Abstract
Protein conformational changes play crucial roles in enabling function. The Carr–Purcell–Meiboom–Gill (CPMG) experiment forms the basis for studying such dynamics when they involve the interconversion between highly populated and sparsely formed states, the latter having lifetimes ranging from ~ 0.5 to ~ 5 ms. Among the suite of experiments that have been developed are those that exploit methyl group probes by recording methyl 1H single quantum (Tugarinov and Kay in J Am Chem Soc 129:9514–9521, 2007) and triple quantum (Yuwen et al. in Angew Chem Int Ed Engl 55:11490–11494, 2016) relaxation dispersion profiles. Here we build upon these by developing a third experiment in which methyl 1H double quantum coherences evolve during a CPMG relaxation element. By fitting single, double, and triple quantum datasets, akin to recording the single quantum dataset at static magnetic fields of Bo, 2Bo and 3Bo, we show that accurate exchange values can be obtained even in cases where exchange rates exceed 10,000 s−1. The utility of the double quantum experiment is demonstrated with a pair of cavity mutants of T4 lysozyme (T4L) with ground and excited states interchanged and with exchange rates differing by fourfold (~ 900 s−1 and ~ 3600 s−1), as well as with a fast-folding domain where the unfolded state lifetime is ~ 80 µs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Conformational dynamics are a ubiquitous feature of proteins and are essential for their function. A description of the conformations that are sampled by a protein, therefore, is important (Austin et al. 1975; Frauenfelder et al. 1991; Karplus and Kuriyan 2005; Rosenzweig et al. 2017; Xie 2002), yet in many cases such a description has proven elusive as key states are often sparsely populated and only transiently formed. In such cases the Carr–Purcell–Meiboom–Gill (CPMG) relaxation dispersion NMR experiment (Carr and Purcell 1954; Meiboom and Gill 1958) is particularly powerful since conformational states that are populated to as low as 0.5% and with lifetimes ranging from ~ 0.5 to 5 ms can be detected and characterized at atomic detail. (Palmer et al. 2001; Sauerwein and Hansen 2015; Sekhar and Kay 2013; Zhuravleva and Korzhnev 2017). Indeed, atomic resolution structural models of minor states have been obtained in some cases from CPMG-based studies of exchanging systems (Bouvignies et al. 2011; Hansen et al. 2008b; Korzhnev et al. 2010; Neudecker et al. 2012; Vallurupalli et al. 2008), and CPMG analyses have provided insights into a diverse range of processes that include protein folding and aggregation, ligand binding and enzyme mechanism (Delaforge et al. 2018; Eisenmesser et al. 2005; Korzhnev et al. 2004, 2010; Kovrigin and Loria 2006; Neudecker et al. 2012; Sugase et al. 2007; Tugarinov et al. 2015). It is noteworthy that detection of such minor, short-lived (so called excited) states by conventional biophysical techniques is most often not possible, and in the case of NMR studies, only peaks derived from the highly-populated ground state conformer are observed.
The power of the CPMG experiment lies in the fact that excited states can be studied indirectly through the visible ground state. This is achieved by quantifying the relaxation of transverse magnetization during the course of a fixed interval where variable numbers of chemical shift refocusing pulses are applied (Mulder et al. 2001b). The resultant curve plotting the effective transverse relaxation rate, R2,eff, versus νCPMG, where νCPMG = 1/(4τCPMG) and 2τCPMG is the spacing between successive refocusing pulses, is referred to as a relaxation dispersion profile that is analyzed using a model of chemical exchange to extract, in the case of a two-site exchange reaction, the exchange rate, kex, and the fractional minor state population, pE, along with absolute values of chemical shift differences |Δω| (rad s−1) between the major and minor state resonances. Often accurate exchange parameters (pE, kex) and |Δω| values cannot be obtained from CPMG data collected at a single static magnetic field but require an analysis of data recorded at two well-separated fields. The improved fitting robustness when multiple field data are analyzed reflects the fact that Δω scales with field strength while the kinetic parameters do not change, reducing correlations between the extracted parameters.
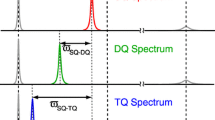
13CH3 methyl groups present an interesting and powerful probe of conformational exchange because it is possible to exploit the three equivalent protons in this spin system to record different classes of 1H CPMG dispersion profiles that may obviate the requirement of multiple field data. For example, we have previously shown that single (SQ) (Tugarinov and Kay 2007) and triple (TQ) quantum (Yuwen et al. 2016) methyl 1H CPMG dispersion profiles can be recorded that depend on Δω (SQ) and 3Δω (TQ), respectively, reflecting the fact that SQ and TQ transitions involve 1 and 3 quanta, respectively, Fig. 1. The TQ experiment increases the sensitivity of the dispersion technique to exchange processes that involve small chemical shift differences between converting states, as well as facilitating the study of faster exchange processes than would otherwise be possible using the more traditional SQ class of experiment (Yuwen et al. 2016). An additional advantage of the TQ experiment in relation to its SQ counterpart is one of sensitivity. As discussed in detail previously, the application of a 1H 180° refocusing train during the CPMG period leads to artifacts in SQ experiments that do not eliminate signal from the 3/2 manifold (Korzhnev et al. 2005). This results from the fact that in methyl groups the fast and slowly relaxing 1H transitions (Figs. 1, 3/2 manifold) are interconverted due to pulse imperfections that lead to non-flat dispersion profiles even in the absence of chemical exchange. As a result an experiment was developed that selects signal from only the spin ½ manifolds (Fig. 1) where such an effect cannot occur (Tugarinov and Kay 2007). The resulting dispersions are artifact-free but the sensitivity of the experiment is reduced considerably to approximately 1/10th that of the TQ CPMG (Yuwen et al. 2016). We were interested, therefore, in developing a double quantum (DQ) scheme that could be used in applications where the sensitivity of the SQ experiment was limiting. As DQ-CPMG profiles are sensitive to chemical shift differences of 2Δω it is expected that a combined analysis of DQ and TQ data sets would provide similar benefits to data obtained at multiple fields. This is of particular importance in NMR laboratories where only a single static magnetic field is available, such as is the case for a subset of the authors of the present paper (ABG and PV). We were also interested in establishing whether it would be possible to extend studies to faster exchange processes by a combined analysis of DQ and TQ CPMG data. Here we present a methyl 1H DQ CPMG pulse scheme and demonstrate its utility using two cavity mutants of T4 lysozyme (T4L, 18 kDa) in which a hydrophobic residue interconverts between conformers that place it either inside or out of the cavity (Bouvignies et al. 2011; Vallurupalli et al. 2016). In the first cavity mutant a Leu at position 99 has been replaced by Ala, referred to as T4L L99A (Baase et al. 2010; Eriksson et al. 1992), while in the second construct the three mutations L99A, G113A, R119P are used (referred to as T4Ltm) (Bouvignies et al. 2011). A third system, the fast-folding peripheral subunit binding domain (PSBD) of the acetyl transferase from Bacillus stearothermophilus (4.7 kDa) that interconverts between folded and unfolded conformations (Vugmeyster et al. 2000) is studied as well. Our results establish that a combined analysis of DQ and TQ CPMG datasets provides more robust measures of exchange parameters for fast exchanging systems as PSBD (kex ~ 12,000 s−1) than when TQ CPMG data from a single field exclusively is analyzed, removing the correlation between pE and Δω values that is otherwise present.
Energy level diagram of an X3 spin system, separating levels according to the net spin angular momentum of each state |j〉 (indicated below) and highlighting 1H SQ, DQ and TQ coherences. SQ (purple, red), DQ (green) and TQ (blue) coherences can be excited within the spin 3/2 manifold. Single quantum transitions relax very differently in the macromolecular limit (Kay and Torchia 1991; Tugarinov et al. 2003, 2006) and have been labeled as SQfast (purple, fast relaxing) and SQslow (red, slow relaxing). ZQ coherences are not depicted but are contained within the |2><2| and |3><3| elements of the 3/2 manifold, and within other elements in each of the ½ manifolds. However, as the DQ coherences are those of interest and these are only generated within the 3/2 manifold, only elements within this manifold need be considered as pulses cannot interconvert density elements in separate manifolds. Each of the levels depicted in the figure is further split due to scalar coupling involving the attached methyl 13C spin
Materials and methods
Sample preparation
U-[2H,15N], Ileδ1-[13CH3], Leu,Val-[13CH3,12CD3]-labeled T4L wild-type (T4L), T4L L99A, T4Ltm and PSBD were expressed in E. coli BL21(DE3) cells grown in 100% D2O M9 media with 1 g L−1 15NH4Cl and 3 g L−1 U-[2H,12C] glucose as the sole nitrogen and carbon sources. Appropriate precursors were added to the media one hour before induction (Goto et al. 1999; Ollerenshaw et al. 2005; Tugarinov and Kay 2005). The proteins were purified by ion exchange chromatography followed by size exclusion chromatography as described previously (Gopalan and Vallurupalli 2018; Vallurupalli et al. 2009). The T4L samples comprised ~ 1.5 mM protein dissolved in a 50 mM NaH2PO4, 25 mM NaCl, 2 mM EDTA, 2 mM NaN3, 10% D2O, pH 5.5 buffer. The concentration of protein in the PSBD sample was under 0.5 mM, dissolved in 50 mM NaH2PO4, 25 mM NaCl, 2 mM EDTA, 2 mM NaN3, 10% D2O, pH 6.5 buffer.
NMR experiments
All experiments were performed on a Bruker Avance III HD 700 MHz spectrometer equipped with a cryogenically cooled probe. Methyl (13CH3) 1H SQ and TQ experiments were performed using published pulse sequences (Tugarinov and Kay 2007; Yuwen et al. 2016), while the DQ CPMG experiment was recorded as described below. A constant-time CPMG relaxation delay (Trelax) of 20 ms was used for methyl 1H SQ, DQ and TQ CPMG experiments recorded on T4L L99A, along with νCPMG values in the range 50–1500 Hz. Trelax was set to 10 ms for the DQ and TQ CPMG experiments recorded on T4Ltm and PSBD samples and 15 ms for the SQ CPMG experiment recorded on the T4Ltm sample with νCPMG values between 100 and 2000 Hz. Amide 1HN and 15N CPMG data were recorded on PSBD at 41 °C to demonstrate the accuracy of exchange parameters extracted from the methyl 1H DQ experiment, as described below. 1HN CPMG data were recorded using a pulse scheme that does not contain a P-element (Ishima and Torchia 2003; Vallurupalli et al. 2011) and the 15N TROSY CPMG experiment (Loria et al. 1999b; Vallurupalli et al. 2007) was used to record amide 15N data. Trelax was set to 15 ms, νCPMG values ranged between 133 and 2467 Hz in the 1HN CPMG experiment and between 67 and 1000 Hz in the 15N CPMG experiment.
Pseudo-3D CPMG datasets were recorded using 12–15 different νCPMG values in an interleaved manner with an inter-scan delay of 2.25 s and 2–3 repetitions for error estimation (Gopalan et al. 2018; Korzhnev et al. 2004). The total acquisition time for each dataset varied between 10 and 16 h except for measurements on the dilute PSBD sample that required up to 27 h.
Data analysis
All NMR data was processed using NMRPipe (Delaglio et al. 1995), spectra were visualized with SPARKY (Goddard and Kneller 2008; Lee et al. 2015) and the program PINT (Ahlner et al. 2013) was used for integrating peaks and computing intensities as a function of νCPMG, I(νCPMG). R2,eff rates were calculated for each 1H or 15N site according to (Mulder et al. 2001b),
where Io is the intensity of the peak in a reference plane recorded without the Trelax delay.
Exchange parameters were obtained from the experimental data via a standard best-fit procedure using the program ChemEx (Bouvignies 2011) that integrates the Bloch–McConnell equations (McConnell 1958) numerically (Korzhnev et al. 2004) to minimize the function \({\chi ^2}=\mathop \sum \nolimits_{{i=1}}^{N} {\left[ {\left( {I_{i}^{{Exp}} - I_{i}^{{Calc}}} \right)/{\sigma _i}} \right]^2}\), where \(I_{i}^{{Exp}}\) are the experimentally measured peak intensities, \(I_{i}^{{Calc}}\) are the intensities calculated by integrating the Bloch-McConnell equations numerically and \({\sigma _i}\) is the uncertainty in the experimentally measured intensity value. Uncertainties in the fitted parameters were estimated using 1000 trials of standard Monte Carlo or bootstrap analysis (Press et al. 1992). In the simplest version of the fitting procedure DQ data is analyzed in a manner similar to SQ data, but with the evolution frequencies and 1JCH (one bond 13C–1H scalar coupling) values doubled (Yuwen et al. 2016) so that magnetization evolves under chemical exchange (\(G{\mathop{\leftrightarrows}\limits^{k_{EG}}_{k_{GE}}}E\)) according to
In Eq. (2) \(M_{{IP}}^{{}}=\frac{1}{2}\left( {H_{+}^{1}H_{+}^{2}+H_{+}^{2}H_{+}^{3}+H_{+}^{3}H_{+}^{1}} \right)\) and \(M_{{AP}}^{{}}={C_Z}\left( {H_{+}^{1}H_{+}^{2}+H_{+}^{2}H_{+}^{3}+H_{+}^{3}H_{+}^{1}} \right)\) are methyl 1H DQ coherences that are in-phase (IP) or anti-phase (AP) with respect to the methyl 13C spin, respectively, ΩG (ΩE) is the difference between the resonance frequency of a spin in the ground (excited) state and the carrier frequency (rad/s), and \(R_{{2\infty ,IP}}^{{DQ}}\), \(R_{{2\infty ,AP}}^{{DQ}}\) are the intrinsic relaxation rates of IP and AP DQ coherences, respectively. In the derivation of Eq. (2) we have not included cross relaxation between IP and AP DQ coherences that arises from intra-methyl 1H–1H, 1H–13C cross-correlated dipolar interactions. Simulations show that such effects are small (see Supporting Information). Fitting parameters for the DQ experiment include the global values kex = kGE + kEG and the fractional population of the excited state pE (pG + pE = 1), with residue specific chemical shift differences Δϖ (ppm) and \(R_{{2\infty ,IP}}^{{DQ}}\) fit as well. In general, methyl 1H DQ CPMG profiles are fit along with SQ and TQ curves that require additional fitting parameters \(R_{{2\infty ,IP}}^{{SQ}}\) and \(R_{{2\infty ,IP}}^{{TQ}}\). The differences in the relaxation rates between the IP and AP DQ and TQ coherences \(\Delta R_{{IP - AP}}^{{DQ}}=R_{{2\infty ,IP}}^{{DQ}} - R_{{2\infty ,AP}}^{{DQ}}\) and \(\Delta R_{{IP - AP}}^{{TQ}}=R_{{2\infty ,IP}}^{{TQ}} - R_{{2\infty ,AP}}^{{TQ}}\) are related to each other according to \(\Delta R_{{IP - AP}}^{{DQ}} \approx \frac{1}{3}{{{\Delta}}}R_{{IP - AP}}^{{TQ}}\). \(\Delta R_{{IP - AP}}^{{TQ}}\) can be measured, as described earlier (Yuwen et al. 2016), from which \(\Delta R_{{IP - AP}}^{{DQ}}\) is obtained using the above relation. As \(\Delta R_{{IP - AP}}^{{TQ}}\) values are small, ~ 0.6 s−1 for T4Ltm at 15 °C and ~ 1.3 s−1 for PSBD at 41 °C, we find that the same exchange parameters are obtained even when \(\Delta R_{{IP - AP}}^{{TQ}}\) (or \(\Delta R_{{IP - AP}}^{{DQ}}~\) in the case when only DQ data is analyzed) is treated as an additional fitting parameter or when it is set to a reasonable value (~ 1 s−1).
Results and discussion
Figure 1 illustrates the energy level diagram for a spin system comprising three equivalent protons, as in a methyl group. For the case of 13CH3 methyls, considered here, each of the energy levels is further split by the spin state of the 13C (not shown). Notably, there are a number of distinct 1H transitions that can be distinguished on the basis of their relaxation properties or frequencies. These include SQ transitions that relax either rapidly (purple) or slowly (red) (Kay and Prestegard 1987; Tugarinov et al. 2003; Werbelow and Marshall 1973a, b) as well as the DQ (green) and TQ (blue) transitions that were mentioned in the Introduction. Previous pulse schemes have isolated either SQslow transitions from the spin ½ manifolds (Tugarinov and Kay 2007) or TQ transitions (Yuwen et al. 2016) in the context of 1H CPMG experiments and we focus here on the corresponding 1H DQ experiment.
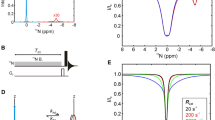
Figure 2 illustrates the methyl 1H DQ CPMG experiment that we have developed. In the brief description of the pulse scheme that follows we label each of the protons as 1, 2 or 3 to distinguish them and follow the trajectory of the coherences of interest throughout the sequence, neglecting the effects of pulse imperfections and spin relaxation. Starting from equilibrium, 1H magnetization, \(H_{z}^{1}+H_{z}^{2}+H_{z}^{3}\) (point A), is subsequently evolved during a refocused INEPT element (Morris and Freeman 1979) to generate \(4\left( {H_{z}^{2}H_{z}^{3}+H_{z}^{1}H_{z}^{3}+H_{z}^{1}H_{z}^{2}} \right){C_z}\) at point B. The subsequent 1H 90° pulse at point C converts the longitudinal order 1H components into an equal combination of ZQ and DQ coherences at the start of the CPMG period that can be readily noted by considering the action of a 1H pulse of phase ϕ1 = x,
Pulse sequence for measurement of methyl 1H DQ CPMG relaxation dispersion profiles on highly deuterated, 13CH3 labeled proteins. Narrow (wide) black bars denote 90° (180°) pulses, applied along the x-axis unless otherwise indicated, while grey bars indicate z-pulsed field gradients. All pulses are applied at the highest possible power level except for the 1H pulses during the CPMG period where a ~ 22 kHz field was used. The 1H carrier is set to the water resonance for the EBURP-1 pulse (Geen and Freeman 1991) at the start of the experiment (shaped pulse) and subsequently moved to the center of the ILV methyl resonance region ~ 0.8 ppm for the rest of the experiment. 2H and 13C carriers are placed at ~ 0.8 ppm and ~ 18 ppm, respectively. The delay τa is set to 1/(41JCH) (2 ms). 13C and 2H WALTZ-16 decoupling (Shaka et al. 1983) elements are applied with B1 field strengths of ~ 1.8 and ~ 0.55 kHz, respectively; 2H decoupling is required only when νCPMG values < 100 Hz are used (Yuwen et al. 2016). Phases are cycled as: ϕ1 = − 45°, 45°; ϕ2 = 45°, 135°; ϕ3 = 2y, 2(− y); ϕ4 = − 45°; ϕ5 = 4x, 4(− x); ϕ6 = x; ϕ7 = x, − x; receiver = 2(x, − x), 2(− x, x). Phases of the CPMG pulses were incremented according to the XY-16 scheme (Gullion et al. 1990) by varying Ψ1 and Ψ2 as described previously (Yuwen et al. 2016). Briefly the phases of the (regular, i.e., not composite) π pulses in the first half of the CPMG train are incremented according to Ψ1 = (x,y,x,y,y,x,y,x,− x,− y,− x,− y,− y,− x,− y,− x) followed by the central πϕ3 pulse, after which Ψ2 is decremented in the reverse order of the increase in Ψ1 and phases y and –y are inverted to reduce off-resonance effects (Hansen et al. 2008a). Thus the scheme is: \({\tau _{{\text{CP}}}}{\Pi _1}{\tau _{{\text{CP}}}}{\tau _{{\text{CP}}}}{\Pi _2}{\tau _{{\text{CP}}}} \ldots {\tau _{{\text{CP}}}}{\Pi _{{\text{k}} - 1}}{\tau _{{\text{CP}}}}{\tau _{{\text{CP}}}}{\Pi _{\text{k}}}{\tau _{{\text{CP}}}}{\Pi _{\Phi 3}}{\tau _{{\text{CP}}}}{\bar {\Pi }_{\text{k}}}{\tau _{{\text{CP}}}}{\tau _{{\text{CP}}}}{\bar {\Pi }_{{\text{k}} - 1}}{\tau _{{\text{CP}}}} \ldots {\tau _{{\text{CP}}}}{\bar {\Pi }_2}{\tau _{{\text{CP}}}}{\tau _{{\text{CP}}}}{\bar {\Pi }_1}{\tau _{{\text{CP}}}}\) where \({\Pi _1}\) is \({\Pi _{\text{x}}}\), \({\Pi _2}\) is \({\Pi _{\text{y}}}\) and so on according to the XY-16 scheme (Ψ1). \({\bar {\Pi }_{\text{k}}}\) is obtained from \({\Pi _{\text{k}}}\) by replacing y with − y and − y with y. 13C t1 evolution is carried out in a shared constant time manner (Grzesiek and Bax 1993; Logan et al. 1993), as indicated. Complex data in the indirect dimension is obtained by incrementing ϕ6 and the receiver in a States-TPPI manner (Marion et al. 1989). Gradients (Strengths in G/cm: durations in µs) are: g0 (10:1000), g1 (12.5:300), g2 (13.5:500), g3 (15:300), g4 (20:500), g5 (25:500), g6 (18.5:300), g7 (− 25:500). Water suppression is achieved via the ~ 8 ms water selective EBURP-1 pulse followed by the g0 gradient to destroy water magnetization at the start of the sequence and by the 3-9-19 WATERGATE scheme during the last INEPT (Sklenar et al. 1993)
The relevant DQ terms \(\left( {H_{+}^{i}H_{+}^{j}+H_{ - }^{i}H_{ - }^{j}} \right){C_z}\) are selected by a two-step cycle of ϕ1 that filters out the ZQ coherences (Bain 1984; Cavanagh et al. 2006; Ernst et al. 1987; Gopalan and Vallurupalli 2018). After the Trelax delay 13C chemical shift is recorded during a t1 interval with the subsequent transfer of magnetization back to 1H for detection. Here, as in the methyl 1H TQ CPMG experiment (Yuwen et al. 2016) and other 1H CPMG experiments (Orekhov et al. 2004; Vallurupalli et al. 2011), we have chosen not to use a P-element (Loria et al. 1999a) to compensate for differential relaxation between DQ coherences that are in- and anti-phase with respect to 13C because the difference between the two rates is sufficiently small (\(\Delta R_{{IP - AP}}^{{DQ}}\) < 1.5 s−1, see Materials and methods section) that it can be readily accounted for in the fitting procedure after it is estimated from \(\Delta R_{{IP - AP}}^{{TQ}}~\) (Yuwen et al. 2016) and because the extra delays introduced by the P-element reduce the sensitivity of the experiment.
The main sources of relaxation of methyl 1H DQ coherences are dipolar interactions with external protons as well as interactions involving intra-methyl 13C–1H and 1H–1H dipole pairs (see Supporting Information). External contributions can be minimized by using a highly 2H-enriched sample where only one of the two prochiral methyl groups of Leu and Val is labeled, as has been done here (see Materials and methods section). The 2H, 12C labelled background ensures the absence of homonuclear 1H–1H scalar couplings that would deteriorate the quality of the 1H dispersion profiles; elimination of 13C–13C scalar couplings is also important to the sensitivity and resolution of the resulting dispersion datasets. Based on similar considerations, an identical labeling scheme has been used for methyl 1H TQ CPMG experiments as well (Yuwen et al. 2016).
As a first test of the pulse scheme we wanted to establish that flat dispersion profiles are obtained for non-exchanging methyl sites. To this end we considered PSBD (~ 5 kDa) and identified eight methyl groups that were not sensitive to the exchange between folded and unfolded forms in this system, as established by flat dispersion curves from the TQ CPMG scheme (\(R_{{2,eff}}^{{TQ}}\left( {100~{\text{Hz}}} \right) - R_{{2,eff}}^{{TQ}}\left( {2000~{\text{Hz}}} \right)\) ≤ 2 s−1). As shown in Fig. 3a dispersion profiles are obtained with root-mean-squared-differences (RMSD) < 0.7 s−1 between experimental R2,eff values and values of the best fit horizontal lines. The replacement of ‘simple’ refocusing 180° pulses (referred to as ‘regular’) with composite pulses also results in flat dispersion profiles. The situation is different when the DQ experiment is recorded on a sample of wild-type T4 lysozyme, T4L (Baase et al. 2010), an 18 kDa protein that has been shown previously not to exchange within the millisecond time window. Although flat dispersion profiles were expected due to the absence of chemical exchange, we found small, non-flat \(\Delta R_{{2,eff}}^{{DQ}}\left( {{\nu _{CPMG}}} \right)\) profiles for all peaks when composite pulses were used in the CPMG train (Fig. 3b). These apparent dispersions were significantly less pronounced with regular refocusing pulses. Simulations show that during a CPMG pulse, starting DQ coherence evolves into diagonal density elements that include ZQ and in addition, while net SQ magnetization does not build up, the density elements connecting the fast-relaxing single-quantum 1, 2 and 3, 4 transitions (Fig. 1) are also non-zero. In Fig. 3c we illustrate the evolution of different density elements during the course of a regular (left) or composite (right) pulse, with groupings of density elements that relax with similar rates considered together. The intrinsic relaxation rates (\(R_{{2\infty }}^{{}}\) that does not include exchange contributions) of the DQ and individual SQ transitions are distinct, as is the relaxation of the on-diagonal density elements, with rates of DQ coherences (\(R_{{2\infty }}^{{DQ}}\) ~ 120 s−1) higher than for ZQ (\(R_{{2\infty }}^{{ZQ}}\) ~ 6 s− 1), SQfast (\(R_{{2\infty }}^{{S{Q_{fast}}}}\) ~ 95 s− 1), SQslow (\(R_{{2\infty }}^{{S{Q_{slow}}}}\) ~ 12.5 s− 1) and TQ (\(R_{{2\infty }}^{{TQ}}\) ~ 60 s− 1) coherences in the case of T4L at 15 °C where the tumbling time, τC is estimated to be ~ 14.3 ns from the measured value of 10.8 ns at 25 °C (Skrynnikov et al. 2001). Thus, R2,eff values become a function of νCPMG and the pulse duration, decreasing for higher νCPMG values as the contribution from the ZQ elements (and the other diagonal density terms) that are generated during the pulse (Fig. 3c) becomes more significant. A similar phenomenon is observed in dispersion profiles recorded using ‘traditional’ SQ CPMG schemes when \({R_{2\infty }} \gg {R_1}\) that in this case results from off-resonance effects (Jiang et al. 2015; Yip and Zuiderweg 2004), leading to an interconversion between transverse and longitudinal magnetization during the course of the pulse; this can be readily taken into account in fitting software by including a basis set consisting of Iz, I+ and I− operators (Hansen et al. 2008a). Similarly, the ‘apparent’ dispersions seen with the composite pulses can be accounted for by using a basis set that consists of all of the density elements in the spin 3/2 manifold, taking into account the appropriate intrinsic relaxation rates for the different terms (See Supporting information and Fig. S1). However, for the T4L data recorded using regular 180° refocusing pulses we find that essentially the same exchange parameters are obtained from an analysis that considers just the evolution of the DQ coherences (Eq. 2) rather than a more elaborate fit that accounts for all the transitions in the spin 3/2 manifold.
Flat methyl 1H DQ CPMG dispersions are obtained in the absence of exchange when regular π pulses are used during the CPMG element. a Flat dispersions at sites where Δϖ ~ 0 ppm in PSBD at 41 °C. RMSD values from the y = 0 line are shown. Here \(\Delta R_{{2,eff}}^{{DQ}}\left( {{\nu _{{\text{CPMG}}}}} \right)=R_{{2,eff}}^{{DQ}}\left( {{\nu _{{\text{CPMG}}}}} \right) - R_{{2,eff}}^{{DQ}}\left( {{\nu _{{\text{CPMG}},{\text{max}}}}} \right)\) where \({\nu _{{\text{CPMG}},{\text{max}}}}\) is the maximum value of \({\nu _{{\text{CPMG}}}}\) used in the experiment. b Apparent dispersions are observed even in the absence of exchange with a dataset recorded on wild-type T4L at 15 °C (τC ~ 14.3 ns) using composite pulses but not when regular π pulses are used. c Evolution of coherences starting from DQ \(\propto \left( {H_{+}^{1}H_{+}^{2}+H_{+}^{2}H_{+}^{3}+H_{+}^{3}H_{+}^{1}+H_{ - }^{1}H_{ - }^{2}+H_{ - }^{2}H_{ - }^{3}+H_{ - }^{3}H_{ - }^{1}} \right)\) during a regular π pulse of x phase and a composite 90−y180x90−y π pulse (Levitt and Freeman 1979). The plots were calculated by propagating the Liouville-von Neumann equation for the spin 3/2 manifold (Fig. 1) using the basis functions for the 4 × 4 density matrix as shown in Fig. 1. In the starting density matrix \(\rho (0)\), the elements corresponding to the DQ transitions are each set to ½, \({\rho _{1,3}}(0)={\rho _{3,1}}(0)={\rho _{2,4}}(0)={\rho _{4,2}}(0)=0.5\), and all other elements are set to 0. Relaxation is not included. Here we define \(M_{{Diag}}^{2}=\rho _{{1,1}}^{2}+\rho _{{2,2}}^{2}+\rho _{{3,3}}^{2}+\rho _{{4,4}}^{2}\), \(M_{{S{Q_{fast}}}}^{2}=\rho _{{1,2}}^{2}+\rho _{{2,1}}^{2}+\rho _{{3,4}}^{2}+\rho _{{4,3}}^{2}\), \(M_{{S{Q_{slow}}}}^{2}=\rho _{{2,3}}^{2}+\rho _{{3,2}}^{2}\), \(M_{{DQ}}^{2}=\rho _{{1,3}}^{2}+\rho _{{3,1}}^{2}+\rho _{{2,4}}^{2}+\rho _{{4,2}}^{2}\), \(M_{{TQ}}^{2}=\rho _{{1,4}}^{2}+\rho _{{4,1}}^{2}~\)and \(\rho _{{i,j}}^{2}=\rho _{{i,j}}^{*}\rho _{{i,j}}^{{}}.\) \(M_{{ZQ}}^{2}\) is subsumed within the \(\rho _{{2,2}}^{2}+\rho _{{3,3}}^{2}\) terms of \(M_{{Diag}}^{2}.~\)Note that \(M_{{Diag}}^{2}~+~M_{{S{Q_{fast}}}}^{2}+M_{{S{Q_{slow}}}}^{2}+M_{{DQ}}^{2}+M_{{TQ}}^{2}\) is constant throughout the pulse. SQslow and TQ coherences are not generated during these pulses
Having established that flat dispersion profiles can be obtained so long as regular CPMG refocusing pulses that are as short as possible are used we next recorded a 1H DQ CPMG dataset on a sample of T4Ltm that has been shown previously to exchange between two compact conformers, including a major state where Phe114 is buried inside a cavity created by the L99A mutation and a minor conformer where the Phe114 sidechain is exposed to solvent (Bouvignies et al. 2011; Vallurupalli et al. 2016). Methyl 1H DQ CPMG dispersion curves from 12 peaks with \(R_{{2,eff}}^{{DQ}}\left( {100~{\text{Hz}}} \right) - R_{{2,eff}}^{{DQ}}\left( {2000~{\text{Hz}}} \right)>12\;{{\text{s}}^{ - 1}}\) were fit globally to a two-state process (Fig. 4a). Exchange parameters kex = 3548 ± 98 s− 1, pE = 4.8 ± 0.1%, extracted from analysis of the DQ CPMG data, are in good agreement with kex = 3775 ± 66 s−1, pE = 4.9 ± 0.1% obtained from a combined analysis of 1H TQ and SQ CPMG datasets. Notably, flat dispersion profiles (Fig. 4a) were obtained when ZQ rather than DQ coherences were selected during the CPMG interval (RMSD < 1.0 s−1). This is to be expected because methyl 1H ZQ elements do not evolve under chemical shift; the result further confirms the robustness of the experiment. Finally, Fig. 4b shows a very good correlation between Δϖ values fit independently from analysis of the DQ or the SQ, TQ CPMG data, confirming that the DQ CPMG experiment is robust in this exchange regime.
Selected methyl 1H CPMG dispersion profiles recorded on T4Ltm at 15 °C. a DQ and ZQ relaxation dispersion profiles; although large DQ dispersions are observed the ZQ profiles are flat. The solid green lines derive from a global fit of the DQ CPMG data to a two-site exchange model. The RMSD between the ZQ CPMG data and the y = 0 line is also shown. b Correlation between Δϖ values obtained from the global analysis of the DQ CPMG dataset and a combined analysis of SQ/TQ CPMG profiles. Signs of Δϖ were obtained previously from exchange induced peak shifts between methyl 1H SQ, DQ and TQ spectra (Gopalan and Vallurupalli 2018). c Site-specific combined analysis of SQ, DQ and TQ methyl 1H CPMG data. d Distribution of the kex and pE values obtained from site-specific fits of the CPMG data as indicated. Site-specific kex and pE values (red) are very similar to the value obtained from a global analysis of all the CPMG data (blue). Data points with the same numbers in each panel derive from the same residue
It is well established that the accuracy of extracted exchange parameters generally improves upon analysis of CPMG profiles recorded at different static magnetic field strengths (Gopalan and Vallurupalli 2018; Korzhnev et al. 2004; Kovrigin et al. 2006; Millet et al. 2000; Vallurupalli et al. 2011). Recording 1H SQ, DQ and TQ CPMG data is analogous to measuring SQ profiles at field strengths of Bo, 2Bo and 3Bo since the energy separation between levels connected by DQ and TQ transitions is two- and three-fold larger than for SQ transitions (Gopalan and Vallurupalli 2018; Yuwen et al. 2016). Notably, when 1H SQ, DQ and TQ CPMG datasets were analyzed together (Fig. 4c) on a per-residue basis the resulting exchange parameters for 7 peaks with |Δϖ| ≥ 0.2 ppm clustered (Fig. 4d), consistent with a two-state chemical exchange process. This is an important result because it reinforces the fact that a comparison of (pE, kex) values obtained on a per-residue basis from analysis of multiple datasets can be used to establish whether a global exchange process is occurring. Often only kex can be obtained reliably from analysis of per-residue CPMG data so that, in these cases, identification of multi-site exchange is based only on variations of kex (Vallurupalli and Kay 2006). Robust estimates of pE values from single-site data improves the identification and characterization of multi-state chemical exchange occurring on the µs to ms timescale.
Next we explored the possibility of using the 1H DQ CPMG experiment to characterize the folding reaction of the PSBD protein domain (Vugmeyster et al. 2000) that folds with a rate of ~ 13,000 s−1 at 41 °C (Gopalan and Vallurupalli 2018). The low concentration of the sample (< 0.5 mM) and the short lifetime of the unfolded minor state precluded recording methyl 1H SQ CPMG data but DQ and TQ CPMG profiles were obtained (Fig. 5a), as the corresponding DQ and TQ CPMG experiments are more sensitive than their SQ counterpart (Yuwen et al. 2016). The population of the minor state was not well defined based on analysis of TQ profiles (Fig. 5b, c) recorded at a single field yet exchange parameters from a combined analysis of DQ and TQ profiles from six sites, kex = 13,280 ± 779 s−1 and pE = 4.0 ± 0.2%, (Fig. 5b, c), could be estimated with confidence. These values are in reasonably good agreement with previously determined parameters, kex = 12,911 ± 300 s−1 and pE = 3.5 ± 0.1%, based on an analysis of amide 1HN, 15N and methyl 1H TQ CPMG experiments along with exchange-induced peak shifts between 15N–1H HSQC, HMQC spectra and between 13C–1H SQ,DQ and SQ,TQ correlation maps recorded on a more concentrated sample (Gopalan and Vallurupalli 2018). The population of the unfolded state and consequently the unfolded state chemical shifts could not be estimated using amide 1HN and 15N CPMG data alone. Yet unfolded state amide 1HN and 15N chemical shifts could be obtained by analyzing the 1HN and 15N CPMG data when kex and pE were fixed to values obtained from the present analysis. The good agreement between experimentally derived Δϖ values and those predicted (Tamiola et al. 2010) for an unfolded state (Fig. 5d) nicely confirms that the pE value obtained from the joint analysis of DQ and TQ CPMG profiles is correct.
Exchange processes on the order of 10,000 s−1 can be studied using a combination of methyl 1H DQ and TQ CPMG experiments. a The two largest experimental TQ and DQ CPMG relaxation dispersion profiles from the fast folding PSBD are shown in blue and green, respectively (41 °C). b 1D reduced χ2 versus pE profiles establishing that the unfolded state population cannot be defined by analysis of the single field TQ CPMG data alone (blue) but that a defined pE value is obtained when the DQ and TQ CPMG data are analyzed together (green). (c) Scatter plot showing the extracted exchange parameters from 1000 bootstrap trials of the TQ CPMG data alone (blue) and in combination with DQ profiles (green). The scatter is reduced to a great extent when the DQ CPMG data is included in the analysis. (d) Differences between predicted unfolded state amide 1HN and 15N chemical shifts (Tamiola et al. 2010) and the corresponding chemical shifts of the folded state compared to Δϖ values obtained by analyzing amide 1HN or 15N CPMG data. In the analysis kex and pE have been fixed to the values obtained from the combined analysis of the methyl 1H DQ and TQ CPMG experiments. The good correlation between the values shows that the exchange parameters obtained from the analysis of DQ and TQ CPMG data are accurate. Signs of the amide 15N Δϖ values were determined previously (Gopalan and Vallurupalli 2018) by analyzing peak shifts between HSQC and HMQC spectra (Skrynnikov et al. 2002)
A limitation of the methyl 1H DQ experiment is that the intrinsic relaxation rates of methyl 1H DQ coherences are large, significantly higher than those of SQslow or TQ coherences (Fig. 6a). This is a consequence of the fact that intra-methyl 1H-1H dipolar interactions do not contribute spectral densities at zero-frequency to the relaxation rates of SQslow or TQ elements but do so for the relaxation of DQ coherences. The higher relaxation rates of DQ coherences do place limits on Trelax delays that can be used in this experiment. For the small protein PSBD \(R_{{2\infty }}^{{DQ}}\) = 19 ± 4 s−1 at 41 °C (τC ~ 2 ns, estimated from molecular weight) and for Trelax = 36 ms there is approximately 50% loss in signal. For T4L, that is approximately 3.5 fold larger than PSBD, \(R_{{2\infty }}^{{DQ}}\) = 87 ± 18 s− 1 at 22 °C (τC ~ 11.7 ns), while \(R_{{2\infty }}^{{S{Q_{slow}}}}\) = 10 ± 3 s−1 and \(R_{{2\infty }}^{{TQ}}\) = 56 ± 16 s−1 (Fig. 6a). In this case the signal to noise ratio (S/N) of the DQ CPMG experiment is 2.9 ± 0.9 times higher than for the SQ CPMG experiment when Trelax is 10 ms (when Trelax=0 the S/N of the DQ experiment is ~ eightfold higher), reflecting the fact that 50% of the starting magnetization is utilized in the DQ CPMG scheme as opposed to only 1/18th in the case of the SQ CPMG (Tugarinov and Kay 2007). For Trelax = 20 ms the S/N of DQ CPMG spectra are only marginally better than for the SQ datasets (1.6 ± 0.7). The significant decay of DQ coherences for Trelax = 20 ms for 20 kDa proteins (such as T4L) suggests that 20 ms may be an upper bound for the constant-time CPMG relaxation delay, limiting studies to exchanging systems with kex > ~ 300 s−1. For larger proteins the utility of the DQ CPMG experiment will be restricted to studies of even faster exchanging systems because of more severe limitations on the maximum size of Trelax. By means of example, Fig. 6a, c plot relaxation rates and relative S/N values for cross peaks from datasets acquired on T4L at 5 °C (τC ~ 19.8 ns). For Trelax = 10 ms the relative S/N between the DQ and SQ CPMG experiments is reduced to 1.3 ± 0.6 (from 2.9 ± 0.9 at 22 °C). As expected, for all of the conditions explored here the TQ CPMG experiment has the highest sensitivity as 75% of the starting magnetization is preserved (neglecting relaxation) and \(R_{{2\infty }}^{{TQ}}<R_{{2\infty }}^{{DQ}}.\)
Relative sensitivities of methyl 1H SQ, DQ and TQ CPMG datasets recorded on a wild-type T4L sample (~ 18 kDa) that does not undergo chemical exchange. a Relaxation rates for DQ coherences are significantly larger than for the TQ and the SQslow coherences. b Relative intensities of peaks in the reference plane (Trelax = 0) and in planes with νCPMG = 2000 Hz and Trelax = 10 or 20 ms for the SQ, DQ and TQ CPMG experiments recorded at 22 °C (τC ~ 11.7 ns). The large sensitivity differences reflect, to a large extent, the fact that only coherences from the spin ½ manifolds are selected in the SQ experiment, whereas 1/2 and 3/4th of the starting 1H magnetization is retained in the DQ and TQ CPMG experiments, respectively, in the absence of relaxation. c As in (b) but for data recorded at 5 °C where the rotational correlation time of T4L is ~ 19.8 ns
The above discussion establishes that the DQ experiment becomes increasingly insensitive as the size of the protein increases and as kex becomes slow since large Trelax values would then be required. We were interested, therefore, in demonstrating the utility of the DQ method for moderate exchange rates, on the order of 1000 s−1, and for protein systems in the 20 kDa molecular mass range. To this end we have recorded a CPMG dataset on T4L L99A (as opposed to T4Ltm discussed above containing three mutations that exchanges much more rapidly). Here exchange is between a major conformer in which Phe114 is exposed outside a cavity in the protein and a minor state in which Phe114 is inserted inside, opposite to T4Ltm where the minor conformer is the one with Phe114 exposed and the major conformer is the one with Phe114 inserted in the cavity as described earlier. An exchange rate of ~ 900 s−1 at 22 °C has been measured previously (Mulder et al. 2001a). Methyl 1H DQ CPMG data were collected with Trelax = 20 ms (Fig. 7a) and the kex = 881 ± 60 s−1, pE = 2.9 ± 0.1% parameters obtained from the subsequent analysis of the data are in good agreement with kex = 943 ± 57 s−1, pE = 2.6 ± 0.1% fit independently from a combined analysis of SQ and TQ data (Fig. 7b, c). Additionally, extracted |Δϖ| values from the different methods are well correlated (Fig. 7d).
Methyl 1H DQ CPMG study of exchange in T4L L99A. a Selected 1H DQ CPMG relaxation dispersion curves at 22 °C with Trelax = 20 ms, along with a global best-fit of the data to a two-site exchange model (solid lines). b Similar reduced χ2 surfaces as a function of kex and pE are obtained from a global analysis of the SQ and TQ CPMG data (left) or the DQ CPMG data (right). c Histograms of kex (left) or pE (right) values from a bootstrap analysis of datasets as indicated. d Linear correlation plot of fitted |Δϖ| values from global analyses of SQ and TQ CPMG data or of DQ CPMG data
Conclusions
We have presented a methyl 1H DQ CPMG relaxation dispersion experiment for studies of conformational exchange using 13CH3-labeled methyl probes in highly deuterated protein systems. The experiment has been used to study proteins exchanging with rates varying between ~ 1000 and ~ 13,000 s−1. Although the DQ CPMG experiment can be used in place of the SQ scheme, based on sensitivity considerations, perhaps its most important feature lies in the complementarity between SQ, DQ and TQ datasets that when fit together leads to a more accurate definition of exchange parameters than when only a single class of 1H CPMG data recorded at a single static magnetic field is analyzed. This combined analysis can be important for identifying multiple exchange processes in the µs−ms time regime because kex and pE values can vary substantially from site to site when there are different exchange process as opposed to a global process where similar kex and pE values are obtained for every site. The faster 1H DQ relaxation rate than for 1H TQ coherences and the lower overall inherent sensitivity of the 1H DQ CPMG scheme relative to the TQ counterpart does suggest, however, that if at least a pair of spectrometers operating at different magnetic fields is available for experiments, recording a pair of 1H TQ datasets to obtain accurate exchange parameters is most likely the best option.
References
Ahlner A, Carlsson M, Jonsson BH, Lundstrom P (2013) PINT: a software for integration of peak volumes and extraction of relaxation rates. J Biomol NMR 56:191–202
Austin RH, Beeson KW, Eisenstein L, Frauenfelder H, Gunsalus IC (1975) Dynamics of ligand binding to myoglobin. Biochemistry 14:5355–5373
Baase WA, Liu L, Tronrud DE, Matthews BW (2010) Lessons from the lysozyme of phage T4. Protein Sci 19:631–641
Bain AD (1984) Coherence levels and coherence pathways in NMR. A simple way to design phase cycling procedures. J Magn Reson 56:418–427
Bouvignies G (2011) ChemEx https://github.com/gbouvignies/chemex
Bouvignies G, Vallurupalli P, Hansen DF, Correia BE, Lange O, Bah A, Vernon RM, Dahlquist FW, Baker D, Kay LE (2011) Solution structure of a minor and transiently formed state of a T4 lysozyme mutant. Nature 477:111–114
Carr HY, Purcell EM (1954) Effects of diffusion on free precession in nuclear magnetic resonance experiments. Phys Rev 94:630–638
Cavanagh J, Fairbrother WJ, Palmer AG, Rance M, Skelton NJ (2006) Protein NMR spectroscopy, principles and practice, 2nd ed. Academic Press, Cambridge
Delaforge E, Kragelj J, Tengo L, Palencia A, Milles S, Bouvignies G, Salvi N, Blackledge M, Jensen MR (2018) Deciphering the dynamic interaction profile of an intrinsically disordered protein by NMR exchange spectroscopy. J Am Chem Soc 140:1148–1158
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Eisenmesser EZ, Millet O, Labeikovsky W, Korzhnev DM, Wolf-Watz M, Bosco DA, Skalicky JJ, Kay LE, Kern D (2005) Intrinsic dynamics of an enzyme underlies catalysis. Nature 438:117–121
Eriksson AE, Baase WA, Wozniak JA, Matthews BW (1992) A cavity-containing mutant of T4 lysozyme is stabilized by buried benzene. Nature 355:371–373
Ernst RR, Bodenhausen G, Wokaun A (1987) Principles of nuclear magnetic resonance in one and two dimensions, 1st ed. Oxford Science Publications, Oxford
Frauenfelder H, Sligar SG, Wolynes PG (1991) The energy landscapes and motions of proteins. Science 254:1598–1603
Geen H, Freeman R (1991) Band-selective radiofrequency Pulses. J Magn Reson 93:93–141
Goddard TD, Kneller DG (2008) SPARKY 3. University of California, San Francisco
Gopalan AB, Vallurupalli P (2018) Measuring the signs of the methyl 1H chemical shift di erences between major and ‘invisible’ minor protein conformational states using methyl 1H multi-quantum spectroscopy. J Biomol NMR 70:187–202
Gopalan AB, Hansen DF, Vallurupalli P (2018) CPMG Experiments for protein minor conformer structure determination. Methods Mol Biol 1688:223–242
Goto NK, Gardner KH, Mueller GA, Willis RC, Kay LE (1999) A robust and cost-effective method for the production of Val, Leu, Ile (delta 1) methyl-protonated 15N-, 13C-, 2H-labeled proteins. J Biomol NMR 13:369–374
Grzesiek S, Bax A (1993) Amino-acid type determination in the sequential assignment procedure of uniformly C-13/N-15 enriched proteins. J Biomol NMR 3:185–204
Gullion T, Baker DB, Conradi MS (1990) New, compensated Carr–Purcell sequences. J Magn Reson 89:479–484
Hansen DF, Vallurupalli P, Kay LE (2008a) An improved 15N relaxation dispersion experiment for the measurement of millisecond time-scale dynamics in proteins. J Phys Chem B 112:5898–5904
Hansen DF, Vallurupalli P, Kay LE (2008b) Using relaxation dispersion NMR spectroscopy to determine structures of excited, invisible protein states. J Biomol NMR 41:113–120
Ishima R, Torchia DA (2003) Extending the range of amide proton relaxation dispersion experiments in proteins using a constant-time relaxation-compensated CPMG approach. J Biomol NMR 25:243–248
Jiang B, Yu BH, Zhang X, Liu ML, Yang DW (2015) A 15N CPMG relaxation dispersion experiment more resistant to resonance offset and pulse imperfection. J Magn Reson 257:1–7
Karplus M, Kuriyan J (2005) Molecular dynamics and protein function. Proc Natl Acad Sci USA 102:6679–6685
Kay LE, Prestegard JH (1987) Methyl group dynamics from relaxation of double quantum filtered NMR signals. Application to deoxycholate. J Am Chem Soc 109:3829–3835
Kay LE, Torchia DA (1991) The effects of dipolar cross correlation on 13C methyl-carbon T1, T2, and NOE measurements in macromolecules. J Magn Reson 95:536–547
Korzhnev DM, Salvatella X, Vendruscolo M, Di Nardo AA, Davidson AR, Dobson CM, Kay LE (2004) Low-populated folding intermediates of Fyn SH3 characterized by relaxation dispersion NMR. Nature 430:586–590
Korzhnev DM, Mittermaier AK, Kay LE (2005) Cross-correlated spin relaxation effects in methyl 1H CPMG-based relaxation dispersion experiments: complications and a simple solution. J Biomol NMR 31:337–342
Korzhnev DM, Religa TL, Banachewicz W, Fersht AR, Kay LE (2010) A transient and low-populated protein-folding intermediate at atomic resolution. Science 329:1312–1316
Kovrigin EL, Loria JP (2006) Characterization of the transition state of functional enzyme dynamics. J Am Chem Soc 128:7724–7725
Kovrigin EL, Kempf JG, Grey MJ, Loria JP (2006) Faithful estimation of dynamics parameters from CPMG relaxation dispersion measurements. J Magn Reson 180:93–104
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Levitt MH, Freeman R (1979) NMR population-inversion using a composite pulse. J Magn Reson 33:473–476
Logan TM, Olejniczak ET, Xu RX, Fesik SW (1993) A general method for assigning NMR spectra of denatured proteins using 3D HC(CO)NH-TOCSY triple resonance experiments. J Biomol NMR 3:225–231
Loria JP, Rance M, Palmer AG (1999a) A relaxation-compensated Carr–Purcell–Meiboom–Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Loria JP, Rance M, Palmer AG (1999b) A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J Biomol NMR 15:151–155
Marion D, Ikura M, Tschudin R, Bax A (1989) Rapid recording of 2D NMR-spectra without phase cycling—application to the study of hydrogen-exchange in proteins. J Magn Reson 85:393–399
McConnell HM (1958) Reaction rates by nuclear magnetic resonance. J Chem Phys 28:430–431
Meiboom S, Gill D (1958) Modified spin-echo method for measuring nuclear relaxation times. Rev Sci Instrum 29:688–691
Millet O, Loria JP, Kroenke CD, Pons M, Palmer AG (2000) The static magnetic field dependence of chemical exchange linebroadening defines the NMR chemical shift time scale. J Am Chem Soc 122:2867–2877
Morris GA, Freeman R (1979) Enhancement of nuclear magnetic- resonance signals by polarization transfer. J Am Chem Soc 101(3):760–762
Mulder FA, Mittermaier A, Hon B, Dahlquist FW, Kay LE (2001a) Studying excited states of proteins by NMR spectroscopy. Nat Struct Biol 8:932–935
Mulder FA, Skrynnikov NR, Hon B, Dahlquist FW, Kay LE (2001b) Measurement of slow (micros-ms) time scale dynamics in protein side chains by (15)N relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:967–975
Neudecker P, Robustelli P, Cavalli A, Walsh P, Lundstrom P, Zarrine-Afsar A, Sharpe S, Vendruscolo M, Kay LE (2012) Structure of an intermediate state in protein folding and aggregation. Science 336:362–366
Ollerenshaw JE, Tugarinov V, Skrynnikov NR, Kay LE (2005) Comparison of 13CH3, 13CH2D, and 13CHD2 methyl labeling strategies in proteins. J Biomol NMR 33:25–41
Orekhov VY, Korzhnev DM, Kay LE (2004) Double- and zero-quantum NMR relaxation dispersion experiments sampling millisecond time scale dynamics in proteins. J Am Chem Soc 126:1886–1891
Palmer AG 3rd, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Press WH, Flannery BP, Teukolsky SA, Vetterling WT (1992) Numerical recipes in C. The art of scientific computing, 2nd edn. Cambridge University Press, Cambridge (UK)
Rosenzweig R, Sekhar A, Nagesh J, Kay LE (2017) Promiscuous binding by Hsp70 results in conformational heterogeneity and fuzzy chaperone-substrate ensembles. Elife 6:e28030. https://doi.org/10.7554/eLife.28030
Sauerwein A, Hansen DF (2015) Relaxation dispersion NMR spectroscopy. In: Berliner L (ed) Protein NMR. Biological magnetic resonance and biomedical applications, vol 32. Springer, Boston, pp 75–132
Sekhar A, Kay LE (2013) NMR paves the way for atomic level descriptions of sparsely populated, transiently formed biomolecular conformers. Proc Natl Acad Sci USA 110:12867–12874
Shaka AJ, Keeler J, Frenkiel T, Freeman R (1983) An improved sequence for broadband decoupling: WALTZ-16. J Magn Reson 52:335–338
Sklenar V, Piotto M, Leppik R, Saudek V (1993) Gradient-tailored water suppression for 1H-15N HSQC experiments optimized to retain full sensitivity. J Magn Reson A 102:241–245
Skrynnikov NR, Mulder FA, Hon B, Dahlquist FW, Kay LE (2001) Probing slow time scale dynamics at methyl-containing side chains in proteins by relaxation dispersion NMR measurements: application to methionine residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:4556–4566
Skrynnikov NR, Dahlquist FW, Kay LE (2002) Reconstructing NMR spectra of “invisible” excited protein states using HSQC and HMQC experiments. J Am Chem Soc 124:12352–12360
Sugase K, Dyson HJ, Wright PE (2007) Mechanism of coupled folding and binding of an intrinsically disordered protein. Nature 447:1021–1025
Tamiola K, Acar B, Mulder FA (2010) Sequence-specific random coil chemical shifts of intrinsically disordered proteins. J Am Chem Soc 132:18000–18003
Tugarinov V, Kay LE (2005) Methyl groups as probes of structure and dynamics in NMR studies of high-molecular-weight proteins. Chembiochem 6:1567–1577
Tugarinov V, Kay LE (2007) Separating degenerate 1H transitions in methyl group probes for single-quantum 1H-CPMG Relaxation Dispersion NMR spectroscopy. J Am Chem Soc 129:9514–9521
Tugarinov V, Hwang PM, Ollerenshaw JE, Kay LE (2003) Cross-correlated relaxation enhanced 1H−13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125:10420–10428
Tugarinov V, Ollerenshaw JE, Kay LE (2006) Dipolar dynamic frequency shifts in multiple-quantum spectra of methyl groups in proteins: correlation with side-chain motion. Magn Reson Chem 44(Spec No):S122–S129
Tugarinov V, Libich DS, Meyer V, Roche J, Clore GM (2015) The energetics of a three-state protein folding system probed by high-pressure relaxation dispersion NMR spectroscopy. Angew Chem Int Ed Engl 54:11157–11161
Vallurupalli P, Kay LE (2006) Complementarity of ensemble and single-molecule measures of protein motion: a relaxation dispersion NMR study of an enzyme complex. Proc Natl Acad Sci USA 103:11910–11915
Vallurupalli P, Hansen DF, Stollar E, Meirovitch E, Kay LE (2007) Measurement of bond vector orientations in invisible excited states of proteins. Proc Natl Acad Sci USA 104:18473–18477
Vallurupalli P, Hansen DF, Kay LE (2008) Structures of invisible, excited protein states by relaxation dispersion NMR spectroscopy. Proc Natl Acad Sci USA 105:11766–11771
Vallurupalli P, Hansen DF, Lundstrom P, Kay LE (2009) CPMG relaxation dispersion NMR experiments measuring glycine 1H alpha and 13C alpha chemical shifts in the ‘invisible’ excited states of proteins. J Biomol NMR 45:45–55
Vallurupalli P, Bouvignies G, Kay LE (2011) Increasing the exchange time-scale that can be probed by CPMG relaxation dispersion NMR. J Phys Chem B 115:14891–14900
Vallurupalli P, Chakrabarti N, Pomes R, Kay LE (2016) Atomistic picture of conformational exchange in a T4 lysozyme cavity mutant: an experiment-guided molecular dynamics study. Chem Sci 7:3602–3613
Vugmeyster L, Kroenke CD, Picart F, Palmer AG, Raleigh DP (2000) 15N R1ρ measurements allow the determination of ultrafast protein folding rates. J Am Chem Soc 122:5387–5388
Werbelow LG, Marshall AG (1973a) Internal-rotation and methyl proton magnetic-relaxation for macromolecules. J Am Chem Soc 95:5132–5134
Werbelow LG, Marshall AG (1973b) Internal-rotation and monexponential methyl nuclear relaxation for macromolecules. J Magn Reson 11:299–313
Xie XS (2002) Single-molecule approach to dispersed kinetics and dynamic disorder: Probing conformational fluctuation and enzymatic dynamics. Journal of Chemical Physics 117:11024–11032
Yip GNB, Zuiderweg ERP (2004) A phase cycle scheme that significantly suppresses offset-dependent artifacts in the R2-CPMG 15N relaxation experiment. J Magn Reson 171:25–36
Yuwen T, Vallurupalli P, Kay LE (2016) Enhancing the sensitivity of CPMG relaxation dispersion to conformational exchange processes by multiple-quantum spectroscopy. Angew Chem Int Ed Engl 55:11490–11494
Zhuravleva A, Korzhnev DM (2017) Protein folding by NMR. Prog Nucl Magn Reson Spectrosc 100:52–77
Acknowledgements
We thank Dr. G. Bouvignies for making available the ChemEx program for analysis of dispersion data along with the source code, the national NMR facility at TIFR, Hyderabad and Dr. Krishna Rao for spectrometer time. The work was supported by generous startup funds from TCIS/TIFRH and Grant ECR/2016/001088 from SERB awarded to PV, by a Canadian Institutes of Health Research (CIHR) Grant to LEK and a CIHR post-doctoral fellowship to TY. LEK holds a Canada Research Chair in Biochemistry.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gopalan, A.B., Yuwen, T., Kay, L.E. et al. A methyl 1H double quantum CPMG experiment to study protein conformational exchange. J Biomol NMR 72, 79–91 (2018). https://doi.org/10.1007/s10858-018-0208-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-018-0208-z