Abstract
It has been almost 30 years since C/EBPß was discovered. Seminal studies have shown that C/EBPß is a master regulator of mammary gland development and has been shown to control and influence proliferation and differentiation through varying mechanisms. The single-exon C/EBPß mRNA yields at least three different protein isoforms which have diverse, specific, context-dependent, and often non-overlapping roles throughout development and breast cancer progression. These roles are dictated by a number of complex factors including: expression levels of other C/EBP family members and their stoichiometry relative to the isoform in question, binding site affinity, post-translational modifications, co-factor expression, and even hormone levels and lactogenic status. Here we summarize the historical work up to the latest findings in the field on C/EBPß in the mammary gland and in breast cancer. With the current emphasis on improving immunotherapy in breast cancer the role of specific C/EBPß isoforms in regulating specific chemokine and cytokine expression and the immune microenvironment will be of increasing interest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
C/EBPß History
The single exon gene C/EBPß is a part of the six member CCAAT enhancer-binding protein gene family which also includes C/EBPα, C/EBPδ, C/EBPε, C/EBPɣ, and C/EBPζ which is now known as C/EBP homologous protein (CHOP). This family shares two highly conserved DNA binding and basic leucine zipper (bZIP) domains in the C-terminus, but differs greatly in the N-terminal regulatory regions [1]. This close homology allows for dimerization not only between isoforms but also between different family members and allows all family members to bind the same DNA sequences. C/EBPß is known as a master regulator of multiple tissue types/states such as autophagy [2], differentiation of the myeloid lineage [3], in the etiology of glioblastoma [4, 5], and as a transcriptional “cooperator” under the control of KDM3A in many cancer types [6]. As eloquently described by Zaret and Carroll, pioneer transcription factors have the ability to bind DNA to act in either a passive or active fashion. If functioning in a passive role, they bind complex regulatory sequences such as those found in enhancer elements long before that enhancer is activated thereby reducing the number of other factors required to bind before activation and thus “priming” that enhancer. If the factors are functioning in an active role they bind exposed regions in closed nucleosomal DNA and either directly recruit other proteins to the region, or indirectly recruit proteins by simply opening up the closed chromatin which allows those factors to find their exposed binding sites [7]. The C/EBPß gene encodes multiple isoforms of the bZIP pioneer transcription factor which have diverse functions in multiple tissues and cell types such as hematopoietic cells [8,9,10,11,12,13,14], hepatocytes [15,16,17], keratinocytes [18], adipocytes [19, 20], female reproductive tissues [21,22,23] and in the mammary gland [24,25,26,27,28]. These studies often have been related to development and tissue differentiation, but there are also many studies examining the regulation of the secretion of chemokines and cytokines. In fact, as discussed below C/EBPß was first identified as Nuclear Factor for IL-6 (NF-IL6), nuclear factors that specifically bound to a response element in the Interleukin- (IL-)6 gene.
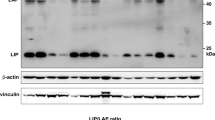
C/EBPß was first described by Isshiki et al.. as NF-IL6 based upon the observation that two distinct proteins bound to a 14 bp palindromic IL-1 response element in the IL-6 promotor [29] in SK-MG-4 human glioblastoma cells. Akira et al. further characterized NF-IL6 and noted that these two proteins were in fact likely different isoforms of the same protein [30]. Furthermore, they suggested that NF-IL6 was part of a family related to C/EBPα, which at the time was called C/EBP. Shortly later, Descombes and Schibler showed that indeed, three different protein isoforms were generated from a single exon gene in rat hepatocytes by leaky ribosomal scanning and that these were likely orthologs of Akira’s NF-IL6. They called the short ~20kd protein, Liver-enriched Inhibitory Protein (LIP) and the ~36kd protein, Liver-enriched transcriptional Activator Protein (LAP). They designated the full-length ~39kd protein LAP* or LAP-FL and found that that isoform was very poor at transactivation of target genes relative to the other two isoforms (Fig. 1). Furthermore, these investigators reported that LIP and LAP can form heterodimers, consistent with previous results showing LAP and C/EBP could also form heterodimers. They also showed that these heterodimers have an antagonistic effect on the expression of the albumin gene, and speculated that LAP accumulation in the rat liver during development might be related to differentiation. Unfortunately, at that time they lacked the experimental tools to directly test this hypothesis [31, 32]. Cao et al. then proposed a new nomenclature for the C/EBP-like proteins due to the clear homologies between what had been termed C/EBP, NF-IL6, and NF-IL6ß. C/EBP became C/EBPα, NF-IL6 became C/EBPß, and NF-IL6ß became C/EBPδ [19]. Here they showed that C/EBP family members were expressed in a number of tissues and could heterodimerize with one another. They also reported the first direct evidence that C/EBPß may play a role in tissue differentiation. In 3T3-L1 cells they observed an accumulation of RNA and protein polypeptides at specific time points during differentiation, and that C/EBPß levels increased after hormone stimulation. These observations led them to speculate that C/EBPß played a role in the terminal differentiation of adipocytes [19] and led to later work by others in this model which demonstrated that the ratio of C/EBPß isoforms was translationally controlled and determined cell fate [33]. Subsequently, direct interactions in cell free systems between C/EBPß and other proteins such as Nuclear Factor Kappa light chain enhancer of activated B cells (NF-κB), the Glucocorticoid Receptor (GR), and the Activator Protein- (AP-)1 factors Jun and Fos were reported [34,35,36,37]. Moreover, investigators began to uncover additional examples of C/EBPß’s involvement in the regulation of inflammatory cytokines and acute phase proteins such as Granulocyte-Colony Stimulating Factor (G-CSF), Tumor Necrosis Factor alpha (TNFα), IL-8, IL-4, and IL-1ß [35, 38,39,40,41,42], however, it took an additional 5 years after first publication before C/EBPß made it to the mammary gland (Fig. 2).
C/EBPß mRNA is translated into multiple protein isoforms. Schematic representation of C/EBPß mRNA and the three protein isoforms resulting from translation. Adapted from Zahnow. PMID:12052253 [111]
There have been a number of mouse models generated to study C/EBPß isoform specific effects in-vivo which have been utilized to make some unique observations in a number of tissues and may have utility for studies in the mammary gland. A C/EBPß∆uORF mouse generated by Wethmar et al, surprisingly revealed that the LAP* translation initiation site was necessary for expression of LIP in the livers and osteoclast populations in C/EBPß∆uORF mice compared to controls. The livers of these animals displayed increases in LAP expression and a complete loss of LIP expression when challenged by Lipopolysaccharide (LPS) [43]. A LIP knock-in (LIPki) mouse has also been made utilizing the endogenous C/EBPß promoter and was shown to have increased susceptibility to tumorigenesis, primarily B cell lymphomas and histiocytic sarcomas, and a negatively dysregulated metabolic profile [44, 45]. Furthermore, reduced expression of LIP in the C/EBPß∆uORF mouse has also been shown to increase lifespan in a similar manner to caloric restriction and Rapamycin treatment, which suggests that C/EBPß may be one target for treating aging related diseases as well [46]. These models highlight the importance of the exquisite balance in the translation of C/EBPß isoforms in-vivo and open the door to better understanding the intricacies of this balance by adapting these models to study the developing mammary gland. However, this most likely will require targeted rather than germline manipulation of C/EBPß isoforms.
C/EBPß Post-Translational Modifications
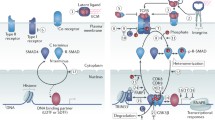
There are a diverse set of post-translational modifications (PTMs) which can be added to different residues within C/EBPß. These modifications can affect protein stability and dimerization, DNA binding, transcriptional output, co-factor association, and/or protein degradation. These modifications include phosphorylation, O-GlcNAcylation, acetylation, ubiquitination, methylations-(mono-, di-, and tri-), citrullination, and also sumoylation [47] (Fig. 3). The function of a limited number of these modifications has been elucidated to date as briefly summarized below. Further studies will be required to fully decipher the PTM code in C/EBPß.
C/EBPß model depicting isoform start sites and relative potential PTM sites. Isoform-specific N-terminus’ are shown above the bar depicting the C/EBPß protein primary structure. Methylation (green plus), phosphorylation (yellow circle), sumoylation (red diamond), and acetylation (pink triangle) are depicted in the relative sites they were identified, however many still require validation and studies to determine their functional importance. Adapted from Dittmar et al.. PMID: 30884312 [47]
Sequential phosphorylation of Thr-188 by cdk2 or MAPK followed by Ser-184 or Thr-179 by GSK3ß on C/EBPß dimers results in a conformational change to a scissor- like motif which allows the complex to bind to C/EBPß consensus sites in the DNA [48, 49]. These PTMs have also been shown to protect C/EBPß from calpain-dependent degradation and ubiquitination [50, 51]. Conversely, O-linked GlcNAcylation of neighboring residues Ser-180 or Ser-181 inhibits phosphorylation of these adjacent residues and subsequent C/EBPß binding to DNA [52].
There are also a number of residues which can be acetylated: Lys-39 acetylation by CBP/P300 increases transcription of target gene promoters linked to a luciferase reporter, while de-acetylation at that same site by HDAC1 reduced transcription [53, 54]. GCN5 and PCAF acetylate a cluster of lysine residues between Lys-98 and Lys-102 which potentiated glucocorticoid-mediated transcription and are required for adipocyte differentiation and inhibit the association of C/EBPß with HDAC1 [55]. Interestingly de-acetylation by HDAC1 at Lys-213, Lys-215, and Lys-216 were shown to be required for STAT5 induced expression of Id-1. HDAC de-acetylation of these sites was shown to stabilize C/EBPß hetero- and homodimers and increase their DNA binding affinities [56]. Thus there can be reciprocal effects of acetylation dependent on the specific site modified.
C/EBPß can also be methylated. Pless et, al showed that methylation, in contrast to acetylation, at Lys-39 by G9a represses C/EBPß dependent transcription [57]. That same group also reported that dimethylation at Arg-3 by PRMT4/CARM1 reduced recruitment of SWI/SNF to C/EBPß and inhibited adipocyte differentiation [58].
There are also multiple sites where C/EBPß can be sumoylated [59,60,61,62,63]. Sumoylation of Lys-132 in mice (Lys-173 in human) by SUMO2/3 is integral to the repression of LAP* induced transcription of cyclin D1 [59, 60]. Furthermore, sumoylation of the transactivation domain of the LAP isoforms lead to a relief of LAP-induced repression of c-myc expression in murine T-lymphocytes but this PTM did not affect expression of IL-4 [62]. Finally, sumoylation of Lys-133 by PIAS1 is essential for adipocyte differentiation by promoting ubiquitination and subsequent degradation of C/EBPß during late-stage adipogenesis [63].
Recently, studies by Dittmar et al. have illustrated just how elegant and complex the PTM code is for C/EBPß. This was accomplished by tiling short C/EBPß peptides with or without PTMs across an array of HeLA cell nuclear extracts. By subjecting those peptide conjugates to mass spectrometry these investigators were able to identify over 1300 different protein interactions. While each interaction must still be validated experimentally, these studies also showed that C/EBPß is at the center of a network of proteins facilitating promoter and enhancer regulation. Furthermore, that C/EBPß assists in many facets of RNA regulation including pre- and post-transcriptional modifications as well as in RNA Polymerase II (PolII) initiation and pausing, splicing, degradation, polyadenylation, and nuclear export [47]. Very little is known about the function of these C/EBPß PTMs during mammary gland development or about their hormonal regulation. With the advent of CRISPR/CAS9 base editing techniques, it should be possible to introduce both gain- and loss- of-function mutations into specific PTM sites in the C/EBPß gene in a high-throughput manner [64, 65].
C/EBPß Protein/Protein Interactions
There is a large body of work showing that C/EBPß isoforms interact with a number of other transcription factors and other co-regulators at a diverse range of genes to alter transcriptional output either positively or negatively [35, 66,67,68,69,70,71,72,73,74,75,76]. For example, the transcriptional repressor YY1 exerts differential effects with LIP on two target genes. At the ß-casein promoter there is a binding site for YY1 which interacts with LIP to negatively regulate transcription in the absence of glucocorticoid signaling [66, 73], LAP has also been shown to interact with cyclin D1 here to drive expression of ß-casein and promote differentiation of mammary epithelial cells [68]. However, in the CXCR4 promoter, LIP but not LAP homodimers relieve YY1 induced transcriptional repression. Furthermore, LIP/LAP heterodimers were shown to have reduced DNA binding affinity at that site compared to LIP homodimers indicating that LIP is required to relieve repression and drive expression of CXCR4 [76].
In the IL-6 and IL-8 promoters there are NF-κB binding sites proximal to C/EBPß binding sites. Here both C/EBPß isoforms were shown to have a stimulatory effect over basal NF-κB mediated expression, but LAP binding increased expression more than LIP [35, 39, 69]. Conversely, CCL2 expression requires LIP rather than LAP and does not rely on a C/EBPß/NF-κB interaction. Here the investigators showed that C/EBPß is likely interacting with an AP-1 family member to drive transcription because deletion of the AP-1 site which is proximal to the C/EBPß binding site reduced CCL2 expression ~20 fold relative to the wild-type promoter while deletion of the NF-κB site had little effect. These authors also showed that the exogenous addition of HIV-1 tat physically interacts with C/EBPß to increase CCL2 expression while the addition of SMAD3 which is activated following TGFß1 stimulation dramatically reduced that interaction and CCL2 expression [77]. Other investigators have also demonstrated that C/EBPß interacts with c-Jun and have shown that a LIP/c-Jun interaction can stimulate expression of PRB significantly more than a LAP/c-Jun interaction [74]. Finally, additional studies also have shown that C/EBPß can activate E2F regulated genes by associating with E2F1 and E2F2. This association then recruits CBP/p300 to acetylate histone 4 which then opens the chromatin to a transcriptionally active state [71].
As shown previously, these protein-protein interactions are often dependent upon specific PTMs which can regulate binding specificity. These marks can be dynamically placed as cell states change which in-turn regulates binding partner affinity. Thus, understanding the regulation of both PTMs and protein-protein interactions will be necessary to fully elucidate the functional importance of either interaction individually.
C/EBPß in the Mammary Gland
C/EBPß isoforms have been characterized during rat and mouse mammary gland development long before they were ever examined in breast cancer. Much of the development of the mammary gland can be attributed to downstream effects of C/EBPß isoforms. Doppler et al. were the first to show that C/EBPß was expressed in an immortalized mouse Mammary Epithelial Cell (MEC) line, HC11, and that it bound specific sequences in the ß-casein gene promoter and enhanced casein gene expression [28]. Subsequently, Raught et al. also reported in HC11 cells the addition of glucocorticoids relieved LIP induced repression of ß-casein gene expression [73] and allowed lactogenesis to occur. These studies indicated that C/EBPß likely associates with GR, and that this association is important during mammary gland development. Another study by these same investigators examined C/EBPß expression across a panel of human and murine tumor models and normal tissues and found that LIP was only expressed at high levels in neoplastic tissue, not the surrounding normal tissue. This suggested that perhaps LIP had a specific role in breast cancer which was different from that in normal development [78]. Subsequently, it was reported by Robinson et al., that C/EBPß was essential for normal mammary gland development by regulating the proliferation and differentiation of the MECs during development and that C/EBPß−/− mammary glands exhibited impaired ductal outgrowth and alveolar defects during pregnancy [24]. In a companion manuscript, Seagroves et al. then showed that C/EBPß was indeed essential for not only ductal morphogenesis and the proliferation of lobuloalveolar secretory units, but also that C/EBPß was required for functional differentiation of these cells as milk production was inhibited or absent in C/EBPß−/− mammary glands [25]. It also was subsequently reported by Gigliotti and DeWille that C/EBPß mRNA expression was influenced by lactation status. In late-pregnant mice LAP expression is high, at parturition LAP levels decline greatly and are almost undetectable until weaning when the gland undergoes involution and LAP levels rise again [79]. C/EBPß was also shown to control cell fate decisions in the developing mammary gland by influencing expression of cell-type specific markers such as PR and potentially IGFII which then drive proliferation and terminal differentiation as determined by Seagroves et al. [26]. In addition, C/EBPß is important for stem cell proliferation and luminal cell fate commitment as reported by Lamarca et al.. These investigators used limiting dilution transplant experiments and Flow Assisted Cytometric Sorting (FACS) analysis to show that C/EBPß−/− MECs regenerate the mammary gland at a substantially reduced frequency as compared to wild-type cells, and that the repopulated glands showed a reduction in luminal progenitor cells and an increase in fully differentiated luminal cells [27]. Studies by Eaton et al. in human MECs (HMECs) showed that LAP1 or LAP* was found exclusively in normal MECs while LAP2 or LAP was found only in dividing normal or neoplastic cells. Their data suggested that LAP was activating genes which push cells to divide and hypothesized that cyclin D1 might be a potential candidate [80]. Liu et al. followed this work and showed that all three C/EBPß isoforms can bind to cyclin D1, however, only LAP* was transactivated by cyclin D1 and was able to drive differentiation as evidenced by the expression of ß-casein and whey acidic protein (WAP) in response to lactogenic hormones in mouse MEC cell lines. Dearth et al. then showed that LAP was the predominantly expressed isoform in culture in HC11 cells and that overexpression of either LIP or LAP resulted in an increase in proliferation in-vitro. However, the cells which overexpressed LIP failed to differentiate after the switch to a differentiation protocol. These studies showed that undifferentiated MECs with a high expression of LIP are poorly differentiated.
Thus in many ways the previous studies have similarities to the hormone receptor (HR) negative breast cancer samples examined earlier by Zahnow et al. [81, 82]. These latter authors also showed that involuted mouse mammary glands from mice with overexpression of rat C/EBPß-LIP driven by a targeted WAP expression construct designated WAP-LIP-WAP contained numerous hyperplasias. Some of these lesions progressed into high-grade neoplasias which indicates that LIP expression supports proliferation and could potentially initiate a growth cascade if cells are exposed to further oncogenic hits [83]. Bundy and Sealy reported that ectopic overexpression of LAP but not LAP* in “normal” MCF10A cells was sufficient to cause a gain in the expression of mesenchymal properties such as anchorage-independent growth and to cause them to undergo an Epithelial-to-Mesenchymal-Transition (EMT) and express canonical EMT markers in 2D cultures in-vitro [84]. LAP overexpression in those same cells conferred an EGF-independent growth advantage, and disrupted lumen formation and acinar structures when cells are grown in 3D cultures in matrigel as reported by Bundy et al. [85]. Conversely Miura et al. found that overexpression of LIP in SCp2 cells drove the EMT phenotype in 3D cultures in-vitro [86]. The induction of EMT in those previous experiments with MCF10A cells, at least in part, was due to LAP-induced transcription of proIL-1ß as reported by Russel et al.. Instead of being cleaved and secreted normally, this variant remained in its pro-form and was translocated to the nucleus where it bound tightly to chromatin in distinct locations that were highly correlated with pro-metastasis gene regions. They also showed that proIL-1ß was expressed in a handful of ER− primary breast cancer samples, two of which were Human Epidermal Growth Factor Receptor 2 (HER2)+ and one was PR+, but the sample size was too small to make any definitive correlations about specific breast cancer subtypes [87]. Overall, these studies show that C/EBPß expression is evident throughout mammary gland development and definitively establish C/EBPß as a master regulator of that process (Fig. 4). However, there are still significant questions that remain to be answered with the development of new technologies.
Expression of C/EBPß during early mammary gland development. A model for expression of markers representing different cell types in the early development of the mammary gland. C/EBPß-expressing cells (pink) are scattered throughout the developing mammary bud, mainly in the central epithelial cells. In the terminal end buds present during ductal elongation, C/EBPß is expressed in both the highly proliferative cap cell layer and the body cells. In a quiescent, mature gland, C/EBPß expression is found in a punctate pattern along the ducts in both the luminal MECs and myoepithelial cells. Expression appears to be localized to the tall columnar-like MECs, rather than the round cuboidal-like MECs that either express PR/ERα (red cells), or are proliferating (green cells). Some MECs along the ducts express none of these markers (dark blue cells). Reproduced and adapted from Grimm and Rosen, PMID:14635794 [112]
C/EBPß in Breast Cancer
Utilizing the ratio of C/EBPß isoforms as specific biomarkers for breast cancer progression appears to be attractive in theory, but has been difficult to put into clinical practice. This is because C/EBPß isoforms have diverse functions and there are currently no good tools to independently examine LIP and LAP isoform expression from small amounts of patient samples. Thus, understanding the pathways and networks that are regulated by C/EBPß in breast cancer and which patients are most likely affected is still very important because FDA approved drugs exist which potentially can modulate the LIP:LAP ratio [88]. Zahnow et al. were the first to explore the role of C/EBPß in breast cancer. By examining quantitative Western blots from human tumors these investigators discovered that high LIP expression was associated with poorly differentiated, highly proliferative, primarily estrogen receptor- (ER-) and progesterone receptor- (PR-) negative disease [82]. Subsequently, Gomis et al. examined C/EBPß isoform expression in patient samples obtained from pleural effusions from patients with multiple metastases. These samples were almost entirely HR+ but some expressed very high levels of LIP. These authors observed that as the ratio of LIP to LAP (LIP:LAP) increased, the cytostatic effects of Transforming Growth Factor ß (TGFß) were abrogated. LAP induced expression of p21CIP1 after TGFß stimulation by associating with a FoxO/Smad protein complex on the p21CIP1 promoter. Conversely LAP inhibited expression of c-MYC by associating with an E2F4/5/Smad complex on the c-MYC promoter. As the LIP:LAP ratio was increased p21CIP1 expression was diminished and c-MYC expression was activated switching TGFß signaling from cytostatic to pro-proliferative. This was a seminal finding since c-MYC has long been known to be a potent oncogene in breast cancer [70, 89].
In an effort to uncover a potential mechanism for LIP overexpression in patients, Baldwin et al. found that epidermal growth factor receptor (EGFR) signaling led to increased phosphorylation of the RNA binding protein CUG-BP1 or CELF1 which can bind two regions between the LAP and LAP* translation initiation sites to essentially block initiation at those sites [90]. These investigators observed that phosphorylation promotes increased binding of CELF1 to C/EBPß mRNA and thus increases expression of LIP [91]. These findings provided the foundation for studies by Arnal-Estape et al. who obtained HER2+ breast cancer samples from pleural effusions and showed that HER2 overexpression led to an increase in HER2 signaling. This, in turn led to an increased LIP:LAP ratio by the same mechanism as that observed previously for EGFR signaling. Furthermore, these authors found that treating HER2+ cell lines with trastuzumab, an FDA approved HER2 targeted monoclonal antibody therapy, decreased the LIP:LAP ratio and reduced markers associated with proliferation and survival [92, 93]. Gustafson et al. then found that Ha-Ras transformed MCF10A cells increased the expression of LIP. Furthermore, a similar effect was observed following either the overexpression of Ha-Ras, or by expression of constitutive isoform specific expression vectors.
In other studies, C/EBPß expression in general reduced the expression of breast tumor suppressor gene SIM2s, but LIP overexpression resulted in the greatest decrease in the SIM2s expression [94]. Park et al. showed that LIP binding to the CXCR4 promoter in human breast cancer cell lines relieved YY1 mediated repression, but that LAP alone could not achieve the same effect. Either LIP/LIP homodimers or LIP/LAP heterodimers were required to displace YY1 to allow for transcription of CXCR4 which is known to modulate breast cancer cell migration [76]. Johansson et al. found that TGFß induced microRNA- (miR-)155 expression in mouse mammary tumor models substantially reduced the level of total C/EBPß expression. This reduction promoted a pro-EMT response to TGFß which was correlated with a loss of expression of junction proteins [95]. The same group went on to suggest that C/EBPß should be viewed as a predictor of Overall Survival (OS) in patients with breast cancer due to the increase in CD45+, CD3+, and CD4+ lymphocytes in a C/EBPß−/− 4T1 mouse mammary tumor model. Interestingly they also noticed an overall downregulation in chemokine expression in the C/EBPß−/− cells as measured by microarray [96].
Willis et al. did an in-silico analysis of publicly available datasets to try to determine a transcription factor footprint common to TNBC searching for druggable targets. Although C/EBPß binding sites were one of the most commonly found footprints, C/EBPß was ultimately excluded as a suitable drug target due to the ubiquitous expression of C/EBPß in most tissues [97]. A drawback to in-silico analyses of C/EBP proteins is that all C/EBP family members can bind the same DNA sequences, thus to determine if the site is indeed specific for C/EBPß these analyses must be overlaid with ChIP data. In another in-silico analysis Jinesh et al. mined publicly available datasets to determine if the expression from the large cluster of miRNAs on chromosome 19 (C19MC) showed any correlation with the different human breast cancer subtypes. They observed that C19MC was positively correlated with basal-like TNBC and that it was tightly correlated with a C/EBPßhigh signature [98].
Recently Li et al. showed that Myeloid Derived Suppressor Cell (MDSC) recruitment to tumors in two TNBC mouse mammary tumor models was significantly decreased after inhibition of aerobic glycolysis. This inhibition resulted in an increase of intracellular AMP which can activate AMPK leading to an increase in autophagy. They showed that after the induction of autophagy, LAP levels decreased and thus LAP target genes G-CSF and Granulocyte-Macrophage Colony Stimulating Factor (GM-CSF) expression decreased as well ultimately resulting in a decrease in MDSC recruitment [99].
Salaroglio et al. also suggested a positive role for LIP in TNBC. Using a panel of human and mouse breast cancer cell lines in-vitro and in-vivo, they found that increasing levels of LIP were able to abrogate doxorubicin resistance by reactivation of immunogenic cell death [100]. Thus, as an increasing number of studies are published it becomes increasingly evident why it is so important to understand C/EBPß biology in breast cancer. Clearly, there are many ways C/EBPß expression can either directly affect tumor cells, or indirectly to affect the surrounding microenvironment. Thus, understanding the complexity of these interactions is vital before attempting to pharmacologically modulate the LIP:LAP ratio.
C/EBPß Regulation of Chemokines and Cytokines
Since it was first discovered, C/EBPß has been associated with cytokine and chemokine expression [29]. The list of discovered chemokines and cytokines whose expression is modulated by C/EBPß is extensive and growing. As mentioned previously, C/EBPß was discovered as a group of proteins which bound to an IL-1 response element in the IL-6 promoter and drove IL-6 and IL-8 expression [29, 30, 35, 39]. These discoveries were followed soon thereafter by a number of groups showing that C/EBPß could regulate IL-1ß [38, 101], IL-4 [42], Chemokine (C-C motif) Ligand (CCL)3 [102], TNFα [103], GM-CSF [104], and G-CSF [40]. It was a few years later before it was shown that IL-10 could be regulated by C/EBPß [105], and shortly after a study was published that showed CCL5 was also likely regulated by C/EBPß, however, these studies did not provide any evidence of direct regulation [106]. Calonge et al. found one site in the Chemokine (C-X-C motif) Ligand (CXCL)12 promoter which interacted with C/EBPß and could activate expression [107]. C/EBPß has also been shown to regulate the expression of Receptor Activator of Nuclear Factor Kappa B Ligand (RANKL). In this case both LIP and LAP were shown to be activators of expression in cooperation with ATF4 [108]. The IL-23 receptor also is also regulated by C/EBPß, although there was no information presented concerning isoform specific roles [109]. Most recently it was shown that C/EBPß can actually regulate the entire M2 macrophage transcriptome [11].
Early studies revealed that C/EBPß−/− mice also had lymphoproliferative diseases and defective helper T-cells likely due to the loss of regulation of a large number of chemokines and cytokines [110]. Although this list of secreted chemokines and cytokines may seem large, there are still many others identified in screens in published studies and our own unpublished studies which remain to be validated. It is likely that C/EBPß isoforms can regulate much of the chemokine/cytokine secretome and that the specific landscape will be significantly modulated by the ratio of specific C/EBPß isoform expression.
From the work presented here it is clear that C/EBPß is indeed a master regulator of multiple tissues and disease states. From the initial discovery of NF-IL6 in cells in-vitro to the more recent studies providing insight mechanistically into the downstream in-vivo consequences of C/EBPß expression in breast cancer, investigators will continue to uncover more aspects of C/EBPß-related regulation. As mentioned above, the complexity of C/EBPß isoform-specific regulation of target gene expression makes predicting expected transcriptional outcomes, and thus biological outcomes immensely challenging. Future studies will be required to better understand these regulatory events.
One significant challenge that has plagued C/EBPß biologists and has made understanding isoform specific effects difficult both at the bench, and in the clinic is a lack of a LIP-specific antibody. In the past, many investigators have tried to generate this reagent, but unfortunately have not yet been successful. With this antibody and the advent of and increased accessibility to single-cell technologies, a more complete understanding of C/EBPß biology in the mammary gland should be forthcoming. One can envision experiments in which data from single-cell RNA sequencing (scRNA-seq) will be combined with single-cell proteomics by mass spectrometry (SCoPE-MS) and single-cell ChIP-seq (scChIP-seq) at various developmental stages in mice which have been engineered to overexpress individual C/EBPß isoforms in a tissue-specific manner. These data may then inform subsequent experiments in breast cancer models to definitively determine whether or not specific C/EBPß isoform expression is prognostic for women with breast cancer.
References
Ramji DP, Foka P. CCAAT/enhancer-binding proteins: structure, function and regulation. Biochem J. 2002;365:561–75 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12006103.
Ma D, Panda S, Lin JD. Temporal orchestration of circadian autophagy rhythm by C/EBPβ. EMBO J. 2011;30:4642–51 Available from: http://emboj.embopress.org/cgi/doi/10.1038/emboj.2011.322.
Pham T, Langmann S, Schwarzfischer L, El Chartouni C, Lichtinger M, Klug M, et al. CCAAT enhancer-binding protein beta regulates constitutive gene expression during late stages of monocyte to macrophage differentiation. J Biol Chem. 2007;282:21924–33 Available from: http://www.ncbi.nlm.nih.gov/pubmed/17540774.
Carro MS, Lim WK, Alvarez MJ, Bollo RJ, Zhao X, Snyder EY, et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature. Nature Publishing Group; 2010;463:318–325. https://doi.org/10.1038/nature08712
Luedi MM, Singh SK, Mosley JC, Hatami M, Gumin J, Sulman EP, et al. A Dexamethasone-regulated Gene Signature Is Prognostic for Poor Survival in Glioblastoma Patients. J Neurosurg Anesthesiol. 2017;29:46–58 Available from: http://www.ncbi.nlm.nih.gov/pubmed/27653222.
Wilson S, Filipp FV. A network of epigenomic and transcriptional cooperation encompassing an epigenomic master regulator in cancer. NPJ Syst Biol Appl. Springer US. 2018;4:24. https://doi.org/10.1038/s41540-018-0061-4.
Zaret KS, Carroll JS. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 2011;25:2227–41 Available from: http://www.ncbi.nlm.nih.gov/pubmed/22056668.
Iyer VV, Kadakia TB, McCabe LR, Schwartz RC. CCAAT/enhancer-binding protein-β has a role in osteoblast proliferation and differentiation. Exp Cell Res. 2004;295:128–37.
Müller C, Kowenz-Leutz E, Grieser-Ade S, Graf T, Leutz A. NF-M (chicken C/EBP beta) induces eosinophilic differentiation and apoptosis in a hematopoietic progenitor cell line. EMBO J. 1995;14:6127–35 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8557032.
Pan Z, Hetherington CJ, Zhang DE. CCAAT/enhancer-binding protein activates the CD14 promoter and mediates transforming growth factor beta signaling in monocyte development. J Biol Chem. 1999;274:23242–8 Available from: http://www.ncbi.nlm.nih.gov/pubmed/10438498.
Lamkin DM, Srivastava S, Bradshaw KP, Betz JE, Muy KB, Wiese AM, et al. C/EBPβ regulates the M2 transcriptome in β-adrenergic-stimulated macrophages. Brain Behav Immun. Elsevier. 2019;80:839–48. https://doi.org/10.1016/j.bbi.2019.05.034.
Yoshioka S, Miura Y, Yao H, Satake S, Hayashi Y, Tamura A, et al. CCAAT/enhancer-binding protein β expressed by bone marrow mesenchymal stromal cells regulates early B-cell lymphopoiesis. Stem Cells. 2014;32:730–40 Available from: http://www.ncbi.nlm.nih.gov/pubmed/24115241.
Kowenz-Leutz E, Leutz A. A C/EBPβ Isoform Recruits the SWI/SNF Complex to Activate Myeloid Genes. Mol Cell. 1999;4:735–43 Available from: http://www.ncbi.nlm.nih.gov/pubmed/10619021.
Smink JJ, Bégay V, Schoenmaker T, Sterneck E, De Vries TJ, Leutz A. Transcription factor C/EBPΒ isoform ratio regulates osteoclastogenesis through MafB. EMBO J. 2009;28:1769–81.
Darlington GJ. Molecular mechanisms of liver development and differentiation. Curr Opin Cell Biol. 1999;11:678–82 Available from: https://linkinghub.elsevier.com/retrieve/pii/S0955067499000356.
Welm AL, Timchenko NA, Darlington GJ. C/EBPalpha regulates generation of C/EBPbeta isoforms through activation of specific proteolytic cleavage. Mol Cell Biol. 1999;19:1695–704 Available from: http://www.ncbi.nlm.nih.gov/pubmed/10022857.
Ferrini JB, Rodrigues E, Dulic V, Pichard-Garcia L, Fabr JM, Blanc P, et al. Expression and DNA-binding activity of C/EBPalpha and C/EBPbeta in human liver and differentiated primary hepatocytes. J Hepatol. 2001;35:170–7 Available from: http://www.ncbi.nlm.nih.gov/pubmed/11580138.
Maytin EV, Habener JF. Transcription factors C/EBP alpha, C/EBP beta, and CHOP (Gadd153) expressed during the differentiation program of keratinocytes in vitro and in vivo. J invest Dermatol. Elsevier Masson SAS. 1998;110:238–46. https://doi.org/10.1046/j.1523-1747.1998.00123.x.
Cao Z, Umek RM, McKnight SL. Regulated expression of three C/EBP isoforms during adipose conversion of 3T3-L1 cells. Genes Dev. 1991;5:1538–52 Available from: http://www.ncbi.nlm.nih.gov/pubmed/1840554.
Manchado C, Yubero P, Viñas O, Iglesias R, Villarroya F, Mampel T, et al. CCAAT/enhancer-binding proteins alpha and beta in brown adipose tissue: evidence for a tissue-specific pattern of expression during development. Biochem J. 1994;302 ( Pt 3:695–700. Available from: http://www.ncbi.nlm.nih.gov/pubmed/7945193.
Sterneck E, Tessarollo L, Johnson PF. An essential role for C/EBPbeta in female reproduction. Genes Dev. 1997;11:2153–62 Available from: http://www.ncbi.nlm.nih.gov/pubmed/9303532.
Fan H-Y, Liu Z, Johnson PF, Richards JS. CCAAT/enhancer-binding proteins (C/EBP)-α and -β are essential for ovulation, luteinization, and the expression of key target genes. Mol Endocrinol. 2011;25:253–68 Available from: http://www.ncbi.nlm.nih.gov/pubmed/21177758.
Wang W, Taylor RN, Bagchi IC, Bagchi MK. Regulation of human endometrial stromal proliferation and differentiation by C/EBPβ involves cyclin E-cdk2 and STAT3. Mol Endocrinol. 2012 [cited 2013 Jun 26];26:2016–30. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23097472.
Robinson GW, Johnson PF, Hennighausen L, Sterneck E. The C/EBPbeta transcription factor regulates epithelial cell proliferation and differentiation in the mammary gland. Genes Dev. 1998;12:1907–16 Available from: http://www.ncbi.nlm.nih.gov/pubmed/9637691.
Seagroves TN, Krnacik S, Raught B, Gay J, Burgess-Beusse B, Darlington GJ, et al. C/EBPbeta , but not C/EBPalpha , is essential for ductal morphogenesis, lobuloalveolar proliferation, and functional differentiation in the mouse mammary gland. Genes Dev. 1998;12:1917–28 Available from: http://www.ncbi.nlm.nih.gov/pubmed/9637692.
Seagroves TN, Lydon JP, Hovey RC, Vonderhaar BK, Rosen JM. C/EBPβ (CCAAT/Enhancer Binding Protein) Controls Cell Fate Determination during Mammary Gland Development. Mol Endocrinol. 2000;14:359–68 Available from: http://www.ncbi.nlm.nih.gov/pubmed/10707954.
LaMarca HL, Visbal AP, Creighton CJ, Liu H, Zhang Y, Behbod F, et al. CCAAT/enhancer binding protein beta regulates stem cell activity and specifies luminal cell fate in the mammary gland. Stem Cells. 2010 [cited 2013 Jun 26];28:535–44. Available from: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=3006225&tool=pmcentrez&rendertype=abstract
Doppler W, Welte T, Philipp S. CCAAT/enhancer-binding protein isoforms beta and delta are expressed in mammary epithelial cells and bind to multiple sites in the beta-casein gene promoter. J Biol Chem. 1995;270:17962–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7629103.
Isshiki H, Akira S, Tanabe O, Nakajima T, Shimamoto T, Hirano T, et al. Constitutive and interleukin-1 (IL-1)-inducible factors interact with the IL-1-responsive element in the IL-6 gene. Mol Cell Biol. 1990;10:2757–64 Available from: http://www.ncbi.nlm.nih.gov/pubmed/2111442.
Akira S, Isshiki H, Sugita T, Tanabe O, Kinoshita S, Nishio Y, et al. A nuclear factor for IL-6 expression (NF-IL6) is a member of a C/EBP family. Embo J. 1990;9:1897–906 http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=551896&tool=pmcentrez&rendertype=abstract.
Descombes P, Chojkier M, Lichtsteiner S, Falvey E, Schibler U. LAP, a novel member of the C/EBP gene family, encodes a liver-enriched transcriptional activator protein. Genes Dev. 1990;4:1541–51 Available from: http://www.ncbi.nlm.nih.gov/pubmed/2253878.
Descombes P, Schibler U. A liver-enriched transcriptional activator protein, LAP, and a transcriptional inhibitory protein, LIP, are translated from the same mRNA. Cell. 1991;67:569–79 Available from: http://www.ncbi.nlm.nih.gov/pubmed/1934061.
Calkhoven CF, Müller C, Leutz A. Translational control of C/EBPalpha and C/EBPbeta isoform expression. Genes Dev. 2000;14:1920–32 Available from: http://www.ncbi.nlm.nih.gov/pubmed/10921906.
LeClair KP, Blanar MA, Sharp PA. The p50 subunit of NF-kappa B associates with the NF-IL6 transcription factor. Proc Natl Acad Sci U S A. 1992;89:8145–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/1518839.
Matsusaka T, Fujikawa K, Nishio Y, Mukaida N, Matsushima K, Kishimoto T, et al. Transcription factors NF-IL6 and NF-kappa B synergistically activate transcription of the inflammatory cytokines, interleukin 6 and interleukin 8. Proc Natl Acad Sci U S A. 1993;90:10193–7 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8234276.
Nishio Y, Isshiki H, Kishimoto T, Akira S. A nuclear factor for interleukin-6 expression (NF-IL6) and the glucocorticoid receptor synergistically activate transcription of the rat alpha 1-acid glycoprotein gene via direct protein-protein interaction. Mol Cell Biol. 1993;13:1854–62 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8441418.
Hsu W, Kerppola TK, Chen PL, Curran T, Chen-Kiang S. Fos and Jun repress transcription activation by NF-IL6 through association at the basic zipper region. Mol Cell Biol. 1994;14:268–76 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8264594.
Zhang Y, Rom WN. Regulation of the interleukin-1 beta (IL-1 beta) gene by mycobacterial components and lipopolysaccharide is mediated by two nuclear factor-IL6 motifs. Mol Cell Biol. 1993;13:3831–7 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7684503.
Stein B, Baldwin AS. Distinct mechanisms for regulation of the interleukin-8 gene involve synergism and cooperativity between C/EBP and NF-kappa B. Mol Cell Biol. 1993;13:7191–8 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8413306.
Dunn SM, Coles LS, Lang RK, Gerondakis S, Vadas MA, Shannon MF. Requirement for nuclear factor (NF)-kappa B p65 and NF-interleukin-6 binding elements in the tumor necrosis factor response region of the granulocyte colony-stimulating factor promoter. Blood. 1994;83:2469–79 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7513199.
Mukaida N, Morita M, Ishikawa Y, Rice N, Okamoto S, Kasahara T, et al. Novel mechanism of glucocorticoid-mediated gene repression. Nuclear factor-kappa B is target for glucocorticoid-mediated interleukin 8 gene repression. J Biol Chem. 1994;269:13289–95 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8175759.
Davydov IV, Krammer PH, Li-Weber M. Nuclear factor-IL6 activates the human IL-4 promoter in T cells. J Immunol. 1995;155:5273–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7594540.
Wethmar K, Bégay V, Smink JJ, Zaragoza K, Wiesenthal V, Dörken B, et al. C/EBPβΔuORF mice - a genetic model for uORF-mediated translational control in mammals. Genes Dev. 2010;24:15–20.
Bégay V, Smink JJ, Loddenkemper C, Zimmermann K, Rudolph C, Scheller M, et al. Deregulation of the endogenous C/EBPβ LIP isoform predisposes to tumorigenesis. J Mol Med. 2015;93:39–49.
Zidek LM, Ackermann T, Hartleben G, Eichwald S, Kortman G, Kiehntopf M, et al. Deficiency in mTORC 1-controlled C/ EBP β - mRNA translation improves metabolic health in mice. EMBO Rep. 2015;16:1022–36.
Müller C, Zidek LM, Ackermann T, de Jong T, Liu P, Kliche V, et al. Reduced expression of C/EBPβ-LIP extends health and lifespan in mice. Elife. 2018;7:1–28.
Dittmar G, Hernandez DP, Kowenz-Leutz E, Kirchner M, Kahlert G, Wesolowski R, et al. PRISMA: Protein Interaction Screen on Peptide Matrix Reveals Interaction Footprints and Modifications- Dependent Interactome of Intrinsically Disordered C/EBPβ. iScience. 2019;13:351–70 Available from: https://linkinghub.elsevier.com/retrieve/pii/S2589004219300604.
Tang Q, Grønborg M, Huang H, Kim J, Otto TC, Pandey A, et al. Sequential phosphorylation of CCAAT enhancer-binding protein beta by MAPK and glycogen synthase kinase 3beta is required for adipogenesis. Proc Natl Acad Sci U S A. 2005;102:9766–71 Available from: http://www.ncbi.nlm.nih.gov/pubmed/15985551.
Kim J, Tang Q, Li X, Lane MD. Effect of phosphorylation and S-S bond-induced dimerization on DNA binding and transcriptional activation by C/EBPbeta. Proc Natl Acad Sci U S A. 2007;104:1800–4 Available from: http://www.ncbi.nlm.nih.gov/pubmed/17264204.
Hattori T, Ohoka N, Inoue Y, Hayashi H, Onozaki K. C/EBP family transcription factors are degraded by the proteasome but stabilized by forming dimer. Oncogene. 2003;22:1273–80 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12618752.
Zhang Y, Li S, Qian S, Zhang Y, Liu Y, Tang Q-Q, et al. Phosphorylation prevents C/EBPβ from the calpain-dependent degradation. Biochem Biophys Res Commun. 2012;419:550–5 Available from: http://www.ncbi.nlm.nih.gov/pubmed/22369944.
Li X, Molina H, Huang H, Zhang Y, Liu M, Qian S, et al. O-linked N-acetylglucosamine modification on CCAAT enhancer-binding protein beta: role during adipocyte differentiation. J Biol Chem. 2009;284:19248–54 Available from: http://www.ncbi.nlm.nih.gov/pubmed/19478079.
Ceseña TI, Cardinaux J-R, Kwok R, Schwartz J. CCAAT/enhancer-binding protein (C/EBP) beta is acetylated at multiple lysines: acetylation of C/EBPbeta at lysine 39 modulates its ability to activate transcription. J Biol Chem. 2007 [cited 2013 Jun 26];282:956–67. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17110376.
Ceseña TI, Cui TX, Subramanian L, Fulton CT, Iñiguez-Lluhí JA, Kwok RPS, et al. Acetylation and deacetylation regulate CCAAT/enhancer binding protein β at K39 in mediating gene transcription. Mol Cell Endocrinol. 2008;289:94–101.
Wiper-Bergeron N, Salem HA, Tomlinson JJ, Wu D, Haché RJG. Glucocorticoid-stimulated preadipocyte differentiation is mediated through acetylation of C/EBPbeta by GCN5. Proc Natl Acad Sci U S A. 2007;104:2703–8 Available from: http://www.ncbi.nlm.nih.gov/pubmed/17301242.
Xu M, Nie L, Kim S, Sun X. STAT5-induced Id-1 transcription involves recruitment of HDAC1 and deacetylation of C/EBPbeta. EMBO J. 2003;22:893–904 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12574125.
Pless O, Kowenz-Leutz E, Knoblich M, Lausen J, Beyermann M, Walsh MJ, et al. G9a-mediated lysine methylation alters the function of CCAAT/enhancer-binding protein-beta. J Biol Chem. 2008;283:26357–63 Available from: http://www.ncbi.nlm.nih.gov/pubmed/1864779.
Kowenz-Leutz E, Pless O, Dittmar G, Knoblich M, Leutz A. Crosstalk between C/EBPbeta phosphorylation, arginine methylation, and SWI/SNF/Mediator implies an indexing transcription factor code. EMBO J . Nature Publishing Group; 2010 [cited 2013 Jun 18];29:1105–15. Available from: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2845275&tool=pmcentrez&rendertype=abstract
Kim J, Cantwell CA, Johnson PF, Pfarr CM, Williams SC. Transcriptional Activity of CCAAT / Enhancer-binding Proteins Is Controlled by a Conserved Inhibitory Domain That Is a Target for Sumoylation *. 2002;277:38037–44.
Eaton EM, Sealy L. Modification of CCAAT/enhancer-binding protein-beta by the small ubiquitin-like modifier (SUMO) family members, SUMO-2 and SUMO-3. J Biol Chem. 2003;278:33416–21 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12810706.
Subramanian L, Benson MD, In JA. A Synergy Control Motif within the Attenuator Domain of CCAAT / Enhancer-binding Protein ␣ Inhibits Transcriptional Synergy through Its PIASy-enhanced Modification by SUMO-1 or SUMO-3 *. 2003;278:9134–41.
Berberich-Siebelt F, Berberich I, Andrulis M, Santner-Nanan B, Jha MK, Klein-Hessling S, et al. SUMOylation interferes with CCAAT/enhancer-binding protein beta-mediated c-myc repression, but not IL-4 activation in T cells. J Immunol. 2006;176:4843–51 Available from: http://www.ncbi.nlm.nih.gov/pubmed/16585579.
Liu Y, Zhang Y, Guo L, Huang H-Y, Zhu H, Huang J, et al. Protein inhibitor of activated STAT 1 (PIAS1) is identified as the SUMO E3 ligase of CCAAT/enhancer-binding protein β (C/EBPβ) during adipogenesis. Mol Cell Biol. 2013;33:4606–17 Available from: http://www.ncbi.nlm.nih.gov/pubmed/24061474.
Cong L, Zhang F. Genome engineering using CRISPR-Cas9 system. Methods Mol Biol. 2015;1239:197–217 Available from: http://www.ncbi.nlm.nih.gov/pubmed/25408407.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23 Available from: http://www.ncbi.nlm.nih.gov/pubmed/23287718.
Kabotyanski EB, Huetter M, Xian W, Rijnkels M, Rosen JM. Integration of prolactin and glucocorticoid signaling at the beta-casein promoter and enhancer by ordered recruitment of specific transcription factors and chromatin modifiers. Mol Endocrinol. 2006;20:2355–68 Available from: http://www.ncbi.nlm.nih.gov/pubmed/16772529.
Christian M, Pohnke Y, Kempf R, Gellersen B, Brosens JJ. Functional association of PR and CCAAT/enhancer-binding protein beta isoforms: promoter-dependent cooperation between PR-B and liver-enriched inhibitory protein, or liver-enriched activatory protein and PR-A in human endometrial stromal cells. Mol Endocrinol. 2002;16:141–54 Available from: http://www.ncbi.nlm.nih.gov/pubmed/11773445.
Liu Q, Boudot A, Ni J, Hennessey T, Beauparlant SL, Rajabi HN, et al. Cyclin D1 and C/EBPβ LAP1 operate in a common pathway to promote mammary epithelial cell differentiation. Mol Cell Biol. 2014;34:3168–79 Available from: http://mcb.asm.org/lookup/doi/10.1128/MCB.00039-14.
Spooner CJ, Guo X, Johnson PF, Schwartz RC. Differential roles of C/EBP beta regulatory domains in specifying MCP-1 and IL-6 transcription. Mol Immunol. 2007;44:1384–92 Available from: http://www.ncbi.nlm.nih.gov/pubmed/16784777.
Gomis RR, Alarcón C, Nadal C, Van Poznak C, Massagué J. C/EBPβ at the core of the TGFβ cytostatic response and its evasion in metastatic breast cancer cells. Cancer Cell . 2006 [cited 2013 Jun 6];10:203–14. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16959612.
Wang H, Larris B, Peiris TH, Zhang L, Le Lay J, Gao Y, et al. C/EBPbeta activates E2F-regulated genes in vivo via recruitment of the coactivator CREB-binding protein/P300. J Biol Chem . 2007 [cited 2013 Jun 26];282:24679–88. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17599912.
Davis CA, Hitz BC, Sloan CA, Chan ET, Davidson JM, Gabdank I, et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res . Oxford University Press; 2018;46:D794–801. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29126249.
Raught B, Liao WS-L, Rosen JM. Developmentally and hormonally regulated CCAAT/enhancer-binding protein isoforms influence beta-casein gene expression. Mol Endocrinol. 1995;9:1223–32 Available from: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=7491114.
Wang W, Do HN, Aupperlee MD, Durairaj S, Flynn EE, Miksicek RJ, et al. C/EBPβ LIP and c-Jun synergize to regulate expression of the murine progesterone receptor. Mol Cell Endocrinol . Elsevier. 2018;477:57–69. https://doi.org/10.1016/j.mce.2018.06.001.
Kabotyanski EB, Rijnkels M, Freeman-Zadrowski C, Buser AC, Edwards DP, Rosen JM. Lactogenic hormonal induction of long distance interactions between beta-casein gene regulatory elements. J Biol Chem. 2009;284:22815–24 Available from: http://www.ncbi.nlm.nih.gov/pubmed/19542223.
Park B, Kook S, Lee S, Jeong J, Brufsky A, Lee B. An isoform of C/EBPβ, LIP, regulates expression of the chemokine receptor CXCR4 and modulates breast cancer cell migration. J Biol Chem. 2013;288:28656–67 Available from: http://www.ncbi.nlm.nih.gov/pubmed/23966000.
Abraham S, Sweet T, Sawaya BE, Rappaport J, Khalili K, Amini S. Cooperative interaction of C/EBP beta and Tat modulates MCP-1 gene transcription in astrocytes. J Neuroimmunol. 2005;160:219–27 Available from: http://www.ncbi.nlm.nih.gov/pubmed/15710476.
Raught B, Gingras AC, James A, Medina D, Sonenberg N, Rosen JM. Expression of a translationally regulated, dominant-negative CCAAT/enhancer-binding protein beta isoform and up-regulation of the eukaryotic translation initiation factor 2alpha are correlated with neoplastic transformation of mammary epithelial cells. Cancer Res. 1996;56:4382–6 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8813130.
Gigliotti AP, DeWille JW. Lactation status influences expression of CCAAT/enhancer binding protein isoform mRNA in the mouse mammary gland. J Cell Physiol. 1998;174:232–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/9428809.
Eaton EM, Hanlon M, Bundy L, Sealy L. Characterization of C/EBPbeta isoforms in normal versus neoplastic mammary epithelial cells. J Cell Physiol. 2001;189:91–105 Available from: http://www.ncbi.nlm.nih.gov/pubmed/11573208.
Dearth LR, Hutt J, Sattler A, Gigliotti A, DeWille J. Expression and function of CCAAT/enhancer binding protein? (C/EBP?) LAP and LIP isoforms in mouse mammary gland, tumors and cultured mammary epithelial cells. J Cell Biochem. 2001;82:357–70 Available from: http://www.ncbi.nlm.nih.gov/pubmed/11500913.
Zahnow CA, Younes P, Laucirica R, Rosen JM. Overexpression of C/EBPbeta-LIP, a naturally occurring, dominant-negative transcription factor, in human breast cancer. J Natl Cancer Inst. 1997;89:1887–91 Available from: https://academic.oup.com/jnci/article-lookup/doi/10.1093/jnci/89.24.1887.
Zahnow CA, Cardiff RD, Laucirica R, Medina D, Rosen JM. A role for CCAAT/enhancer binding protein beta-liver-enriched inhibitory protein in mammary epithelial cell proliferation. Cancer Res. 2001;61:261–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/11196172.
Bundy LM, Sealy L. CCAAT/enhancer binding protein beta (C/EBPbeta)-2 transforms normal mammary epithelial cells and induces epithelial to mesenchymal transition in culture. Oncogene. 2003;22:869–83 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12584567.
Bundy L, Wells S, Sealy L. C/EBPbeta-2 confers EGF-independent growth and disrupts the normal acinar architecture of human mammary epithelial cells. Mol Cancer. 2005;4:43 Available from: http://www.ncbi.nlm.nih.gov/pubmed/16371159.
Miura Y, Hagiwara N, Radisky DC, Hirai Y. CCAAT/enhancer binding protein beta (C/EBPβ) isoform balance as a regulator of epithelial-mesenchymal transition in mouse mammary epithelial cells. Exp Cell Res . Elsevier. 2014;327:146–55. https://doi.org/10.1016/j.yexcr.2014.05.019.
Russell A, Boone B, Jiang A, Sealy L. Genomic profiling of C/EBPβ2 transformed mammary epithelial cells: a role for nuclear interleukin-1β. Cancer Biol Ther. 2010;10:509–19 Available from: http://www.ncbi.nlm.nih.gov/pubmed/21057224.
Jundt F, Raetzel N, Müller C, Calkhoven CF, Kley K, Mathas S, et al. A rapamycin derivative (everolimus) controls proliferation through down-regulation of truncated CCAAT enhancer binding protein {beta} and NF-{kappa}B activity in Hodgkin and anaplastic large cell lymphomas. Blood . 2005 [cited 2014 mar 17];106:1801–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15886325.
Benz CC, Scott GK, Santos GF, Smith HS. Expression of c-myc, c-Ha-ras1, and c-erbB-2 proto-oncogenes in normal and malignant human breast epithelial cells. J Natl Cancer Inst. 1989;81:1704–9 Available from: http://www.ncbi.nlm.nih.gov/pubmed/2572702.
Timchenko NA, Welm AL, Lu X, Timchenko LT. CUG repeat binding protein (CUGBP1) interacts with the 5. 1999;27:1–9. Available from: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC148737/pdf/274517.pdf%5Cnpapers3://publication/uuid/4D31EAFB-579C-444A-9E80-43A9663E4B5A
Baldwin BR, Timchenko NA, Zahnow CA. Epidermal growth factor receptor stimulation activates the RNA binding protein CUG-BP1 and increases expression of C/EBPbeta-LIP in mammary epithelial cells. Mol Cell Biol. 2004;24:3682–91 Available from: http://www.ncbi.nlm.nih.gov/pubmed/15082764.
Baselga J, Norton L, Albanell J, Kim YM, Mendelsohn J. Recombinant humanized anti-HER2 antibody (Herceptin) enhances the antitumor activity of paclitaxel and doxorubicin against HER2/neu overexpressing human breast cancer xenografts. Cancer Res. 1998;58:2825–31 Available from: http://www.ncbi.nlm.nih.gov/pubmed/9661897.
Arnal-Estapé A, Tarragona M, Morales M, Guiu M, Nadal C, Massagué J, et al. HER2 silences tumor suppression in breast cancer cells by switching expression of C/EBPß isoforms. Cancer Res . 2010 [cited 2013 Jun 6];70:9927–36. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21098707.
Gustafson TL, Wellberg E, Laffin B, Schilling L, Metz RP, Zahnow CA, et al. Ha-Ras transformation of MCF10A cells leads to repression of Singleminded-2s through NOTCH and C/EBPbeta. Oncogene. 2009;28:1561–8 Available from: http://www.ncbi.nlm.nih.gov/pubmed/19169276.
Johansson J, Berg T, Kurzejamska E, Pang M, Tabor V, Jansson M, et al. MiR-155-mediated loss of C/EBPβ shifts the TGF-β response from growth inhibition to epithelial-mesenchymal transition, invasion and metastasis in breast cancer. Oncogene . Nature Publishing Group. 2013;32:5614–24. https://doi.org/10.1038/onc.2013.322.
Kurzejamska E, Johansson J, Jirström K, Prakash V, Ananthaseshan S, Boon L, et al. C/EBPβ expression is an independent predictor of overall survival in breast cancer patients by MHCII/CD4-dependent mechanism of metastasis formation. Oncogenesis. 2014;3:e125 http://www.nature.com/articles/oncsis201438.
Willis S, De P, Dey N, Long B, Young B, Sparano JA, et al. Enriched transcription factor signatures in triple negative breast cancer indicates possible targeted therapies with existing drugs. Meta gene . Elsevier B.V. 2015;4:129–41. https://doi.org/10.1016/j.mgene.2015.04.002.
Jinesh GG, Flores ER, Brohl AS. Chromosome 19 miRNA cluster and CEBPB expression specifically mark and potentially drive triple negative breast cancers. Ahmad A, editor. PLoS One . 2018;13:e0206008. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30335837.
Li W, Tanikawa T, Kryczek I, Xia H, Li G, Wu K, et al. Aerobic Glycolysis Controls Myeloid-Derived Suppressor Cells and Tumor Immunity via a Specific CEBPB Isoform in Triple-Negative Breast Cancer. Cell Metab. 2018;28:87–103.e6 Available from: http://www.ncbi.nlm.nih.gov/pubmed/29805099.
Salaroglio IC, Gazzano E, Abdullrahman A, Mungo E, Castella B, Abd-Elrahman GEFA-E, et al. Increasing intratumor C/EBP-β LIP and nitric oxide levels overcome resistance to doxorubicin in triple negative breast cancer. J Exp Clin Cancer Res . Journal of Experimental & Clinical Cancer Research; 2018;37:286. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30482226.
Tsukada J, Saito K, Waterman WR, Webb AC, Auron PE. Transcription factors NF-IL6 and CREB recognize a common essential site in the human prointerleukin 1 beta gene. Mol Cell Biol. 1994;14:7285–97 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7935442.
Grove M, Plumb M. C/EBP, NF-kappa B, and c-Ets family members and transcriptional regulation of the cell-specific and inducible macrophage inflammatory protein 1 alpha immediate-early gene. Mol Cell Biol. 1993;13:5276–89 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8355682.
Pope RM, Leutz A, Ness SA. C/EBP beta regulation of the tumor necrosis factor alpha gene. J Clin Invest. 1994;94:1449–55 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7929820.
Nomiyama H, Hieshima K, Hirokawa K, Hattori T, Takatsuki K, Miura R. Characterization of cytokine LD78 gene promoters: positive and negative transcriptional factors bind to a negative regulatory element common to LD78, interleukin-3, and granulocyte-macrophage colony-stimulating factor gene promoters. Mol Cell Biol. 1993;13:2787–801 Available from: http://www.ncbi.nlm.nih.gov/pubmed/8474441.
Brenner S, Prösch S, Schenke-Layland K, Riese U, Gausmann U, Platzer C. cAMP-induced Interleukin-10 promoter activation depends on CCAAT/enhancer-binding protein expression and monocytic differentiation. J Biol Chem. 2003;278:5597–604 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12493739.
Fernández N, Renedo M, García-Rodríguez C, Sánchez CM. Activation of monocytic cells through Fc gamma receptors induces the expression of macrophage-inflammatory protein (MIP)-1 alpha, MIP-1 beta, and RANTES. J Immunol. 2002;169:3321–8 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12218153.
Calonge E, Alonso-Lobo JM, Escandón C, González N, Bermejo M, Santiago B, et al. c/EBPbeta is a major regulatory element driving transcriptional activation of the CXCL12 promoter. J Mol Biol . Elsevier B.V. 2010;396:463–72. https://doi.org/10.1016/j.jmb.2009.11.064.
Tsushima H, Okazaki K, Ishihara K, Ushijima T, Iwamoto Y. CCAAT/enhancer-binding protein β promotes receptor activator of nuclear factor-kappa-B ligand (RANKL) expression and osteoclast formation in the synovium in rheumatoid arthritis. Arthritis Res Ther. 2015;17:–31 Available from: http://www.ncbi.nlm.nih.gov/pubmed/25811130.
Simpson-Abelson MR, Hernandez-Mir G, Childs EE, Cruz JA, Poholek AC, Chattopadhyay A, et al. CCAAT/enhancer-binding protein β promotes pathogenesis of EAE. Cytokine . Elsevier Ltd. 2017;92:24–32. https://doi.org/10.1016/j.cyto.2017.01.005.
Screpanti I, Romani L, Musiani P, Modesti A, Fattori E, Lazzaro D, et al. Lymphoproliferative disorder and imbalanced T-helper response in C/EBP beta-deficient mice. EMBO J. 1995;14:1932–41 Available from: http://www.ncbi.nlm.nih.gov/pubmed/7744000.
Zahnow CA. CCAAT/enhancer binding proteins in normal mammary development and breast cancer. Breast Cancer Res . BioMed Central. 2002;4:113–21 Available from: http://www.ncbi.nlm.nih.gov/pubmed/12052253.
Grimm SL, Rosen JM. The Role of C/EBPβ in Mammary Gland Development and Breast Cancer. J Mammary Gland Biol Neoplasia. 2003;8:191–204 Available from: http://www.ncbi.nlm.nih.gov/pubmed/14635794.
Acknowledgements
These studies were supported by grant CA16303 from the National Cancer Institute. We would like to thank Dr. Sandy Grimm for generously providing us with the figure illustrating C/EBPß expression in the mammary gland throughout development. While we tried to be as comprehensive as possible, we apologize to those investigators whose work we were unable to cite due to space limitations.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Spike, A.J., Rosen, J.M. C/EBPß Isoform Specific Gene Regulation: It’s a Lot more Complicated than you Think!. J Mammary Gland Biol Neoplasia 25, 1–12 (2020). https://doi.org/10.1007/s10911-020-09444-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10911-020-09444-5