Abstract
Carr-Purcell-Meiboom-Gill (CPMG) relaxation dispersion NMR spectroscopy has emerged as a powerful tool for quantifying the kinetics and thermodynamics of millisecond exchange processes between a major, populated ground state and one or more minor, low populated and often invisible ‘excited’ conformers. Analysis of CPMG data-sets also provides the magnitudes of the chemical shift difference(s) between exchanging states (|Δϖ|), that inform on the structural properties of the excited state(s). The sign of Δϖ is, however, not available from CPMG data. Here we present one-dimensional NMR experiments for measuring the signs of 1HN and 13Cα Δϖ values using weak off-resonance R 1ρ relaxation measurements, extending the spin-lock approach beyond previous applications focusing on the signs of 15N and 1Hα shift differences. The accuracy of the method is established by using an exchanging system where the invisible, excited state can be converted to the visible, ground state by altering conditions so that the signs of Δϖ values obtained from the spin-lock approach can be validated with those measured directly. Further, the spin-lock experiments are compared with the established H(S/M)QC approach for measuring the signs of chemical shift differences. For the Abp1p and Fyn SH3 domains considered here it is found that while H(S/M)QC measurements provide signs for more residues than the spin-lock data, the two different methodologies are complementary, so that combining both approaches frequently produces signs for more residues than when the H(S/M)QC method is used alone.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NMR spectroscopy has emerged as a powerful tool for investigating the role of protein dynamics in a variety of biologically important processes that include allostery, ligand binding, catalysis, and molecular recognition (Boehr et al. 2006; Frederick et al. 2007; Henzler-Wildman and Kern 2007; Kay et al. 1998; Popovych et al. 2006; Sugase et al. 2007). Particularly significant in this regard is the fact that motion can be probed over a broad spectrum of time-scales and in a site-specific manner (Ishima and Torchia 2000; Mittermaier and Kay 2006; Palmer et al. 1996) so that a very detailed description of dynamics is, in principle, possible. Over the course of the past several decades a large number of different experiments (Igumenova et al. 2006; Mittermaier and Kay 2006; Palmer et al. 2005) and labeling schemes (Goto and Kay 2000; Kainosho et al. 2006; LeMaster 1999; Tugarinov and Kay 2004; Zhang et al. 2006) have been developed, optimized for different classes of biomolecules and for the investigation of dynamics in different time-regimes. One particular area that has generated significant recent interest is the study of millisecond (ms) time-scale dynamics, which can be quantified using CPMG relaxation dispersion experiments (Palmer et al. 2001). Although the basic ideas behind this approach date back over 50 years (Carr and Purcell 1954; Meiboom and Gill 1958), the successful application of the CPMG method to complex systems such as biomolecules had to await developments in both pulse sequence methodology and labeling technology (Loria et al. 1999). In the case of protein applications it is now possible to measure backbone 1HN (Ishima and Torchia 2003), 15N (Loria et al. 1999; Tollinger et al. 2001), 13Cα (Hansen et al. 2008b), 1Hα (Lundstrom et al. 2009a) and 13CO (Ishima et al. 2004; Lundstrom et al. 2008) as well as side-chain 13Cβ (Lundstrom et al. 2009b) and methyl 13C (Lundstrom et al. 2007b; Skrynnikov et al. 2001) CPMG dispersion profiles for systems in exchange between states that include a major conformation and one or more minor conformers so long as their populations are greater than ≈0.5% with lifetimes between ≈0.5 and 10 ms (Palmer et al. 2001).
The utility of the CPMG relaxation dispersion experiment is several-fold. First, populations of exchanging conformers can be obtained along with rates of exchange (Palmer et al. 2001), so that in cases where experiments are performed as a function of temperature and/or pressure it is possible to generate a detailed one-dimensional energy landscape for the system under study (Korzhnev et al. 2004, 2006). Second, absolute values of chemical shift differences between exchanging states, |Δϖ|, are extracted from fits of dispersion profiles. In cases where the signs of Δϖ are available, the chemical shifts of the excited states can be obtained and exploited for structural studies of these conformers (Vallurupalli et al. 2008). This is particularly significant when one considers that these excited states are generally populated at low levels and only very transiently so that they are invisible both to most other NMR methods and to other biophysical techniques in general.
A number of approaches for obtaining the signs of the chemical shift differences that are conspicuously missing from CPMG dispersion experiments have been developed. In one type of experiment, suggested by Skrynnikov et al. (2002), peak positions in the indirect dimensions of HSQC and HMQC data-sets recorded at several static magnetic fields are compared to isolate the sign information for 15N and 13C Δϖ values (referred to in what follows as the H(S/M)QC method). A second approach, termed CEESY, is based on very similar principles except that the sign information is encoded in the relative peak intensities of a pair of data-sets (van Ingen et al. 2006). This approach has been applied to obtain the sign of 15N and 1HN Δϖ values. Finally, a third method measures selective R 1ρ relaxation rates as a function of spin-lock offset (Auer et al. 2009; Hansen et al. 2009; Korzhnev et al. 2003, 2005; Massi et al. 2004; Trott and Palmer 2002), which can be a sensitive reporter of sign information as well. To date, applications of R 1ρ methods have focused on both 15N (Korzhnev et al. 2005) and 1Hα (Auer et al. 2009) spins. Recognizing the importance of chemical shifts in structural studies of excited states, the goal of the present work is to extend the methodology further to include both 13Cα and 1HN nuclei as well. With the development of several different experimental schemes for measuring such values it is important to establish the limitations and strengths of each approach. With this in mind results from R 1ρ and H(S/M)QC approaches are compared and an evaluation of the relative merits of each set of experiments is presented.
Materials and methods
Sample preparation
An NMR sample of U-[15N,2H] Abp1p SH3 domain was prepared as described in detail previously (Vallurupalli et al. 2007). The final protein concentration was ≈1.5 mM, in a buffer consisting of 50 mM sodium phosphate, 100 mM NaCl, 1 mM EDTA, 1 mM NaN3, 90% H2O/10% D2O, pH = 7.0. Ark1p peptide, which binds the SH3 domain (Haynes et al. 2007), was expressed and purified as described previously (Vallurupalli et al. 2007) and added to the Abp1p SH3 domain sample to produce a complex with 2.5 ± 0.1% mole fraction bound, as established by 15N CPMG relaxation dispersion experiments (Hansen et al. 2008a). 15N (Korzhnev et al. 2005) and 1HN R 1ρ experiments, along with 15N–1HN H(S/M)QC data-sets (Skrynnikov et al. 2002) were measured on this sample. A sample of the A39V/N53P/V55L Fyn SH3 domain was prepared as U-13C, ≈50% 2H, by protein over-expression in M9 minimal media with 50% D2O and 3 g/l [13C6,2H7]-glucose, as described in detail previously (Neudecker et al. 2006). The sample was 1.0 mM in protein, 0.2 mM EDTA, 0.05% NaN3, 50 mM sodium phosphate, pD 7.0, 100% D2O and was used to record 1Hα R 1ρ rates. 13Cα R 1ρ and H(S/M)QC spectra were recorded on a second A39V/N53P/V55L Fyn SH3 domain sample (1.0 mM) which was generated using [2-13C]-glucose as the sole carbon source to produce isolated 13Cα nuclei, as described previously (Lundstrom et al. 2007a).
NMR spectroscopy
All measurements were carried out at 25°C (Abp1p SH3/Ark1p peptide) or 20°C (A39V/N53P/V55L Fyn SH3) using Varian Inova spectrometers operating at 1H resonance frequencies of 500 and 800 MHz and equipped with triple-resonance room-temperature probes. Residue selective 1D R 1ρ experiments were recorded at 800 MHz. For each residue known to be affected by exchange two decay curves were recorded, corresponding to \( R_{1\rho }^{ \pm } \) (see below), with relaxation delays T up to 60 ms (1Hα, 13Cα) or 100 ms (1HN, 15N). For 15N/1HN R 1ρ measurements each of the 9 points comprising a single decay curve (\( R_{1\rho }^{ + } \) or \( R_{1\rho }^{ - } \)) was recorded in 1.8/2.35 min, resulting in a total measurement time of 1.1/1.4 h per residue (duplicates for each of the 9 points were obtained). R 1ρ values for 13Cα/1Hα were based on 7 time points (duplicates for only 2 of the time points) each requiring 11.3/4.7 min of acquisition time for a net time of 3.4/1.4 h for the complete set of \( R_{1\rho }^{ \pm } \) decays curves.
Signs of 15N and 13Cα shift differences were also obtained from 15N–1HN and 13Cα–1Hα HSQC and HMQC data-sets which were recorded at 500 and 800 MHz in 3 and 2.5 h, respectively. Each data-set was measured in duplicate. Data were processed and analyzed with the nmrPipe/nmrDraw suite of programs (Delaglio et al. 1995) and NMRViewJ (Johnson and Blevins 1994). Simulation and numerical data interpretation were carried out using in-house programs written in Matlab (MathWorks, Inc.).
Statistical analysis
A number of statistical tests have been used to evaluate whether differences in measured \( R_{1\rho }^{ \pm } \) rates are statistically significant, and hence whether the sign of the chemical shift difference between exchanging states can be extracted from the data (see below). The first approach uses a Student t-test analysis (Zar 1984) in which R 1ρ rates are extracted from fits of experimental data to a straight line
and \( R_{1\rho }^{ \pm } \) values compared according to
where
is the variance of the difference in the slopes \( R_{1\rho }^{ \pm } \). The value \( s_{yx}^{2} \) is, in turn, given by
where x i,± are the relaxation time delays (T values in the pulse schemes of Figs. 1 and 6) used to record the decay curves for measuring \( R_{1\rho }^{ \pm } \) comprised of n + and n − points, y i,± are the corresponding \( \ln \left[ {{\frac{{I^{ \pm } (T)}}{{I^{ \pm } (0)}}}} \right] \) values and n + + n − − 4 = ν is the number of degrees of freedom. Calculated Student t-values were compared with two-sided t-test statistics at the 95% confidence level (for example, t 0.05(2),14 = 2.15 for ν = 14 degrees of freedom, which is germane for the measurements here). For t-values such that t > t α(2),ν the \( R_{1\rho }^{ \pm } \) rates are significantly different with confidence level >(1 − α).
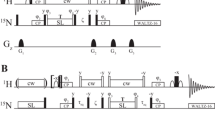
Pulse scheme for measuring 1HN off-resonance R 1ρ relaxation rates in U-15N labeled proteins. 1H and 15N carrier frequencies are placed initially on resonance for the peak of interest; subsequently the 1H carrier is jumped to the position of the spin-lock field immediately prior to the pulse of phase ϕ4 and then to the water line prior to application of gradient 5. All solid pulses have flip angles of 90° and are applied along the x-axis, unless indicated otherwise. The shaped 1H pulses are water selective (Piotto et al. 1992). Simultaneous 1H/15N spin-lock fields (90 Hz) are applied for durations of 1/|JNH| (between points a, b and c, d) where JNH is the one-bond 1HN–15N scalar coupling constant (≈−90 Hz). Immediately after gradient 1 1H purge pulses are applied (17 kHz) for durations of 4 ms (x-axis) and 2 ms (y-axis) to eliminate residual water signal. 1H pulses of phases ϕ4/ϕ5 are applied with a flip angle θ such that tanθ = ω1/|δ G |, using (δ G ,ω1) optimized as described in the text. During the spin-lock period an on-resonance 15N continuous-wave decoupling field of 1.1 kHz is applied to eliminate scalar coupling modulations as well as cross-correlation effects between 1H–15N dipolar and 1H CSA interactions (Peng and Wagner 1992). The delay τeq is set to 5 ms to ensure that the magnetization from each of the exchanging states corresponds faithfully to the equilibrium distribution (Korzhnev et al. 2005). 15N decoupling during acquisition is achieved with a WALTZ-16 field (Shaka et al. 1983). The phase cycle is: ϕ1 = (y,−y), ϕ2 = 2(x), 2(−x), ϕ3 = 4(x), 4(−x), ϕ6 = x, 2(−x), x, −x, 2(x), −x. For the spin-lock carrier upfield(downfield) of the ground state resonance ϕ4 = y(−y) and ϕ5 = −y(y) (on Varian spectrometers). Gradient strengths and durations are (ms,G/cm): G0 = (1,7.5), G1 = (0.5,10), G2 = (0.8,15), G3 = (0.6,−4), G4 = (0.3,−4), G5 = (0.5,8), G6 = (0.4,−20)
A second method has also been used to evaluate whether \( R_{1\rho }^{ \pm } \) rates are distinct, based on the F-test statistic (Zar 1984). Here the intensities of the decay curves I +(T) and I −(T), from which \( R_{1\rho }^{ + } \) and \( R_{1\rho }^{ - } \) are obtained, respectively, are used to generate \( y_{e}^{\prime } (T) = I^{ + } (T)/I^{ - } (T) \). The function \( y_{e}^{\prime } (T) \) is subsequently fit to (1) a horizontal line \( y_{c}^{\prime } = k \) and (2) an exponential \( y_{c}^{\prime } = a\exp (bT) \) and χ2 values calculated for each fit according to
where σ i is the error of each data point, computed as the root mean squared deviations of peak volumes from duplicate measurements. Values of χ 2 obtained from the two models (‘linear’ vs. ‘exponential’) are compared using F-test statistics. A cut-off of p < 2.5% (1 − p > 97.5%) was used to establish whether \( R_{1\rho }^{ \pm } \) differ. Both the t-test and F-test give consistent results for the protein systems studied here.
Results and discussion
Measuring signs of Δϖ by low power off-resonance R 1ρ measurements
For a two-site exchange process involving the interconversion of ground (G) and excited (E) states, \( G\mathop \rightleftharpoons \limits_{{k_{EG} }}^{{k_{GE} }} E \), with the population of the ground state, p G , greatly exceeding that of the excited state, p E , the rotating frame relaxation rate of a nucleus attached to state G is given by (Trott and Palmer 2002)
where \( R_{1} \) and \( R_{2}^{o} \) are longitudinal and intrinsic transverse relaxation rates, respectively, and R ex sin2 θ is the contribution to R 1ρ from chemical exchange. In (5.2) Δω = Ω E − Ω G (rad/sec), where Ω i is the chemical shift of the nucleus in state i, k ex = k GE + k EG , θ = arctan(ω 1/δ G ), ω 1 is the strength of the applied field, \( \omega_{E,eff}^{2} = \omega_{1}^{2} + \delta_{E}^{2} \) and δ E = Ω E − Ω SL , δ G = Ω G − Ω SL are the resonance offsets from the spin-lock carrier for states E and G, respectively. Note that the maximum in R ex occurs when the spin-lock field is resonant with the frequency of the minor state (5.2) and that R ex measured for δ G ≈ −Δω (i.e., Ω SL ≈ Ω E ) is larger than the corresponding value obtained for δ G ≈ Δω. Thus, by recording a pair of R 1ρ values, corresponding to δ G ≈ ± Δω it is possible to establish the sign of Δω since the rate measured for δ G ≈ −Δω will be larger than for δ G ≈ Δω (Korzhnev et al. 2003, 2005; Massi et al. 2004; Trott and Palmer 2002).
Experiments for measurement of the signs of Δω using the R 1ρ approach have been developed previously for both 15N (Korzhnev et al. 2005) and 1Hα (Auer et al. 2009) nuclei and are similar to schemes described by Boulat and Bodenhausen for the measurement of 1H relaxation rates in proteins (Boulat and Bodenhausen 1993) and Hansen and Al-Hashimi for quantifying 13C R 1ρ values in nucleic acids (Hansen et al. 2009). The experiments make use of very weak spin-lock fields that are optimized for each spin examined (see below) and are thus best implemented in one-dimensional mode, resolution permitting. Analogous experiments, exploiting the improved resolution of two-dimensional spectroscopy, and using larger spin-lock fields have also been described (Korzhnev et al. 2003; Massi et al. 2004).
Figure 1 shows the pulse sequence that has been designed for the measurement of 1HN R 1ρ values (and hence for determination of the sign of 1HN Δϖ; in what follows ϖ is in units of ppm) using very weak spin-lock fields, following very closely on approaches published for 15N and 1Hα nuclei (Auer et al. 2009; Korzhnev et al. 2005). Briefly, isolation of the resonance in question is achieved by selective Hartmann-Hahn magnetization transfers from 1HN to 15N (a to b) and back (c to d) using matched spin-lock fields of strength 90 Hz (Chiarparin et al. 1998; Pelupessy and Chiarparin 2000); for spin-locks of this magnitude correlations outside ≈1.5|JNH| from the positions of the 1H/15N radio-frequency carriers are suppressed. The resonance of interest is subsequently aligned along the effective magnetic field by a θ-pulse just prior to point e, followed by a relaxation delay of duration T, during which time the R 1ρ decay rate is measured. A pair of relaxation curves are obtained, corresponding to δ G ≈ ±Δω, from which the sign of Δϖ is established (see below). In order to maximize the difference between R 1ρ values recorded for δ G ≈ +Δω and δ G ≈ −Δω, (referred to in what follows as \( R_{1\rho }^{ + } \) and \( R_{1\rho }^{ - } \), respectively) and hence increase the sensitivity of the experiment, optimal spin-lock offset (δ G ) and field strength (ω 1) values are calculated for each residue by maximizing
as described previously (Auer et al. 2009). In (6) Δ is the difference in decay curves measured for δ G ≈ ±Δω at time T, normalized to T = 0 and T corresponds to the maximum time value used in experiments (either 50 or 100 ms in the applications considered here) with
We have chosen to validate the methodology using an Abp1p SH3 domain–ligand exchanging system described in detail in previous publications (Hansen et al. 2008b; Vallurupalli et al. 2007, 2008), with only a small mole-fraction of ligand added, p E = 2.5%, k ex = 300 s−1, 25°C (as established by CPMG relaxation dispersion). The addition of small amounts of ligand renders the bound conformer the invisible, ‘excited’ state whose chemical shifts can be quantified by CPMG (magnitude) and R 1ρ (sign) relaxation experiments. The signs of Δϖ obtained using the R 1ρ approach can then be directly compared with those measured from spectra of apo and fully ligand-bound conformers as a rigorous test of the method. Figure 2 illustrates a number of examples, focusing on both selective 1HN and 15N R 1ρ experiments. Optimized values of (δ G ,ω 1) were calculated using the exchange parameters from CPMG measurements listed above and maximizing Δ of (6). The first column in Fig. 2 shows selected regions of 2D 15N–1HN correlation maps centered on A12 (a), N28 (b) and D34 (c) with the position of the excited state correlation indicated by the green dot. The 1D spectra obtained from the 15N and 1HN selective R 1ρ experiments recorded on these particular residues are shown as insets in the figures and the positions of the spin-lock fields are denoted by arrows (orange-upfield and red-downfield for 1HN and purple-upfield, blue-downfield for 15N). The 1HN and 15N decay curves in columns 2 and 3, respectively, establish that for the most part distinct \( R_{1\rho }^{ \pm } \) values are measured from which the signs of Δϖ are readily obtained. For example, for A12 the red curve decays more rapidly than the orange, establishing that the sign of Δω = Ω E − Ω G is positive, consistent with expectations from the direct measurements of chemical shifts of apo- and ligand-saturated SH3 domain (green peak). For N28 \( R_{1\rho }^{ + } ({}^{ 1}{\text{H}}^{\text{N}} ) \approx R_{1\rho }^{ - } ({}^{ 1}{\text{H}}^{\text{N}} ) \) and it is not possible to obtain the sign of 1HN Δω (ambiguity indicated by the pair of green peaks in the spectrum).
Selected regions of 15N–1HN HSQC spectra centered on A12 (a), N28 (b) and D34 (c) of the Abp1p SH3 domain, 2.5% Ark1p peptide, 25°C, 800 MHz. The green dots correspond to positions of the excited state correlations (two dots if the 1HN sign could not be extracted from R 1ρ measurements). 1D spectra obtained from the pulse scheme of Fig. 1 are shown as insets. Arrows indicate the positions of the spin-lock fields that are used to generate the \( R_{1\rho }^{ \pm } \) decay curves for 1HN (orange, red; 2nd column) and 15N (purple, blue; 3rd column) nuclei of A12, N28 and D34. Optimized (|δ G /(2π)|,ν 1) values for measurements at 800 MHz calculated from (6, 7) along with k ex = 300 s−1, p E = 2.5% and the value of |Δϖ| from CPMG experiments are listed
For many residues with 1HN, 15N chemical shifts such that |\( \Updelta \varpi_{{H^{N} }} \)| > 0.035 ppm, |\( \Updelta \varpi_{N} \)| > 0.3 ppm, substantial differences in \( R_{1\rho }^{ \pm } \) rates are measured (see Fig. 2) and it is straightforward to obtain the desired sign. In cases where values of Δϖ are small, leading to similar \( R_{1\rho }^{ \pm } \) rates we have employed a number of statistical tests, including a Student t-test analysis and an F-test (Zar 1984), to help with the selection of the correct sign, as outlined in “Materials and Methods”. Shown in Fig. 3 are plots of \( I^{ \pm } (T) \) (a–c), along with \( y_{e}^{\prime } (T) = I^{ + } (T)/I^{ - } (T) \) and the best fit horizontal line (d–f) for a number of residues. Note that panels (d–f) provide a complementary visual approach for assessing the data; only in cases where \( y_{e}^{\prime } (T) \) is ‘distinct’ from the horizontal line does it follow that \( R_{1\rho }^{ + } \ne R_{1\rho }^{ - } \). In order to provide a ‘frame of reference’ for what constitutes ‘distinct’ we have chosen residues where \( R_{1\rho }^{ \pm } \) (1HN) values are different with essentially 1 − p = 100% certainty (N16; a,d), ≈97% certainty (E30; b,e) and <85% certainty (F20; c,f), based on an F-test analysis, with t-values of 50.2, 2.4 and 1.0, respectively. Note that a t-value of 2.15 corresponds to a confidence level of 95% for the 14 degrees of freedom in each fit.
1HN\( R_{1\rho }^{ \pm } \) decay curves (\( I^{ + } (T),I^{ - } (T) \)) for residues N16 (a), E30 (b) and F20 (c) of the Abp1p SH3 domain (Abp1p SH3 domain/Ark1p peptide exchanging system), 25°C, 800 MHz, listing calculated Student t-values along with 1 − p values calculated from an F-test analysis. Also shown are \( y^{\prime}_{e} (T) = I^{ + } (T)/I^{ - } (T) \) and the best fit horizontal line for N16 (d), E30 (e) and F20 (f)
Because the Abp1p SH3 domain–peptide exchanging system has been chosen, for which the signs of Δϖ are known a priori, it is possible to establish some general guidelines that are helpful for selecting correct signs from the R 1ρ methodology discussed above. (1) For values of t < 2.15 (t-test) or 1 − p < 97.5% (F-test) the data must be disregarded even in the few cases where on the basis of inspection of decay curves one might be tempted to choose a sign. Our experience for the several cases in the Abp1p SH3 domain that are in this category is that erroneous signs can be obtained if the data is analyzed by inspection. (2) For borderline cases where (1 − p)-values are on the order of 97.5% (or t values on the order of 2.15 for the decay curves here), each data-set must be inspected carefully and only in cases where all of the data points from one curve are either above or below those of the second decay profile should signs be extracted and they must be treated with skepticism. (3) For values of t > 3.5 (t-test) or 1 − p > 99.5% (F-test) accurate values of the sign are obtained.
Comparison of the signs isolated from the R 1ρ method, following the criteria listed above, with the ‘correct’ values measured directly from spectra showed that all of the signs for the 29 1HN values that were obtained in confidence were correct, with Δϖ values ≥0.03 ppm quantified. In the case of 15N, signs could be obtained for 26 residues using the statistical test criteria described in “Materials and Methods” and for all but one residue |Δϖ| values were larger than 0.15 ppm. Interestingly, for L57, for which |Δϖ| = 0.1 ppm, a wrong sign was obtained. We are uncertain as to why this is the case, however, it is worth noting that simulations establish that a statistically significant difference between \( R_{1\rho }^{ \pm } \) values should not have been obtained for this residue with the exchange parameters of the Abp1p SH3 domain–peptide system used here.
In addition to cross-validating the R 1ρ experiments by establishing how many of the determined signs are correct, as discussed above, it is also possible to compare experimental and calculated |R 1ρ | rates. Figure 4 shows the correlation between calculated and measured |ΔR 1ρ | values (a,1HN; b,15N), with the former given by
Correlation between measured |ΔR 1ρ | values (Y-axis) and |ΔR 1ρ | calculated according to (8) (X-axis) for 1HN (a) and 15N (b) nuclei of the Abp1p SH3 domain. Calculated values use \( \Updelta \varpi_{{H^{N} }} \) and \( \Updelta \varpi_{N} \) shift differences based on direct measurements of apo- and fully ligated protein and values of k ex and p E extracted from CPMG measurements. A large error is noted for N53 that arises from the weak peak intensity for this residue. The lines indicated are the best fits from linear regression
In (8) values of (p E , k ex ) from CPMG measurements were used, with values of Δω obtained from direct measurement of shift differences. Notably, ΔR 1ρ is independent of R 1 and R 02 . As can be seen in Fig. 4, the correlation is high, providing further confidence in the methodology. It is worth emphasizing that cross-validation is possible even in cases where the signs of Δω are not known a priori, since the calculated values of |ΔR 1ρ | are derived solely from parameters that are available from CPMG relaxation dispersion experiments.
The experimental results obtained on the SH3 domain–ligand exchanging system presented here establish the robustness of the methodology, at least in this case. It is of interest, however, to explore the sensitivity of the approach to the full range of exchange parameters that might in general be encountered in studies of excited protein states. Figure 5 plots Δ values of (6) (T = 100 ms) as a function of (p E , k ex , |Δϖ|). For reference, a plot of Δ vs. |Δϖ| is presented in Fig. 5a using values of (p E , k ex ) = (2.5%, 300 s−1) that are germane for the Abp1p SH3 system, with a horizontal line at Δ = 4%, corresponding to a threshold above which differences in \( R_{1\rho }^{ \pm } \) values have been found to be reliable in this case. As p E increases values of Δ become larger so that it is easier to measure signs of shift differences, as expected (Fig. 5b). Further, as k ex increases, it becomes increasingly difficult to obtain the signs of small shift differences, but easier for large Δϖ values, Fig. 5c. Thus, the R 1ρ methodology is optimal for systems with large p E and relatively small k ex values. In cases where temperature dependent CPMG data is available it is worthwhile simulating similar curves to those in Fig. 5a to establish the best set of conditions to record experiments. It should be noted that values of Δ do depend on R 1 and R 02 , with transverse relaxation rates available from CPMG dispersion profiles recorded at high νCPMG. Here we have used values of R 1 = 1 s−1, R 02 (1HN) = 10 s−1, R 02 (15N) = 10 s−1, R 02 (1Hα) = 35 s−1 and R 02 (13Cα) = 35 s−1 (approximate values obtained for the A39V/N53P/V55L Fyn SH3 domain, 20°C, see below).
a Values of Δ (6, T = 100 ms) vs. |Δϖ| calculated for exchange parameters of the Abp1p-Ark1p system (k ex = 300 s−1, p E = 2.5%) for 1HN (red) and 15N (blue) nuclei. b Δ vs. \( |\Updelta \varpi_{{H^{N} }} | \) for k ex = 300 s−1 as a function of p E . c Δ vs.\( |\Updelta \varpi_{{H^{N} }} | \) for p E = 2.5% as a function of k ex . The horizontal line at Δ = 4% is the threshold below which it is not possible to obtain signs of chemical shift differences on the protein systems examined here
In addition to the 1HN R 1ρ scheme presented here and the corresponding 15N and 1Hα experiments published previously (Auer et al. 2009; Korzhnev et al. 2005), we have also developed an analogous sequence for obtaining the signs of 13Cα Δϖ values, Fig. 6. This scheme, along with the 1Hα version, is applied to a protein folding ‘reaction’ involving an on-pathway folding intermediate of the A39V/N53P/V55L Fyn SH3 domain (Neudecker et al. 2006), where the intermediate state (p E = 1.4%) exchanges with the folded (ground) conformer with a rate of k ex = 780 s−1 at 20°C. 1Hα and 13Cα \( R_{1\rho }^{ \pm } \) decay curves for E33 and T47 are shown in Fig. 7, illustrating the quality of data that can be obtained on this system. Of the 45 residues for which 13Cα dispersion profiles could be quantified, signs were obtained for 19 with values of \( \Updelta \varpi_{{C^{\alpha } }} \) down to 0.5 ppm. All the signs obtained are in agreement with those measured using the H(S/M)QC approach of Skrynnikov et al. (Skrynnikov et al. 2002). Finally, signs for \( \Updelta \varpi_{{H^{\alpha } }} \) were obtained for 19 residues with shift differences as low as 0.17 ppm (Auer et al. 2009).
Pulse sequence for measuring 13Cα off-resonance R 1ρ relaxation rates in proteins labeled with 13C at the Cα position. 1H and 13C carrier frequencies are placed initially on resonance (1Hα and 13Cα positions) for the cross-peak of interest; subsequently the 13C carrier is jumped to the position of the spin-lock field immediately prior to the pulse of phase ϕ3 and then back on-resonance prior to point e. All solid pulses have flip angles of 90° and are applied along the x-axis, unless indicated otherwise. Simultaneous 1H/13C spin-lock fields (140 Hz) are applied for durations of 1/JCH (between points a, b and again between points e, f) where JCH is the one-bond 1Hα–13Cα scalar coupling constant (≈140 Hz). Immediately after gradient 1 1H purge pulses are applied (17 kHz) for durations of 2 ms (x-axis) and 1 ms (y-axis) to eliminate the residual water signal. The delay τeq is set to 5 ms. 1H pulses of phases ϕ3/ϕ4 are applied with a flip angle θ such that tanθ = ω 1/|δ G |, using (δ G ,ω 1) optimized as described in the text. During the spin-lock period an on-resonance 1H continuous-wave decoupling field of 13.8 kHz is applied to eliminate scalar coupling modulations and cross-correlation effects between 1H–13C dipolar and 13C CSA interactions (Peng and Wagner 1992). 15N decoupling during the spin-lock and 13C decoupling during acquisition are achieved with WALTZ-16 fields (Shaka et al. 1983). The phase cycle is: ϕ1 = (y,−y), ϕ2 = 2(x), 2(−x), ϕ5 = 4(x), 4(−x), ϕ6 = x, 2(−x), x−x, 2(x), −x. For the spin-lock carrier upfield(downfield) of the ground state resonance ϕ3 = y(−y) and ϕ4 = −y(y) (on Varian spectrometers). Gradient strengths and durations are (ms,G/cm): G0 = (1,7.5), G1 = (0.5,10), G2 = (0.8,15), G3 = (0.6,−4), G4 = (0.2,−4)
Selected regions of 13Cα–1Hα HSQC spectra centered on residues E33 and T47 of the A39V/N53P/V55L Fyn SH3 domain, 20°C, 800 MHz along with \( R_{1\rho }^{ \pm } \) measured for 1Hα (orange, red) and 13Cα (purple, blue) nuclei. Optimized (|δ G /(2π)|,ν 1) values for measurements at 800 MHz calculated from (6, 7) along with k ex = 780 s−1, p E = 1.4% and the value of |Δϖ| from CPMG experiments are listed. The peak flanking the E33 line in the 1D spectrum is due to M−1 and D35; the separation is sufficiently large such that there is not a problem with quantification of the E33 peak. Other details are as in Fig. 2
Comparison of low power off-resonance R 1ρ and H(S/M)QC measurements for the determination of the sign of Δϖ
To date the most common approach for measuring the signs of 15N or 13C shift differences between exchanging states involves comparison of HSQC and HMQC data-sets recorded at a number of static magnetic fields (Skrynnikov et al. 2002). More specifically, Skrynnikov et al. (2002) have shown that the difference in peak positions (ppm) in HSQC spectra recorded at a pair of fields, \( B_{0}^{(i)} \) and \( B_{0}^{(ii)} \) \( ({B_{0}^{(i)} < B_{0}^{(ii)}}) \), is given by
with a shift (ppm) also noted between peaks in HSQC and HMQC data-sets recorded at a single magnetic field,
where \( \tilde{\xi }_{X} = \Updelta \varpi_{X} /k_{EG} \) and it is assumed that nuclei in states G and E have the same intrinsic transverse relaxation rates. It can be shown that \( \tilde{\sigma }_{X} \) and \( \Updelta \varpi_{X} \) have the same sign, as do \( \tilde{\Upomega }_{X} \) and \( \Updelta \varpi_{X} \) for the most part (but see below), facilitating extraction of the sign of Δϖ X (Skrynnikov et al. 2002). It is of considerable interest to compare the R 1ρ method with the more established methodology in cases were both are applicable. Figure 8a shows values of Δ (6) as a function of \( \Updelta \varpi_{{C^{\alpha } }} \) for a number of (p E , k ex ) values, including those measured for the A39V/N53P/V55L Fyn SH3 domain. The 4% threshold (horizontal line) above which meaningful differences in \( R_{1\rho }^{ \pm } \) can be measured is shown. It is worth noting that for exchange parameters of (p E , k ex ) = (1.4%, 780 s−1), signs are predicted to be measurable in cases where \( \Updelta \varpi_{{C^{\alpha } }} \)> 0.5 ppm, as observed experimentally. By means of comparison, plots of differences in peak positions in HSQC spectra recorded at 500 and 800 MHz, \( \tilde{\sigma }_{{C^{\alpha } }} \), for a number of (p E , k ex ) values are shown in Fig. 8b, where \( \tilde{\sigma }_{{C^{\alpha } }} =\) \( \varpi_{{C^{\alpha } }} (500{\text{ MHz)}}- \varpi_{{C^{\alpha } }} (800{\text{ MHz)}} \), while in Fig. 8c a contour plot of \( \tilde{\Upomega }_{{C^{\alpha } }} \) as a function of \( (\Updelta \varpi_{{C^{\alpha } }} ,\Updelta \varpi_{{H^{\alpha } }} ) \) is presented for (p E , k ex ) = (1.4%,780 s−1), 500 MHz. Shown also in Fig. 8b are horizontal lines at ±2 ppb, the experimentally determined lower limit for measuring accurate \( \tilde{\sigma }_{{C^{\alpha } }} \)values for the Fyn SH3 domain based on the reproducibility of ‘exchange-free’ peak positions in HSQC spectra. It is clear from a comparison of the plots, focusing on (p E , k ex ) = (1.4%,780 s−1), that measurements of R 1ρ and \( \tilde{\sigma } \) methods are sensitive to the sign of Δϖ only for relatively large chemical shift differences, on the order of greater than 0.5 ppm in this case. By contrast, the measurement of \( \tilde{\Upomega }_{{C^{\alpha } }} \) provides accurate sign information for much smaller \( \Updelta \varpi_{{C^{\alpha } }} \) values (≈0.14 ppm), although only if \( \Updelta \varpi_{{H^{\alpha } }} \) ≠ 0. An additional advantage with the \( \tilde{\Upomega }_{X} \) measurements is that because spectra recorded at a single magnetic field are compared the reproducibility of peak positions is very high; for the Fyn SH3 domain ‘exchange free peaks’ were within 0.5 ppb in HSQC/HMQC spectra recorded at the same field so that \( \tilde{\Upomega }_{X} \) values as low as 1 ppb are significant. A (small) disadvantage with the approach, however, is that there is a region in \( (\Updelta \varpi_{{C^{\alpha } }} ,\Updelta \varpi_{{H^{\alpha } }} ) \) space where negative values of \( \tilde{\Upomega }_{{C^{\alpha } }} \) correspond to positive \( \Updelta \varpi_{{C^{\alpha } }} \)and vice versa, indicated in Fig. 8c by shading. It should be noted that in this region values of \( \tilde{\Upomega }_{{C^{\alpha } }} \) are typically very small (<0.5 ppb in the example considered here), making it hard to extract reliable shift differences in any event. Often an indication that values of \( (\Updelta \varpi_{{C^{\alpha } }} ,\Updelta \varpi_{{H^{\alpha } }} ) \) lie in a ‘problem’ region is that opposite signs are obtained from measurement of \( \tilde{\sigma }_{{C^{\alpha } }} \) and \( \tilde{\Upomega }_{{C^{\alpha } }} \). However, so long as the magnitudes of \( \Updelta \varpi_{{C^{\alpha } }} \)and \( \Updelta \varpi_{{H^{\alpha } }} \)are known, along with the exchange parameters, it is possible to establish the reliability of the method through simulation (10).
Simulations of Δ (a, 6) and \( \tilde{\sigma }_{{C^{\alpha } }} \) (b, 9) as a function of \( \Updelta \varpi_{{C^{\alpha } }} \) for a number of (k ex [s−1]/p E [%]) values: black (780/1.4), orange (300/1.4), red (1,500/1.4), purple (780/3), blue (780/6). c Contour plot of \( \tilde{\Upomega }_{{{\text{C}}^{\alpha } }} \) as a function of \( (\Updelta \varpi_{{C^{\alpha } }} ,\Updelta \varpi_{{H^{\alpha } }} ) \) (contours are in ppb) for (p E , k ex ) = (1.4%, 780 s−1), 500 MHz. The gray shaded areas denote regions where negative values of \( \tilde{\Upomega }_{{{\text{C}}^{\alpha } }} \) correspond to positive \( \Updelta \varpi_{{{\text{C}}^{\alpha } }} \) and vice versa. d–f Corresponding simulations for k ex = 200 s−1 and p E = 1.4%. For |\( \Updelta \varpi_{{{\text{H}}^{\alpha } }} \)| > 0.3 ppm, the sign of \( \Updelta \varpi_{{{\text{C}}^{\alpha } }} \) can be obtained from a comparison of \( R_{1\rho }^{ \pm } \) rates (d), while the sign is likely to be difficult to obtain from the H(S/M)QC approach (e, f)
Of the 46 |Δϖ N| values that were quantified from fits of CPMG dispersion profiles recorded on the Abp1p-ligand exchanging system, signs have been obtained for 28 residues from measurement of \( \tilde{\sigma }_{N} \) and \( \tilde{\Upomega }_{N} \). The signs of chemical shift differences, |Δϖ N|, as low as 0.06 ppm could be determined. By comparison, 25 signs (|Δϖ N| > 0.15 ppm) were obtained from the R 1ρ method. Notably, 4 signs could be obtained only using the R 1ρ approach since for these residues \( \Updelta \varpi_{{H^{N} }} \) ≈ 0, preventing quantitation from a comparison of HSQC and HMQC data-sets, while 7 signs were only available from the H(S/M)QC methodology (see supporting information). Similar trends are noted from 13Cα measurements on the A39V/N53P/V55L Fyn SH3 domain. Out of a total of 45 |\( \Updelta \varpi_{{C^{\alpha } }} \)| values quantified by CPMG relaxation dispersion experiments, signs were obtained for 34 \( \Updelta \varpi_{{C^{\alpha } }} \) values larger than 0.3 ppm from a combined \( \tilde{\sigma }_{{C^{\alpha } }} \), \( \tilde{\Upomega }_{{C^{\alpha } }} \) analysis, while signs were available for 19 residues with \( \Updelta \varpi_{{C^{\alpha } }} \) > 0.5 ppm based on the R 1ρ method (see supporting information).
The experimental results establish that in general the H(S/M)QC method is preferred over the R 1ρ approach, at least for obtaining signs of chemical shift differences of heteronuclei. However, the methods are complementary. For example, simulations establish that over a reasonable range of k ex values that are normally relevant for systems studied by CPMG dispersion experiments \( \tilde{\Upomega }_{X} \) values increase with exchange rate (see supporting information), while the sensitivity of the off-resonance spin-lock method decreases, at least for relatively small chemical shift differences (Fig. 8a). By contrast, a strength of the R 1ρ approach is that the differences in \( R_{1\rho }^{ \pm } \) values increase with Δϖ, while both \( \tilde{\sigma }_{{}} \)and \( \tilde{\Upomega }_{{}} \) values have definite maxima, depending on the exchange parameters (Fig. 8a–c). Figure 8d–f presents an example of an exchanging system where the sign information is forthcoming from R 1ρ measurements but potentially not from H(S/M)QC spectra. Here (p E , k ex ) = (1.4%,200 s−1) and so long as |\( \Updelta \varpi_{{C^{\alpha } }} \)| > 0.3 ppm, the sign of \( \Updelta \varpi_{{C^{\alpha } }} \) can be obtained from a comparison of \( R_{1\rho }^{ \pm } \) rates, Fig. 8d. The very small \( \tilde{\sigma }_{{C^{\alpha } }} \)values obtained in this case (<1 ppb), Fig. 8e, preclude sign extraction from HSQC spectra and likewise the small values of \( \tilde{\Upomega }_{{C^{\alpha } }} \) < 2ppb could very well complicate the determination of sign from this method as well, Fig. 8f. Certainly for |\( \Updelta \varpi_{{C^{\alpha } }} \)| > 1 ppm the sign information is not available from any combination of HMQC and HSQC data-sets. It is also noteworthy that while the H(S/M)QC approach is the first choice for 15N and 13C nuclei, the R 1ρ method is now commonly used in our laboratory for extracting signs of 1HN and 1Hα shift differences. Previously, we have compared double- and zero-quantum 15N–1HN relaxation dispersion profiles to obtain the relative signs of \( \Updelta \varpi_{{H^{N} }} \)and \( \Updelta \varpi_{N} \) from which the sign of \( \Updelta \varpi_{{H^{N} }} \) can be obtained so long as (1) the sign of \( \Updelta \varpi_{N} \) is known and (2) that \( \Updelta \varpi_{N} \) is sufficiently large that differences in the profiles are present in the first place (Orekhov et al. 2004). Extraction of robust sign information for \( \Updelta \varpi_{{H^{N} }} \)from \( R_{1\rho }^{ \pm } \) measurements is not dependent on the value of \( \Updelta \varpi_{N} \). Finally, the spin-lock approach remains the most convenient method for measuring signs of \( \Updelta \varpi_{{H^{\alpha } }} \), especially since the same sample used to record 1Hα CPMG dispersion profiles can be used in this case as well (Auer et al. 2009).
In summary, we have presented pulse schemes for the measurement of 1HN and 13Cα \( R_{1\rho }^{ \pm } \) relaxation rates, complementing schemes previously published for 15N and 1Hα (Auer et al. 2009; Korzhnev et al. 2005). The methodology is shown to be robust, although for 15N and 13Cα at least, fewer signs are available than from the H(S/M)QC approach (Skrynnikov et al. 2002). In addition, the H(S/M)QC method is less time consuming and the information is available in the form of 2D data-sets, as opposed to the 1D spectra recorded here, that leads to increased numbers of residues that can be quantified. We therefore suggest that sign determination for 15N and 13Cα be carried out using \( \tilde{\sigma }_{X} \) and \( \tilde{\Upomega }_{X} \) values from HSQC and HMQC data-sets recorded at no fewer than a pair of static magnetic fields. In some cases where data might be ambiguous or for exchanging systems for which the sign information is difficult to extract from the H(S/M)QC method (low k ex and p E ), \( R_{1\rho }^{ \pm } \)measurements can readily be performed. For measurement of signs of 1HN and 1Hα shift differences the R 1ρ approach has been very useful for the small protein systems that we are currently studying. It is clear that chemical shifts will play a critical role in structural studies of invisible, excited states. The development of a number of robust approaches to extract such information is thus an important goal as a prerequisite for the in-depth characterization of these biologically important conformers.
Supporting material
Tables of signed Δϖ values for the Abp1p-Ark1p exchanging system (15N, 1HN) and the A39V/N53P/V55L Fyn SH3 domain (13Cα, 1Hα). Figures of contour plots of \( \tilde{\Upomega }_{X} \) for different values of exchange parameters and static magnetic fields. Pulse sequence code for measurement of 1HN and 13Cα R 1ρ rates.
References
Auer R, Neudecker P, Muhandiram DR, Lundstrom P, Hansen DF, Konrat R, Kay LE (2009) Measuring the signs of 1H(alpha) chemical shift differences between ground and excited protein states by off-resonance spin-lock R(1rho) NMR spectroscopy. J Am Chem Soc 131:10832–10833
Boehr DD, McElheny D, Dyson HJ, Wright PE (2006) The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313:1638–1642
Boulat B, Bodenhausen G (1993) Measurement of proton relaxation rates in proteins. J Biomol NMR 3:335–348
Carr HY, Purcell EM (1954) Effects of diffusion on free precession in nuclear magnetic resonance experiments. Phys Rev 54:630–638
Chiarparin E, Pelupessy P, Bodenhausen G (1998) Selective cross-polarization in solution state NMR. Mol Phys 95:767–769
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Frederick KK, Marlow MS, Valentine KG, Wand AJ (2007) Conformational entropy in molecular recognition by proteins. Nature 448:325–329
Goto NK, Kay LE (2000) New developments in isotope labeling strategies for protein solution NMR spectroscopy. Curr Opin Struct Biol 10:585–592
Hansen DF, Vallurupalli P, Kay LE (2008a) An improved 15N relaxation dispersion experiment for the measurement of millisecond time-scale dynamics in proteins. J Phys Chem B 112:5898–5904
Hansen DF, Vallurupalli P, Lundstrom P, Neudecker P, Kay LE (2008b) Probing chemical shifts of invisible states of proteins with relaxation dispersion NMR spectroscopy: how well can we do? J Am Chem Soc 130:2667–2675
Hansen AL, Nikolova EN, Casiano-Negroni A, Al-Hashimi HM (2009) Extending the range of microsecond-to-millisecond chemical exchange detected in labeled and unlabeled nucleic acids by selective carbon R(1rho) NMR spectroscopy. J Am Chem Soc 131:3818–3819
Haynes J, Garcia B, Stollar EJ, Rath A, Andrews BJ, Davidson AR (2007) The biologically relevant targets and binding affinity requirements for the function of the yeast actin-binding protein 1 SRC-homology 3 domain vary with genetic context. Genetics 176:193–208
Henzler-Wildman K, Kern D (2007) Dynamic personalities of proteins. Nature 450:964–972
Igumenova TI, Frederick KK, Wand AJ (2006) Characterization of the fast dynamics of protein amino acid side chains using NMR relaxation in solution. Chem Rev 106:1672–1699
Ishima R, Torchia DA (2000) Protein dynamics from NMR. Nat Struct Biol 7:740–743
Ishima R, Torchia D (2003) Extending the range of amide proton relaxation dispersion experiments in proteins using a constant-time relaxation-compensated CPMG approach. J Biomol NMR 25:243–248
Ishima R, Baber J, Louis JM, Torchia DA (2004) Carbonyl carbon transverse relaxation dispersion measurements and ms-micros timescale motion in a protein hydrogen bond network. J Biomol NMR 29:187–198
Johnson BA, Blevins RA (1994) NMRView: a computer program for the visualization and analysis of NMR data. J Biomol NMR 4:603–614
Kainosho M, Torizawa T, Iwashita Y, Terauchi T, Mei Ono A, Guntert P (2006) Optimal isotope labelling for NMR protein structure determinations. Nature 440:52–57
Kay LE, Muhandiram DR, Wolf G, Shoelson SE, Forman-Kay JD (1998) Correlation between binding and dynamics at SH2 domain interfaces. Nat Struct Biol 5:156–163
Korzhnev DM, Orekhov VY, Dahlquist FW, Kay LE (2003) Off-resonance R1rho relaxation outside of the fast exchange limit: an experimental study of a cavity mutant of T4 lysozyme. J Biomol NMR 26:39–48
Korzhnev DM, Salvatella X, Vendruscolo M, Di Nardo AA, Davidson AR, Dobson CM, Kay LE (2004) Low-populated folding intermediates of Fyn SH3 characterized by relaxation dispersion NMR. Nature 430:586–590
Korzhnev DM, Orekhov VY, Kay LE (2005) Off-resonance R(1rho) NMR studies of exchange dynamics in proteins with low spin-lock fields: an application to a Fyn SH3 domain. J Am Chem Soc 127:713–721
Korzhnev DM, Bezsonova I, Evanics F, Taulier N, Zhou Z, Bai Y, Chalikian TV, Prosser RS, Kay LE (2006) Probing the transition state ensemble of a protein folding reaction by pressure-dependent NMR relaxation dispersion. J Am Chem Soc 128:5262–5269
LeMaster DM (1999) NMR relaxation order parameter analysis of the dynamics of protein side chains. J Am Chem Soc 121:1726–1742
Loria JP, Rance M, Palmer AG (1999) A relaxation compensated CPMG sequence for characterizing chemical exchange. J Am Chem Soc 121:2331–2332
Lundstrom P, Teilum K, Carstensen T, Bezsonova I, Wiesner S, Hansen DF, Religa TL, Akke M, Kay LE (2007a) Fractional 13C enrichment of isolated carbons using [1–13C]- or [2–13C]-glucose facilitates the accurate measurement of dynamics at backbone Calpha and side-chain methyl positions in proteins. J Biomol NMR 38:199–212
Lundstrom P, Vallurupalli P, Religa TL, Dahlquist FW, Kay LE (2007b) A single-quantum methyl 13C-relaxation dispersion experiment with improved sensitivity. J Biomol NMR 38:79–88
Lundstrom P, Hansen DF, Kay LE (2008) Measurement of carbonyl chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy: comparison between uniformly and selectively (13)C labeled samples. J Biomol NMR 42:35–47
Lundstrom P, Hansen DF, Vallurupalli P, Kay LE (2009a) Accurate measurement of alpha proton chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy. J Am Chem Soc 131:1915–1926
Lundstrom P, Lin H, Kay LE (2009b) Measuring 13Cb chemical shifts of invisible excited states in proteins by relaxation dispersion NMR spectroscopy. J Biomol NMR 44:139–145
Massi F, Johnson E, Wang C, Rance M, Palmer AG 3rd (2004) NMR R1 rho rotating-frame relaxation with weak radio frequency fields. J Am Chem Soc 126:2247–2256
Meiboom S, Gill D (1958) Modified spin-echo method for measuring nuclear magnetic relaxation times. Rev Sci Instrum 29:688–691
Mittermaier A, Kay LE (2006) New tools provide new insights in NMR studies of protein dynamics. Science 312:224–228
Neudecker P, Zarrine-Afsar A, Choy WY, Muhandiram DR, Davidson AR, Kay LE (2006) Identification of a collapsed intermediate with non-native long-range interactions on the folding pathway of a pair of Fyn SH3 domain mutants by NMR relaxation dispersion spectroscopy. J Mol Biol 363:958–976
Orekhov VY, Korzhnev DM, Kay LE (2004) Double- and zero-quantum NMR relaxation dispersion experiments sampling millisecond time scale dynamics in proteins. J Am Chem Soc 126:1886–1891
Palmer AG, Williams J, McDermott A (1996) Nuclear magnetic resonance studies of biopolymer dynamics. J Phys Chem 100:13293–13310
Palmer AG, Kroenke CD, Loria JP (2001) NMR methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Palmer AG, Grey MJ, Wang C (2005) Solution NMR spin relaxation methods for characterizing chemical exchange in high-molecular-weight systems. Methods Enzymol 394:430–465
Pelupessy P, Chiarparin E (2000) Hartmann-Hahn polarization transfer in liquids: an ideal tool for selective experiments. Concepts Magn Reson 12:103–124
Peng JW, Wagner G (1992) Mapping of spectral density functions using heteronuclear NMR relaxation measurements. J Magn Reson 98:308–332
Piotto M, Saudek V, Sklenar V (1992) Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J Biomol NMR 2:661–665
Popovych N, Sun S, Ebright RH, Kalodimos CG (2006) Dynamically driven protein allostery. Nat Struct Mol Biol 13:831–838
Shaka AJ, Keeler J, Frenkiel T, Freeman R (1983) An improved sequence for broadband decoupling: WALTZ-16. J Magn Reson 52:335–338
Skrynnikov NR, Mulder FAA, Hon B, Dahlquist FW, Kay LE (2001) Probing slow time scale dynamics at methyl-containing side chains in proteins by relaxation dispersion NMR measurements: application to methionine residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:4556–4566
Skrynnikov NR, Dahlquist FW, Kay LE (2002) Reconstructing NMR spectra of “invisible” excited protein states using HSQC and HMQC experiments. J Am Chem Soc 124:12352–12360
Sugase K, Dyson HJ, Wright PE (2007) Mechanism of coupled folding and binding of an intrinsically disordered protein. Nature 447:1021–1024
Tollinger M, Skrynnikov NR, Mulder FAA, Forman-Kay JD, Kay LE (2001) Slow dynamics in folded and unfolded states of an SH3 domain. J Am Chem Soc 123:11341–11352
Trott O, Palmer AG 3rd (2002) R1rho relaxation outside of the fast-exchange limit. J Magn Reson 154:157–160
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172
Vallurupalli P, Hansen DF, Stollar EJ, Meirovitch E, Kay LE (2007) Measurement of bond vector orientations in invisible excited states of proteins. Proc Natl Acad Sci USA 104:18473–18477
Vallurupalli P, Hansen DF, Kay LE (2008) Structures of invisible, excited protein states by relaxation dispersion NMR spectroscopy. Proc Natl Acad Sci U S A 105:11766–11771
van Ingen H, Vuister GW, Wijmenga S, Tessari M (2006) CEESY: characterizing the conformation of unobservable protein states. J Am Chem Soc 128:3856–3857
Zar ZH (1984) Biostatistical analysis. Prentice-Hall, Englewood Cliffs
Zhang Q, Sun X, Watt ED, Al-Hashimi HM (2006) Resolving the motional modes that code for RNA adaptation. Science 311:653–656
Acknowledgments
R.A. is a recipient of a DOC-fFORTE-fellowship of the Austrian Academy of Sciences. P.N. and D.F.H. are recipients of postdoctoral fellowships from the Canadian Institutes of Health Research (CIHR). This work was supported by a grant from the CIHR. L.E.K. holds a Canada Research Chair in Biochemistry.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Auer, R., Hansen, D.F., Neudecker, P. et al. Measurement of signs of chemical shift differences between ground and excited protein states: a comparison between H(S/M)QC and R 1ρ methods. J Biomol NMR 46, 205–216 (2010). https://doi.org/10.1007/s10858-009-9394-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-009-9394-z