Abstract
Patients that are exposed to biotechnology-derived therapeutics often develop antibodies to the therapeutic, the magnitude of which is assessed by measuring antibody titers. A statistical approach for analyzing antibody titer data conditional on seroconversion is presented. The proposed method is to first transform the antibody titer data based on a geometric series using a common ratio of 2 and a scale factor of 50 and then analyze the exponent using a zero-inflated or hurdle model assuming a Poisson or negative binomial distribution with random effects to account for patient heterogeneity. Patient specific covariates can be used to model the probability of developing an antibody response, i.e., seroconversion, as well as the magnitude of the antibody titer itself. The method was illustrated using antibody titer data from 87 male seroconverted Fabry patients receiving Fabrazyme®. Titers from five clinical trials were collected over 276 weeks of therapy with anti-Fabrazyme IgG titers ranging from 100 to 409,600 after exclusion of seronegative patients. The best model to explain seroconversion was a zero-inflated Poisson (ZIP) model where cumulative dose (under a constant dose regimen of dosing every 2 weeks) influenced the probability of seroconversion. There was an 80% chance of seroconversion when the cumulative dose reached 210 mg (90% confidence interval: 194–226 mg). No difference in antibody titers was noted between Japanese or Western patients. Once seroconverted, antibody titers did not remain constant but decreased in an exponential manner from an initial magnitude to a new lower steady-state value. The expected titer after the new steady-state titer had been achieved was 870 (90% CI: 630–1109). The half-life to the new steady-state value after seroconversion was 44 weeks (90% CI: 17–70 weeks). Time to seroconversion did not appear to be correlated with titer at the time of seroconversion. The method can be adequately used to model antibody titer data.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein therapeutics, including monoclonal antibodies, enzymes, proteins, and peptides, play an increasing role in the pharmaceutical marketplace. Despite the fact that many proteins use sequence homologies identical to or near identical their endogenous counterparts, the development and magnitude of immune response, i.e., anti-protein antibodies, continues to be of concern. Such antibodies can be of no consequence, can neutralize the therapeutic effect of the exogenous protein, or can enhance the effect of the exogenous protein. Sometimes the clinical effects of such proteins are life-threatening and require acute intervention. van Regenmortel et al. [1] present a review on the biological mechanisms of protein immunogenicity.
Not all patients form anti-protein antibodies to an exogenously administered protein. Schellekens [2] reviewed the factors involved in immunogenicity, which include sequence variation, degree of glycosylation, formulation, contaminants and impurities, route of administration, dose, length of treatment, assay technologies, patient characteristics, and unknown factors. Quantification of the degree of immunogenicity is defined using a tiered approach [3]. In the first tier, whether or not a patient has seroconverted, i.e., formed antibodies to the protein, is assessed using an immunoassay format. Second tier assays further characterize the nature of these antibodies in seroconverted patients. It is the analysis of data arising from the first tier that is of interest in this manuscript.
Once it is determined that a patient has seroconverted, an immunoassay, such as enzyme-linked immuno-sorbent assay (ELISA), is often used to assess the extent of the immune response to the therapeutic protein. Antibody titers provide a measure of the degree of antibodies present in an individual and are often expressed as the largest serum dilution that still gives a positive result. For example, a titer of 800 indicates the test still shows positive when the sample is diluted 800-fold, i.e., 1 part patient serum to 799 parts diluent while at greater dilutions the test is negative. Antibody titers are a surrogate for the magnitude of the immune response and are considered “quasi-quantitative” in nature as they reflect antibody protein concentration as well as antibody affinity. Antibody titers are influenced by the assay conditions and sensitivity. Of interest would be to determine what factors, e.g., weight, dose, etc., are predictive of the degree of antibody titer in a patient. To do so in a quantitative manner requires statistical modeling, which has not been reported in the literature to date. The purpose of this communication is to report a new methodology for the analysis of antibody titers collected as part of a clinical development program conditional on a patient seroconverting.
Methodology
The development of specific antibodies to a foreign protein by an otherwise immunologically naïve individual is called seroconversion. The magnitude of antibody response to foreign protein is assessed using antibody titers, which when using a 2-fold dilution scheme starting with the minimum dilution 1/100 take the form <100, 100, 200, 400, 800, 1600, etc. Modeling such data is not immediately intuitive because they are not exactly continuous and are more informative than a typical ordinal data type. Therefore, models based on continuous random variables are not applicable and, although ordinal models may be applicable, loss of information and potential loss of power would be expected to occur in the analysis. The problem is further complicated when it is realized that some patients that are exposed to foreign proteins never develop antibodies; hence, the population consists of seroconverters and nonseroconverters. One question of interest is what are the patient-specific factors that are important in quantifying the magnitude of antibody response in those patients that develop antibodies and what patient-specific factors distinguish seroconverters from nonseroconverters? The purpose of this report is to address the first problem—the conditional seroconversion problem. Further research is needed to solve the second problem, as will be shown.
The conditional seroconversion issue can be solved by realizing that antibody titer data take the form of a geometric sequence. All geometric sequences can be expressed as
where Y is the variable, a is the scale factor, b is the common ratio, and n is the exponent. For arbitrary detection limit denoted as LOQ, the scale factor (a) equals LOQ/2 and the common ratio (b) equals 2. For example, for an assay having a detection limit of 100, antibody titers can be expressed using a scale factor of 50 and a common ratio of 2, i.e.,100 can be expressed as 50 × 21, 200 as 50 × 22, 400 as 50 × 23, etc. Antibody titers less than the detection limit are problematic as they are technically left-censored. However, if it can be assumed that n = 0 for antibody titers less than the LOQ, then the problem becomes finding a suitable model for a series of positive integers or count data.
Count data is usually fit by either the Poisson or negative binomial distribution. The Poisson distribution has the form
where p(Y = y) is the probability the realized value of Y equals y and λ is the population mean. With the Poisson distribution the variance of the distribution equals the mean. To account for situations where the variance is larger or smaller than the mean, the negative binomial distribution is used. The negative binomial distribution takes the form
where Γ(.) is the gamma function. Estimates of λ and k can be done using maximum likelihood by maximizing the joint probability distribution.
Fitting count data typically runs into two problems, however, an over-excess of zeros and a variance that does not equal the mean (overdispersion or underdispersion) [4]. Analysis of the antibody titer data from clinical data clearly show an excess of zeros and possible overdispersion at most of the time points examined. To account for an excess of zeros, two types of models may be used, a zero-inflated model or a hurdle model [4]. The zero-inflated Poisson (ZIP) distribution is
where p is the probability of an excess zero and λ is the mean of the non-zero responses that can be modeled using a log-link function λ = exp(θz). In Eq. 4, p, the probability of seroconversion, can be modeled as a logistic function
where x is a set of predictor variables. It should be noted that the same or a different set of predictors can be used to model p and λ; they are not necessarily the same.
To account for overdispersion (where the variance exceeds the mean) or underdispersion (where the variance is less than the mean) the negative binomial distribution be used. The zero-inflated negative binomial (ZINB) distribution can be written as
where k is the overdispersion parameter. As k → ∞, the negative binomial part of the distribution approaches the Poisson distribution in the limit, i.e.,
The ZIP model accounts for excess zeros only, while the ZINB model allows for both excess zeros and overdispersion.
A competing model to account for excess zeros is the hurdle model [6]. Hurdle models are thought of as a two-part model. The first part is the typical Bernoulli outcome, the patient experiences the event or doesn’t experience the event. The second part is that once the patient experiences the event, a truncated-at-zero count model is used to model the positive, nonzero outcomes. Hence, the hurdle model is a model in which separate and distinct processes are used to account for the zero and non-zero responses. The basic form of the hurdle model is
where f1(.) is the probability distribution function (pdf) of the Bernoulli part of the model and governs the hurdle part of the model, f1(0) is the probability of a zero response for the first pdf, and f2(.) is the pdf of the response once the hurdle has been crossed. For the Poisson hurdle model, the model can be written as
Further complicating matters is that antibody titer data are longitudinal in nature since in clinical trials antibody titers are measured serially over the course of the trial. Therefore, it is also necessary to account for between-patient heterogeneity in both the probability of the event and the mean non-zero response since not all patients will seroconvert at the same time nor will they have the same magnitude of titer after seroconversion. The inclusion of between-patient variability in the model can be accounted for modeling p and λ as a random effect in the model. Assuming that p and λ can be modeled as a simple linear model of the predictors x and z, respectively, p and λ can be expressed as
where p(Y = 0)i now represents the patient-specific probability of a zero-response or threshold for the ith patient from seronegative to seropositive, λi is the mean patient-specific non-zero response, and the ηs are the patient-specific deviations from the population mean parameters that are independent, identically normally distributed with mean 0 and variance ω2. Predicted values can then be defined as
In this manner seroconversion is viewed as crossing some threshold. Below the threshold, patients have no titers but above the threshold, titers to the foreign protein develop and are dependent on λ.
Application: analysis of anti-Fabrazyme titers
Trial designs
Fabrazyme® (recombinant human α-galactosidase A enzyme, r-hαGAL) is an enzyme replacement therapy approved for the treatment of patients with Fabry disease, an X-linked lysosomal storage disorder with multisystemic effects. Patients on chronic administration of Fabrazyme can develop IgG antibodies to Fabrazyme. Of interest would be to determine those patient characteristics that might influence the development of antibodies to Fabrazyme.
For this analysis, antibody titer data were pooled from 5 clinical trials with Fabrazyme:
-
Study AGAL-1-002-98 was a Phase 3 multicenter, placebo-controlled, double-blind, randomized study of the safety and efficacy of r-hαGAL in male or female patients with Fabry disease 16 years or older having no prior treatment with Fabrazyme having low endogenous enzyme activity. Patients received approximately 1 mg/kg (0.9–1.1 mg/kg) of Fabrazyme or placebo every 2 weeks for 20 weeks for a total of 11 infusions of study medication. Patients in Study AGAL-1-002-98 were continued into an open-label extension study (Study AGAL-005-99) wherein patients received approximately 1 mg/kg Fabrazyme (0.9–1.1 mg/kg) every other week for 54 months.
-
Study AGAL-016-01 was an open-label, multi-center, multi-national study designed to examine the safety, efficacy and pharmacokinetics of Fabrazyme therapy in pediatric patients 7–15 years old with Fabry disease. Fabrazyme was administered 1.0 mg/kg every 2 weeks for 48 weeks.
-
Study AGAL-017-01 was an open-label, multi-center study designed to examine the safety, efficacy and pharmacokinetics of 1.0 mg/kg Fabrazyme followed by low dose maintenance therapy (0.3 mg/kg) in male patients ≥16 years old with Fabry disease. Patients received Fabrazyme 1.0 mg/kg every 2 weeks for approximately 6 months followed by Fabrazyme 0.3 mg/kg every 2 weeks for approximately 18 months.
-
Study AGAL-007-99 was a multi-center, open label study of the safety and efficacy of Fabrazyme in male Japanese patients with Fabry disease 16 years or older with no prior treatment with Fabrazyme. Patients received approximately 1 mg/kg (0.9–1.1 mg/kg) Fabrazyme intravenously every 2 weeks for 20 weeks.
These studies comprise a range of ages (pediatric and adults) and races (Japanese vs. Western) with somewhat similar dose intensities.
Antibody titer analysis
Antibody titers were measured using an r-hαGAL specific ELISA and were confirmed by a radioimmunoprecipitation (RIP) assay [7]. Two-fold serial dilutions of patient sera at an initial dilution of 1/100 were analyzed to obtain an endpoint titer. All assays utilized an enzyme conjugated monoclonal antibody to human IgG as the detector reagent, except for Studies AGAL-1-002-98, AGAL-005-99 and AGAL-007-99 which utilized an enzyme conjugated polyclonal antibody specific for human IgG. Patients with reactivity in the screening ELISA and a positive RIP confirmatory assay were classified as seroconverters, otherwise the patients did not develop antibodies and were not seroconverters. Patients that developed an antibody response were further analyzed in ELISA for endpoint titers.
Statistical analysis
Because of the potential problem regarding the distribution of the random effects when seroconverters (which are assumed normally distributed) were analyzed with non-seroconverters (which are assumed to be degenerate centered on 0), only seroconverters were analyzed. Further, females were excluded from the analysis female Fabry patients are heterozygous and have significantly more residual endogenous enzyme activity compared to male Fabry patients and therefore the degree of antibody formation would be expected to be different between genders. Also, only three females were in the dataset and with such a small sample size testing for sex effects would not be entirely valid.
All data were analyzed using the NLMIXED procedure in SAS for Windows (Version 8.02, SAS Institute, Cary NC). The marginal likelihood was estimated using Gaussian quadrature with 15–20 quadrature points and a quadrature tolerance of 0.002. Optimization was done using the Newton–Raphson algorithm.
To analyze the data, a base model was first established. The base models tested were Poisson, negative binomial, ZIP, ZINB, Poisson hurdle, and negative binomial hurdle model. The model with the best Akaike Information Criteria (AIC) was used for further model development with additional covariates. To allow patients to eventually seroconvert at some time t, the base models used cumulative Fabrazyme dose as the only covariate in the model. All models minimized without errors. The AIC for the models were 9145.6, 8741.9, 6324.1, 6324.6, 6815.9, and 6595.2, respectively. The best model based on the AIC was the ZIP model. The ZINB model was close to the ZIP model but the estimate of k, the dispersion parameter, was 14046. As k → ∞, the negative binomial distribution becomes more Poisson-like. Hence, based on the AIC and value of the dispersion parameter, the model used for further development was the ZIP model with random effects.
Next, covariates were added to the ZIP model in a forward stepwise manner. The following covariates were tested on p: weight, race, age, assay effect, and study. The effect of the assay was treated as a constant for those patients analyzed using the monoclonal antibody once seroconversion occurred. The same covariates were tested on λ with the addition of average dose and time since seroconversion. Time since seroconversion was modeled as a linear function, quadratic function, and exponentially declining function. At each stage, each covariate was tested singly for significance. The most significant covariate at each stage was treated as the base model and the process was repeated with the remaining covariates. This process was repeated until no further covariates could be added to the model. A significance level of 0.01 based on the likelihood ratio test (LRT) was used. At the final stage, correlation among the random effects was tested by including a covariance term between η1 and η2. Goodness of fit was tested by residual analysis.
Results
A total of six seroconverters were removed prior to analysis because of inadequate sample collection and inadequate characterization of antibody titers after seroconversion. Two patients had all zero titers except for a single occasion when both patients had titers of 100. Two patients had only two observations in the entire study period. Two patients had a prolonged period of time when no antibody titers were measured and it was during this period of time that the patients seroconverted. Of the patients in the database, there were only 3 females. Since females typically have a different disease progression than males and are expected, because of their residual endogenous enzyme protein and activity, to generate antibody responses (i.e. antibody titers) differently than males, it was decided that including females in this study would possibly bias the model and that testing for patient sex as a covariate would not be an option in this analysis. Hence, after removal of females from the dataset, a total of 1,826 observations from 96 patients collected over a 276 week period remained.
Of the 96 patients, 87 seroconverted (91%) and 9 (9%) did not. Of the 87 seroconverted patients, the median number of observations per subject was 18 with a range of 5–34. Table 1 presents a demographic summary by study and across studies for seroconverters.
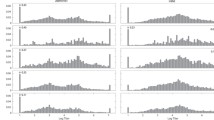
Figure 1 presents a scatter plot of the number of observations per week of therapy. The number of observations peaked at baseline where all patients had a baseline sample collected and declined to about 3–4 samples by 96 weeks therapy. Figure 2 presents a scatter plot of median antibody titers in patients that seroconverted by week of therapy for the first year of therapy. The median time to seroconversion was 8 weeks with a range of 2–26 weeks which translated to a median cumulative dose of 204 mg and corresponding range of 50–724.5 mg. In patients that eventually seroconverted, antibody titers after seroconversion ranged from 100 to 409,600. In general, titers appeared to peak to approximately Week 15 and then appeared to decline to a new steady-state value. Figure 3 presents a histogram of antibody titers after transformation. The observed exponents ranged from 0 to 12 with a distinct bimodal distribution centered at zero and six or seven.
The final model for the 87 seroconverters was a ZIP model of the form
where CDOSE was the cumulative dose administered in mg and DT was the time in weeks since seroconversion. Table 2 presents the estimates for the model parameters in Eq. 12. All parameters were precisely estimated with coefficients of variation <30%.
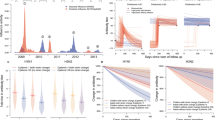
Figures 4 and 5 present the goodness of fit and residual plots under the final model. The model did an adequate job of predicting observed exponents with no obvious bias in the goodness of fit. The residuals were not exactly normally distributed (Anderson–Darling test: 14.6, P < 0.005) but were centered at zero and were symmetric around zero. No obvious outliers were discerned. Figures 6 and 7 present the individual goodness of fit plots in the analysis domain and in the original domain for six randomly chosen patients. The ZIP model with random effects adequately described the change from antibody free to seroconversion in those patients that eventually seroconvert. In sum, no gross deviations from the model assumptions were observed and the ZIP model appeared to do an adequate job at capturing the characteristics of the data.
The model identified cumulative dose as the only important predictor of seroconversion. Figure 8 presents the probability of seroconversion for each individual patient that seroconverted as a function of cumulative dose. There was an 80% chance of seroconversion when the cumulative dose reached 209 mg (90% confidence interval (CI): 194–226 mg). With a dose of 1 mg/kg to a 70 kg adult, seroconversion is expected to occur about three doses or 6 weeks after initiation of therapy.
After seroconversion, no factors were predictive of the degree of antibody titer, including race (Japanese vs. Western) and assay. For a typical patient who seroconverted the expected titer at the time of seroconversion was 2,795. Note that the model does not generate titers of the form 1/2, 1/4, 1/8, etc., but rather generates a more continuous response because the model for λ is continuous in nature. Antibody titers did not remain constant over time but decreased in an exponential manner from an initial magnitude to a new lower steady-state value. Models that tested time after seroconversion as a predictor in the model, using a linear or quadratic model, were not as effective at describing titers after seroconversion as an exponential model. After 1 year of therapy, titers were expected to be 1383 (90% CI: 1127–1640) in those patients that seroconverted. The expected titer after the new steady-state titer has been achieved was 870 (90% CI: 630–1110). The half-life to the new steady-state titer after seroconversion was 44 weeks (90% CI: 17–70 weeks). Lastly, time to seroconversion did not appear to be correlated with magnitude of seroconversion.
Discussion
This analysis presents for the first time a method to rigorously analyze antibody titer data from a statistical point of view. The method is to first transform the titer data based on a geometric series using a common ratio of two and a scale factor of 50 and then analyze the exponent using a zero-inflated or hurdle model assuming a Poisson or negative binomial distribution. These models have the advantage in that both the probability of seroconversion and expected titer can be modeled using a function that includes patient-specific covariates. To account for between-patient variability, random effects are included in the model and to account for the overdispersion in the data, either a zero-inflated or hurdle model can be used.
The factors identified as influencing titers are plausible and have a certain degree of face-validity to them. The fact that cumulative dose, under a fixed dosing regimen of dosing every other week, was identified as influencing the probability of seroconversion makes sense since in the genetic basis for disease there are patients that produce an abnormal enzyme and the enzyme replacement therapy exposes patients to the normal protein to which they are not naturally fully tolerized. Therefore repeated exposure to a conformationally different protein than their endogenous protein would suggest that formation of an antibody response seems likely. Further, dose and frequency of administration are known to influence seroconversion.
It should be pointed out, however, that cumulative dose is confounded with other factors that might be in fact be the real mechanism. These include time on therapy or number of infusions. It is impossible from these study designs to determine whether in fact these factors are more plausible. Also, that cumulative dose was the only factor identified as affecting the probability of seroconversion does not mean that it was the only factor. Other factors not assessed in this analysis, such as type of mutation, might affect seroconversion.
The other factor affecting the magnitude of titer was the time from seroconversion. Patients initially have their highest titers soon after seroconversion and then over time have a reduced response and lower titer with repeated Fabrazyme administration.
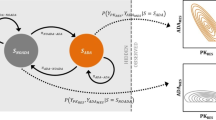
One limitation of the model is that it is conditional on a patient being identified as a seroconverter. The model does not classify patients as seroconverters or non-seroconverters; the analyst must identify those seroconverted patients a priori. In theory, these conditional models can be expanded to develop joint models that model both seroconverters and nonseroconverters. This problem is similar to cure models of disease wherein some patients do not respond to treatment and some do [8]. Let psi be the probability a patient seroconverts, U be a binary value to denote a seroconverter (U = 1) and nonseroconverter (U = 0), z be the set of parameters that differentiate seroconverters from nonseroconverters, and θ be the model parameters associated with x, then the full model for the ZIP model is
Similar expressions can be written for their negative binomial and hurdle model counterparts. The problem with modeling seroconverters with nonseroconverters simultaneously in this manner is that these models are mixture models that violate the assumption of the random effects being normally distributed. The mixture would necessarily consist of a random distribution (seroconverters) and a degenerate distribution with mass on zero (non-seroconverters). In other words, nonseroconverters by definition should have η = 0 for all random effects in the model but seroconverters have η ≠ 0. Current software based on maximum likelihood estimation cannot accommodate this situation. Such a model would require complex likelihoods involving mixture distributions that the NLMIXED procedure in SAS is not capable of handling. One possibility, however, is to use a Bayesian non-likelihood-based approach.
Besides being able to identify patient-specific factors that might influence antibody titers, the model also has other uses. Sometimes in a clinical trial patients have missing titer values (perhaps the patient did not show up for a blood draw) and that information is needed for interpretation of efficacy or safety data. Rather than interpolating the missing value, the model could be used to impute the antibody titers after drug administration. Under this scheme, the assay used to measure the titers is defined a priori. A random draw from a normal distribution is made with mean 0 and standard deviation σ(p). Using Eq. 12 and based on the cumulative dose the patient received at the time of the missing value, as well as the random draw that is substituted for η1, the probability of seroconversion is determined. If the value exceeds 0.5 the patient is classified as a seroconverter. Otherwise the patient is not a seroconverter and the titer is imputed to be 0. If the patient is a seroconverter, a random draw from a normal distribution having mean 0 and standard deviation σ(λ) is made as is another random draw from a normal distribution having mean 0 and standard deviation σ(θ1). Then using Eq. 12 the expected value λ is calculated and transformed to a titer value using 50 × 2λ. In this manner, imputed titers can be used for further analysis.
The geometric series conversion of antibody titer data is also useful in other analyses in which antibody titer might influence the dependent variable. Because of the geometric progression of antibody titer data their use as covariates in linear models is problematic because they are not continuous. Antibody titers can be treated as categorical variables but this loses the ordinal nature of the data. A better method is to use the exponent from the geometric series transformation in the linear model as a continuous covariate and while, although the integers are not exactly continuous, they are much more so than titers in the original domain.
In summary, a method was developed to analyze antibody titers that allow identification of those patient characteristics that may influence the formation of these antibodies conditional on the patient having seroconverted. The method is based on a transformation that allows standardized mixed effect generalized linear models to do the parameter estimation. One downside of the method is that prediction of who seroconverts and who doesn’t seroconvert cannot be discerned; the method is conditional on knowing who seroconverts and who doesn’t. One limitation of the method is its degree of empiricism and future research will look at using the transformation in combination with a mechanistic model for seroconversion. The advantage of the method is that no special algorithms are needed and that the transformation can be used to generate a value that can be used as a covariate in other types of analyses.
References
van Regenmortel MHV, Boven K, Bader F (2005) Immunogenicity of biopharmaceuticals: an example from erythropoetin. BioPharm Int 18:36–50
Schellekens H (2002) Bioequivalence and the immunogenicity of biopharmaceuticals. Nat Rev Drug Dis 1:457–462
Richards SM (2002) Immunologic considerations for enzyme replacement therapy in the treatment of lysosomal storage disorders. Clin Appl Immunol Rev 2:241–253
Cameron AC, Trivedi PK (1998) Regression analysis of count data. Cambridge University Press, Cambridge
Rose CE, Martin SW, Wannemuehler KA, Plikaytis BD (2006) On the use of zero-inflated and Hurdle models for modeling vaccine adverse event count data. J Biopharm Stat 16:463–481
Kianifard F, Gallo PP (1995) Poisson regression analysis in clinical research. J Biopharm Stat 5:115–129
Wilcox WR, Banikazemi M, Guffon N, Waldek S, Lee P, Linthorst GE, Desnick RJ, Germain DP, for the International Fabry Disease Study Group (2004) Long-term safety and efficacy of enzyme replacement therapy for Fabry disease. Am J Hum Genet 75:65–74
Corbiere F, Joly P (2007) A SAS macro for parametric and semiparametric mixture cure models. Comput Methods Programs Biomed 85:173–180
Acknowledgments
All authors are employees or past employees of Genzyme and hold stock in the company.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bonate, P.L., Sung, C., Welch, K. et al. Conditional modeling of antibody titers using a zero-inflated poisson random effects model: application to Fabrazyme® . J Pharmacokinet Pharmacodyn 36, 443–459 (2009). https://doi.org/10.1007/s10928-009-9132-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-009-9132-x