Abstract
The specific protonation of valine and leucine methyl groups in proteins is typically achieved by overexpressing proteins in M9/D2O medium supplemented with either labeled α-ketoisovalerate for the labeling of the four prochiral methyl groups or with 2-acetolactate for the stereospecific labeling of the valine and leucine side chains. However, when these labeling schemes are applied to large protein assemblies, significant overlap between the correlations of the valine and leucine methyl groups occurs, hampering the analysis of 2D methyl-TROSY spectra. Analysis of the leucine and valine biosynthesis pathways revealed that the incorporation of labeled precursors in the leucine pathway can be inhibited by the addition of exogenous l-leucine-d10. We exploited this property to label stereospecifically the pro-R and pro-S methyl groups of valine with minimal scrambling to the leucine residues. This new labeling protocol was applied to the 468 kDa homododecameric peptidase TET2 to decrease the complexity of its NMR spectra. All of the pro-S valine methyl resonances of TET2 were assigned by combining mutagenesis with this innovative labeling approach. The assignments were transferred to the pro-R groups using an optimally labeled sample and a set of triple resonance experiments. This improved labeling scheme enables us to overcome the main limitation of overcrowding in the NMR spectra of prochiral methyl groups, which is a prerequisite for the site-specific measurement of the structural and dynamic parameters or for the study of interactions in very large protein assemblies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Progress in the NMR spectroscopy of high molecular weight proteins has been strongly connected to the development of new isotopic labeling schemes, in particular, the expression of selectively methyl-protonated perdeuterated proteins (Gardner and Kay 1997). The objective of these labeling approaches is to produce highly deuterated (i.e., >98 %) proteins with targeted [13CH3]-labeling at residue-specific methyl sites. Labeling protocols in which specifically [1H,13C]-methyl labeled biosynthetic precursors are added as the sole proton source in a perdeuterated culture medium can provide a high level of methyl protonation without detectable isotopic scrambling. The combination of the selective protonation of methyl groups in fully perdeuterated proteins with optimized methyl spectroscopy (Tugarinov et al. 2003; Amero et al. 2009) has enabled structural studies of large proteins and the dynamics of protein assemblies of up to 1 MDa to be probed by solution NMR techniques (Tugarinov et al. 2005; Gelis et al. 2007; Hiller et al. 2008; Sprangers and Kay 2007). Over the past 15 years, a variety of strategies for the selective labeling of the eight different types of methyl groups in proteins have been proposed (Tugarinov et al. 2006; Sprangers et al. 2007; Ruschak and Kay 2010; Plevin and Boisbouvier 2012). Initial methyl-labeling schemes used α-keto-acids as the precursors in the production of δ1-methyl protonated isoleucine (Gardner and Kay 1997) or for the simultaneous labeling of the two prochiral methyl groups of valine and leucine (Goto et al. 1999; Hajduk et al. 2000; Gross et al. 2003). These first protocols were later complemented by new schemes to specifically label methyl groups of methionine (Gelis et al. 2007; Fischer et al. 2007), alanine (Isaacson et al. 2007; Ayala et al. 2009; Godoy-Ruiz et al. 2010), isoleucine-γ2 (Ruschak and Kay 2010; Ayala et al. 2012) and threonine (Velyvis et al. 2012), which allowed the acquisition of high resolution spectra and the detection of structural and dynamic parameters in large molecular assemblies.
Most of these precursors allow independent labeling of each type of methyl side chain. The leucine and valine labeling schemes are more complex because the isopropyl group present in both side chains is generated before the two pathways diverge. Consequently, the labeling techniques directed towards these residues result in equal labeling of both residues. Labeling schemes based on α-ketoisovalerate led to the observation of four types of methyl groups sharing a common spectral window. For high molecular weight proteins, this labeling strategy can result in an overcrowded [1H,13C]-correlation spectra due to the sheer number of NMR-visible methyl probes. Furthermore, the first generation of α-ketoisovalerate precursors proved inefficient for studying large protein assemblies because the intense intra-residue 1H–1H dipolar interactions between prochiral methyl groups limited the sensitivity of NMR experiments. The introduction of [13CH3/CD3]-α-ketoisovalerate protonated on a single methyl (Tugarinov and Kay 2004; Lichtenecker et al. 2004) has been shown to enhance the resolution and sensitivity of the methyl-TROSY spectra of large proteins, despite the 50 % incorporation reduction of “NMR visible” 13CH3 isotopomers in the prochiral groups. However, α-ketoisovalerate is only available as a racemic mixture of pro-S and pro-R compounds. Therefore, two signals are still detected for each leucine and valine residue in the methyl-TROSY spectra. More recently, stereospecific labeling of the prochiral methyl groups of leucine and valine was achieved using specific methyl-labeled 2-acetolactate, a precursor involved in the early steps of the leucine and valine biogenesis pathway (Gans et al. 2010). The enzymatic rearrangement of the two methyl groups in labeled acetolactate during leucine and valine biosynthesis occurs in a stereospecific fashion such that the methyl group substituent at position 2 becomes the pro-S methyl group and the methyl at position 4 becomes the pro-R methyl group. While the aforementioned labeling scheme allows the study of symmetrical protein assemblies composed of low molecular weight subunits, the large number of overlaps between the methyl groups of valine and leucine still precludes the analysis of the NMR spectra for larger protein complexes. Recently, it was shown that the methyl resonances of leucine and valine could be distinguished using spectral properties of the corresponding 13Cα/β atoms (Hu et al. 2012). Here, we introduce a straightforward labeling scheme to incorporate stereospecific 13CH3 isotopomers into valine residues without labeling the corresponding leucine groups. The protocol, based on the simultaneous incorporation of labeled acetolactate and deuterated l-leucine, offers a significant simplification of [13C,1H]-methyl TROSY spectra. This new labeling scheme has been applied to the 468 kDa homododecameric peptidase TET2 and has allowed the complete assignment of the valine methyl resonances by combining mutagenesis, innovative labeling and adapted triple resonance experiments. As demonstrated here, this new method will be particularly useful for NMR studies of very large biomolecular assemblies.
Materials and methods
Precursor synthesis
2-Hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) and 2-hydroxy-2-[2H3]methyl-3-oxo-[1,2,3,4-tetra-13C]butanoate (pro-R acetolactate-13C4) were obtained from NMR-Bio (www.nmr-bio.com). 2-Hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H],[3,4-di-13C]butanoate (pro-Y acetolactate-13C3) was made as described previously (Gans et al. 2010) using ethyl [3,4-13C2]-3-oxobutanoate and 13CH3I (Sigma-Aldrich).
Optimization of the incorporation of leucine into overexpressed protein
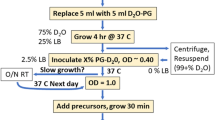
The initial experiments to determine the level of leucine incorporation into overexpressed proteins were performed using ubiquitin as a model system. Escherichia coli BL21(DE3) cells were transformed with a pET41c plasmid carrying the human His-tagged ubiquitin (pET41c-His-Ubi) gene, and the transformants were grown in M9/D2O medium containing 1 g/L 15ND4Cl and 2 g/L d-glucose-d7. When the optical density (O.D.) at 600 nm reached 0.8, a solution containing the labeled acetolactate precursors and the non-labeled l-leucine was added. After an additional 1 h, protein expression was induced by the addition of IPTG to a final concentration of 1 mM. The induced culture was grown for 3 h at 37 °C. The ubiquitin was then purified by Ni–NTA (Qiagen) chromatography in a single step.
The optimal quantity of l-leucine required to achieve near complete incorporation into the overexpressed protein was assessed in a series of cultures (50 mL each) in which different amounts of non-labeled l-leucine were added 1 h prior to induction to final concentrations of 0, 5, 10, 15, 20 and 60 mg/L in addition to a saturating quantity of the leucine and valine precursor pro-S acetolactate-13C (final concentration of 300 mg/L). l-methionine (methyl-13C) (Sigma-Aldrich) at 125 mg/L was used as an internal reference. The level of the non-labeled l-leucine incorporation into the purified protein was monitored using a 13C-HSQC experiment (figure S.1). When unlabeled leucine is incorporated into the overexpressed protein, the leucine-[13CH3]pro-S residues are replaced by unlabeled amino acids. The quantification was performed by comparing the volume of the leucine pro-S methyl group signals to the methionine methyl signals from 2D 13C-HSQC.
Production and purification of specifically methyl labeled TET2
Escherichia coli BL21-CodonPlus®(DE3)-RIL cells transformed with a pET-41c plasmid encoding TET2 were progressively adapted in three stages over 24 h to M9/D2O medium containing 1 g/L 15 ND4Cl and 2 g/L d-glucose-d7 (Sigma-Aldrich). In the final culture, the bacteria were grown at 37 °C in M9 medium prepared with 99.85 % D2O (Eurisotop). When the O.D. at 600 nm reached 0.8, a solution containing the labeled precursors was added. The precursor solution added per liter of culture medium contained:
-
125 mg of 2-oxo-3-[2H]-3-[2H3]methyl-4-[13C]-butanoate (α-ketoisovalerate) for the production of the U-[2H, 15N, 12C], Leu/Val-[13CH3/12CD3] TET2 sample (Tugarinov et al. 2006).
-
300 mg of 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) for the production of the U-[2H, 15N, 12C], Leu/Val-[13CH3]pro-S TET2 sample (Gans et al. 2010).
-
300 mg of 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) and 40 mg of l-leucine-d10 (Sigma-Aldrich) for the production of the U-[2H, 15N, 12C], Val-[13CH3]pro-S TET2 sample.
-
300 mg of 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) with 40 mg of l-leucine-d10 followed by the addition of 60 mg of 2-oxo-3-[2H2]-4-[13C]-butanoate (α-ketobutyrate; Gardner and Kay 1997) for the production of the U-[2H, 15N, 12C], Ile-[13CH3]δ1, Val-[13CH3]pro-S TET2 sample.
-
300 mg of 2-hydroxy-2-[2H3]methyl-3-oxo-[1,2,3,4-tetra-13C]butanoate (pro-R acetolactate-13C4) and 40 mg of l-leucine-d10 for the production of the U-[2H, 15N, 12C], Val-[2,3-2H2; 1,2,3-13C3; [13C1H3]pro-R/[12C2H3]pro-S] TET2 sample.
-
300 mg of 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H],[3,4-di-13C]butanoate (pro-Y acetolactate-13C3) and 40 mg of l-leucine-d10 for the production of the U-[2H, 15N, 12C], Val-[2,3-2H2; 3-13C; [13C2H3]pro-R/[13C1H3]pro-S] TET2 sample.
One hour after the addition of the precursors, TET2 expression was induced by the addition of IPTG to a final concentration of 0.5 mM. The induced culture grew for 4 h at 37 °C before harvesting. TET2 was purified using one anion exchange chromatography step (Resource Q 6 mL, GE Healthcare) followed by a size exclusion chromatography step (HiLoad 16/60 Superdex 200 pg, GE Healthcare). The final yield generally reached 20 mg/L methyl-specific protonated TET2. The protein was concentrated in 250 μL of buffered D2O (20 mM Tris (pH 7.4 uncorrected) and 20 mM NaCl) at a concentration of approximately 40–80 μM of TET2 dodecamer (~0.5–1 mM of monomer).
Production and purification of TET2 mutants
The constructs containing valine to alanine single point mutations were generated by an automated molecular biology platform (RoBioMol—Institut de Biologie Structurale, Jean-Pierre Ebel) using an automated PCR-based protocol adapted from the QuikChange site-directed mutagenesis method (Amero et al. 2011). The library of mutants was expressed in parallel using 24-well DeepWell plates. Each TET2 mutant was produced in 10 mL of M9/D2O medium supplemented with pro-S acetolactate-13C and l-leucine-d10, following the protocol described above. The cells were lysed by the addition of BugBuster® lysis buffer, and the plate containing the crude extracts was heated at 85 °C for 15 min. After centrifugation of the 24-well DeepWell plates at 4,000 rpm for 30 min, the labeled proteins were purified in parallel from supernatant fractions using a 96-well filter plate containing Q-Sepharose resin (GE Healthcare). The TET2 mutants were resuspended in 20 mM Tris (pH 7.4 uncorrected) and 20 mM NaCl (in D2O) to a final concentration of ~4 μM of TET2 dodecamer (~50 μM of monomer). Each sample (60 μL) was loaded in a 2.5 mm Shigemi tube placed coaxially to a regular 5 mm NMR tube used as a sample holder.
NMR spectroscopy
2D HSQC NMR spectra of ubiquitin were recorded at 37 °C on an Agilent DirectDrive spectrometer operating at a proton frequency of 600 MHz equipped with a cryogenic triple resonance probe head. All of the NMR spectra for the TET2 samples were recorded at 50 °C on an Agilent DirectDrive spectrometer operating at a proton frequency of 800 MHz equipped with a cryogenic triple resonance probe head. For the assignments of the methyl resonances using the SeSAM strategy (Amero et al. 2011), the duration of each 2D SOFAST-methyl-TROSY NMR experiment (Amero et al. 2009) was adjusted depending on the final concentration of the purified protein (experimental time ranging from 0.5 to 2 h maximum per sample). The angle of the proton excitation pulse was set to 30°, and the recycling delay was optimized to 0.8 s to enhance sensitivity.
A 3D HMQC-NOESY experiment was recorded over 64 h with a 1 mM U-[2H, 15N, 12C], Ile-[13CH3]δ1, Val-[13CH3]pro-S sample of TET2 with a NOE mixing time of 400 ms (which corresponds to the optimal NOE mixing time determined from the build-up of the cross-peak intensities in a series of short 2D NOESY spectra). The experiment was collected with 12 scans per increment and a maximum acquisition time of 20 ms in both the 13C and 1H indirect dimension. The 3D COSY-based “out-and-back” HCC (HC(C)C relay) experiments (Tugarinov and Kay 2003; Ayala et al. 2009; Ayala et al. 2012) were acquired in 11 h (44 h) with 0.5 mM U-[2H, 15N, 12C], Val-[2,3-2H2; 1,2,3-13C3; [13C1H3]pro-R/[12C2H3]pro-S] and 0.5 mM U-[2H, 15N, 12C], Val-[2,3-2H2; 3-13C; [13C2H3]pro-R/[13C1H3]pro-S] (U-[2H, 15N, 12C], Val-[2,3-2H2; 3-13C; [13C2H3]pro-R/[13C1H3]pro-S]) TET2 samples, and the experimental data were collected with 4 scans (16 scans) per increment and a maximum acquisition time of 11 and 12 ms in the two indirect carbon dimensions. All of the data were processed and analyzed using nmrPipe/nmrDraw (Delaglio et al. 1995) and CCPN software (Vranken et al. 2005).
Results and discussion
Leucine incorporation
The use of regioselectively labeled acetolactate improves the quality of spectra by reducing the number of resonances by a factor of two (Gans et al. 2010). However, for high molecular weight proteins containing many valine and leucine residues, a further reduction of overlaps is a prerequisite for the unambiguous assignment and analysis of complex NMR spectra. Leucine could be the choice candidate for this specific labeling because the metabolic pathway connecting α-keto-isovalerate (the immediate precursor of valine) to Leucine is irreversible (Fig. 1a). The direct addition of [2H,13C]-labeled l-leucine or its corresponding precursors will allow selective labeling of the leucines without scrambling to the valines. Recently, Lichtenecker et al. (2013) demonstrated that the addition of 2-oxo-3-[2H3]-4-[2H]-4-methyl[2H3]-5-[13C]-pentanoate (or α-ketoisocaproate) in M9/D2O culture medium can be used to directly label the methyl groups of leucine. Furthermore, the authors reported the preparation of a racemic mixture of α-ketoisocaproate precursors leading to the non-stereospecific labeling of either pro-S or pro-R methyl groups, where the occupancy level for each prochiral site was limited to 50 %. This decreased incorporation reduces the intensities of the structurally meaningful long range NOEs by a factor of 4. Because leucine and valine residues are equivalently abundant in proteins, this labeling scheme failed to significantly reduce the spectral overlap compared to the stereospecific labeling of valine and leucine residues using acetolactate precursors (Gans et al. 2010). In other words, the valine resonances suppressed by this labeling scheme are replaced by a similar number of leucine pro-R methyl groups overlapping with leucine pro-S resonances. Therefore, deuterated l-leucine (or its corresponding precursor) stereospecifically labeled on a single methyl group is required for the optimal incorporation and improvement of the NMR spectra of large proteins. Such a complex stereoselective synthesis of l-leucine was first reported by Kainosho et al. (2006) for the in vitro production of an optimally labeled protein. A similar approach is described by the same group in an accompanying article (Miyanoiri et al. 2013) regarding the in vivo labeling of the valine and leucine prochiral methyl groups of Malate Synthase G (MSG; Howard et al. 2000) overexpressed in E. coli.
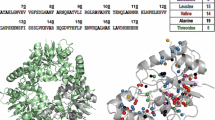
Incorporation pathway of acetolactate into the valine and leucine residues in E. coli. a The stereochemistry of the different intermediates in the leucine and valine biogenesis pathway is displayed (assuming incorporation of pro-S acetolactate-13C in D2O-based M9 culture medium). The enzymes responsible for catalyzing each reaction are indicated by EC number. EC 1.1.1.86: ketol-acid reductoisomerase; EC 4.2.1.9: dihydroxy-acid dehydratase; EC 2.6.1.42: branched-chain amino acid aminotransferase; EC 2.3.3.13: 2-isopropylmalate synthase; EC 4.2.1.33: 3-isopropylmalate dehydratase; and EC 1.1.1.85: 3-isopropylmalate dehydrogenase. Further information on the leucine and valine metabolic pathway can be found online: http://www.genome.jp/kegg/. b The decrease in the 13C1H3 isotopomers incorporation at the leucine pro-S methyl group position in overexpressed ubiquitin as a function of the added amount of exogenous unlabeled leucine. Different amounts of non-labeled l-leucine were added to the M9/D2O medium 1 h prior to induction to final concentrations of 0, 5, 10, 15, 20 and 60 mg/L, together with a saturating quantity of the leucine/valine precursor, pro-S acetolactate-13C (300 mg/L) and 125 mg/L l-methionine (methyl-13C). The decrease in the leucine pro-S methyl group signal is analyzed using the average volume of the methionine ε methyl group signals as a reference. A level of incorporation into the leucine side chains less than 5 % was obtained by adding 20 mg of leucine per liter of M9/D2O culture medium
Due to the complexity of this stereoselective synthesis and the associated costs of the labeled l-leucine or its precursors, we explored an alternative strategy to label only the valine methyl groups using more accessible precursors: labeled acetolactate and perdeuterated l-leucine. The addition of exogenous l-leucine not only allows dilution of the endogenous 13C-labeled l-leucine but also has an inhibitory effect on 2-isopropylmalate synthase (EC 2.3.3.13) (De Carvalho et al. 2005), catalyzing the conversion of α-ketoisovalerate to 2-isopropylmalate (the first specific step of the l-leucine biosynthetic pathway, Fig. 1). Therefore, the addition of exogenous l-leucine is expected to strongly reduce the flux of 13C-carbon in this pathway and to prevent the incorporation of labeled acetolactate into l-leucine (Fig. 1a). The effect of the addition of exogenous unlabeled l-leucine on the level of the incorporation of 13C into the leucine pro-S groups is reported on the Fig. 1b. To this end, ubiquitin was overexpressed in M9/D2O medium supplemented with a saturating amount of pro-S acetolactate-13C (300 mg/L) and various concentrations of unlabeled l-leucine. Nearly complete inhibition of incorporation (incorporation level ≤2 %) of pro-S acetolactate-13C was achieved for concentrations of unlabeled l-leucine ≥20 mg/L. Furthermore, no change was observed in the stereospecificity of the valine methyl group labeling (figure S.1). Surprisingly, we observed a slight (approximately 5–7 %) decrease in the valine signal, which was dependent on the exogenous quantity of added leucine (data not shown). However, by adding U-[13C] l-leucine in deuterated minimal medium containing U-[12C, 2H]-glucose with unlabeled acetolactate, we did not detect isotopic scrambling of the added l-leucine into l-valine (figure S.2). This result indicates that the weak unlabeling of valine residues observed when adding l-leucine is not due to incorporation into valine of exogeneous l-leucine or the corresponding degradation products, but is most likely related to a weak feedback inhibition of valine biosynthesis by excess l-leucine. Despite not identifying the exact metabolic pathway involved in the regulation of the labeled valine biosynthesis, we postulate that the greater than 90 % incorporation level of the 13C1H3 isotopomer into the valine pro-S groups obtained with the protocol described in this paper, along with the residual labeling of the corresponding leucine methyl groups being reduced to less of 2 %, would be sufficient for most biomolecular NMR applications.
Application of the labeling to TET2
This labeling strategy was then applied to the TET2 protein, a 468 kDa homododecameric aminopeptidase of 468 kDa involved in polypeptide degradation in the hyperthermophilic Archaea Pyrococcus horikoshii. This protein contains 37 valines and 23 leucines. The methyl groups of these 2 amino acid types have similar chemical shift dispersions, resulting in significant overlap between them. Figure 2 shows a comparison of the 13C-HMQC spectra of TET2 obtained for the non-stereospecific labeling of the leucine and valine methyl groups using an α-ketoisovalerate precursor (Fig. 2a; Tugarinov and Kay 2004), the stereospecific labeling of the leucine and valine pro-S methyl groups using acetolactate (Fig. 2b; Gans et al. 2010) and this new labeling protocol combining the acetolactate precursor with l-leucine-d10 for the stereospecific labeling of valine pro-S methyl groups only (Fig. 2c). Although the use of labeled acetolactate alone already decreased the number of signals twofold, many leucine and valine methyl resonances still overlapped (Fig. 2b), hampering spectral analysis. The specific labeling of the valine pro-S methyl groups enables a significant decrease in the number of signals and overlaps in the central part of the spectrum (Fig. 2b, c). The improvement of the resolution achieved in the 2D methyl-TROSY spectra using this specific labeling protocol allowed the observation of 32 individual correlations out of the 37 expected valine signals. Thus, the quality of the spectrum is sufficiently high to initiate assignment and NMR analyses using this improved labeling scheme.
Comparison of HMQC spectra recorded on specifically methyl-labeled TET2 samples (468 kDa). The 2D methyl-TROSY NMR spectra were recorded at 50 °C in D2O buffer (20 mM NaCl and 20 mM Tris, pH 7.4) on an NMR spectrometer operating at a proton frequency of 800 MHz. An enlarged picture of the central part of the spectrum is displayed in the bottom right corner of each panel. a U-[2H, 15N, 12C], Leu/Val-[13C1H3/12CD3] TET2 with non-stereospecific [13C1H3]-methyl labeling prepared using racemic 3-[2H3]methyl-3-[2H]-4-[13C]-α-ketoisovalerate (Tugarinov et al. 2006). b U-[2H, 15N, 12C], Leu/Val-[13C1H3]pro-S TET2 with stereospecific labeling prepared using 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) (Gans et al. 2010). c U-[2H, 15N, 12C], Val-[13C1H3]pro-S TET2 with stereospecific labeling of only valine residues prepared using 2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H]butanoate (pro-S acetolactate-13C) and l-leucine-d10
Assignment of TET2 valine pro-S methyl groups using the SeSAM approach
While efficient methods can be used to connect methyl resonances to sequentially assigned backbone nuclei in medium-size proteins, the stereospecific assignment of prochiral methyl groups remains a difficult step. Moreover, the size of the TET2 particle (468 kDa) impedes sequential assignments using classical approaches relying on magnetization transfer along the backbone and the side chain bonds. Recently, we have established an efficient approach based on the systematic mutagenesis of methyl containing residues for the sequence specific assignment of methyl groups in large proteins (Amero et al. 2011). This technique, named SeSAM for Sequence-Specific Assignment of methyl groups by Mutagenesis, is based on using site-directed mutagenesis to individually “turn off” the NMR signal of each methyl-containing amino acid in the target protein and thereby provide a sequence-specific resonance assignment. Conceptually, assignment-by-mutagenesis is straightforward. In practice, however, the overlap of resonances and the occurrence of secondary chemical shift changes (Sprangers et al. 2007; Amero et al. 2011) can make the analysis more difficult. The resonance perturbations of non-mutated methyl-containing residues are likely to occur but can be minimized using conservative mutations. In the first round of this approach, all of the spectra with only one missing peak are considered for a straightforward assignment of a first set of resonances. Then, more complex spectra affected by the secondary chemical shift perturbations are analyzed taking into consideration the first set of unambiguous assignments, the 3D structure of the protein assembly and the entire set of spectra. Because it cross-validates the results several times, considering the full library of single-site mutations greatly simplifies the process of resonance assignment.
Here, we systematically mutated each valine residue into an alanine. The stereospecific labeling of a single methyl group (valine pro-S) instead of the labeling of 4 methyl groups (valine and leucine) using α-ketoisovalerate reduces, up to a factor of 4, peak overlapping as well as the number of correlations affected by secondary chemical shift. Therefore, in most cases, the disappearance of only one signal unambiguously provides the assignment of the corresponding valine. For TET2, 24 of the 37 valines were directly assigned from the library of mutants. Examples of these assignments are presented for valine 76 and valine 193 in Fig. 3a, b, respectively, where the mutant spectra are shown superimposed on the WT spectrum. In the remaining spectra, the disappearance of the signal upon mutation was accompanied by small changes in the chemical shift of a few additional correlations (see, for example, the V15A mutant spectrum in Fig. 3c). Peak movements that do not directly concern the mutated resonance can complicate the process of obtaining a sequence-specific assignment from a single experiment, especially in an overcrowded region of the spectrum. Nonetheless, secondary chemical shift perturbations reflect modifications in the local electronic environment and can therefore provide complementary information that can be used to confirm the proposed assignment. Ambiguous assignments can therefore be readily cross-validated using structurally close and previously unambiguously assigned methyl groups. Analysis of the secondary chemical shift for the complete library of mutants with regards to the proximity predicted from the TET2 structure (PDB code: 1Y0R) allowed unambiguous assignment of eight supplementary valine resonances. The remaining valines correspond to weak methyl resonances or signals located in the overlapped area of the spectrum. These five remaining resonances, corresponding to valine residues 21, 46, 187, 236 and 334, were assigned based on the detection of intermethyl NOE correlations as described below. The completely assigned spectrum corresponding to the valine pro-S methyl groups of the TET2 protein is shown in Fig. 3d, and the proton and carbon-13 chemical shift values are listed in table S.1.
Assignment of TET2 pro-S valine methyl groups. The 2D SOFAST-methyl-TROSY spectra displayed in this figure were recorded using the valine to alanine mutant samples of the U-[2H, 15N, 12C], Val-[13C1H3]pro-S TET2 protein assembly. a The mutant spectra of V76A, b V193A and c V15A and the d WT spectrum of TET2 annotated with residue-specific assignments of the valine pro-S methyl groups. Each mutant spectrum extract (red) was overlaid with the reference spectrum of the native particle (black). The arrow in panel c indicates a secondary chemical shift perturbation
Cross-validation of the assignments using methyl–methyl NOEs
We have demonstrated that the use of the specific labeling of methyl groups in a small perdeuterated protein allowed detection of long-range NOEs between methyl groups separated by up to 12 Å (Sounier et al. 2007). For a protein the size of TET2, NOEs between methyl groups separated by up to 7 Å are still expected to be detectable. Therefore, if the crystal structure of a large assembly is available, the detection of NOE correlations between remote methyl probes could be used to confirm and extend the assignments. However, the analysis of the TET2 X-ray structure (Borissenko and Groll 2005) indicates that the expected number of NOE connectivities between valine pro-S methyl groups is reduced to approximately 50. Such a small number of putative NOE restraints is too low to allow cross-validation of the assignment. Consequently, to increase the number of NOE connectivities detected, we combined the labeling of the valine pro-S groups with isoleucine-δ1 methyl probes. The resonances of these methyl groups have distinct chemical shifts from valine resonances (figure S.3), avoiding overlapping. These methyl group resonances were previously assigned using the SeSAM approach (Amero et al. 2011).
A total of 292 NOE cross-peaks can be detected between the 71 labeled valine and isoleucine methyl probes, corresponding to an average of 4 connectivities for each methyl group. All of the observed NOE connectivities between assigned methyl probes correspond to the predicted NOE based on the theoretical methyl–methyl distance extracted from the crystal structure. We were unable to detect NOE connectivities involving leucine resonances or valine pro-R methyl groups, illustrating the specificity of the reported labeling strategy. Figure 4 shows examples of 2D planes extracted from the 3D 13C-HMQC-NOESY experiments at the 13C frequencies of the valine 105 pro-S, valine 87 pro-S and isoleucine 141 δ1 methyl groups. Each of these three residues presents correlations with the other two as well as with supplementary methyl groups. This unambiguous NOE network (as illustrated on the TET2 structure in Fig. 4) allows reliable cross-validation of the assignment based on mutagenesis. As already mentioned, some valine methyl groups remained unassigned after the SeSAM protocol. These residues have been assigned by analyzing the TET2 structure and looking for unassigned NOE correlations with their neighboring residues. Examples are shown in figure S.4 for the valine 46 and valine 21 methyl groups.
Cross-validation of the valine methyl group assignments. a The 2D extracts of the 3D 13C-HMQC–NOESY spectrum of the U-[2H, 15N, 12C], Ile-[13CH3]δ1, Val-[13CH3]pro-S TET2 sample. The planes were extracted at the 13C frequencies of valine 87 (NOE connectivity annotated in blue), valine 105 (red) and isoleucine 141 (green). b The NOEs observed in the three planes are displayed on the 3D structure of TET2 (PDB code: 1Y0R). Lines represent the detected NOE connections between the methyl groups, where each color is related to the corresponding residues annotated in the depicted 2D extracts
The assignment of such high quality 2D correlation spectra is a prerequisite for the measurement of structural and dynamic parameters or for the study of interactions. We have already shown that methyl-specific labeling can be used to detect the interaction between large protein assemblies, such as TET2, and small molecules (Amero et al. 2011). In this previous study, isoleucine-δ1 and alanine-β methyl probes were used to characterize the perturbations induced by the addition of amastatin, an inhibitor of aminopeptidases. The assigned valine pro-S groups can be used to map the binding site of this inhibitor more precisely. As shown in Fig. 5a, the addition of a saturating amount of amastatin significantly modified the position of the methyl resonances of several valine residues. Together with chemical shift changes previously detected for isoleucine-δ1 and alanine-β methyl groups (Amero et al. 2011), these new data better defined the interaction surface of the inhibitor in the TET2 internal cavity (Fig. 5b) and agreed with the TET2/amastatin structure previously resolved by X-ray crystallography (Borissenko and Groll 2005).
Characterization of the amastatin binding surface of TET2 by NMR spectroscopy. Left the overlay of a region of the 2D 1H–13C SOFAST-methyl-TROSY spectra (Amero et al. 2009) of free (black) and amastatin-bound (red) TET2 particles (Borissenko and Groll 2005) specifically labeled on the valine pro-S methyl groups. Right the binding-induced chemical shift perturbations mapped onto the inner surface of the TET2 dodecamer in complex with amastatin. Only an expansion of the TET2 region involved in the interaction with amastatin is displayed. The chemical shift perturbation data obtained for the valine residues were combined with previously obtained data for alanine and isoleucine methyl groups (Amero et al. 2011). The molecular surface corresponding to these residues is color-coded in blue (no perturbation) and red (shifted resonances). We observed a moderate chemical shift perturbation for the valine 161 resonance upon addition of amastatin, but this residue is located outside of the structure zoom displayed on the right panel
Transfer of assignments from TET2 valine pro-S to pro-R methyl groups
The stereospecific assignment of the valine pro-R methyl groups could be potentially obtained using the SeSAM approach. However, because the valine pro-S methyl groups were already assigned, it was more attractive to use these previous assignments as a starting point to identify the corresponding pro-R methyl resonances. The connection between both valine methyl resonances can be achieved using through-space NOE transfer provided that the protein sample is simultaneously [13C1H3]-labeled on both prochiral methyl groups, as previously reported (Sprangers and Kay 2007). This approach is limited by the significant overlap (Fig. 2a) from the presence of both pro-R and pro-S methyl signals in the NMR spectra and the extensive line broadening in the 1H-dimension due to the intense intra-residue methyl–methyl 1H–1H dipolar interaction (Tugarinov and Kay 2004). To preserve the high resolution of the NMR spectra (Fig. 2c), we preferred the exploitation of the intra-residue network of scalar couplings to connect both prochiral methyl group signals.
Indeed, the frequencies of the valine pro-S and pro-R methyl groups can be connected to Cβ resonances using a combination of two different TET2 samples labeled with different labeled acetolactate precursors and simple 3D ‘out-and-back’ NMR experiments. Sample 1 (SaI) was obtained by incorporating pro-R acetolactate-13C4 (2-hydroxy-2-[2H3]methyl-3-oxo-[1,2,3,4-tetra-13C]butanoate) to produce a TET2 sample with the valine residues containing a linear chain of 13C spins connecting the backbone atoms to the pro-R 13C1H3 groups (Fig. 6). A second sample (SaII) was obtained using pro-Y acetolactate-13C3 (2-hydroxy-2-[13C]methyl-3-oxo-4,4,4-tri-[2H],[3,4-di-13C]butanoate). For SaII, only the isopropyl end of the valine side chains is 13C-labeled, and only the pro-S methyl group is protonated (Fig. 6).
3D ‘out-and-back’ HCC and HC(C)C experiments used to transfer the assignment between prochiral methyl groups. Each valine-labeling scheme is described in the upper part of the figure. The three boxes represent the 3D extracts corresponding to the correlation detected for valine-310. The experiments used for the assignment transfer are a a 3D HCC experiment using the U-[2H, 15N, 12C], Val-[2,3-2H2; 1,2,3-13C3; [13C1H3]pro-R/[12C2H3]pro-S] TET2 sample prepared with pro-R acetolactate-13C4 and l-leucine-d10; b a 3D HCC experiment using the U-[2H, 15N, 12C], Val-[2,3-2H2; 3-13C; [13C2H3]pro-R/[13C1H3]pro-S TET2 sample prepared with pro-Y acetolactate-13C3 and l-leucine-d10; and c a 3D HC(C)C experiment using the U-[2H, 15N, 12C], Val-[2,3-2H2; 3-13C; [13C2H3]pro-R/[13C1H3]pro-S TET2 sample prepared with pro-Y acetolactate-13C3 and l-leucine-d10. The corresponding magnetization transfers are reported on the valine structure (at the top of each panel) with colored arrows. The assignment process is described in the lower part of the figure by the arrows showing the connection between the sampled frequencies in each of the three experiments. All of the spectra were recorded at 50 °C in D2O on an NMR spectrometer operating at a proton frequency of 800 MHz
The 3D COSY-based ‘out-and-back’ HCC experiments (Tugarinov and Kay 2003; Ayala et al. 2009, 2012) were collected using both SaI and SaII samples for the correlation of the resonances of each methyl group to the β carbon resonances. A common frequency can be used to connect prochiral methyl groups belonging to the same residues. However, in the case of TET2, which contains 37 valines, the superimpositions of some of the Cβ signals impede unambiguous assignments for 16 valines. We then used the SaII sample to collect a 3D ‘out-and-back’ HC(C)C-relay experiment to obtain another frequency to resolve the ambiguities. This experiment correlates the 13C resonances of the pro-R groups to the 13C and 1H resonances of the pro-S methyl groups. Together with the connection to the Cβ resonances, these supplementary connectivities linking together prochiral methyl groups allowed the reliable transfer of the sequence-specific assignment obtained for the pro-S methyls to all of the pro-R groups. This assignment protocol is illustrated in Fig. 6, which displays the 2D extracts corresponding to the methyl group chemical shifts for valine 310. The assignment of the pro-R methyl groups of the TET2 assembly is detailed in figure S.5, and the corresponding 1H and 13C chemical shift values are listed in table S.1.
Conclusion
A stereospecific labeling scheme of the valine methyl groups in large proteins was developed based on the simultaneous introduction of specifically labeled 2-acetolactate and perdeuterated l-leucine into the culture medium. This protocol enables efficient inhibition of the incorporation of labeled methyl groups into the leucine side chains (scrambling was reduced to less than 2 %), while the stereospecific incorporation of the 13CH3 isotopomers in valine is preserved (at approximately 95 %). This innovative labeling scheme was applied to the 468 kDa homododecameric TET2 protein to give a significant reduction in the number of overlaps between the resonances of the prochiral methyl groups. The resulting improvement of the 2D methyl TROSY spectra resolution allowed the assignment of all 37 valine methyl resonances of TET2 through a combination of mutagenesis, innovative labeling and adapted triple resonance experiments. This robust and simple labeling strategy will be particularly useful for the detection of meaningful structural and dynamic parameters or for the study of interactions in large oligomeric proteins containing many leucine and valine residues.
References
Amero C, Schanda P, Durá MA, Ayala I, Marion D, Franzetti B, Brutscher B, Boisbouvier J (2009) Fast two-dimensional NMR spectroscopy of high molecular weight protein assemblies. J Am Chem Soc 131:3448–3449
Amero C, Durá MA, Noirclerc-Savoye M, Perollier A, Gallet B, Plevin MJ, Vernet T, Franzetti B, Boisbouvier J (2011) A systematic mutagenesis-driven strategy for site-resolved NMR studies of supramolecular assemblies. J Biomol NMR 50:229–236
Ayala I, Sounier R, Usé N, Gans P, Boisbouvier J (2009) An efficient protocol for the complete incorporation of methyl-protonated alanine in perdeuterated protein. J Biomol NMR 43:111–119
Ayala I, Hamelin O, Amero C, Pessey O, Plevin MJ, Gans P, Boisbouvier J (2012) An optimized isotopic labelling strategy of isoleucine-γ2 methyl groups for solution NMR studies of high molecular weight proteins. Chem Commun 48:1434–1436
Borissenko L, Groll M (2005) Crystal structure of TET protease reveals complementary protein degradation pathways in prokaryotes. J Mol Biol 346:1207–1219
De Carvalho LPS, Argyrou A, Blanchard JS (2005) Slow-onset feedback inhibition: α inhibition of mycobacterium tuberculosis α-isopropylmalate synthase by l-leucine. J Am Chem Soc 127:10004–10005
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Fischer M, Kloiber K, Häusler J, Ledolter K, Konrat R, Schmid W (2007) Synthesis of a 13C methyl group labeled methionine precursor as a useful tool for simplifying protein structural analysis by NMR spectroscopy. Chem Biochem 8:610–612
Gans P, Hamelin O, Sounier R, Ayala I, Durá MA, Amero CD, Noirclerc-Savoye M, Franzetti B, Plevin MJ, Boisbouvier J (2010) Stereospecific isotopic labeling of methyl groups for NMR spectroscopic studies of high molecular weight proteins. Ang Chem Int Ed 49:1958–1962
Gardner KH, Kay LE (1997) Production and incorporation of 15N, 13C, 2H (1H-δ1 methyl) isoleucine into proteins for multidimensional NMR studies. J Am Chem Soc 119:7599–7600
Gelis I, Bonvin AMJJ, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG (2007) Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131:756–769
Godoy-Ruiz R, Guo C, Tugarinov V (2010) Alanine methyl groups as NMR probes of molecular structure and dynamics in high-molecular-weight proteins. J Am Chem Soc 132:18340–18350
Goto N, Gardner K, Mueller G, Willis R, Kay L (1999) A robust and cost-effective method for the production of Val, Leu, Ile (δ1) methyl-protonated 15N-, 13C-, 2H-labeled proteins. J Biomol NMR 13:369–374
Gross JD, Gelev VM, Wagner G (2003) A sensitive and robust method for obtaining intermolecular NOEs between side chains in large protein complexes. J Biomol NMR 25:235–242
Hajduk PJ, Augeri DJ, Mack J, Mendoza R, Yang J, Betz SF, Fesik SW (2000) NMR-based screening of proteins containing 13C-labeled methyl groups. J Am Chem Soc 122:7898–7904
Hiller S, Garces RG, Malia TJ, Orekhov VY, Colombini M, Wagner G (2008) Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 321:1206–1210
Howard BR, Endrizzi JA, Remington SJ (2000) Crystal structure of Escherichia coli malate synthase G complexed with magnesium and glyoxylate at 2.0 Å resolution: mechanistic implications. Biochemistry 39:3156–3168
Hu W, Namanja AT, Wong S, Chen Y (2012) Selective editing of Val and Leu methyl groups in high molecular weight protein NMR. J Biomol NMR 53:113–124
Isaacson RL, Simpson PJ, Liu M, Cota E, Zhang X, Freemont P, Matthews S (2007) A new labeling method for methyl transverse relaxation-optimized spectroscopy NMR spectra of alanine residues. J Am Chem Soc 129:15428–15429
Kainosho M, Torizawa T, Iwashita Y, Terauchi T, Mei Ono A, Güntert P (2006) Optimal isotope labelling for NMR protein structure determinations. Nature 440:52–57
Lichtenecker R, Ludwiczek ML, Schmid W, Konrat R (2004) Simplification of protein NOESY spectra using bioorganic precursor synthesis and NMR spectral editing. J Am Chem Soc 126:5348–5349
Lichtenecker RJ, Coudevylle N, Konrat R, Schmid W (2013) Selective isotope labelling of leucine residues by using α-ketoacid precursor compounds. Chem Biochem 14:818–821
Miyanoiri Y, Takeda M, Okuma K, Ono AM, Terauchi T, Kainosho M (2013) Differential isotope-labeling for Leu and Val residues in a protein by E. coli cellular expression using stereo-specifically methyl labeled amino acids. J Biomol NMR. doi:10.1007/s10858-013-9784-0
Plevin MJ, Boisbouvier J (2012) Isotope-labelling of methyl groups for NMR studies of large proteins. In: Clore GM, Potts J (eds) Recent developments in biomolecular NMR. Royal Society of Chemistry 1–24. doi:10.1039/9781849735391-00001
Ruschak A, Kay L (2010) Methyl groups as probes of supra-molecular structure, dynamics and function. J Biomol NMR 46:75–87
Sounier R, Blanchard L, Wu Z, Boisbouvier J (2007) High-accuracy distance measurement between remote methyls in specifically protonated proteins. J Am Chem Soc 129:472–473
Sprangers R, Kay LE (2007) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445:618–622
Sprangers R, Velyvis A, Kay LE (2007) Solution NMR of supramolecular complexes: providing new insights into function. Nat Meth 4:697–703
Tugarinov V, Kay LE (2003) Ile, Leu, and Val methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J Am Chem Soc 125:13868–13878
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172
Tugarinov V, Hwang PM, Ollerenshaw JE, Kay LE (2003) Cross-correlated relaxation enhanced 1H–13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125:10420–10428
Tugarinov V, Choy W-Y, Orekhov VY, Kay LE (2005) Solution NMR-derived global fold of a monomeric 82-kDa enzyme. Proc Natl Acad Sci USA 102:622–627
Tugarinov V, Kanelis V, Kay LE (2006) Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat Protoc 1:749–754
Velyvis A, Ruschak AM, Kay LE (2012) An economical method for production of 2H, 13CH3-threonine for solution NMR studies of large protein complexes: application to the 670 kDa proteasome. PLoS One 7:e43725
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Acknowledgments
We would like to thank Drs. M. Kainosho, P. Macek, M. Plevin, P. Schanda, A. Sivertsen, Mrs I. Ayala and R. Kerfah, as well as Mr. T. Ogden, for stimulating discussions, Dr. D. Marion for help in processing the NMR spectra and Drs. T. Vernet and M. Noirclerc-Savoye for the preparation of the library of mutants. We thank Dr. B. Franzetti for providing clones of TET2. This work used the RoBioMol, high-field NMR and the isotopic labeling facilities at the Grenoble Instruct Centre (ISBG; UMS 3518 CNRS-CEA-UJF-EMBL) with support from FRISBI (ANR-10-INSB-05-02) and GRAL (ANR-10-LABX-49-01) within the Grenoble Partnership for Structural Biology (PSB). The research leading to these results has received funding from the European Research Council under the European Community’s Seventh Framework Program FP7/2007-2013 Grant Agreement no. 260887.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10858_2013_9785_MOESM1_ESM.pdf
The NMR spectra used to characterize the incorporation of exogenous l-leucine in overexpressed ubiquitin, as well as the table and the spectra corresponding to the assignment of the TET2 valine methyl groups, are available online: http://springerlink.bibliotecabuap.elogim.com/journal/volumesAndIssues/10858 (PDF 2499 kb)
Rights and permissions
About this article
Cite this article
Mas, G., Crublet, E., Hamelin, O. et al. Specific labeling and assignment strategies of valine methyl groups for NMR studies of high molecular weight proteins. J Biomol NMR 57, 251–262 (2013). https://doi.org/10.1007/s10858-013-9785-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-013-9785-z