Abstract
Copper oxide (CuO) NPs offer promising applications and green synthesis is widely preferred over synthetic methods because of its merits. The past literatures show the preparation of copper nanoparticles from plants extract of leaves but not from the enzymes. But here the greener method of CuO nanoparticles which has been derived from enzymes extract of Citrus aurantifolia is reported. When the enzyme powder obtained from leaf extracts of Citrus aurantifolia was treated with copper sulphate, copper oxide nanoparticles were produced. Such nanoparticles prepared are characterized by UV– Vis, FT-IR, XRD, HR-TEM and scanning electron microscope studies. On the other hand, antibacterial activity against K. pneumonia and S. aureus, was evaluated. Results showed sensibility of K. pneumonia and S. aureus and the antibacterial property improves infections due to drug-resistant bacteria’s.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
These days, microbes are skilled to coordinate to hassled circumstance as well as figure varied instrument of barrier since of enormous exploitation of anti-microbials, which prompt a reflective concern in therapeutic troubles [1]. The most widely recognized risky multidrug-safe pathogens are Acineto bacter baumannii, E.coli, penicillin-safe Streptococcus pneumoniae, Klebsiella pneumoniae, vancomycin-safe Enterococcus, methicillin-safe S. aureus, and broadly medicate safe Mycobacterium tuberculosis [2]. In this way, consideration is attracted to treatment rules and the utilization of novel antimicrobial operators with better bactericidal properties [3].The unavoidable task in the applied science is the communication of living molecules and the inorganic precursors [4]. In this circumstance, nanotechnology has concentrated on the improvement of nanomaterials with the capacity to communicate with organic elements that are pathogenic to people. In future, nanostructure engineering will be involved to judiciously design and optimize the performance [5]. In chemical reactions, the primary steps towards greener approach are the use of non-toxic and non-volatile solvent as well as conservation of energy without debris [6]. Hence, the replacement of organic solvents by phytochemicals present in plants is an important step in green chemistry. Among these, preparation of metal nanoparticles has pulled in logical intrigue, in view of the bactericidal properties appeared by metals like gold, copper, and silver which showed fascinating antimicrobial properties [4]. The inorganic compounds which are in nano range indicate outstanding action against bacteria because of the ratio of proportion of increase in availability of surface to the dimension [5]. Copper nanoparticles have colossal probability to replace silver nanoparticles which are costly and hence have gained immense attention in research as well as industry [7]. Among the accessible diverse green science strategies for the preparation of CuO-NPs, plant based preparation has been pulled in for their non-harmful and easiest strategy because lately, there has been incredible worry over the numerous genuine ecological issues [8]. As of late green preparation of CuO-NPs by various plants, for example, Phyllanthusamarus [9], Gloriosasuperba [10] and a lot more plants has been accounted for. Phyto- compounds intervened integrated CuO-NPs showed potential antibacterial action. Acharyulu et al. [9] announced green incorporated CuO-NPs demonstrated huge antibacterial movement against multidrug safe bacterial pathogens.
Copper compounds have been used to treat cancer and several diseases for thousands of years [11]. Copper compounds may contain similar properties to arsenic trioxide, which is another anti-tumor compound [12]. Copper is a well- known heavy metal that is toxic to mammalian cells. However, with the advancement of nanotechnology, the NPs can target specific cells with lesser side effects. Even though the copper compounds have strong therapeutic potential, no study has been reported at the in vitro level.
Several bioactive compounds were identified in C.aurantifolia, including ascorbic acid, flavonoids, limonoids, coumarins, and phytosterols, and were investigated [13]. Among the flavonoids, hesperidin was found to be the most abundant [14] and acts as anticancer agent through prostaglandin and inhibits chemical carcinogenesis [15]. The other flavonoid reported in limes is rutin (quercetin-3-rutinoside) [16]. The synthesis of CuO nanoparticles from the leaf extract of C.aurantifolia has been extensively studied for various anticancer studies but no work has been carried out on synthesis of CuO nanoparticles from the enzyme extract of C.aurantifolia and anticancer activity against SK MEL 28 cell line. Based on this reason we selected only enzyme mediated CuO NPs synthesized from C.aurantifolia in the anticancer studies and antibacterial movement against S.Pneumoniae and S.Aureus.
Materials and Methods
Materials
All the chemical reagents used in this experiment were of ultra pure grade.
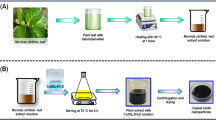
Preparation of Enzymes Extracts
Plant tissues contain a wide scope of proteins, which differ extraordinarily in their properties, and require explicit conditions for their extraction and cleansing. It is along these lines unrealistic to suggest a solitary convention for extraction of all plant proteins. Step by step protein extraction conventions are given below. Three step by step protein extraction conventions are given. The initial two are for the extraction of the enzymically active proteins involved with Rubisco (EC 4.1.1.39) and nitrate reductase (EC 1.6.6.1), from vegetative tissues. Rubisco is a hexadec-americ protein (eight subunits of approx 50–60 kDa and eight subunits of 12–20 kDa) with a molecular weight of 500,000, which catalyzes the obsession of carbon in the chloroplast stroma. It regularly speaks to fifty percent of the whole chloroplast protein and is perceived as the vast protein on the planet. Conversely, the unpredictable catalyst nitrate reductase (NR), which has a molecular weight of approx 200,000, is available in plant tissues at < 5 mg/kg new weight [17]. This low plenitude, joined with vulnerability to proteolysis and loss of functional prosthetic group during extraction and cleansing, frequently prompts an extremely low recuperation of the enzyme.
Extraction of Enzymically Active Preparations from Leaf Tissues
In this strategy [17], it is insisted that the extraction methodology and extraction buffers utilized are significant in influencing the initial rate and total activities of the enzyme. It is additionally significant for initial activity estimations to keep up the extract at a temperature of 2 °C.
-
i.
Cut the 3 weeks old leaves into 1-cm lengths and homogenize in an ice cold buffer using a ratio of 6:1.
-
ii.
Filter and afterward include adequate solid (NH4)2S04 to give 35%) saturation.
-
iii.
Centrifuge the suspension at 20,000 g for 15 min and then dispose of the pellet.
Add further solid (NH4)2S04 to increase the saturation by fifty percent. After centrifugation disintegrate the pellet in EDTA, Magnesium chloride, polyvinyl pyrrolidone (PVP), dithiothreitol (DTT), glycerol and protease inhibitors at pH 8.0. Buffers help to keep up pH since bursting of the cells discharges numerous organic acids. EDTA is incorporated to chelate heavy metals which might be available in traces in reagents. Mg2+ is incorporated for the extraction medium since numerous enzymes are subjected to it for their action. PVP binds phenolic compounds present in the plant tissue separates and shields the enzymes from inactivation. Thiol groups present in the enzymes are kept without losing electrons by including DTT, mercapto ethanol, ascorbate or reduced glutathione. To avoid the proteolytic cleavage by the proteolytic enzymes during enzyme extraction, protease inhibitors like phenyl methyl sulfonyl fluoride (PMSF) are normally included. These techniques help in getting stable enzymes from the plants.
Synthesis of Copper Oxide Nanoparticles
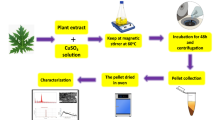
In a typical reaction mixture, 100 ml of aqueous 1 mM copper sulphate dehydrate CuSO4.5H2O was treated with 10 ml of above mentioned aqueous enzyme extract of Citrus aurantifolia and stirred magnetically at room temperature for about 4 h. Here, C.aurantifolia enzyme extracts was used as a repeatable agent to control the resulting particle size. The quality of the enzyme extract can be controlled by maintaining the pH. The nanoparticles formed were collected and dried at 60 °C in an oven. The dried product was subjected to calcination in a muffle furnace at 400 °C for 2 h to remove any attached water molecules to the product.
Characterization
The phase identification was done by powder XRD (Bruker D8 advance). The functional group was analysed using FTIR (Shimadzu). The microscopical analysis of the samples was carried out by a scanning electron microscope (SEM; Jeol 6390LA/ OXFORD XMX N) equipped with energy-dispersive spectroscopy (EDS), and the powder sample was mixed with ethanol to analyze the sample through TEM (JEM-2100; JEOL).
Characterization of Copper Oxide Nanoparticles
The absorption peak for copper oxide nanoparticles was identified by UV–Vis spectrophotometer (Shimadzu) and the functional groups by FTIR (Shimadzu). The shape and grain size were characterized by using SEM (Hitachi).
Anti Bacterial Tests
The antibacterial spectrum of copper oxide nanoparticles, Cu-Citrus aurantifolia enzyme nanoparticle composite samples were determined against the test bacteria by disc well—diffusion method on an agar plate [18]. Briefly incubated cultures of bacteria were swabbed uniformly on the individual plates using sterile cotton swabs on the Muller Hinton Agar, 50 μl of copper oxide nanoparticles and Cu-Citrus aurantifolia enzyme nanoparticle composite samples with different concentration of 1,2,3,5% were loaded in sterile discs and incubated for analysis by looking at the percentage of area of inhibition.
Anti Cancer Tests
The anticancer assay was done by MTT assay method [19]. The results were expressed as IC50 which is the concentration of the sample required to inhibit 50% of MTT concentration.
Results and Discussion
Structural and Morphological Study
The phase and nature of the sample was analysed by XRD analysis. In Fig. 1,the peaks at 35.58°, 38.68° and 49.12° are assigned to the (111) (002) and (202) which represent the reflection lines of monoclinic CuO NPs [20] and the lattice parameter a = 4.683 Å. The average size of the crystal was calculated from XRD data using the formula given by Scherrer as follows
where D—crystallite size, k—wavelength of the X-ray, h—angle of diffraction and β—full width at half maximum.
(FWHM) are equated. The average size (D) of copper nanoparticles is found to be 18 nm. Williamson– Hall method (Fig. 2) was used to find the lattice strain using the modified Scherrer equation.
W–H plot of β cos θ Vs 4εsinθ shows the information about microstrain. The plot shows it has a non zero slope and the calculated value of microstrain for the prepared monoclinic CuO nanoparticles is 0.53527.
Dislocation Density (d)
The dislocation density (d) is given by the below equation.
The number of unit cells (n) is estimated from
In the Eq. (4), V is the volume of the cell unit.
Morphology Index
In XRD, FWHM is used to calculate the Morphology Index (MI). MI is obtained using the equation below [21].
where FWHMh is the highest FWHM value obtained from peaks, FWHMp is a value of particular peaks of FWHM.
The MI values are calculated in Table 1.
FT-IR Analysis
The FTIR spectra of enzyme extract as well as CuO NPs are shown in Fig. 2. The FTIR result of extract showed strong broad peaks at 3736 cm–1which is corresponding to O–H stretching, 3257 cm−1 and 3445 cm−1due to aromatic OH vibrations, 2922 and 2851 cm−1 peak appeared for presence of many methylene groups, 2360 cm−1for C≡C stretching, 1799 cm −1 due to cyclic anhydride, 1744 cm−1 due to –C=O- stretching, 1150 cm−1corresponds to the –C- O-C bond of carboxylate group, 883 cm−1 is due to unsaturated –CH bending (Fig. 3b). The FTIR result of CuO NPs (Fig. 3a) showed stretching and vibration peaks at 3446 cm-1 due to aromatic OH vibration, 2076 due to C≡C functional group, 1635 cm −1 due to C=O stretching vibration of primary amines, 1383 cm-1 due to 30 carbon atoms and 1113 cm-1 due to carboxylate group respectively. Therefore, after addition of copper sulphate to the enzyme extract of Citrus aurantifolia, the C≡C peak at 2360 cm−1 was reduced to 2076 cm−1 and the C–O–C bond of carboxylate peak of the enzyme extract at 1402 cm−1 was reduced to 1113 cm−1 and it can be seen in FTIR of CuO NPs (Fig. 3a). Thus it can be concluded that the unsaturated bonds of alkynes and carboxylate group of enzymes extract of Citrus aurantifolia were mainly responsible for capping and stabilization of CuO NPs.
Morphological Analysis
Figure 4 shows the SEM images at different magnifications and EDAX observations of the nanomaterial. The surface morphology of the synthesized CuO nanoparticles was rice shaped structure examined by SEM analysis. The materials prepared were inspected for impurities and structure. The major element was identified as copper along with carbon and oxygen as impurities. The higher content of oxygen about 41.2% was attributed to phytochemicals present in enzyme extract [22]. These serves as a confirmation for organic substrates attached to the copper nanoparticles along with the three small peaks present at 2.0, 2.5 and 2.7 due to P, S and Cl respectively intended for the similar cause.
TEM analysis in Fig. 5a–d at different magnifications showed the nanoparticles were without clusters, globular and crystalline. The average particle sizes analyzed on the basis of TEM scale were found to be in the range of 4.08 ± 1.53 nm as it can be seen in the histogram of Fig. 5e. The TEM image and the histogram also show a narrow size distribution of the nanoparticles; this could be corroborated by Dynamic Light Scattering measurements.
Linear Optical Studies
The pattern of CuO NPs indicates the occurrence of a precipitate at the underneath of the flask which is brown in color. The extract of enzymes of Citrus aurantifolia with 0.5 M CuSO4 solution began to change color 30 min into the reaction and turned completely dark brown after 1 h. A blank test devoid of enzyme extract was colorless indicating no precipitates of CuO NPs.
The Copper oxide nanoparticles yielded a strong absorption band at 335 nm but no such peak was observed with the enzyme extract and it has been well highlighted in Fig. 6. This strong absorption band is due to the transport of electrons from the valence band to the conduction band.
Using the Tauc plot method, the band gap energy is calculated by plotting Energy (eV) vs (αhν)2(eVcm−1)2. The band gap energy of CuO NPs was found to be 3.35 eV as in Fig. 6 which is greater than the bulk CuO due to quantum size defects of semiconductors. There are various factors that influence the band gap namely crystalline size, presence of metal ions as an impurity, or lattice defects (e.g. Cu+1 and O vacancies). As the molar concentration of copper ions increases the band gap value increases because of vibrant transitions involving 3d levels in Cu2+ as well as the strong sp-d exchange interactions between the band electrons and the localized ‘d’ electrons of the dopant [23].
Antibacterial Activity
The anti-bacterial study of copper oxide nanoparticles was established against Klebsiella pneumoniae (gram negative) and S.Aureus (gram positive) as shown in Fig. 7. Maximum activity was seen against Klebsiella pneumoniae and minimum activity was seen against S.Aureus. The antibacterial activity provided by our nanoparticles on Klebsiella pneumoniae and S. aureus were almost similar to the activity provided by the standard. The zone of inhibition values are calculated in Table 2. As per zone of inhibition in Fig. 8, the CuO NPs exhibited more inhibition towards gram negative than gram positive bacteria because of the contrast in their mechanisms. There is a distinction in shape of the cell wall of bacteria as gram negative cell wall comprises of single peptide glycan layer while the gram positive has many peptide glycan layers [24]. Since Cu+ ions and the peptide glycans are oppositely charged, firm binding takes place between them [25]. Large numbers of Cu+ ions are permitted to the plasma membrane of K.Pneumoniae because of the oppositely charged nature [26]. Size, shape and agglomeration pattern of the nanoparticles rely upon amount of phyto chemicals present in medicinal plants. This may be helpful in reducing, capping and stabilization of nanoparticles to limit the size with spherical shape [27]. In this study it is found that Cu NPs synthesized from enzymes of the leaf extract display smaller size particles with circular shape. This is because of the plant Citrus aurantifolia which is a good source for different phyto chemicals especially phenols as reported in previous studies [28].
Anticancer Activity
The cytotoxic effect of enzyme extract from Citrus aurantifolia on SK MEL 28 cells was examined by the MTT assay method. SK MEL 28 cells were treated with different concentrations of enzyme extract for 24 h and are depicted in Fig. 9a–e. The values in Table 3 and Fig. 10 demonstrates that the enzyme extract significantly reduced the viability of the cultured SK MEL 28 cells by 79%, 49%, 27%, 14.5% and 10% at the concentrations 100, 50, 25, 12.5 and 6.25 µg/ml respectively. The half maximal inhibitory concentration [IC50] was counted as the concentration required inhibiting the growth of SK MEL 28 cancer cells in the culture medium by 50%. The IC50 of CuO-NPs was found to be 56.6 μg/ml as shown in Fig. 9f. The IC50 was calculated based on the linear regression.
The significant cytotoxic effect recorded for CuO-NPs is attributed to ion leaching shown by most of the transition metal NPs [29]. At pH 7, the NPs are stable but when these NPs enter into the cancer cell microenvironment at acidic pH, the stability of NPs decrease and these NPs released maximum Cu2+ ions by ion leaching. So, as the dose increases the concentration of intracellular Cu2+ ions are increased in acidic pH which affects the generation of ROS in cancer cells. Similar result was obtained from Chattopadhyay et al. [30]. In our study, the cancer cells was reduced by 79% (Fig. 9e) after the treatment with enzymes of C. aurantifolia derived CuONPs.
Reduced Glutathione sustains a pivotal role to balance ROS (reactive oxygen species) and execution of apoptosis. Now a days in case of cancer therapy, ROS is the promising targeted approach for several drugs [31, 32]. In our metal based nano therapy, the balancing of ROS was perturbed by their redox properties. When oxidation of the reduced glutathione level takes place it is oxidized glutathione and reacts with several proteins using thiol group and assembles inside the cells [33].
Caspase is the key regulatory proteins to activate and implement the process of apoptosis [34] by two pathways namely intrinsic and extrinsic pathways [35]. Thus, stimulation of apoptosis in cancer cell by copper oxide nanoparticles is the vital mechanism [36].
Conclusion
We have demonstrated a simple biological approach to fabricate highly stable copper oxide nanoparticles by green method from the enzymes of Citrus aurantifolia enzyme extract. The antibacterial properties of copper oxide nanoparticle was documented using disk diffusion method and the bactericidal effect of copper nanoparticles has been attributed to the ratio of area available at the apex to volume and small size which let them to interact very thoroughly with microbial membranes. They have exposed more activity on gram-negative than gram-positive bacteria and the zone of inhibition was increasing with the increase of nanoparticle concentration. This approach with compactness, shorter duration, ambient temperature and economical friendly would lead to variety of genesis of nanoparticles. However industrial preparation of nanoparticles would require more research improving the industrial know-how of plant-based synthesis method.
References
M.Maheshwari, I.Ahmad, and A.S.Althubiani, (2016). J Glob Antimicrob. 6, 142–149.
M.N.Alekshun and S.B.Levy (2007). Cell 128, 1037–1050.
J.López-Esparza, L.F.Espinosa-Cristóbal, A.Donohue-Cornejo and S.Y.Reyes-López (2016). Ind Eng Chem Res 55, 12532–12538.
L.Giussani, G.Tabacchi, S.Coluccia, and E.Fois (2019). Int. J. Mol. Sci 20, 2965.
W.Iqbal , B.Yang, X.Zhao, M.Rauf, I.M.A.Mohamed, J.Zhang and Y.Mao (2020). Catal. Sci. Technol 10, 549-559.
S.Magdassi, M.Grouchko and A.Kamyshny (2010). Materials 3: 4626–4638.
A.Rakshit, S.Chowdhury, A.Acharjee, I.Datta, K.Dome, S.Biswas, S.S.Bhattacharyya and B.Saha (2020). Res Chem Intermediat 46, 2559–2578.
J.Singh, G.Kaur, and M.Rawat (2016). J Bioelectron Nanotechnol 1, 1-9.
N.P.S.Acharyulu, R.S.Dubey, P.Kollu, V.Swaminadham, R.L.Kalyani, and S.V.N.Pammi (2014). IJERT 3, 639-641
H.R.Naika, K.Lingaraju, K.Manjunath., D.Kumar, G.Nagaraju, D.Suresh, and H.Nagabhushana (2015). J Taibah Univ Sci 9, 7–12.
K.M. Hajra., J.R. Liu (2004). Apoptosis 6, 691-704.
Z.Y.Wang, Z.Chen (2000). Lancet Oncol 1, 101-106
Y.C.Wang, Y.C.Chuang, Y.H.Ku (2007). Food Chem 102, 1163–1171.
S.Kawaii, Y.Tomono, E.Katase, K.Ogawa, M.Yano (1999). J. Agric. Food Chem 47, 3565–3571.
D.Kupfer, W.H.Bulger (1987). Fed. Proc 46, 1864–1869.
A.Casas, M.Gonzalez de Buitrago, E.Primo (1975). Rev. Agroquim. Tecnol. Aliment 15, 77–88
P.R.Shewry and R.J.Fido, in Protein Extraction from Plant Tissues. (Humana press, New Jersey, 1996), p.24, 25.
N.Abou-Zeid, A.Waly, N.Kandile, A.Rushdy, M.El-Sheikh, and Ibrahim H (2011). Carbohydr. Polym 84, 223–230.
S.Vijayakumar, and S.Ganesan (2012). J Nanomater 2012, 1–9.
M.Qasem, R.El Kurdi, and D.Patra (2020). Chemistry Select 5, 1694–1704
A. Sahai and N. Goswami (2014). Physica E. 58, 130–137.
M.Valodkar, R.N.Jadeja, M.C.Thounaojam, R.V.Devkar, and S.Thakore (2011). Mater. Chem. Phys. 128, 83–89
L.Arun, C.Karthikeyan, D.Philip, D.Dhayanithi, N.V.Giridharan & C.Unni (2018). Opt. Quantum Electron.50, 414
Q.Li, S.Mahendra, D.Y.Lyon, L.Brunet, M.V.Liga, D.Li, and P.J.J.Alvarez (2008). Water Res 42, 4591–4602.
H. Raja Naika, K. Lingaraju, K. Manjunath, Danith Kumar, G. Nagaraju, D.Suresh and H. Nagabhushana (2015). J Taibah Univ Sci 1,7-12
A.L.Koch, S.W.Woeste (1992). J. Bacteriol 17,4811–4819
Saif S, Tahir A, Asim T, Chen Y (2016). Nanomaterials 6, 209
R.Sivaraj, P.K.Rahman, S.M.Rahman, P.Rajiv, S.Narendhran, and R.Venckatesh (2014). Spectrochim. Acta Part A: Mole. Biomol.Spectro 129, 255-258.
L.D. Pacho´n, G. Rothenberg (2008). Appl. Organomet. Chem 22, 288–299.
S. Chattopadhyay, S.K. Dash, S. Tripathy, P. Pramanik, S. Roy (2015). J. Biol. Inorg. Chem 20, 123–141.
S. Choi, C. Filotto, M. Bisanzo, S. Delaney, D. Lagasee, J.L. Whitworth, A. Jusko, C.R. Li, N.A. Wood, J. Willingham, A. Schwenker, K. Spaulding (1998). Inorg. Chem 37, 2500–2504.
M.J. Clarke (2003). Coord. Chem. Rev. 236, 209–233.
M. Valko, C.J. Rhodes, J. Moncol, M. Izakovic, M. Mazur (2006). Chem. Biol. Interact 160, 1–40.
M.O. Hengartner (2000). Nature 407, 770–776.
K.M. Debatin (2004). Cancer Immunol. Immunother 53, 153–159.
P.C.Nagajyothi, P.Muthuraman, T.V.M.Sreekanth, D.H.Kim, J.Shim (2017). Arabian J Chem 10, 215-225.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vincent, J., Lekha, N.C. Bio-Engineered Copper Oxide Nanoparticles Using Citrus Aurantifolia Enzyme Extract and its Anticancer Activity. J Clust Sci 33, 45–53 (2022). https://doi.org/10.1007/s10876-020-01940-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10876-020-01940-2