents: détentes cordiale and contraire. J Immunol 173:2211–2215
Belkaid Y, Piccirillo CA, Mendez S, Shevach EM, Sacks DL (2002) CD4+ CD25 + regulatory T cells control Leishmania major persistence and immunity. Nature 420:502–507
Suzuki Y, Sher A, Yap G, Park D, Neyer LE, Liesenfeld O, Fort M, Kang H, Gufwoli E (2000) IL-10 is required for prevention of necrosis in the small intestine and mortality in both genetically resistant BALB/c and susceptible C57BL/6 mice following peroral infection with Toxoplasma gondii. J Immunol 164:5375–5382
Germain RN (2012) Maintaining system homeostasis: the third law of Newtonian immunology. Nat Immunol 13:902–906
Burke MA, Morel BF, Oriss TB, Bray J, McCarthy SA, Morel PA (1997) Modeling the proliferative response of T cells to IL-2 and IL-4. Cell Immunol 178:42–52
Morel BF, Burke MA, Kalagnanam JR, McCarthy SA, Tweardy DJ, Morel PA (1996) Making sense of the combined effect of interleukin-2 and interleukin-4 on lymphocytes using a mathematical model. Bull Mathemat Biol 58:569–594
Morel PA (1988) Mathematical modeling of immunological reactions. Frontiers Biosci 3:338–347
Morel PA, Ta’asan S, Morel BF, Kirschner DE, Flynn JL (2006) New insights into mathematical modeling of the immune system. Immunol Res 36:157–165. doi:10.1385/IR:36:1:157
Cantrell DA, Smith KA (1984) The interleukin-2 T-cell system: a new cell growth model. Science 224:1312–1316
Schorle H, Holtschke T, Hunig T, Schimpl A, Horak I (1991) Development and function of T cells in mice rendered interleukin-2 deficient by gene targeting. Nature 352:621–624
Sadlack B, Merz H, Schorle H, Schimpl A, Feller AC, Horak I (1993) Ulcerative colitis-like disease in mice with a disrupted interleukin-2 gene. Cell 75:253–261. doi:10.1016/0092-8674(93)80067-O
Malek TR (2003) The main function of IL-2 is to promote the development of T regulatory cells. J Leukocyte Biol 74:961–965. doi:10.1189/jlb.0603272
Furtado GC, Curotto de Lafaille MA, Kutchukhidze N, Lafaille JJ (2002) Interleukin 2 signaling is required for CD4(+) regulatory T cell function. J Exp Med 196:851–857
Tyson JJ, Novak B (2010) Functional motifs in biochemical reaction networks. Annu Rev Phys Chem 61:219–240. doi:10.1146/annurev.physchem.012809.103457
Zinkernagel RM, Doherty PC (1975) H-2 compatability requirement for T-cell-mediated lysis of target cells infected with lymphocytic choriomeningitis virus. Different cytotoxic T-cell specificities are associated with structures coded for in H-2K or H-2D. J Exp Med 141:1427–1436. doi:10.1084/jem.141.6.1427
Bjorkman PJ, Saper MA, Samraoui B, Bennett WS, Strominger JL, Wiley DC (1987) Structure of the human class I histocompatibility antigen, HLA-A2. Nature 329:506–512
Bjorkman PJ, Saper MA, Samraoui B, Bennett WS, Strominger JL, Wiley DC (1987) The foreign antigen binding site and T cell recognition regions of class I histocompatibility antigens. Nature 329:512–518
Ding Y-H, Smith KJ, Garboczi DN, Utz U, Biddison WE, Wiley DC (1998) Two human T cell receptors bind in a similar diagonal mode to the HLA-A2/Tax peptide complex using different TCR amino acids. Immunity 8:403–411. doi:10.1016/S1074-7613(00)80546-4
Speir JA, Garcia KC, Brunmark A, Degano M, Peterson PA, Teyton L, Wilson IA (1998) Structural basis of 2C TCR allorecognition of H-2Ld peptide complexes. Immunity 8:553–562. doi:10.1016/S1074-7613(00)80560-9
Garcia KC, Tallquist MD, Pease LR, Brunmark A, Scott CA, Degano M, Stura EA, Peterson PA, Wilson IA, Teyton L (1997) αβ T cell receptor interactions with syngeneic and allogeneic ligands: affinity measurements and crystallization. Proc Natl Acad Sci 94:13838–13843
Margulies DH, Plaksin D, Khilko SN, Jelonek MT (1996) Studying interactions involving the T-cell antigen receptor by surface plasmon resonance. Curr Opin Immunol 8:262–270
Hogquist KA, Baldwin TA, Jameson SC (2005) Central tolerance: learning self-control in the thymus. Nat Rev Immunol 5:772–782
Yates AJ (2014) Theories and quantification of thymic selection. Front Immunol 5:13. doi:10.3389/fimmu.2014.00013
Johnson PL, Yates AJ, Goronzy JJ, Antia R (2012) Peripheral selection rather than thymic involution explains sudden contraction in naive CD4 T-cell diversity with age. Proc Natl Acad Sci USA 109:21432–21437. doi:10.1073/pnas.1209283110
Johnson PL, Goronzy JJ, Antia R (2014) A population biological approach to understanding the maintenance and loss of the T cell repertoire during aging. Immunology. doi:10.1111/imm.12244
Van Den Berg HA, Rand DA, Burroughs NJ (2001) A reliable and safe T cell repertoire based on low-affinity T cell receptors. J Theor Biol 209:465–486. doi:10.1006/jtbi.2001.2281
Hapuarachchi T, Lewis J, Callard RE (2013) A mechanistic model for naive CD4 T cell homeostasis in healthy adults and children. Front Immunol 4:366. doi:10.3389/fimmu.2013.00366
Sawicka M, Stritesky GL, Reynolds J, Abourashchi N, Lythe G, Molina-Paris C, Hogquist KA (2014) From pre-DP, post-DP, SP4, and SP8 thymocyte cell counts to a dynamical model of cortical and medullary selection. Front Immunol 5:19. doi:10.3389/fimmu.2014.00019
Sinclair C, Bains I, Yates AJ, Seddon B (2013) Asymmetric thymocyte death underlies the CD4:CD8 T-cell ratio in the adaptive immune system. Proc Natl Acad Sci USA 110:E2905–E2914. doi:10.1073/pnas.1304859110
Mehr R, Globerson A, Perelson AS (1995) Modeling positive and negative selection and differentiation processes in the thymus. J Theor Biol 175:103–126. doi:10.1006/jtbi.1995.0124
Thomas-Vaslin V, Altes HK, de Boer RJ, Klatzmann D (2008) Comprehensive assessment and mathematical modeling of T cell population dynamics and homeostasis. J Immunol 180:2240–2250
Stritesky GL, Xing Y, Erickson JR, Kalekar LA, Wang X, Mueller DL, Jameson SC, Hogquist KA (2013) Murine thymic selection quantified using a unique method to capture deleted T cells. Proc Natl Acad Sci USA 110:4679–4684. doi:10.1073/pnas.1217532110
Bhakta NR, Oh DY, Lewis RS (2005) Calcium oscillations regulate thymocyte motility during positive selection in the three-dimensional thymic environment. Nat Immunol 6:143–151. http://www.nature.com/ni/journal/v6/n2/suppinfo/ni1161_S1.html
Liu X, Bosselut R (2004) Duration of TCR signaling controls CD4-CD8 lineage differentiation in vivo. Nat Immunol 5:280–288. doi:10.1038/ni1040
Palmer E, Naeher D (2009) Affinity threshold for thymic selection through a T-cell receptor-co-receptor zipper. Nat Rev Immunol 9:207–213
Mason D (1998) A very high level of crossreactivity is an essential feature of the T-cell receptor. Immunol Today 19:395–404
Zarnitsyna VI, Evavold BD, Schoettle LN, Blattman JN, Antia R (2013) Estimating the Diversity, Completeness, and Cross-Reactivity of the T Cell Repertoire. Front Immunol 4:485. doi:10.3389/fimmu.2013.00485
Hoerter JA, Brzostek J, Artyomov MN, Abel SM, Casas J, Rybakin V, Ampudia J, Lotz C, Connolly JM, Chakraborty AK, Gould KG, Gascoigne NR (2013) Coreceptor affinity for MHC defines peptide specificity requirements for TCR interaction with coagonist peptide-MHC. J Exp Med 210:1807–1821. doi:10.1084/jem.20122528
Yachi PP, Ampudia J, Gascoigne NRJ, Zal T (2005) Nonstimulatory peptides contribute to antigen-induced CD8-T cell receptor interaction at the immunological synapse. Nat Immunol 6:785–792. http://www.nature.com/ni/journal/v6/n8/suppinfo/ni1220_S1.html
Su LF, Davis MM (2013) Antiviral memory phenotype T cells in unexposed adults. Immunol Rev 255:95–109. doi:10.1111/imr.12095
Su LF, Kidd BA, Han A, Kotzin JJ, Davis MM (2013) Virus-specific CD4(+) memory-phenotype T cells are abundant in unexposed adults. Immunity 38:373–383. doi:10.1016/j.immuni.2012.10.021
Moon JJ, Chu HH, Pepper M, McSorley SJ, Jameson SC, Kedl Ross M, Jenkins MK (2007) Naive CD4+ T cell frequency varies for different epitopes and predicts repertoire diversity and response magnitude. Immunity 27:203–213. doi:10.1016/j.immuni.2007.07.007
Jenkins MK, Moon JJ (2012) The role of naive T cell precursor frequency and recruitment in dictating immune response magnitude. J Immunol 188:4135–4140. doi:10.4049/jimmunol.1102661
Six A, Mariotti-Ferrandiz ME, Chaara W, Magadan S, Pham HP, Lefranc MP, Mora T, Thomas-Vaslin V, Walczak AM, Boudinot P (2013) The past, present, and future of immune repertoire biology: the rise of next-generation repertoire analysis. Front Immunol 4:413. doi:10.3389/fimmu.2013.00413
Pannetier C, Cochet M, Darche S, Casrouge A, Zoller M, Kourilsky P (1993) The sizes of the CDR3 hypervariable regions of the murine T-cell receptor beta chains vary as a function of the recombined germ-line segments. Proc Natl Acad Sci USA 90:4319–4323
He M, Tomfohr JK, Devlin BH, Sarzotti M, Markert ML, Kepler TB (2005) SpA: web-accessible spectratype analysis: data management, statistical analysis and visualization. Bioinformatics 21:3697–3699. doi:10.1093/bioinformatics/bti600
Kepler TB, He M, Tomfohr JK, Devlin BH, Sarzotti M, Markert ML (2005) Statistical analysis of antigen receptor spectratype data. Bioinformatics 21:3394–3400. doi:10.1093/bioinformatics/bti539
Miqueu P, Guillet M, Degauque N, Dore JC, Soulillou JP, Brouard S (2007) Statistical analysis of CDR3 length distributions for the assessment of T and B cell repertoire biases. Mol Immunol 44:1057–1064. doi:10.1016/j.molimm.2006.06.026
Arstila TP, Casrouge A, Baron V, Even J, Kanellopoulos J, Kourilsky P (1999) A direct estimate of the human alphabeta T cell receptor diversity. Science 286:958–961
Turchaninova MA, Britanova OV, Bolotin DA, Shugay M, Putintseva EV, Staroverov DB, Sharonov G, Shcherbo D, Zvyagin IV, Mamedov IZ, Linnemann C, Schumacher TN, Chudakov DM (2013) Pairing of T-cell receptor chains via emulsion PCR. Eur J Immunol 43:2507–2515. doi:10.1002/eji.201343453
Michaeli M, Barak M, Hazanov L, Noga H, Mehr R (2013) Automated analysis of immunoglobulin genes from high-throughput sequencing: life without a template. J Clin Bioinform 3:15
Shapiro E, Biezuner T, Linnarsson S (2013) Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat Rev Genet 14:618–630. doi:10.1038/nrg3542
Newell EW, Davis MM (2014) Beyond model antigens: high-dimensional methods for the analysis of antigen-specific T cells. Nat Biotechnol 32:149–157. doi:10.1038/nbt.2783
Wang C, Sanders CM, Yang Q, Schroeder HW, Wang E, Babrzadeh F, Gharizadeh B, Myers RM, Hudson JR, Davis RW, Han J (2010) High throughput sequencing reveals a complex pattern of dynamic interrelationships among human T cell subsets. Proc Natl Acad Sci 107:1518–1523. doi:10.1073/pnas.0913939107
Fohse L, Suffner J, Suhre K, Wahl B, Lindner C, Lee CW, Schmitz S, Haas JD, Lamprecht S, Koenecke C, Bleich A, Hammerling GJ, Malissen B, Suerbaum S, Forster R, Prinz I (2011) High TCR diversity ensures optimal function and homeostasis of Foxp3+ regulatory T cells. Eur J Immunol 41:3101–3113. doi:10.1002/eji.201141986
Wing JB, Sakaguchi S (2011) TCR diversity and Treg cells, sometimes more is more. Eur J Immunol 41:3097–3100. doi:10.1002/eji.201142115
Ferreira C, Singh Y, Furmanski AL, Wong FS, Garden OA, Dyson J (2009) Non-obese diabetic mice select a low-diversity repertoire of natural regulatory T cells. Proc Natl Acad Sci USA 106:8320–8325. doi:10.1073/pnas.0808493106
Caridade M, Oliveira VG, Agua-Doce A, Graca L, Ribeiro RM (2013) The fate of CD4+ T cells under tolerance-inducing stimulation: a modeling perspective. Immunol Cell Biol 91:652–660. doi:10.1038/icb.2013.63
Gowans JL, Knight EJ (1964) The route of re-circulation of lymphocytes in the rat. Proc R Soc Lond B 159:257–282
von Andrian UH, Mempel TR (2003) Homing and cellular traffic in lymph nodes. Nat Rev Immunol 3:867–878. doi:10.1038/nri1222
Kirberg J, Berns A, von Boehmer H (1997) Peripheral T cell survival requires continual ligation of the T cell receptor to major histocompatibility complex-encoded molecules. J Exp Med 186:1269–1275
Martin B, Becourt C, Bienvenu B, Lucas B (2006) Self-recognition is crucial for maintaining the peripheral CD4+ T-cell pool in a nonlymphopenic environment. Blood 108:270–277. doi:10.1182/blood-2006-01-0017
Viret C, Wong FS, Janeway CA Jr (1999) Designing and maintaining the mature TCR repertoire: the continuum of self-peptide:self-MHC complex recognition. Immunity 10:559–568
Sprent J (1973) Circulating T and B lymphocytes of the mouse. I. Migratory properties. Cell Immunol 7:10–39
Smith ME, Ford WL (1983) The recirculating lymphocyte pool of the rat: a systematic description of the migratory behaviour of recirculating lymphocytes. Immunology 49:83–94
Stekel DJ, Parker CE, Nowak MA (1997) A model of lymphocyte recirculation. Immunol Today 18:216–221
Miller MJ, Wei SH, Parker I, Cahalan MD (2002) Two-photon imaging of lymphocyte motility and antigen response in intact lymph node. Science 296:1869–1873. doi:10.1126/science.1070051
Sumen C, Mempel TR, Mazo IB, von Andrian UH (2004) Intravital microscopy: visualizing immunity in context. Immunity 21:315–329. doi:10.1016/j.immuni.2004.08.006
Mandl JN, Liou R, Klauschen F, Vrisekoop N, Monteiro JP, Yates AJ, Huang AY, Germain RN (2012) Quantification of lymph node transit times reveals differences in antigen surveillance strategies of naïve CD4+ and CD8+ T cells. Proc Natl Acad Sci 109:18036–18041. doi:10.1073/pnas.1211717109
Gottschalk RA, Hathorn MM, Beuneu H, Corse E, Dustin ML, Altan-Bonnet G, Allison JP (2012) Distinct influences of peptide-MHC quality and quantity on in vivo T-cell responses. Proc Natl Acad Sci USA 109:881–886. doi:10.1073/pnas.1119763109
Gong C, Linderman JJ, Kirschner D (2014) Harnessing the heterogeneity of T cell differentiation fate to fine-tune generation of effector and memory T cells. Front Immunol 5:57. doi:10.3389/fimmu.2014.00057
Lee M, Mandl JN, Germain RN, Yates AJ (2012) The race for the prize: T-cell trafficking strategies for optimal surveillance. Blood 120:1432–1438. doi:10.1182/blood-2012-04-424655
Gong C, Mattila JT, Miller M, Flynn JL, Linderman JJ, Kirschner D (2013) Predicting lymph node output efficiency using systems biology. J Theor Biol 335:169–184. doi:10.1016/j.jtbi.2013.06.016
Mirsky HP, Miller MJ, Linderman JJ, Kirschner DE (2011) Systems biology approaches for understanding cellular mechanisms of immunity in lymph nodes during infection. J Theor Biol 287:160–170. doi:10.1016/j.jtbi.2011.06.037
van der Merwe PA, Dushek O (2011) Mechanisms for T cell receptor triggering. Nat Rev Immunol 11:47–55. doi:10.1038/nri2887
Chakraborty AK, Das J (2010) Pairing computation with experimentation: a powerful coupling for understanding T cell signalling. Nat Rev Immunol 10:59–71. doi:10.1038/nri2688
Valitutti S, Muller S, Cella M, Padovan E, Lanzavecchia A (1995) Serial triggering of many T-cell receptors by a few peptide-MHC complexes. Nature 375:148–151. doi:10.1038/375148a0
Goldstein B, Faeder JR, Hlavacek WS (2004) Mathematical and computational models of immune-receptor signalling. Nat Rev Immunol 4:445–456
McKeithan TW (1995) Kinetic proofreading in T-cell receptor signal transduction. Proc Natl Acad Sci USA 92:5042–5046
Rabinowitz JD, Beeson C, Lyons DS, Davis MM, McConnell HM (1996) Kinetic discrimination in T-cell activation. Proc Natl Acad Sci USA 93:1401–1405
Sloan-Lancaster J, Allen PM (1996) Altered peptide ligand–induced partial T cell activation: molecular mechanisms and role in T cell biology. Annu Rev Immunol 14:1–27. doi:10.1146/annurev.immunol.14.1.1
Gascoigne NR, Zal T, Alam SM (2001) T-cell receptor binding kinetics in T-cell development and activation. Expert Rev Mol Med 2001:1–17. doi:10.1017/S1462399401002502
Tkach K, Altan-Bonnet G (2013) T cell responses to antigen: hasty proposals resolved through long engagements. Curr Opin Immunol 25:120–125. doi:10.1016/j.coi.2012.12.001
Altan-Bonnet G, Germain RN (2005) Modeling T cell antigen discrimination based on feedback control of digital ERK responses. PLoS Biol 3:e356. doi:10.1371/journal.pbio.0030356
Das J, Ho M, Zikherman J, Govern C, Yang M, Weiss A, Chakraborty AK, Roose JP (2009) Digital signaling and hysteresis characterize ras activation in lymphoid cells. Cell 136:337–351. doi:10.1016/j.cell.2008.11.051
Schmeitz C, Hernandez-Vargas EA, Fliegert R, Guse AH, Meyer-Hermann M (2013) A mathematical model of T lymphocyte calcium dynamics derived from single transmembrane protein properties. Front Immunol 4:277. doi:10.3389/fimmu.2013.00277
Monks CRF, Freiberg BA, Kupfer H, Sciaky N, Kupfer A (1998) Three-dimensional segregation of supramolecular activation clusters in T cells. Nature 395:82–86
Xie J, Tato CM, Davis MM (2013) How the immune system talks to itself: the varied role of synapses. Immunol Rev 251:65–79. doi:10.1111/imr.12017
Dustin ML, Depoil D (2011) New insights into the T cell synapse from single molecule techniques. Nat Rev Immunol 11:672–684. doi:10.1038/nri3066
Basak S, Behar M, Hoffmann A (2012) Lessons from mathematically modeling the NF-kappaB pathway. Immunol Rev 246:221–238. doi:10.1111/j.1600-065X.2011.01092.x
Behar M, Barken D, Werner SL, Hoffmann A (2013) The dynamics of signaling as a pharmacological target. Cell 155:448–461. doi:10.1016/j.cell.2013.09.018
Ramshaw IA, Eidinger D (1979) The in vitro induction of T cells which mediate delayed-type hypersensitivity toward horse red blood cells. Cell Immunol 42:42–47
Bretscher PA, Wei G, Menon JN, Bielefeldt-Ohmann H (1992) Establishment of stable, cell-mediated immunity that makes “susceptible” mice resistant to Leishmania major. Science 257:539–542
Constant SL, Bottomly K (1997) Induction of Th1 and Th2 CD4+ T cell responses: the alternative approaches. Annu Rev Immunol 15:297–322
Hosken NA, Shibuya K, Heath AW, Murphy KM, O’Garra A (1995) The effect of antigen dose on CD4+ T helper cell phenotype development in a T cell receptor-alpha beta-transgenic model. J Exp Med 182:1579–1584
Zhu J, Yamane H, Paul WE (2010) Differentiation of effector CD4 T cell populations (*). Annu Rev Immunol 28:445–489. doi:10.1146/annurev-immunol-030409-101212
Hong T, Xing J, Li L, Tyson JJ (2011) A mathematical model for the reciprocal differentiation of T helper 17 cells and induced regulatory T cells. PLoS Comput Biol 7:e1002122. doi:10.1371/journal.pcbi.1002122
Mendoza L (2006) A network model for the control of the differentiation process in Th cells. Biosystems 84:101–114. doi:10.1016/j.biosystems.2005.10.004
Naldi A, Carneiro J, Chaouiya C, Thieffry D (2010) Diversity and plasticity of Th cell types predicted from regulatory network modelling. PLoS Comput Biol 6:e1000912. doi:10.1371/journal.pcbi.1000912
van den Ham HJ, de Boer RJ (2012) Cell division curtails helper phenotype plasticity and expedites helper T-cell differentiation. Immunol Cell Biol 90:860–868. doi:10.1038/icb.2012.23
Turner MS, Kane LP, Morel PA (2009) Dominant role of antigen dose in CD4+ Foxp3+ regulatory T cell induction and expansion. J Immunol 183:4895–4903. doi:10.4049/jimmunol.0901459
Daniel C, Weigmann B, Bronson R, von Boehmer H (2011) Prevention of type 1 diabetes in mice by tolerogenic vaccination with a strong agonist insulin mimetope. J Exp Med 208:1501–1510. doi:10.1084/jem.20110574
Gottschalk RA, Corse E, Allison JP (2010) TCR ligand density and affinity determine peripheral induction of Foxp3 in vivo. J Exp Med 207:1701–1711. doi:10.1084/jem.20091999
Kretschmer K, Apostolou I, Hawiger D, Khazaie K, Nussenzweig MC, von Boehmer H (2005) Inducing and expanding regulatory T cell populations by foreign antigen. Nat Immunol 6:1219–1227
Haxhinasto S, Mathis D, Benoist C (2008) The AKT-mTOR axis regulates de novo differentiation of CD4+ Foxp3+ cells. J Exp Med 205:565–574
Sauer S, Bruno L, Hertweck A, Finlay D, Leleu M, Spivakov M, Knight ZA, Cobb BS, Cantrell D, O’Connor E, Shokat KM, Fisher AG, Merkenschlager M (2008) T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc Natl Acad Sci USA 105:7797–7802
Turner MS, Isse K, Turnquist HR, Morel PA (2014) Low TCR signal strength induces combined expansion of Th2 and regulatory T cell populations that protect mice from the development of type 1 diabetes. Diabetologia 57(7):1428–1436
Miskov-Zivanov N, Turner MS, Kane LP, Morel PA, Faeder JR (2013) The duration of T cell stimulation is a critical determinant of cell fate and plasticity. Sci Signal 6:ra97. doi:10.1126/scisignal.2004217
Wang RS, Saadatpour A, Albert R (2012) Boolean modeling in systems biology: an overview of methodology and applications. Phys Biol 9:055001. doi:10.1088/1478-3975/9/5/055001
Feinerman O, Veiga J, Dorfman JR, Germain RN, Altan-Bonnet G (2008) Variability and robustness in T cell activation from regulated heterogeneity in protein levels. Science 321:1081–1084. doi:10.1126/science.1158013
Liston A, Gray DH (2014) Homeostatic control of regulatory T cell diversity. Nat Rev Immunol 14:154–165. doi:10.1038/nri3605
Lan RY, Selmi C, Gershwin ME (2008) The regulatory, inflammatory, and T cell programming roles of interleukin-2 (IL-2). J Autoimmun 31:7–12. doi:10.1016/j.jaut.2008.03.002
Feinerman O, Jentsch G, Tkach KE, Coward JW, Hathorn MM, Sneddon MW, Emonet T, Smith KA, Altan-Bonnet G (2010) Single-cell quantification of IL-2 response by effector and regulatory T cells reveals critical plasticity in immune response. Mol Syst Biol 6:437. doi:10.1038/msb.2010.90
Tang Q, Adams JY, Penaranda C, Melli K, Piaggio E, Sgouroudis E, Piccirillo CA, Salomon BL, Bluestone JA (2008) Central role of defective interleukin-2 production in the triggering of islet autoimmune destruction. Immunity 28:687–697. doi:10.1016/j.immuni.2008.03.016
Dendrou CA, Wicker LS (2008) The IL-2/CD25 pathway determines susceptibility to T1D in humans and NOD mice. J Clin Immunol 28:685–696. doi:10.1007/s10875-008-9237-9
Boyman O, Surh CD, Sprent J (2006) Potential use of IL-2/anti-IL-2 antibody immune complexes for the treatment of cancer and autoimmune disease. Expert opinion on biological therapy 6:1323–1331. doi:10.1517/14712598.6.12.1323
Garcia-Martinez K, Leon K (2010) Modeling the role of IL-2 in the interplay between CD4+ helper and regulatory T cells: assessing general dynamical properties. J Theor Biol 262:720–732. doi:10.1016/j.jtbi.2009.10.025 S0022-5193(09)00506-2 [pii]
Leon K, Garcia-Martinez K, Carmenate T (2013) Mathematical models of the impact of IL2 modulation therapies on T Cell dynamics. Front Immunol 4:439. doi:10.3389/fimmu.2013.00439
Butler TC, Kardar M, Chakraborty AK (2013) Quorum sensing allows T cells to discriminate between self and nonself. Proc Natl Acad Sci USA 110:11833–11838. doi:10.1073/pnas.1222467110
Mamchak AA, Manenkova Y, Leconet W, Zheng Y, Chan JR, Stokes CL, Shoda LK, von Herrath M, Bresson D (2012) Preexisting autoantibodies predict efficacy of oral insulin to cure autoimmune diabetes in combination with anti-CD3. Diabetes 61:1490–1499. doi:10.2337/db11-1304
Bresson D, Togher L, Rodrigo E, Chen Y, Bluestone JA, Herold KC, von Herrath M (2006) Anti-CD3 and nasal proinsulin combination therapy enhances remission from recent-onset autoimmune diabetes by inducing Tregs. J Clin Investig 116:1371–1381. doi:10.1172/jci27191
Chatenoud L, Thervet E, Primo J, Bach JF (1994) Anti-CD3 antibody induces long term remission of overt autoimmunity in nonobese diabetic mice. Proc Natl Acad Sci USA 91:123–127
Fernandes P, Jain P, Moita C (2012) Training experimental biologists in bioinformatics. Advances in bioinformatics 2012:672749. doi:10.1155/2012/672749
Loman N, Watson M (2013) So you want to be a computational biologist? Nat Biotechnol 31:996–998. doi:10.1038/nbt.2740
Faeder JR, Hlavacek WS, Reischl I, Blinov ML, Metzger H, Redondo A, Wofsy C, Goldstein B (2003) Investigation of early events in Fc epsilon RI-mediated signaling using a detailed mathematical model. J Immunol 170:3769–3781
Lipniacki T, Hat B, Faeder JR, Hlavacek WS (2008) Stochastic effects and bistability in T cell receptor signaling. J Theor Biol 254:110–122. doi:10.1016/j.jtbi.2008.05.001
Chylek LA, Harris LA, Tung CS, Faeder JR, Lopez CF, Hlavacek WS (2013) Rule-based modeling: a computational approach for studying biomolecular site dynamics in cell signaling systems. Wiley Interdiscip Rev Syst Biol Med. doi:10.1002/wsbm.1245
Saez-Rodriguez J, Simeoni L, Lindquist JA, Hemenway R, Bommhardt U, Arndt B, Haus U-U, Weismantel R, Gilles ED, Klamt S, Schraven B (2007) A logical model provides insights into T cell receptor signaling. PLoS Comput Biol 3:e163. doi:10.1371/journal.pcbi.0030163
Klamt S, Saez-Rodriguez J, Lindquist JA, Simeoni L, Gilles ED (2006) A methodology for the structural and functional analysis of signaling and regulatory networks. BMC Bioinform 7:56. doi:10.1186/1471-2105-7-56
Narang V, Decraene J, Wong SY, Aiswarya BS, Wasem AR, Leong SR, Gouaillard A (2012) Systems immunology: a survey of modeling formalisms, applications and simulation tools. Immunol Res 53:251–265. doi:10.1007/s12026-012-8305-7
Germain RN, Meier-Schellersheim M, Nita-Lazar A, Fraser IDC (2011) Systems biology in immunology: a computational modeling perspective. Annu Rev Immunol 29:527–585. doi:10.1146/annurev-immunol-030409-101317
Kirschner DE, Chang ST, Riggs TW, Perry N, Linderman JJ (2007) Toward a multiscale model of antigen presentation in immunity. Immunol Rev 216:93–118. doi:10.1111/j.1600-065X.2007.00490.x
Sadot A, Sarbu S, Kesseli J, Amir-Kroll H, Zhang W, Nykter M, Shmulevich I (2013) Information-theoretic analysis of the dynamics of an executable biological model. PLoS ONE 8:e59303. doi:10.1371/journal.pone.0059303
Faeder JR, Blinov ML, Hlavacek WS (2009) Rule-based modeling of biochemical systems with BioNetGen. Methods Mol Biol 500:113–167. doi:10.1007/978-1-59745-525-1_5
Zhang F, Angermann BR, Meier-Schellersheim M (2013) The simmune modeler visual interface for creating signaling networks based on bi-molecular interactions. Bioinformatics 29:1229–1230. doi:10.1093/bioinformatics/btt134
Lizée G, Overwijk WW, Radvanyi L, Gao J, Sharma P, Hwu P (2013) Harnessing the power of the immune system to target cancer. Annu Rev Med 64:71–90. doi:10.1146/annurev-med-112311-083918
Acknowledgments
JRF and NMZ acknowledge funding from NIH grant P41 GM103712 and NSF Expeditions in Computing Grant (award 0926181). WFH was supported by NIH Grant T32 AI089443.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Morel, P.A., Faeder, J.R., Hawse, W.F. et al. Modeling the T cell immune response: a fascinating challenge. J Pharmacokinet Pharmacodyn 41, 401–413 (2014). https://doi.org/10.1007/s10928-014-9376-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-014-9376-y