Abstract
Pseudocontact shifts (PCSs) and residual dipolar couplings (RDCs) arising from the presence of paramagnetic metal ions in proteins as well as RDCs due to partial orientation induced by external orienting media are nowadays routinely measured as a part of the NMR characterization of biologically relevant systems. PCSs and RDCs are becoming more and more popular as restraints (1) to determine and/or refine protein structures in solution, (2) to monitor the extent of conformational heterogeneity in systems composed of rigid domains which can reorient with respect to one another, and (3) to obtain structural information in protein–protein complexes. The use of both PCSs and RDCs proceeds through the determination of the anisotropy tensors which are at the origin of these NMR observables. A new user-friendly web tool, called FANTEN (Finding ANisotropy TENsors), has been developed for the determination of the anisotropy tensors related to PCSs and RDCs and has been made freely available through the WeNMR (http://fanten-enmr.cerm.unifi.it:8080) gateway. The program has many new features not available in other existing programs, among which the possibility of a joint analysis of several sets of PCS and RDC data and the possibility to perform rigid body minimizations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pseudocontact shifts (PCSs) and residual dipolar couplings (RDCs) from self alignment arise in paramagnetic molecules in case the magnetic susceptibility tensor associated with the paramagnetic center is anisotropic. RDCs can also be induced in the absence of paramagnetic centers by the presence of an anisotropic alignment medium. PCSs and RDCs contain structural information that proved very helpful for solving protein structures (Gochin and Roder 1995; Banci et al. 1996, 1998; Bertini et al. 2001; Gaponenko et al. 2004; Diaz-Moreno et al. 2005; Jensen et al. 2006; Schmitz et al. 2012; Yagi et al. 2013b). They have been thus included as structural restraints (Balayssac et al. 2006; Banci et al. 2004) in the most commonly used programs for protein structure determination from NMR data [CYANA (Güntert 2004) and Xplor-NIH (Schwieters et al. 2003)] and more recently for structural refinement in combination with X-ray data (Rinaldelli et al. 2014) using the program REFMAC5 (Murshudov et al. 1997). PCS and RDC values depend on the coordinates of the nuclei and on the orientation of internuclear vectors between coupled nuclei, respectively, with respect to the unique magnetic susceptibility anisotropy tensor associated to the paramagnetic center present in the molecule (Bertini et al. 2002, 2011b).
In order to translate PCS and RDC data into structural information, the magnetic susceptibility anisotropy tensor should thus be determined. Conversely, once the protein structure is known, PCSs and RDCs can be used to calculate the tensor. Such a calculation can be performed by several available programs. In our lab, we have developed a series of programs (FANTASIA (Finding ANisotropy Tensors: A Simplex Approach), FANTALIN, FANTAORIENT) (Banci et al. 1996, 1998). Huber and co-workers have developed NUMBAT (Schmitz et al. 2008) to obtain the magnetic susceptibility anisotropy tensor from PCSs and protein structures. A python-based library, PyParaTool, has been recently developed, which encompasses the use of PCS and RDC (http://comp-bio.anu.edu.au/mscook/PPT/). RDCs may also arise in the absence of paramagnetic ions by molecular partial orientation achieved in the presence of orienting devices in solution (Tjandra and Bax 1997; Chou et al. 2001; Tolman et al. 2001; Prestegard et al. 2004; Chill et al. 2007; Lange et al. 2008; Grishaev et al. 2008), or by molecular anisotropy (Zhang et al. 2007), with the difference that they are not coupled to PCSs, as the latter are not present in diamagnetic systems. In this case the relevant tensor is the alignment tensor of the molecule, which depends both on the biomolecule and on the alignment medium. Command-line software packages (PALES and PATI) are available to fit RDCs arising from external alignment media and to give an estimate of the alignment tensor due to either steric effects or electrostatic effects or both (Zweckstetter and Bax 2000; Zweckstetter 2008; Berlin et al. 2009, 2010, 2011).
In practice, in structure calculations, PCSs and RDCs are used (1) to calculate the magnetic susceptibility anisotropy tensors in best agreement with a tentative protein structure, using either programs of the FANTASIA family or NUMBAT, and (2) to calculate the protein structures with minimal restraints violations in agreement with the tentative magnetic susceptibility anisotropy tensors, using appropriate versions of CYANA (Balayssac et al. 2006) or Xplor-NIH (Banci et al. 2004). These calculations are repeated iteratively, until convergence of the tensors is achieved, usually in few steps if the restraints are enough to reconstruct the protein fold. PCS-Rosetta have been also employed to obtain simultaneously the protein structure and an accurate fit of the tensor, without the use of an external software for tensor determination (Schmitz et al. 2012; Yagi et al. 2013b). PCSs and RDCs also proved useful for the refinement of protein structures starting from available models (Gochin and Roder 1995; Bertini et al. 2004, 2009, 2012a). In these cases, the models could be used to determine the first estimate of the magnetic susceptibility anisotropy tensors. The same procedure can of course be followed if RDCs from external alignment are used: the tensor can be fit (by PALES or PATI), and the iterations needed for structural determination can be as well performed with Xplor-NIH or CYANA. Also in this case an existing model (Chou et al. 2001) can be used to determine an estimate of the alignment tensor, of course to some degree of approximation.
It was also shown that RDCs and PCS can be used to screen the PDB for proteins which have a partial homology, based on the 3D arrangement of secondary structure elements as given by PCSs and/or RDCs (Meiler et al. 2000). Other most used programs available for the analysis of PCS and RDC data are reported in Table 1.
PCSs and/or RDCs are even more precious structural restraints in the case of proteins composed of multiple rigid domains, the structure of which is known but not the relative orientation, or of protein–protein complexes, when the structure of each protein is known but again their relative orientation is not known. In these cases, a rigid body minimization can locate the position of the different domains or molecules with respect to one another in order to reproduce the experimental PCS and RDC data (Al-Hashimi et al. 2000; Clore 2000; Dosset et al. 2001; Valafar and Prestegard 2004; Pintacuda et al. 2006; Hulsker et al. 2008; Simon et al. 2010; Bertini et al. 2011b). A rigid body molecular docking program, PATIDOCK (Berlin et al. 2011), was developed to recover the structure of a molecular complex from the three-dimensional structure of the individual components and the experimental RDCs arising from the steric alignment, due to the anisotropy of the molecular shape, in the presence of external orienting devices.
Due to the quadratic mathematical form of these restraints, more than one set of PCSs and RDCs are needed to remove degeneracies in the solutions (Dosset et al. 2001; Longinetti et al. 2002; Bertini et al. 2002; Fragai et al. 2013). These multiple sets can be obtained by alternative replacement of the paramagnetic ion with other paramagnetic ions with different magnetic susceptibility anisotropy tensors or by attaching, once at a time, paramagnetic tags in different points (Bertini et al. 2001; Pintacuda et al. 2006; Otting 2010; Liu et al. 2012). Finally, PCSs and RDCs can provide information on interdomain mobility in multidomain proteins or internal mobility in protein–protein complexes, whenever internal consistency of all PCSs and RDCs arising from each paramagnetic metal with its own magnetic susceptibility anisotropy tensor cannot be achieved by assuming a single rigid structure (Gardner et al. 2005; Longinetti et al. 2006; Bertini et al. 2007, 2010, 2012b; Russo et al. 2013; Ravera et al. 2014; Andralojc et al. 2014).
The advantages offered by paramagnetic restraints are more and more exploited thanks to the development of paramagnetic tags that can be attached to diamagnetic proteins, so that PCSs and RDCs can be easily measured also for molecules without a natural metal binding site (Wöhnert et al. 2003; Rodriguez-Castañeda et al. 2006; Su et al. 2006; John and Otting 2007; Su et al. 2008a; Keizers et al. 2008; Su et al. 2008b; Zhuang et al. 2008; Häussinger et al. 2009; Su and Otting 2010; Hass et al. 2010; Man et al. 2010; Das Gupta et al. 2011; Liu et al. 2012; Cerofolini et al. 2013; Yagi et al. 2013a; Gempf et al. 2013; Kobashigawa et al. 2012; Saio et al. 2011; Watanabe et al. 2010; Loh et al. 2013; Swarbrick et al. 2011a, 2011b).
Due to the increasingly wide use of paramagnetic restraints and diamagnetic RDCs for structural purposes we found it useful to make available a new web-based interface, called FANTEN (Finding ANisotropy TENsor) for the simultaneous fit of PCSs and RDCs to protein structures in order to determine the anisotropy tensors. FANTEN accepts as experimental data both PCSs and RDCs, and can perform a minimization using several sets of data either independently or globally; it can also be used to perform a rigid body minimization, when data are available for different rigid domains. This user-friendly web interface is based on a framework (Bertini et al. 2011a) developed within the WeNMR project (Wassenaar et al. 2012), and can be accessed from the open access web site (http://fanten-enmr.cerm.unifi.it:8080). Data can be provided either in the Xplor-NIH format (also available through the WeNMR web pages) or in the PARAMAGNETIC CYANA format (Balayssac et al. 2006), and thus the program can be easily used in combination with these programs.
Methods
PCSs, arising in the presence of a paramagnetic metal, depend on the magnetic susceptibility anisotropy tensor and on the nuclear coordinates according to the following equation, written in the so-called linear form (McConnell and Robertson 1958; Kurland and McGarvey 1970; Kemple et al. 1988; Bertini et al. 2002):
where x, y, z are the coordinates of the nucleus when the metal ion sits at the origin of the coordinate system, r is the distance between the observed nucleus and the metal ion and χ ij are the components of the symmetric susceptibility anisotropy tensor. Paramagnetic RDCs arising in the presence of the same metal depend on the same χ tensor components and on the orientation of the dipole–dipole coupled nuclei according the following equation (Tolman et al. 1995; Bertini et al. 2002):
where
r AB is the distance between the two coupled nuclei A and B, and S LS is the model-free order parameter, introduced to take into account some average local mobility of the coupled nuclei vectors. Other symbols have the usual meaning. RDCs do not depend on the position of the coupled nuclei with respect to the metal ion.
Diamagnetic RDCs are described by an equation with the same form as that for paramagnetic RDCs (Eq. 2):
where A ij are the components of the molecular alignment tensor.
In Eqs. 1–3, PCSs and RDCs are reported as linear functions of the five linearly independent components of the traceless χ or A tensors. However, it is common to describe these tensors by reporting two anisotropy values and the three Euler angles describing the rotations needed to pass from the reference frame in which the nuclear coordinates are provided to the frame in which the χ or A tensors are diagonal. These values can be determined after diagonalization of the χ or A tensors. The magnetic susceptibility anisotropy values are defined by the axial and rhombic components provided by
and
where χ ii are the components of the χ tensor in the frame where it is diagonal. Analogously, the anisotropies of the alignment tensor are described by the fraction of alignment along the z axis (A) and by the rhombicity (R), (Zweckstetter 2008)
where A ii are the components of the A tensor in the frame where it is diagonal. Alternatively, the anisotropy values can be described by the maximum RDC induced for a specific nuclear pair, i.e. the N–NH pair (D NH) (Tjandra and Bax 1997),
and by the rhombicity R.
Due to the dependence of Eqs. (1) and (2) on the nuclear coordinates, the axes of the χ tensor are not uniquely determined. Using a standard convention, they are defined in such a way that |χyy| ≤ |χxx| ≤ |χzz| and |Δχrh| ≤ 2/3 |Δχax| (Banci et al. 1996).
The agreement of calculated and experimental PCSs/RDCs is described by the Q-factors (Cornilescu et al. 1998), defined as
The estimation of the error of the values determined through the best fit procedures is implemented using the bootstrap Monte Carlo method, which consists in calculating the standard deviation of the values obtained after multiple removals of a user-defined number of randomly selected data.
The web interface is based on Pylons (www.pylonsproject.org) web application framework (Bertini et al. 2011a), and library functions has been implemented using Python 2.6 (www.python.org). The following Python packages were used: scipy, numpy (Oliphant 2007), lmfit (http://lmfit.github.io/lmfit-py/), and P3D (Fufezan and Specht 2009).
Results
The program FANTEN performs a linear fit of the five independent components of the traceless χ tensor (χ xx , χ zz , χ xz , χ yz , χ xy ; with χ yy = −χ zz − χ xx ) in order to obtain the best agreement of experimental PCSs and/or paramagnetic RDCs against the protein structure, when the coordinates of the metal are included in the structure (if not, the metal coordinates are three additional parameters to be determined in the minimization procedure, see later); it determines the five components of the A tensor in best agreement against the protein structure when diamagnetic RDCs are provided. Users can provide one protein structure or an ensemble thereof (in PDB format) and multiple sets of PCSs and/or paramagnetic RDCs or diamagnetic RDCs (both in PARAMAGNETIC CYANA format or in the format used in the paramagnetic routines for Xplor-NIH) referred to the same or different metals or orienting devices. The interface for data upload is shown in Fig. 1a.
The calculations can be performed using three different interfaces (Custom Interface, Smart Interface and Rigid Body Minimization), designed and optimized for specific applications with different levels of complexity.
Custom Interface
FANTEN allows for a fully customized calculation of the tensor: from the user-provided PDB file it is possible to select the model and the chain for which the calculation should be performed, and the associated PCS and/or RDC datasets (Fig. 2; Fig. S1). For any dataset, the user can define, if known, (1) metal position, (2) tensor orientation, and (3) anisotropy parameters, or any combination thereof.
-
1.
In case the coordinates of the paramagnetic metal are provided in the PDB file, the metal can be simply selected from the list of the metals present in the PDB, with the possibility of keeping its coordinates fixed (by selecting “Fix position to this metal”) or refining them (by selecting “Fit position around this metal”) during the calculation. Information on the position of the paramagnetic metal can also be provided as distance restraints from any atom present in the PDB, so that during the fit the metal position can be restrained in a user-specified volume of space. These restraints may be available from knowledge of the metal binding site or protein tagging position and may be useful to increase the speed of the calculations. In case the paramagnetic metal position is completely unknown, a grid search can be performed starting from points in the proximity of the protein surface or from atoms constituting its backbone. The minimization is performed using eight different starting points for the metal position, corresponding to the vertices of a box containing the protein or randomly choosing eight atoms belonging to the protein backbone, respectively.
-
2.
The tensor orientation can be fixed in three different ways: by providing the Euler angles (in the preferred convention), the eigenvectors of the tensor expressed in its non-diagonal matrix form, or by selecting the tensor origin and axes orientations as defined by pseudoatoms included in the uploaded PDB. The Euler angles convention is defined by a 4-character string, in which the first character defines if rotations are applied to static (s) or rotating (r) frame, whereas the remaining three characters specify the axes about which the three consecutive rotations occur. The resulting transformation matrix represents the rotation that brings the reference system to coincide with the principal axis system of the anisotropy tensor (passive rotation) (Dosset et al. 2013). As example, in the “szyz” convention, where three consecutive rotations of α, β and γ about the fixed initial z, y and z axes, respectively, of the reference frame occur, the rotation matrix is
$$ R = \left( {\begin{array}{*{20}l} {\cos \alpha \cos \beta \cos \gamma - \sin \alpha \sin \gamma } \hfill & { - \sin \alpha \cos \beta \cos \gamma - \cos \alpha \sin \gamma } \hfill & {\sin \beta \cos \gamma } \hfill \\ {\cos \alpha \cos \beta \sin \gamma + \sin \alpha \cos \gamma } \hfill & { - \sin \alpha \cos \beta \sin \gamma + \cos \alpha \cos \gamma } \hfill & {\sin \beta \sin \gamma } \hfill \\ { - \cos \alpha \sin \beta } \hfill & {\sin \alpha \sin \beta } \hfill & {\cos \beta } \hfill \\ \end{array} } \right) $$whereas in the “rzyz” convention, where three consecutive rotations of α, β and γ about the z axis, the new y’ axis and the new z’’ axis, respectively, occur, the rotation matrix is
$$ R = \left( {\begin{array}{*{20}l} {\cos \alpha \cos \beta \cos \gamma - \sin \alpha \sin \gamma } \hfill & { - \cos \alpha \cos \beta \sin \gamma - \sin \alpha \cos \gamma } \hfill & {\cos \alpha \sin \beta } \hfill \\ {\sin \alpha \cos \beta \cos \gamma + \cos \alpha \sin \gamma } \hfill & { - \sin \alpha \cos \beta \sin \gamma + \cos \alpha \cos \gamma } \hfill & {\sin \alpha \sin \beta } \hfill \\ { - \sin \beta \cos \gamma } \hfill & {\sin \beta \sin \gamma } \hfill & {\cos \beta } \hfill \\ \end{array} } \right) $$ -
3.
Users may also choose to fix the anisotropy values in case the latter are known from e.g. analogous constructs (Keizers and Ubbink 2011; Skinner et al. 2013) or previous calculations performed for the same paramagnetic metal and protein domain by providing them through the web interface. This can be useful to obtain a preliminary orientation of the tensor which can then be left free for refinement during structure calculations.
Multiple datasets of PCSs and paramagnetic RDCs referring to different metals can be provided for the minimization against a protein structure. In this case a different anisotropy tensor in best agreement with both PCSs and RDCs related to the same paramagnetic metal is calculated for each metal. The positions of the different metals can be independent or constrained to coincide. The different datasets may in fact arise from the presence of different paramagnetic ions alternatively substituted in the same position: in these cases, the different datasets must be associated to different metals (because the associated anisotropy tensors are different), and the different metals can be either constrained to have the same coordinates or restrained individually using the same distance restraints. In case that the metals are constrained to have the same coordinates, the restraints on the metal position selected for the first dataset are automatically applied to all other datasets, with the additional constraint that the best fit position must be unique for all metals.
The program can also calculate PCSs and/or RDCs, without performing any minimization, if the anisotropy parameters are provided as input together with a single PDB file containing the protein structure and the pseudoatoms defining the anisotropy tensor, as obtained from PARAMAGNETIC CYANA or Xplor-NIH.
The distances of coupled atoms, r AB in Eqs. 2 and 3, can be (1) determined from the coordinates of the protein atoms, (2) fixed to default values, or (3) fixed to values preset by the user. Also the temperature and the model-free order parameter (S LS ) can be fixed by the user. In case calculations are performed using both PCSs and RDCs to determine the same anisotropy tensor, their relative contribution can be controlled through a weight constant.
The Custom Interface permits the optimization of metal coordinates, anisotropy parameters and/or Euler angles defining the orientation of the anisotropy tensor. The minimization is performed using the Levenberg–Marquardt algorithm, available from SciPy library, which also permits the inclusion of constraints on the parameters to be optimized in the minimization procedure. A least-squares approach is used to minimize the norm of the vector with components provided by the product of the differences between experimental and back-calculated data and the corresponding weights. The latter are provided by the product between the specific weights provided in the uploaded files (if present) and the global weights provided through the web interface in the case that both PCSs and RDCs are fitted simultaneously. The default values for the global weights for PCSs and RDCs are 1 and the ratio between the norm of vector with components equal to the experimental values of RDCs and of PCSs, respectively. The target function which is minimized is
where the index l runs over the two classes of restraints (PCSs and RDCs), the index i on all experimental data of each class, w i is the weight of the i-restraint and w′ l is the weight of the l-class restraints.
Once the anisotropy tensor has been calculated through the best fit minimization, the quality of the fit can be evaluated at a glance by inspection of the correlation plots, which show the agreement between experimental and back-calculated PCSs and RDCs, as shown in Fig. 1b. These interactive plots allow the user to identify outliers by positioning the pointer directly on the data point. Moreover, the agreement between experimental and back-calculated data is quantified and reported on the top left of the plots, in terms of Q-factors of PCSs and RDCs (QPCS and QRDC).
The calculation provides the anisotropy parameters (Δχ ax and Δχ rh) and the Euler angles (in the “szyz” convention or in any user-selected convention) defining the main coordinate frame of the anisotropy tensor with respect to the frame of the protein coordinates. The tensor matrix and the eigenvectors providing the main axes of the tensor, i.e. the frame where χ is diagonal, are also provided, together with the tensor representation as a triad of pseudoatoms representing unit vectors in PDB format. The latter can be easily copied and pasted in the uploaded file for graphical visualization. Experimental and back-calculated data are also given at the bottom of the page in tabular form.
When RDCs arising from partial alignment induced by external orienting media are provided, the results of the calculation are given in terms of alignment tensors. As mentioned in the methods section, the size of the tensors is provided by their magnitude A and rhombicity R; for the sake of completeness also the maximum DNH value is given (Fig. S2).
Simultaneous calculation of all tensors referring to several datasets can be performed after the set-up of each of them. The program automatically generates a summary of all calculated tensors, including the plots showing the agreement between experimental and calculated data for the different tensors, so that they can be easily compared (Fig. S3).
All results from the fit procedure and the associated parameters can be downloaded as text files.
Smart Interface
This interface can be either used as a simplified stand-alone interface or, after having run the custom interface, to exploit its additional graphical features or for the calculation of the Monte Carlo error (Fig. 3). The use of this simplified interface is recommended to obtain the best fit anisotropy tensor(s) when the position of the paramagnetic metals is known (or previously determined through the Custom Interface) or when diamagnetic RDCs should be analyzed. Since the metal position does not need to be determined, the Smart Interface permits a more efficient calculation of the anisotropy tensor, which can be estimated directly using Eqs. 1–3. In this case the first derivate can be easily computed and used in a Gauss–Newton optimization procedure. As for the Custom Interface, the procedure minimizes the squared residuals between experimental and back-calculated data, weighted by the product between the specific weights provided in the uploaded files (if present) and the global weights provided through the web interface in the case that both PCSs and RDCs are fitted simultaneously. Again, a model-free order parameter, the value of which can be adjusted in the General Parameters frame of the interface, is used as in Eqs. 2 and 3. Moreover, this interface allows the users to perform the minimization in a single step even if multiple datasets of PCSs and/or RDCs are provided in input, as described above (Fig. 4a).
a The Smart Interface permits to upload multiple PCSs and/or RDCs datasets and determine the anisotropy tensors in a single step. A single anisotropy tensor in best agreement with both PCSs and RDCs can be determined, at will, through the “joint fitting” option. b The isoPCS surface (with threshold equal to 1 ppm) superimposed to the protein chain can be visualized through an integrated JSmol applet
If a single protein structure is provided as input, the errors on the best fit anisotropy values and on the values of the Euler angles defining the orientation of the anisotropy tensors can be estimated through a bootstrap Monte Carlo approach, in which the user may decide the percentage of retained experimental data used during the calculation as well as the number of iterations (Fig. S4). When PCSs are used for the calculation, the isoPCS surface can be generated and visualized (with threshold equal to 1 ppm) superimposed to the protein chain selected for the calculation by an integrated JSmol applet, as shown in Fig. 4b. The isosurface is downloadable as a cube file that can be imported by PyMol and UCSF Chimera.
In case a structural ensemble is provided as input, a tensor for each model contained in the PDB is estimated, and Δχax and Δχrh values, the Euler angles and the tensor matrices are reported in terms of average and standard deviation obtained from the different models. The results of the individual fits are also given and available for download. A plot showing the agreement between experimental and back-calculated PCSs and RDCs for all the models is provided in the interface, with the possibility of visualizing the data referred to the different models by simply clicking/unclicking of the corresponding checkbox (Fig. S5).
Rigid body minimization
In case of systems composed of multiple domains or subunits the structures of which are rigid and available but with unknown relative positions, PCSs and RDCs arising in the presence of paramagnetic metal(s) in one subunit can be used to determine the relative position of the other subunit(s). As already seen, PCSs and/or RDCs in one subunit can easily provide the anisotropy tensor(s); since PCSs and/or RDCs in another subunit should also yield the same tensor(s) (provided the overall system is rigid), the superposition of the tensors determined from the fit of the data measured for the two subunits permits to reconstruct their relative position (or orientation if only RDCs are used).
This interface has the same structure for data upload of the previous interfaces. After the upload of the first molecular structure (Subunit A) and the related experimental datasets, a second molecular structure (Subunit B) and the corresponding PCS/RDC datasets can be provided (Fig. S6). The datasets for the two subunits corresponding to the same tensors can be paired and associated to the corresponding metals, the coordinates of which must be provided in the PDB file related to Subunit A. After having determined the anisotropy tensors from the data referred to Subunit A (see Smart Interface for the algorithm used), the algorithm automatically performs a grid search to determine the metal position and tensor orientations with respect to the coordinates of Subunit B (see Custom Interface for the grid search algorithm). The algorithm maximizes the agreement between the experimental and the back-calculated data referred to Subunit B, using the anisotropy values previously determined for Subunit A. Subunit B is then translated and rotated rigidly with respect to Subunit A in order to superimpose the anisotropy tensors of the two subunits. In this way the relative position of the two subunits is recovered. A further estimate of the paramagnetic tensor based on the resulting structure, i.e. the structure comprising both Subunit A and the rototranslated Subunit B, is finally carried out. As in the previously described interfaces, FANTEN provides as output the anisotropy parameters (Δχ ax and Δχ rh), the values of the Euler angles, the tensor matrices and the eigenvectors providing the main axes of the tensor together with the plots showing the agreement between experimental and back-calculated data for both Subunit A and Subunit B (Fig. S7). The applied transformation can be written as
where X and X′ are the coordinates of Subunit B before and after the transformation, respectively, T2 is the vector of the metal coordinates, M is the rotation matrix, and T1 the translation vector. T1, T2 and M are provided as output.
The results of the fit, the parameters defining the calculated tensors as well as the structure comprising Subunit A and the rototranslated Subunit B are available for download.
When all PCS/RDC data refer to a single paramagnetic metal, the position of Subunit B is not univocally determined, but four degenerate solutions are possible (Tolman et al. 1995; Fragai et al. 2013). Indeed, once one of the four degenerate solutions is obtained, the other three are automatically generated by changing the sign of two out of three nuclear coordinates, when the latter are expressed in the anisotropy tensor frame centered on the metal ion. All the possible solutions are available for download.
Finally, if only RDCs are provided, only the relative orientation of the two subunits can be determined; in this case the relative position of Subunit B with respect to Subunit A is unchanged.
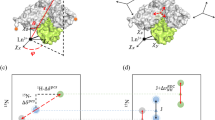
Figure 5 shows the results of the rigid body minimization performed for the two-domain protein calmodulin when bound to a peptide from the death-associated protein kinase. The position of the C-terminal domain of the protein with respect to the N-terminal domain is determined using PCSs and RDCs measured for Tb(III) or for three lanthanides (Tb(III), Tm(III) and Yb(III); Bertini et al. 2009).
A rigid body minimization can be performed after determination of the anisotropy tensor for the Subunit A, shown in gray (a). The position of the subunit B, in yellow, is uniquely determined if at least two datasets with non-parallel anisotropy axes are provided (b). If a single dataset is provided, the four degenerate solutions are shown (c)
FANTEN is hosted in the WeNMR gateway, which is a worldwide e-infrastructure for NMR and structural biology (Wassenaar et al. 2012) jointly maintained by the NMR facilities of Florence, Frankfurt and Utrecht.
Conclusions
A new, user-friendly, web-based interface, FANTEN, for the analysis of PCSs and RDCs against structural models has been made available. It allows the users to determine the anisotropy tensors through a joint analysis of these restraints. The agreement of the experimental data with the structural model can be monitored from the quality of the best fit of the experimental data.
In structural calculation protocols, the PCSs and RDCs files used in either PARAMAGNETIC CYANA or Xplor-NIH can be given as input to FANTEN together with the PDB file of the protein produced by these programs, so that the anisotropy tensors (and possibly the metal coordinates) can be obtained and used for further PARAMAGNETIC CYANA or Xplor-NIH calculations, until convergence is reached. Alternatively, FANTEN can be used to predict PCS and RDC data from the coordinates of the protein structure and the anisotropy tensor either provided as input or determined from the available data.
FANTEN can be run from the open access web site http://fanten-enmr.cerm.unifi.it:8080 through the WeNMR gateway.
References
Al-Hashimi HM, Valafar H, Terrell M, Zartler ER, Eidsness MK, Prestegard JH (2000) Variation of molecular alignment as a means of resolving orientational ambiguities in protein structures from dipolar couplings. J Magn Reson 143:402–406
Andralojc W, Luchinat C, Parigi G, Ravera E (2014) Exploring regions of conformational space occupied by two-domain proteins. J Phys Chem B 118:10576–10587
Balayssac S, Bertini I, Luchinat C, Parigi G, Piccioli M (2006) 13C direct detected NMR increases the detectability of residual dipolar couplings. J Am Chem Soc 128:15042–15043
Banci L, Bertini I, Bren KL, Cremonini MA, Gray HB, Luchinat C, Turano P (1996) The use of pseudocontact shifts to refine solution structures of paramagnetic metalloproteins: Met80Ala cyano-cytochrome c as an example. J Biol Inorg Chem 1:117–126
Banci L, Bertini I, Huber JG, Luchinat C, Rosato A (1998) Partial orientation of oxidized and reduced cytochrome b5 at high magnetic fields: magnetic susceptibility anisotropy contributions and consequences for protein solution structure determination. J Am Chem Soc 120:12903–12909
Banci L, Bertini I, Cavallaro G, Giachetti A, Luchinat C, Parigi G (2004) Paramagnetism-based restraints for Xplor-NIH. J Biomol NMR 28:249–261
Berlin K, O’Leary DP, Fushman D (2009) Improvement and analysis of computational methods for prediction of residual dipolar couplings. J Magn Reson 201:25–33
Berlin K, O’Leary DP, Fushman D (2010) Structural assembly of molecular complexes based on residual dipolar couplings. J Am Chem Soc 132:8961–8972
Berlin K, O’Leary DP, Fushman D (2011) Fast approximations of the rotational diffusion tensor and their application to structural assembly of molecular complexes. Proteins 79:2268–2281
Bertini I, Donaire A, Jiménez B, Luchinat C, Parigi G, Piccioli M, Poggi L (2001) Paramagnetism-based versus classical constraints: an analysis of the solution structure of Ca Ln Calbindin D9k. J Biomol NMR 21:85–98
Bertini I, Luchinat C, Parigi G (2002) Magnetic susceptibility in paramagnetic NMR. Prog NMR Spectrosc 40:249–273
Bertini I, Faraone-Mennella J, Gray BH, Luchinat C, Parigi G, Winkler JR (2004) NMR-validated structural model for oxidized Rhodopseudomonas palustris cytochrome c556. J Biol Inorg Chem 9:224–230
Bertini I, Gupta YK, Luchinat C, Parigi G, Peana M, Sgheri L, Yuan J (2007) Paramagnetism-based NMR restraints provide maximum allowed probabilities for the different conformations of partially independent protein domains. J Am Chem Soc 129:12786–12794
Bertini I, Kursula P, Luchinat C, Parigi G, Vahokoski J, Willmans M, Yuan J (2009) Accurate solution structures of proteins from X-ray data and minimal set of NMR data: calmodulin peptide complexes as examples. J Am Chem Soc 131:5134–5144
Bertini I, Giachetti A, Luchinat C, Parigi G, Petoukhov MV, Pierattelli R, Ravera E, Svergun DI (2010) Conformational space of flexible biological macromolecules from average data. J Am Chem Soc 132:13553–13558
Bertini I, Case DA, Ferella L, Giachetti A, Rosato A (2011a) A grid-enable web portal for NMR structure refinement with AMBER. Bioinformatics 27:2384–2390
Bertini I, Luchinat C, Parigi G (2011b) Moving the frontiers in solution solid state bioNMR. A celebration of Harry Gray’s 75th birthday. Coord Chem Rev 255:649–663
Bertini I, Calderone V, Cerofolini L, Fragai M, Geraldes CFGC, Hermann P, Luchinat C, Parigi G, Teixeira JMC (2012a) The catalytic domain of MMP-1 studied through tagged lanthanides. Dedicated to Prof. A. V. Xavier. FEBS Lett 586:557–567
Bertini I, Ferella L, Luchinat C, Parigi G, Petoukhov MV, Ravera E, Rosato A, Svergun DI (2012b) MaxOcc: a web portal for maximum occurence analysis. J Biomol NMR 53:271–280
Bryson M, Tian F, Prestegard JH, Valafar H (2008) REDCRAFT: a tool for simultaneous characterization of protein backbone structure and motion from RDC data. J Magn Reson 191:322–334
Cerofolini L, Fields GB, Fragai M, Geraldes CFGC, Luchinat C, Parigi G, Ravera E, Svergun DI, Teixeira JMC (2013) Examination of matrix metalloproteinase-1 (MMP-1) in solution: a preference for the pre-collagenolysis state. J Biol Chem 288:30659–30671
Chill JH, Louis JM, Delaglio F, Bax A (2007) Local and global structure of the monomeric subunit of the potassium channel KcsA probed by NMR. Biochim Biophys Acta 1768:3260–3270
Chou JJ, Li S, Klee CB, Bax A (2001) Solution structure of Ca2+ calmodulin reveals flexible hand-like properties of its domains. Nat Struct Biol 8:990–997
Clore GM (2000) Accurate and rapid docking of protein–protein complexes on the basis of intermolecular nuclear overhauser enhancement data and dipolar couplings by rigid body minimization. Proc Natl Acad Sci USA 97:9021–9025
Cornilescu G, Marquardt J, Ottiger M, Bax A (1998) Validation of protein structure from anisotropic carbonyl chemical shifts in a dilute liquid crystalline phase. J Am Chem Soc 120:6836–6837
Das Gupta S, Hu X, Keizers PHJ, Liu W-M, Luchinat C, Nagulapalli M, Overhand M, Parigi G, Sgheri L, Ubbink M (2011) Narrowing the conformational space sampled by two-domain proteins with paramagnetic probes in both domains. J Biomol NMR 51:253–263
Diaz-Moreno I, Diaz-Quintana A, De la Rosa MA, Ubbink M (2005) Structure of the complex between plastocyanin and cytochrome f from the cyanobacterium nostoc Sp. PCC 7119 as determined by paramagnetic NMR. J Biol Chem 280:18908–18915
Dosset P, Hus JC, Marion D, Blackledge M (2001) A novel interactive tool for rigid-body modeling of multi-domain macromolecules using residual dipolar couplings. J Biomol NMR 20:223–231
Dosset P, Barthe P, Cohen-Gonsaud M, Roumestand C, Déméné H (2013) Equivalence between Euler angle conventions for the description of tensorial interactions in liquid NMR: application to different software programs. J Biomol NMR 57:305–311
Fragai M, Luchinat C, Parigi G, Ravera E (2013) Conformational freedom of metalloproteins revealed by paramagnetism-assisted NMR. Coord Chem Rev 257:2652–2667
Fufezan C, Specht M (2009) p3d–Python module for structural bioinformatics. BMC bioinform 10:258
Gaponenko V, Sarma SP, Altieri AS, Horita DA, Li J, Byrd RA (2004) Improving the accuracy of NMR structures of large proteins using pseudocontact shifts as long/range restraints. J Biomol NMR 28:205–212
Gardner RJ, Longinetti M, Sgheri L (2005) Reconstruction of orientations of a moving protein domain from paramagnetic data. Inverse Probl 21:879–898
Gempf KL, Butler SJ, Funk AM, Parker D (2013) Direct and selective tagging of cysteine residues in peptides and proteins with 4-nitropyridyl lanthanide complexes. Chem Commun (Camb) 49:9104–9106
Gochin M, Roder H (1995) Protein structure refinement based on paramagnetic NMR shifts. applications to wild-type and mutants forms of cytochrome c. Protein Sci 4:296–305
Grishaev A, Tugarinov V, Kay LE, Trewhella J, Bax A (2008) Refined solution structure of the 82-kDa enzyme malate synthase G from joint NMR and synchrotron SAXS restraints. J Biomol NMR 40:95–106
Güntert P (2004) Automated NMR structure calculation with CYANA. Methods Mol Biol 278:353–378
Hass MAS, Keizers PHJ, Blok A, Hiruma Y, Ubbink M (2010) Validation of a lanthanide tag for the analysis of protein dynamics by paramagnetic NMR spectroscopy. J Am Chem Soc 132:9952–9953
Häussinger D, Huang J, Grzesiek S (2009) DOTA-M8: an extremely rigid, high-affinity lanthanide chelating tag for PCS NMR spectroscopy. J Am Chem Soc 131:14761–14767
Hulsker R, Baranova MV, Bullerjahn GS, Ubbink M (2008) Dynamics in the transient complex of plastocyanin-cytochrome f from Prochlorothrix hollandica. J Am Chem Soc 130:1985–1991
Jensen MR, Hansen DF, Ayna U, Dagil R, Hass MA, Christensen HE, Led JJ (2006) On the use of pseudocontact shifts in the structure determination of metalloproteins. Magn Reson Chem 44:294–301
John M, Otting G (2007) Strategies for measurements of pseudocontact shifts in protein NMR spectroscopy. ChemPhysChem 8:2309–2313
John M, Schmitz C, Park AY, Dixon NE, Huber T, Otting G (2007) Sequence-specific and stereospecific assignment of methyl groups using paramagnetic lanthanides. J Am Chem Soc 129:13749–13757
Keizers PHJ, Ubbink M (2011) Paramagnetic tagging for protein structure and dynamics analysis. Prog Nucl Magn Reson Spectrosc 58:88–96
Keizers PHJ, Saragliadis A, Hiruma Y, Overhand M, Ubbink M (2008) Design, synthesis, and evaluation of a lanthanide chelating protein probe: CLaNP-5 yields predictable paramagnetic effects independent of environment. J Am Chem Soc 130:14802–14812
Kemple MD, Ray BD, Lipkowitz KB, Prendergast FG, Rao BDN (1988) The use of lanthanides for solution structure determination of biomolecules by NMR: evaluation of methodology with EDTA derivatives as model systems. J Am Chem Soc 110:8275–8287
Kobashigawa Y, Saio T, Ushio M, Sekiguchi M, Yokochi M, Ogura K, Inagaki F (2012) Convenient method for resolving degeneracies due to symmetry of the magnetic susceptibility tensor and its application to pseudo contact shift-based protein-protein complex structure determination. J Biomol NMR 53:53–63
Kurland RJ, McGarvey BR (1970) Isotropic NMR shifts in transition metal complexes: calculation of the Fermi contact and pseudocontact terms. J Magn Reson 2:286–301
Lange OF, Lakomek N-A, Farès C, Schröder GF, Walter KFA, Becker S, Meiler J, Grubmüller H, Griesinger C, de Groot BL (2008) Recognition dynamics up to microseconds revealed from an RDC-derived ubiquitin ensemble in solution. Science 320:1471–1475
Liu WM, Keizers PH, Hass MA, Blok A, Timmer M, Sarris AJ, Overhand M, Ubbink M (2012) A pH-sensitive, colorful, lanthanide-chelating paramagnetic NMR probe. J Am Chem Soc 134:17306–17313
Loh CT, Ozawa K, Tuck KL, Barlow N, Huber T, Otting G, Graham B (2013) Lanthanide tags for site-specific ligation to an unnatural amino acid and generation of pseudocontact shifts in proteins. Bioconjugate Chem 24:260–268
Longinetti M, Parigi G, Sgheri L (2002) Uniqueness and degeneracy in the localization of rigid domains in paramagnetic proteins. J Phys A: Math Gen 35:8153–8169
Longinetti M, Luchinat C, Parigi G, Sgheri L (2006) Efficient determination of the most favored orientations of protein domains from paramagnetic NMR data. Inverse Probl 22:1485–1502
Man B, Su XC, Liang H, Simonsen S, Huber T, Messerle BA, Otting G (2010) 3-Mercapto-2,6-pyridinedicarboxylic acid: a small lanthanide-binding tag for protein studies by NMR spectroscopy. Chem Eur J 16:3827–3832
McConnell HM, Robertson RE (1958) Isotropic nuclear resonance shifts. J Chem Phys 29:1361–1365
Meiler J, Peti W, Griesinger C (2000) DipoCoup: a versatile program for 3D-structure homology comparison based on residual dipolar couplings and pseudocontact shifts. J Biomol NMR 17:283–294
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr 53:240–255
Navarro-Vazquez A. (2012) MSpin-RDC. A program for the use of residual dipolar couplings for structure elucidation of small molecules. Magn Reson Chem 50:S73–S79
Oliphant TE (2007) Python for scientific computing. Comput Sci Eng 9:10–20
Otting G (2010) Protein NMR using paramagnetic ions. Annu Rev Biophys 39:387–405
Ozenne V, Bauer F, Salmon L, Huang JR, Jensen MR, Segard S, Bernadó P, Charavay C, Blackledge M (2012) Flexible-meccano: a tool for the generation of explicit ensemble descriptions of intrinsically disordered proteins and their associated experimental observables. Bioinformatics 28:1463–1470
Pintacuda G, Keniry MA, Huber T, Park AY, Dixon NE, Otting G (2004) Fast structure/based assignment of 15N HSQC spectra of selectively 15N labeled paramagnetic proteins. J Am Chem Soc 126:2963–2970
Pintacuda G, Park AY, Keniry MA, Dixonj NE, Otting G (2006) Lanthanide labeling offers fast NMR approach to 3D structure determinations of protein–protein complexes. J Am Chem Soc 128:3696–3702
Prestegard JH, Bougault CM, Kishore AI (2004) Residual dipolar couplings in structure determination of biomolecules. Chem Rev 104:3519–3540
Ravera E, Salmon L, Fragai M, Parigi G, Al-Hashimi HM, Luchinat C (2014) Insights into domain-domain motions in proteins and RNA from solution NMR. Acc Chem Res 47:3118–3126
Rinaldelli M, Ravera E, Calderone V, Parigi G, Murshudov GN, Luchinat C (2014) Simultaneous use of solution NMR and X-ray data REFMAC5 for joint refinement/detection of structural differences. Acta Crystallogr D D70:958–967
Rodriguez-Castañeda F, Haberz P, Leonov A, Griesinger C (2006) Paramagnetic tagging of diamagnetic proteins for solution NMR. Magn Reson Chem 44:S10–S16
Russo L, Maestre-Martinez M, Wolff S, Becker S, Griesinger C (2013) Interdomain dynamics explored by paramagnetic NMR. J Am Chem Soc 135:17111–17120
Saio T, Ogura K, Shimizu K, Yokochi M, Burke TR Jr, Inagaki F (2011) An NMR strategy for fragment-based ligand screening utilizing a paramagnetic lanthanide probe. J Biomol NMR 51:395–408
Schmitz C, Bonvin AM (2011) Protein–protein HADDocking using exclusively pseudocontact shifts. J Biomol NMR 50:263–266
Schmitz C, John M, Park AY, Dixon NE, Otting G, Pintacuda G, Huber T (2006) Efficient chi-tensor determination and NH assignment of paramagnetic proteins. J Biomol NMR 35:79–87
Schmitz C, Stanton-Cook MJ, Su XC, Otting G, Huber T (2008) Numbat: an interactive software tool for fitting Δχ-tensors to molecular coordinates using pseudocontact shifts. J Biomol NMR 41:179–189
Schmitz C, Vernon R, Otting G, Baker D, Huber T (2012) Protein structure determination from pseudocontact shifts using ROSETTA. J Mol Biol 416:668–677
Schwieters CD, Kuszewski J, Tjandra N, Clore GM (2003) The Xplor-NIH NMR molecular structure determination package. J Magn Reson 160:65–73
Simon B, Madl T, Mackereth CD, Nilges M, Sattler M (2010) An efficient protocol for NMR-spectroscopy-based structure determination of protein complexes in solution. Angew Chem Int Ed 49:1967–1970
Skinner SP, Moshev M, Hass MAS, Ubbink M (2013) PARAssign—paramagnetic NMR assignments of protein nuclei on the basis of pseudocontact shifts. J Biomol NMR 55:379–389
Su XC, Otting G (2010) Paramagnetic labelling of proteins and oligonucleotides for NMR. J Biomol NMR 46:101–112
Su XC, Huber T, Dixon NE, Otting G (2006) Site-specific labelling of proteins with a rigid lanthanide-binding tag. ChemBioChem 7:1599–1604
Su XC, Man B, Beeren S, Liang H, Simonsen S, Schmitz C, Huber T, Messerle BA, Otting G (2008a) A dipicolinic acid tag for rigid lanthanide tagging of proteins and paramagnetic NMR spectroscopy. J Am Chem Soc 130:10486–10487
Su XC, McAndrew K, Huber T, Otting G (2008b) Lanthanide-binding peptides for NMR measurements of residual dipolar couplings and paramagnetic effects from multiple angles. J Am Chem Soc 130:1681–1687
Swarbrick JD, Ung P, Chhabra S, Graham B (2011a) An iminodiacetic acid based lanthanide binding tag for paramagnetic exchange NMR spectroscopy. Angew Chem Int Ed Engl 50:4403–4406
Swarbrick JD, Ung P, Su XC, Maleckis A, Chhabra S, Huber T, Otting G, Graham B (2011b) Engineering of a bis-chelator motif into a protein alpha-helix for rigid lanthanide binding and paramagnetic NMR spectroscopy. Chem Commun (Camb) 47:7368–7370
Tjandra N, Bax A (1997) Direct measurement of distances and angles in biomolecules by NMR in a diluite liquid crystalline medium. Science 278:1111–1114
Tolman JR, Flanagan JM, Kennedy MA, Prestegard JH (1995) Nuclear magnetic dipole interactions in field-oriented proteins: information for structure determination in solution. Proc Natl Acad Sci USA 92:9279–9283
Tolman JR, Al-Hashimi HM, Kay LE, Prestegard JH (2001) Structural and dynamic analysis of residual dipolar coupling data for proteins. J Am Chem Soc 123:1416–1424
Valafar H, Prestegard JH (2004) REDCAT: a residual dipolar coupling analysis tool. J Magn Reson 167:228–241
Wassenaar TA, van Dijk M, Loureiro-Ferreira N, van der Schot G, de Vries SJ, Schmitz C, van der Zwan J, Boelens R, Giachetti A, Ferella L, Rosato A, Bertini I, Herrmann T, Jonker HRA, Bagaria A, Jaravine V, Guntert P, Schwalbe H, Vranken WF, Doreleijers JF, Vriend G, Vuister GW, Franke D, Kikhney A, Svergun DI, Fogh RH, Ionides J, Laue ED, Spronk C, Jurksa S, Verlato M, Badoer S, DalPra S, Mazzucato M, Frizziero E, Bonvin AMJJ (2012) WeNMR: structural Biology on the grid. J Grid Computing 10:743–767
Watanabe Y, Hiraoka W, Igarashi M, Ito K, Shimoyama Y, Horiuchi M, Yamamori T, Yasui H, Kuwabara M, Inagaki F, Inanami O (2010) A novel copper(II) coordination at His186 in full-length murine prion protein. Biochem Biophys Res Commun 394:522–528
Wei Y, Werner MH (2006) iDC: a comprehensive toolkit for the analysis of residual dipolar couplings for macromolecular structure determination. J Biomol 35:17–25
Wöhnert J, Franz KJ, Nitz M, Imperiali B, Schwalbe H (2003) Protein alignment by a coexpressed lanthanide-binding tag for the measurement of residual dipolar couplings. J Am Chem Soc 125:13338–13339
Yagi H, Maleckis A, Otting G (2013a) A systematic study of labelling an alpha-helix in a protein with a lanthanide using IDA-SH or NTA-SH tags. J Biomol NMR 55:157–166
Yagi H, Pilla KB, Maleckis A, Graham B, Huber T, Otting G (2013b) Three-dimensional protein fold determination from backbone amide pseudocontact shifts generated by lanthanide tags at multiple sites. Structure 21:883–890
Zhang Q, Stelzer AC, Fisher CK, Al-Hashimi HM (2007) Visualizing spatially correlated dynamics that directs RNA conformational transitions. Nature 450:1263–1267
Zhuang T, Lee HS, Imperiali B, Prestegard JH (2008) Structure determination of a Galectin-3-carbohydrate complex using paramagnetism-based NMR constraints. Protein Sci 17:1220–1231
Zweckstetter M (2008) NMR: prediction of molecular alignment from structure using the PALES software. Nat Protoc 3:679–690
Zweckstetter M, Bax A (2000) Prediction of sterically induced alignment in a dilute liquid crystalline phase: aid to protein structure determination by NMR. J Am Chem Soc 122:3791–3792
Author information
Authors and Affiliations
Corresponding authors
Additional information
Mauro Rinaldelli and Azzurra Carlon have contributed equally to this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rinaldelli, M., Carlon, A., Ravera, E. et al. FANTEN: a new web-based interface for the analysis of magnetic anisotropy-induced NMR data. J Biomol NMR 61, 21–34 (2015). https://doi.org/10.1007/s10858-014-9877-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-014-9877-4