Abstract
Selective methyl labeling combined with HMQC spectroscopy that exploits a TROSY effect in 13CH3 spin systems has significantly extended the utility of solution NMR spectroscopy in studies of high molecular weight particles. Herein we compare the utility of 13CH3- versus 13CHD2-labeling of Ile, Leu, Val probes in supra-molecular systems through quantification of relative signal-to-noise ratios in optimized spectra of highly deuterated, 13CH3- and 13CHD2-labeled samples of the half proteasome (α7α7, 360 kDa). It is shown that the sensitivity of spectra recorded on Ile, Leu, Val 13CH3-labeled samples is between 1.5 and 2 fold higher than the corresponding data sets obtained on α7α7 with 13CHD2 probes. Thus, labeling of supra-molecules with 13CH3 isotopomers remains the method of choice, but in applications where 13CHD2 moieties are required, sensitivity will in general not be limiting.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the past several years it has become increasingly clear that methyl groups are very valuable NMR probes of supra-molecular structure and dynamics (Tugarinov et al. 2004). A significant number of applications involving suitably methyl-labeled protein systems ranging in molecular weight up to 1 MDa have emerged (Amero et al. 2009; Gelis et al. 2007; Hamel and Dahlquist 2005; Isaacson et al. 2007; Kreishman-Deitrick et al. 2005; Religa et al. 2010; Sprangers et al. 2005; Sprangers and Kay 2007; Velyvis et al. 2009; Velyvis et al. 2007), providing important insight into function and complementing existing data derived from X-ray or cryo-EM analyses. All of these NMR studies are based on experiments utilizing HMQC ‘building blocks’ (Bax et al. 1983; Mueller 1979) that exploit a TROSY effect in which cancellation of intra-methyl dipolar interactions occurs (Tugarinov et al. 2003). By producing samples with 13CH3 isotopomers and high levels of deuteration at all other positions it is possible to obtain spectra of high resolution and sensitivity that can then be used in a wide range of studies (Tugarinov and Kay 2005a). Our laboratory has shown that in cases where isopropyl methyl groups are utilized (such as for Leu and Val) it is best to label samples as U-[2H]-Leu,Val-[13CH3,12CD3] so that only one of the two methyl groups is NMR active, thus minimizing inter-methyl relaxation that would otherwise deteriorate the quality of the data (Tugarinov and Kay 2004). To date most applications have made use of either U-[2H], Ile-[13CH3 δ1] or U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] labeling, although both Met-13CH3 (Fischer et al. 2007; Gelis et al. 2007) and Ala-13CH3 (Amero et al. 2009; Isaacson et al. 2007) methyl groups are also sometimes employed.
As described in detail elsewhere, intra-methyl dipolar relaxation during the t1 period of the 13C-1H HMQC scheme is completely eliminated for a coherence transfer pathway that involves 50% of the magnetization and 1H-1H dipolar relaxation contributions for this pathway are also removed during t2 (Tugarinov et al. 2003). Several years ago we were interested in establishing whether labeling using 13CHD2 moieties would also give rise to high quality spectra since relaxation from 1H-1H dipoles would be eliminated de facto. We showed, in the case of highly deuterated malate synthase G (MSG, 723 residues, 80 kDa) produced only with Ile-[13CH3 δ1] probes, that significant increases in spectral sensitivity (approximately three-fold, see below) could be achieved when methyl groups of the 13CH3 variety are employed versus 13CHD2 isotopomers (Ollerenshaw et al. 2005). Although high quality Ile 13C-1H spectra were obtained using 13CHD2 probes, the decrease in the number of 1H spins and the concomitant increase in 1H T1 relaxation times relative to a protein where Ile residues are labeled 13CH3 results in a significant negative effect on signal-to-noise (s/n) in data sets.
The MSG study focused only on an Ile labeled protein whose size, while certainly large by the standards of several years ago, must be considered small relative to the molecular machines that are the focus of some NMR efforts now. With this in mind, herein we revisit 13CH3 versus 13CHD2 labeling using samples of the half proteasome, comprised of two heptameric rings of identical α-subunits (α7α7) with a molecular weight of 360 kDa (120 ns correlation time at 50°C (Sprangers and Kay 2007)) that is more ‘in line’ with the supra-molecular systems that currently are under study by a number of groups. Specifically, we are interested in comparing the quality of 13C-1H correlation maps recorded on samples with Ile, Leu and Val probes, since the increased number of labels is often necessary to get good ‘coverage’ over the entire molecule. To this end U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] and U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 samples have been prepared. A priori it is not clear which of these two labeling schemes would be preferred. As we show below, high quality spectra can be recorded of samples where both of the isopropyl methyl groups of Leu and Val are labeled 13CHD2, an advantage relative to the 13CH3 labeling scheme where high resolution data sets are best recorded if labeling is restricted to only 1 of the 2 methyl groups (Tugarinov and Kay 2004). Moreover, the larger 1H density associated with a 13CHD2-labeling scheme that includes all Ile, Leu and Val probes, especially in the case where both (proximal) prochiral methyls are 13CHD2 would be expected to reduce longitudinal relaxation times and hence improve spectral sensitivity (relative to labeling specific to Ile, for example). Based on a careful analysis of spectra we show here that significant differences in sensitivity, on the order of a factor of 1.5–2, are obtained in favor of 13CH3-probes when Ile-[δ1], Leu, Val labeled α7α7 is considered. These results establish that 13CH3 methyl labeling remains the method of choice in studies of supra-molecular systems, but that in applications where 13CHD2 labeling is preferred, such as in studies of methyl dynamics via 13C relaxation (Ishima et al. 1999; Tugarinov and Kay 2005b), sensitivity loses will not be prohibitive.
Materials and methods
U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] and U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 samples were generated as described previously (Sprangers and Kay 2007). Samples comprising 0.6 mM protein were prepared in 100% D2O, 25 mM potassium phosphate pH 6.8, 50 mM NaCl, 1 mM EDTA, 0.03% NaN3. In order to quantify the relative amounts of protein in each of the samples Met-13CH3 was added to growth media approximately 1 h prior to induction of protein over-expression (100 mg/L) to produce samples that included Met-methyl labeling as well.
In order to establish which of the possible 13C-1H correlation spectra would be optimal for recording spectra of Ile, Leu, Val 13CHD2-labeled proteins we have evaluated the relative s/n ratios of correlations in unenhanced-HSQC (Bodenhausen and Rubin 1980), PEP-enhanced HSQC (Cavanagh and Rance 1993) and TROSY-HSQC (Pervushin et al. 1997) data sets. Spectra were recorded with relaxation delays of 2 s, and with acquisition times of (30 ms, 64 ms) in (t1,t2), followed by a two-fold extension of the t1 domain using linear prediction (Zhu and Bax 1992). Relative s/n values were calculated for each peak in a pair of data sets as \( {\frac{{N^{j} S^{i} }}{{N^{i} S^{j} }}} \)where S i is a peak intensity and N i is the noise floor in data set i. In a similar manner, a number of experiments were evaluated for optimal s/n for Ile, Leu, Val-13CH3 labeling. These included a standard HMQC [excitation pulse of 90° (Bax et al. 1983; Mueller 1979)] as well as SOFAST HMQC [excitation pulse 60°, either methyl selective or non-selective (Amero et al. 2009)]. Data sets were recorded with relaxation delays of 1 s (standard HMQC) or 0.5 s (SOFAST), as discussed in the text, with (t1,t2) = (30 ms, 64 ms) and with the number of transients adjusted so that net acquisition times were identical. Time domain spectra were doubled by linear prediction (t1) and subsequently processed identically. Data sets recorded on 13CH3 and 13CHD2 samples were normalized for small differences in protein concentrations by quantifying s/n ratios of Met correlations in HMQC spectra obtained with relaxation delays of either 4 or 8 s. Identical ratios were obtained for both recovery delays. Based on analysis of s/n values, SOFAST and PEP-HSQC where chosen as the ‘optimal’ experiments for recording correlation maps of 13CH3- and 13CHD2-labeled α7α7, respectively. SOFAST and PEP-HSQC spectra were obtained for comparing s/n ratios, using acquisition parameters as described above, with equal net recording times (1.7 h/spectrum). In all of the above analyses s/n values were quantified only for well-resolved correlations; in total 16, 30, and 29 Ile, Leu and Val methyl groups were used.
Relaxation rates of the relevant coherences for each of the 13C-1H correlation data sets were measured using pulse schemes that follow directly from those described previously with very minimal modifications (Ollerenshaw et al. 2005). Values of 1H R1 (13CH3, 13CHD2), 13C single-quantum R2 (13CHD2), 13C single-quantum TROSY component R2 (13CHD2), 1H-13C slowly relaxing multiple-quantum relaxation rate (13CH3), 1H single-quantum R2 (13CHD2), 1H single-quantum TROSY component R2 (13CHD2) and 1H slowly relaxing transverse rate (13CH3) were quantified on a per-residue basis. Values of all rates for α7α7 (50°C, 800 MHz), separated according to residue type (Ile, Leu and Val), are reported in Supporting Information (SI).
Results and discussion
A quantitative analysis of the relative s/n of correlations in data sets recorded on two separate samples can only be performed if protein concentrations are known accurately. In the studies reported here this was achieved by labeling all samples with 1H,13Cε-Met and comparing volumes of Met methyl correlations in spectra recorded under fully relaxed conditions. Such an analysis established that U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] and U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 samples used here differed in protein concentration by only 2% and this difference was corrected for in all sensitivity ratios reported herein.
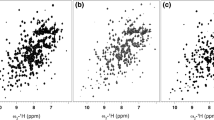
Figure 1 shows selected regions from (a) 13C-1H HMQC (13CH3 probes) and (b) sensitivity enhanced 13C-1H HSQC (13CHD2) maps of α7α7, 800 MHz, 50°C (sample concentrations of 0.6 mM in monomer; measuring times of 1.7 h using a room-temperature probe). It is clear that both data sets are of very high quality so that the relative sensitivities of correlations can be accurately quantified. Before this can be accomplished, however, it is first necessary to establish which of several different possible 13C-1H correlation experiments is the most sensitive for each of the labeled samples. In the case of 13CHD2-α7α7 we have considered (i) unenhanced HSQC (Bodenhausen and Rubin 1980), (ii) PEP-HSQC (Cavanagh and Rance 1993) and (iii) TROSY-HSQC (Pervushin et al. 1997). The PEP scheme was recorded without gradient coherence transfer selection as this requires the insertion of several additional small delays into the pulse scheme to account for the coherence selection gradients that may decrease the sensitivity of the experiment slightly.
Contour plots of selected regions of a 13C-1H HMQC (13CH3 probes) and b sensitivity enhanced 13C-1H HSQC (13CHD2) maps of α7α7, 800 MHz, 50°C (recorded on a system with a room temperature probe-head in 1.7 h/spectrum). Traces for selected residues are shown. Data sets are plotted at the same noise floor
Figure 2 shows the relative s/n ratios of correlations in each of the recorded data sets. The PEP scheme is the most sensitive but it is important to note that the relative sensitivities of the experiments will depend somewhat on the experimental parameters. Measured 13C and 1H transverse relaxation rates are presented in Supporting Information that provides some guide as to optimal selection of acquisition parameters. As described in “Materials and methods” we have chosen a t1 acquisition time of 30 ms that is subsequently doubled by linear prediction. This is a reasonable compromise between sensitivity and resolution, with average 13C single quantum T2 values ranging between 25 and 40 ms (13CHD2), 1H-13C multiple-quantum values between 30 and 40 ms (13CH3) and 13C TROSY T2 values of approximately 60 ms (13CHD2, 800 MHz). Resultant spectra are of sufficiently high resolution to resolve the great majority of correlations. Clearly the relative sensitivity of the TROSY experiment grows with increasing t1 acquisition times, although out to 50 ms there are still sensitivity advantages for the PEP method for Ile residues, with similar s/n ratios for both Leu/Val (PEP HSQC vs. TROSY), at least at 800 MHz. It is noteworthy, however, that as static magnetic fields increase the TROSY approach likely will be the method of choice for recording spectra of proteins prepared with 13CHD2 methyl labeling, especially in cases where resolution becomes critical.
Histograms plotting the relative s/n of Ile, Leu and Val correlations in 13C-1H correlation spectra of U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 (800 MHz, 50°C). a Comparison of PEP-HSQC versus unenhanced HSQC data sets and b PEP-HSQC versus TROSY-HSQC correlation maps. Details of processing and analysis are provided in “Materials and methods”. The dotted vertical lines delineate average ratios
An advantage with the 13CHD2 label is that both isopropyl methyls of Val and Leu can be NMR active without compromising the high quality of 13C, 1H correlation maps (see Fig. 1b). The increased 1H density around each of the methyl probes does lead to decreases in 1H transverse relaxation times (from ~37 ms for Ile to 25 ms for Leu/Val, see Fig. S1e) and hence attenuation of sensitivity. However, a decided advantage is the increased 1H longitudinal relaxation rates arising from the proximal prochiral 13CHD2 pairs for Leu/Val that increases sensitivity per unit measurement time for these residues. This is illustrated in Fig. 3 where histograms of 1H longitudinal relaxation times of Ile, Leu and Val residues of α7α7 are indicated, showing significantly decreased Leu/Val 1H T1 values relative to Ile. Some of this decrease results from the fact that, in general, Ile residues are inherently more dynamic than Leu/Val; for example the rotation of methyl groups about their three-fold symmetry axis is faster for Ile than for Leu/Val (Tugarinov and Kay 2005b), leading to larger Ile 1H T1 values. However, a substantial fraction of the difference between Ile and Leu/Val longitudinal relaxation rates most certainly arises from proximity of methyl probes to adjacent protons (other methyl groups). Also displayed in Fig. 3 are histograms of 1H T1 values recorded on a U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] α7α7 sample. These are considerably shorter for 13CH3 methyl groups, as expected, a decided advantage for the 13CH3-labeling approach.
Recently, Amero et al. (2009) have published SOFAST methyl-TROSY experiments that optimize s/n per measurement time in HMQC spectra recorded on 13CH3-labeled proteins. We have compared standard HMQC data sets (relaxation delay 1 s) with SOFAST spectra recorded with the flip angle, θ, of the excitation pulse (90° in the ‘typical experiment’) set to 60° (relaxation delay of 0.5 s) using the pulse scheme of Boisbouvier, Brutscher and coworkers (Amero et al. 2009). Clearly, the optimal value for the relaxation delay depends on both θ and 1H longitudinal relaxation times, and with the significant differences in average 1H T1 values for Ile (1.34 s, 13CH3) and Leu,Val (0.84 s, 13CH3) in α7α7 (Fig. 3) no single value can optimize sensitivity for all residues. Because the signals from Ile residues are inherently two-fold greater than from Leu,Val (non-stereospecific 13CH3,12CD3 isopropyl labeling reduces the effective concentration of methyl labels for Leu/Val by twofold) we have chosen to optimize s/n in the Leu/Val region of the spectrum by using the values listed above. It is noteworthy, however, that these settings attenuate signals from Ile residues only marginally—by ≤5%—relative to their maximum possible values.
We have also compared data sets obtained where the θ = 60° pulse is either selective for the methyl region of the spectrum or not. In the former case protons outside of the methyl region are not excited and their equilibrium polarization can serve to (slightly) enhance recovery of methyl magnetization (Amero et al. 2009; Pervushin et al. 2002). Figure 4 shows that relatively small gains in sensitivity (≈4–5%) are obtained with the SOFAST approach relative to the standard experiment and that essentially no benefits arise from using a methyl selective excitation pulse. The sensitivity benefits from SOFAST are somewhat smaller than the calculated gains of close to 10% and considerably less than the 30% that Amero et al. (2009) have measured for a U-[2H], Ile-[13CH3 δ1] TET2 sample (correlation time of 315 ns). The fact that no gain is achieved by using a selective excitation pulse indicates that the α7α7 sample used is very highly deuterated, as expected.
Histograms of relative s/n values of Ile, Leu and Val 13C-1H correlations in spectra recorded on U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] α7α7 (800 MHz, 50°C) based on a comparison of HMQC (relaxation delay of 1 s) and SOFAST HMQC (relaxation delay of 0.5 s) data sets. In a correlations from HMQC and SOFAST HMQC (60° non-selective excitation pulse) are compared, while in b correlations from SOFAST HMQC data sets with a selective or non-selective excitation pulse (60°) are analyzed
Sensitivity ratios, summarized in Figs. 2 and 4, indicate that the ‘best’ pulse schemes for recording correlation maps of U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] and U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 under the experimental conditions used here are SOFAST HMQC and PEP-HSQC, respectively. We have therefore compared relative s/n values for 13CH3- and 13CHD2-labeled proteins using these optimal experiments, as summarized in Fig. 5. Sensitivity gains with the 13CH3-label vary between 1.5 to close to 2, on average. These enhancements are somewhat less than what has been previously reported based on studies of Ile-δ1 labeled MSG (≈threefold), although in that case the HMQC (methyl-TROSY) scheme (13CH3) was compared with the unenhanced HSQC (13CHD2) and both data sets were recorded with a relaxation delay of 1.5 s that favors the 13CH3-labeled sample. Indeed when data sets are recorded on Ile-δ1 α7α7 using the same pulse schemes and acquisition parameters as for MSG the intensity ratio increases to 2.9 ± 0.5, consistent with previous results.
Relative s/n values of correlations in ‘optimal’ 13C-1H correlation spectra recorded on samples of U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,12CD3] and U-[2H], Ile-[13CHD2 δ1], Leu,Val-[13CHD2,13CHD2] α7α7 (800 MHz, 50°C), separated according to amino acid type. Each of the data sets was recorded in 1.7 h
In summary, based on the studies of α7α7 reported here, U-[2H], Ile-[13CH3 δ1], Leu,Val-[13CH3,13CD3] labeling remains the best approach for recording high sensitivity methyl data sets of very high molecular weight proteins. With the development of precursors for specific 13CH3-labeling at either the pro-R or pro-S (Gans et al., 2010) methyl positions of Leu/Val, sensitivity of the HMQC experiments will increase further by two-fold for these residues and spectral complexity will be halved, although a pair of data sets will be required to measure all possible Leu/Val methyl correlations. It is also clear from this work that sensitivity losses associated with 13CHD2 labeling are not prohibitive so that a number of applications that are best performed on AX or ‘AX-like’ spin systems, including 13C and 1H relaxation studies, are certainly feasible, even when supra-molecular systems are considered.
References
Amero C, Schanda P, Dura MA, Ayala I, Marion D, Franzetti B, Brutscher B, Boisbouvier J (2009) Fast two-dimensional NMR spectroscopy of high molecular weight protein assemblies. J Am Chem Soc 131:3448–3449
Bax A, Griffey RH, Hawkings BL (1983) Correlation of proton and nitrogen-15 chemical shifts by multiple quantum NMR. J Magn Reson 55:301–315
Bodenhausen G, Rubin DJ (1980) Natural abundance nitrogen-15 NMR by enhanced heteronuclear spectroscopy. Chem Phys Lett 69:185–189
Cavanagh J, Rance M (1993) Sensitivity-enhancement NMR techniques for the study of biomolecules. Ann Reports NMR Spectrosc 27:1–58
Fischer M, Kloiber K, Hausler J, Ledolter K, Konrat R, Schmid W (2007) Synthesis of a 13C-methyl-group-labeled methionine precursor as a useful tool for simplifying protein structural analysis by NMR spectroscopy. Chembiochem 8:610–612
Gans P, Hamelin O, Sounier R, Ayala I, Dura MA, Amero C, Noirclerc-Savoye M, Franzetti B, Plevin MJ, Boisbouvier J (2010) Stereospecific isotopic labeling of methyl groups for NMR spectroscopic studies of high molecular weight proteins. Angew Chem Int Ed. doi:10.1002/anie.200905660
Gelis I, Bonvin AM, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG (2007) Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131:756–769
Hamel DJ, Dahlquist FW (2005) The contact interface of a 120 kD CheA-CheW complex by methyl TROSY interaction spectroscopy. J Am Chem Soc 127:9676–9677
Isaacson RL, Simpson PJ, Liu M, Cota E, Zhang X, Freemont P, Matthews S (2007) A new labeling method for methyl transverse relaxation-optimized spectroscopy NMR spectra of alanine residues. J Am Chem Soc 129:15428–15429
Ishima R, Louis JM, Torchia DA (1999) Transverse C-13 relaxation of CHD2 methyl isotopomers to detect slow conformational changes of protein side chains. J Am Chem Soc 121:11589–11590
Kreishman-Deitrick M, Goley ED, Burdine L, Denison C, Egile C, Li R, Murali N, Kodadek TJ, Welch MD, Rosen MK (2005) NMR analyses of the activation of the Arp2/3 complex by neuronal Wiskott–Aldrich syndrome protein. Biochemistry 44:15247–15256
Mueller L (1979) Sensitivity enhanced detection of weak nuclei using heteronuclear multiple quantum coherence. J Am Chem Soc 101:4481–4484
Ollerenshaw JE, Tugarinov V, Skrynnikov NR, Kay LE (2005) Comparison of 13CH3, 13CH2D, and 13CHD2 methyl labeling strategies in proteins. J Biomol NMR 33:25–41
Pervushin K, Riek R, Wider G, Wüthrich K (1997) Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci USA 94:12366–12371
Pervushin K, Vogeli B, Eletsky A (2002) Longitudinal (1)H relaxation optimization in TROSY NMR spectroscopy. J Am Chem Soc 124:12898–12902
Religa TL, Sprangers R, Kay LE (2010) Dynamic regulation of archaeal proteasome gate opening as studied by TROSY NMR. Science 328:98–102
Sprangers R, Kay LE (2007) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445:618–622
Sprangers R, Gribun A, Hwang PM, Houry WA, Kay LE (2005) Quantitative NMR spectroscopy of supramolecular complexes: dynamic side pores in ClpP are important for product release. Proc Natl Acad Sci U S A 102:16678–16683
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172
Tugarinov V, Kay LE (2005a) Methyl groups as probes of structure and dynamics in NMR studies of high-molecular-weight proteins. Chembiochem 6:1567–1577
Tugarinov V, Kay LE (2005b) Quantitative 13C and 2H NMR relaxation studies of the 723-residue enzyme malate synthase G reveal a dynamic binding interface. Biochemistry 44:15970–15977
Tugarinov V, Hwang P, Ollerenshaw J, Kay LE (2003) Cross-correlated relaxation enhanced 1H–13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125:10420–10428
Tugarinov V, Hwang PM, Kay LE (2004) Nuclear magnetic resonance spectroscopy of high-molecular-weight proteins. Annu Rev Biochem 73:107–146
Velyvis A, Yang YR, Schachman HK, Kay LE (2007) A solution NMR study showing that active site ligands and nucleotides directly purturb the allosteric equilibrium in aspartate transcarbamolyase. Proc Natl Acad Sci U S A 104:8815–8820
Velyvis A, Schachman HK, Kay LE (2009) Application of methyl-TROSY NMR to test allosteric models describing effects of nucleotide binding to aspartate transcarbamoylase. J Mol Biol 387:540–547
Zhu G, Bax A (1992) Two-dimensional linear prediction for signals truncated in both dimensions. J Magn Reson 98:192–199
Acknowledgments
T.L.R. acknowledges The European Molecular Biology Organization (ALTF 827-2006) and The Canadian Institutes of Health Research (CIHR) for postdoctoral fellowships. L.E.K. holds a Canada Research Chair in Biochemistry. This work was supported by a grant from the CIHR.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Religa, T.L., Kay, L.E. Optimal methyl labeling for studies of supra-molecular systems. J Biomol NMR 47, 163–169 (2010). https://doi.org/10.1007/s10858-010-9419-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-010-9419-7