Abstract
We present the performance of blind predictions of water—cyclohexane distribution coefficients for 53 drug-like compounds in the SAMPL5 challenge by three methods currently in use within our group. Two of them utilize QMPFF3 and ARROW, polarizable force-fields of varying complexity, and the third uses the General Amber Force-Field (GAFF). The polarizable FF’s are implemented in an in-house MD package, Arbalest. We find that when we had time to parametrize the functional groups with care (batch 0), the polarizable force-fields outperformed the non-polarizable one. Conversely, on the full set of 53 compounds, GAFF performed better than both QMPFF3 and ARROW. We also describe the torsion-restrain method we used to improve sampling of molecular conformational space and thus the overall accuracy of prediction. The SAMPL5 challenge highlighted several drawbacks of our force-fields, such as our significant systematic over-estimation of hydrophobic interactions, specifically for alkanes and aromatic rings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The prevailing opinion of the computational community is that polarizable force-fields [1–19] are a necessary direction in the development of molecular modeling [20–23]. At the same time, pairwise non-polarizable force-fields [24–31], the old workhorses of the field, still offer the best and most consistent performance. The advantage is certainly due to the large disparity in development and testing time, but likely also to one or more fundamental shortcomings. As InterX Inc. is developing several polarizable FF’s [8], the distribution coefficient component of SAMPL5 was a terrific opportunity to gauge our progress. We would like to thank the organizers (and the participants) of the challenge for a fabulous and extremely useful cooperative scientific experiment.
Previous four SAMPL challenges since 2008 requested blind prediction of experimental hydration free energies [32–35]. While this test is critical for validation of force-fields and molecular simulation methodologies in water, describing ligands’ interaction with alkanes is an equally important test of molecular modeling. SAMPL5 [36] expanded the hydration challenge by asking participants to blindly predict distribution coefficients (difference in free energy of solvation) between cyclohexane and water for 53 drug-like molecules [37]. Distribution coefficients are not only an excellent metric of the capability and accuracy of modeling, they are also valuable in themselves as they are associated with important pharmacological properties, e.g. drug uptake in lipid bilayers.
Intermolecular potentials (QMPFF3 and ARROW)
Because the main goal of our participation was to compare the various force-fields currently employed by our group, we shall start this report by briefly describing them. A detailed description of the Quantum Mechanical Polarizable Force field (QMPFF3) including the functional form has been provided in various articles [6–8, 38]. The Accurate Representation of Angstrom World (ARROW) variant of QMPFF3 will be fully described in a future publication.
The total energy consists of four components: Electrostatics, Exchange, Dispersion and Induction (henceforth denoted as ES, EX, DS, and IND). The nuclei are represented by point charges. The ES and EX electron–electron interactions are multipolar up to L = 2 (quadrupole) and are represented by interaction of diffuse clouds. The multipolar expansion is limited to the reference frame provided by the bond(s). The ES penetration effect is modelled by cloud–cloud penetration, and the EX interaction is proportional to cloud–cloud overlap which is exponentially decreasing with distance between atoms. DS is modeled by Tang-Toennies functions of power R−6 and R−8 terms and is damped. Electrostatic induction is represented by a shift of diffuse dipoles and includes an exchange correction; and the internal energetic cost of polarization is modeled by an an-harmonic anisotropic spring. The 1–4 interactions are accounted for in full strength. The functional forms of bonded interactions are identical to those in the Merck Molecular Force Field (MMFF94) [39].
The ARROW variant carries a slightly more complex functional form than QMPFF3; namely a more nuanced description of multipolar atomic shapes, virtual bonds for terminal atoms, and a charge delocalization interaction [40]. In parametrization, ARROW relies on different quantum mechanical framework (largely DFT-SAPT [41–43] because of its natural decomposition of energies that corresponds to our ES, EX, DS, IND partitioning and its implementation in MOLPRO [44]), and has an expanded parameter set. Figure 1 visualizes the superior representation by ARROW of the electrostatic component of energy compared with non-polarizable force-fields for 1,3,5-Triazine.
An illustration of the ARROW Electrostatic Potential (ESP) fitting for the 1,3,5-triazine molecule, that is part of SAMPL5 molecule 27. (Left) QM (MP2/aqz) ESP map (center) difference between GAFF (based on AM1-BCC charges) and QM ESP maps (right) difference between ARROW and QM ESP maps. 1,3,5-Triazine is a neutral molecule without a net dipole moment, but it has a quadrupole moment with main components in QM [− 1.75, −1.75, 3.5]·0.1 Å2 q. The GAFF representation has an almost opposite quadrupole moment [1.54, 1.54, −3.08]·0.1 Å2 q. Introduction of explicit quadrupoles in ARROW permits a reproduction of the ESP map with almost no error. X,Y-axes on plots are in Angstroms, color bar units are in kcal/mol/q. All maps are plotted at 2 van derWaals radii surface
The determination of force field parameters is still in flux and will be fully described in a future publication as well; we will limit ourselves to a few general comments here. Our guiding philosophy is to derive the force field fully from ab initio calculations, which is advantageous when describing SAMPL5 drug-like molecules for which very little to no experimental data exists. We used a variety of different QM data to fit the non-bonded interactions: monomer properties (e.g. electrostatic potential maps (ESP) see e.g. Figure 1), dipoles, quadrupoles, polarizability tensors and interaction of molecules with charges. We also employed a large collection of homogeneous dimers, a smaller set of heterogeneous dimers, and a still smaller set of multimers.
For SAMPL5 we partitioned the candidate molecules into roughly 50 fragments of different functional groups using a procedure generously described as ‘human intelligence’; a fuller investigation of transferability and separability of functional groups is planned for the near future. In the interest of speed and time we substituted propane for cyclohexane in QM calculations; the consequences of this are under investigation. Most of the training dimers were generated from fragment-water and fragment-propane MD simulations at normal conditions (T = 298 K, P = 1 atm). The resulting conformations were then pruned by clustering close relatives, leaving approximately one to two hundred dimers in each collection. To better fit the repulsive wall of the potential, we took ~ 30 % of closest MD dimers and contracted them towards each other, thus making an additional 4 dimers per each closest dimer. For all clustered fragment—water H2O and fragment—propane C3H8 systems we calculated the energy and its components via DFT-SAPT at aug-cc-pVTZ and aug-cc-pVQZ level and extrapolated the dispersion interaction to the Complete Basis Set (CBS) limit [45]. Our total interaction energy at CBS level therefore consists of all interaction parts at aug-cc-pVQZ level plus dispersion at estimated CBS level. In addition to extrapolation to CBS level we corrected our total CBS energy by the difference between CCSD(T)/aug-cc-pVDZ and DFT-SAPT/aug-cc-pVDZ to provide better QM accuracy.
The bonded parameters were benchmarked at the df-MP2/aTZ level in MOLPRO [44] using step-wise displacements from equilibrium for bond-stretch, angle-bend, stretch-bend and dihedrals.
Simulation details
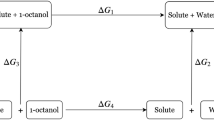
The partition coefficients were estimated from the difference in solvation free energies of the solute in the neutral state in water and cyclohexane at infinite dilution. For species that may undergo ionization in aqueous phase, we applied a pKa correction:
where log D is the distribution coefficient, \(\Delta G_{solvation}\) is the free energy of solvation of molecule in cyclohexane, \(\Delta G_{hydration}\) is the free energy of solvation of molecule in water; both units in kcal/mol. R is the Universal Gas constant in kcal/mol/K and T is temperature = 298 K. The pKa values were determined from publicly available website: https://epoch.uky.edu/ace/public/pKa.jsp. We do not expect accuracy of this pKa estimator be better than 1 pKa units.
Before running simulations we analyzed potential tautomers in water solution for all SAMPL molecules with B3LYP/atz method with COSMO implicit solvent dielectric constant ε = 80. All analyzed tautomers are presented in supplementary information Table S3. We found only 1 tautomer for SAMPL50 molecule that has a structure different that organizers suggested (see Fig. S2), and used this tautomeric version for this molecule. A priori we were not able to judge the accuracy of experimental data (i.e. water dragging effects, dimerization of solute in solvents, etc.) therefore we assume them to be negligible.
Solute, water, and cyclohexane were described by the polarizable non-bonded parameters and valence parameters of the QMPFF3 and ARROW force-fields described in the previous section, as well as those of the General Amber Force-Field. For QMPFF3 submissions we have had some parameters ready before SAMPL assessment, which in turn was based but not equal to published parameters [7]. In addition, during SAMPL assessment for QMPFF3 we derived a special set of parameters for bromine, cyano group, sulfone derivatives, thiophene, oxazoles, etc. For ARROW parameters we have parameterized a significant portion of most frequently occurring functional groups (such as aliphatic, aromatic carbons, ethers, esters, aromatic nitrogen, etc.), but was not able to prepare parameters for all of functional groups. Because ARROW is the superset of QMPFF3 we have put QMPFF3 distribution coefficients instead of missing ARROW numbers, that is contribute to approximately to 30 % of ARROW submission.
Because our polarizable runs were rather short (500 ps) we did not expect an adequate sampling of the conformational states of the solute. To compensate we chose the most energetically favorable structures obtained from the much longer 50 ns of isothermal-isobaric ensemble simulations at 298 K and 1 atm in cyclohexane using Generalized Amber Force Field (GAFF) (parameters made available from the SAMPL5 website) as the starting structures for the SAMPL5 molecules. We placed each minimum energy configuration in a solvated box with a single solute molecule in each solvent. We constructed the unit box to be at least 40 Å per side, which required at least 2124 molecules of water, and 352 of cyclohexane. For water we ran isothermal-isobaric ensemble (NPT) molecular dynamics simulations, with temperature control provided by a six-chain 0.5 ps relaxation time Nose–Hoover thermostat [46, 47] at T = 298 K, and pressure control by a Berendsen thermostat [48] at 1 atm reference pressure with a time constant for relaxation of 0.5 ps and compressibility set at 0.45 GPa−1. For two largest SAMPL5 molecules—83 and 92—we used a larger simulation cell of 45 × 45 × 45 Å3 to avoid interactions with solute in periodic boundary regions. Cyclohexane simulations were identical to water, except the compressibility of cyclohexane was set to 0.114 GPa−1 which resulted in a liquid density of 0.75 g/cc, in good agreement with experimental density of 0.77 g/cc. For computational efficiency all interactions were truncated by a group-based cutoff at 13 Å. We used in-house tools (Arbalest code suite) and various Octave/Matlab scripts to setup the initial configuration and subsequent post-processing of generated data. The difference in free energy between the two states of a system was obtained via the coupling parameter approach of thermodynamic integration. The solute was gradually annihilated through 10 intermediate lambda states, and the interactions were switched off closely following the mutation protocol for protein–ligand complexes [49]. We simulated each solvated system separately at each lambda value by (1) first minimizing the system in Arbalest using the steepest descent algorithm, (2) running a 500 ps MD production phase at each lambda value using a Berendsen barostat and Nose–Hoover thermostat with relaxation times of 0.5 ps and 1 ps, respectively. We typically discarded the first 50 ps of simulation to achieve convergent equilibration. In cases where convergence was suspect longer simulation 1 ns were employed, it was done particularly for the following molecules: 7, 13, 17, 21, 46, 58, 63, 65, 83, 84, 88, 92. We used cubic spline interpolation for smooth integration of dH/dL values to obtain the final solvation energy (and hence the predicted partition coefficients) using Eq. 1. The statistical errors (SEM) of the run was determined not by multiple runs, but by analysis of correlation times, such description found in the supplemental information of the cited article [49].
Accounting for torsional flexibility in Log D calculations for GAFF-TR model
In addition to QMPFF3 and ARROW based calculations (Table 1) we ran a baseline set of calculations with GAFF using GROMACS [50]. Most of the drug-like compounds in SAMPL5 set contain multiple rotatable bonds, frequently with high torsional energy barriers, therefore making the equilibration of torsional degrees of freedom in TI solvation free energy calculations slow. Calculations with non-polarizable GAFF force field are almost 2 orders of magnitude faster than those with QMPFF3 and ARROW that allowed us to employ better convergence techniques and help to choose better initial geometries for more expensive calculations with polarizable force fields.
We performed log D calculations of SAMPL5 molecules with GAFF using starting geometries given by the organizers (denoted “init geom” in Table 2). We also ran calculations starting with the “most probable” ligand conformations in cyclohexane solvent (denoted “opt geom” in Table 2). The “most probable” ligand conformations were found as follows: (1) Torsional values \(\varTheta_{i}^{0}\) with highest probability density were determined from long (50 ns) MD simulations of ligands in cyclohexane (2) MD snapshots of ligands with smallest RMSD of torsions from \(\varTheta_{i}^{0}\) values were selected as the “most probable” conformations. Calculations with initial starting geometries showed large deviations in computed log D values from calculations started with most probable conformations for some of the compounds e.g. SAMPL5_017 compound (which have an internal hydrogen bond in the optimal geometry) and SAMPL5_020 compound (which has a flipped HNCN torsion in the optimal geometry compared to the initial geometry).
To ensure adequate sampling of the torsional degrees of freedom for GAFF calculations we also employed the following methodology. Thermodynamic integration calculations of solvation free energies of test molecules in water and cyclohexane were performed with applied torsional restraints with a functional form:
where θ i i-th torsional angle of the molecule, \(\theta_{i}^{0}\)—restrained value of the i-th torsion (the most probable torsional value in MD simulations of the ligand in cyclohexane). A rather rigid torsional force constant K = 200 kJ/mol/rad2 ensures that the torsional angle θ i stays within interval of 2Δ around \(\theta_{i}^{0}\) value (Δ = 30 deg). 16 λ points were used in solvation free energy TI calculations (first 6 λ points were used to switch off coulomb interactions and 10 λ points to switch off VdW interactions). For each λ-state we had run 500 ps trajectories. As torsional angles of the ligands were restrained in a relatively narrow range of values with no significant torsional barriers in these intervals 500 ps trajectory were deemed sufficient for convergence.
However, to compute free energy of transfer of the molecule from water to cyclohexane we need to account for the free energy cost of applying restraints (3) in water and cyclohexane:
Here \(\Delta G_{wat \to cxn}^{restr} = \Delta G_{wat \to vac}^{restr} - \Delta G_{cxn \to vac}^{restr}\) is the “restrained” free energy of transfer of the molecule from water to cyclohexane, computed as difference of de-solvation free energies of the molecule from water \(\left( {\Delta G_{wat \to vac}^{restr} } \right)\) and from cyclohexane (\(\Delta G_{cxn \to vac}^{restr}\)) using thermodynamic integration method with torsional restraints (3) applied.
We obtained the free energy cost of applying restraints in water \(\Delta G_{wat}^{unrestr \to restr}\) and in cyclohexane \(\Delta G_{cxn}^{unrestr \to restr}\) by two methods:
(1) running long (50 ns) unrestrained MD trajectories of the ligand in water and cyclohexane
where \(P_{restr wat}\) and \(P_{restr cxn}\) – probabilities of all torsions of the ligand to be in the “restrained” space (\(\uptheta_{i}^{0} - \Delta \le\uptheta_{i} \le\uptheta_{i}^{0} + \Delta\)) in the unrestrained MD calculations in water and cyclohexane correspondingly. \(P_{restr wat}\) and \(P_{restr cxn}\) were computed from the ratio of MD snapshots having all torsions satisfying (\(\theta_{i}^{0} - {{\Delta }} \le \theta_{i} \le \theta_{i}^{0} + {{\Delta }}\)) condition.
(2) computing free energy profiles of torsional degrees of freedom using WHAM/Umbrella Sampling. Umbrella sampling simulations (100 ps per umbrella) were run with harmonic restraints applied to individual torsions with a harmonic constant of 100 kcal/rad*rad and equilibrium positions of restraining potential separated by 3 degrees for neighboring umbrellas. Torsional free energy profiles G(Θi) were obtained applying WHAM technique to torsional distributions obtained in Umbrella simulations. Corrections for torsional space restraining were computed using (5) with \(P_{restr wat}\) and \(P_{restr cxn}\) computed from torsional free energy profiles:
For majority of the ligands restraining corrections \({{\Delta }}G_{wat}^{unrestr \to restr} - {{\Delta }}G_{cxn}^{unrestr \to restr}\) computed by two methods were close. In our submitted GAFF Log D results we chose restraining correction method dependent on the ligand structure. Method 1 based on long molecular dynamics was considered more accurate for most of the compounds as it takes into account correlation in the dynamics of different torsions in the molecule. Method 2 based on WHAM/Umbrella sampling calculations [51, 52] was assumed more accurate for ligands with very large torsional barriers (such as SAMPLE_048 compound) that were not sampled during 50 ns MD simulations.
The technique described here improved theoretical predictions for distributions coefficients when compared to unrestricted calculations. We will use the GAFF-TR shorthand for this method.
Results
Validation
Prior to running the SAMPL5 partition challenge molecules we tested the existing version of the simpler polarizable FF, QMPFF3 [7, 8] on how well it predicts the partition coefficient for neutral amino-acid analogues: methane, propane, isobutane, methylimidazole, methylindole, p-cresol, toluene, ethanol, methanol, acetamide, propionamide, butylamide, acetic acid, propionic acid, methanethiol and methyl-ethylsulfide. The results are shown in Fig. 2, along with the corresponding data for GROMOS96 [53] and OPLS-AA [54] in comparison to experiment [55]. QMPFF3 performance here is very satisfactory (mean absolute error (MAE) of the deviation from experimental values equaling to 1.08) especially considering that the FF contains practically no adjustments to experimental data. For OPLS-AA, the MAE of 0.82 is the lowest in comparison to the other two force fields. While good, Fig. 2 also suggests that QMPFF3 parameters have a systematic tendency to be overly hydrophobic, especially for alkyl side chains. The origin of this systematic shift is not clear to us at the time of this writing. In an attempt to devise an ad-hoc fix for the SAMPL5 challenge, we submitted additional sets of predictions which include hydrophobic correction for alkane groups with 0.37 logD units per CH2 groups and 2.28 logD units per phenyl group (submission numbers 58 and 65 with pKa correction and without, correspondingly, see Table 3). Experimental data shows that our hydrophobic correction did not improve overall result. We did not validate ARROW parameters alongside QMPFF3 as they were not yet available.
Blind prediction
Moving on to the blind prediction, the overall results for all three of our approaches (Fig. 3, shown along with the overall winner, COSMO-RS [56]) look significantly worse than the validation in Fig. 2. The predictions are more scattered and show a systematic error over the whole set.
Comparison of our predictions for water-cyclohexane distribution coefficient based on the QMPFF3-pKa, ARROW-pKa, and GAFF-TR to experiment for 13 (Batch 0) SAMPL5 molecules. Also shown for comparison are the predictions for COSMO-RS. The correlation coefficient R for QMPFF3-pKa, ARROW-pKa, GAFF-TR and COSMO-RS are 0.88, 0.86, 0.79, and 0.9 respectively
Because our methods are still in development, the amount of work we ended up doing was likely significantly more than that of the average participant of SAMPL. We had to produce several needed parameter types for QMPFF3, the full set of parameters for ARROW, as well as many other tasks. Consequently, we spent the most time and care on batch 0 as it was put forth as a small but representative subset of the total challenge. The predictions of our three methods along with COSMO-RS for this representative and required set of 13 compounds are in Fig. 3. For QMPFF3-pKa, ARROW-pKa and GAFF-TR the MAEs are 1.94, 2.17 and 1.85 respectively; and the Kendall’s τ for the methods are 0.857, 0.875, 0.828 (See Table 3). The purpose is, of course, the journey, yet we are not satisfied with the results.
Moving on from absolute performance, we were curious to gauge how our methods compare to each other and also to those of other participants. The predicted values’ range is almost double that of the experimental ones, which suggests a systematic (slope) error; released results show that others’ MD methods suffer from the same bias as well. Whenever this occurs we prefer to consider relational measures such as Kendall’s τ rather than only the absolute ones, such as AUE or MAE. Different errors metrics for batch 0 and total set is summarized in Table 3. Based on τ, the ARROW-pKa submission is better than QMPFF3-pKa which, in turn, is better than GAFF-TR. Additionally, the ARROW-pKa submission is the best MD-based prediction method in this set if judged by Kendall’s τ; only QM-based methods do better.
For the full set of 53 molecules the picture is roughly similar but worse (Fig. 4). The numbers are MAE of 3.35, 2.88 and 2.32 again, respectively for ARROW-pKa, QMPFF3-pKa and GAFF-TR, so the order of performance is now reversed. (See Fig. 3; Tables 1, 3). Of note is the fact that QM methods, specifically COSMO-RS, performed noticeably better than the next best Force-Field challenger (us) both in batch 0 and in the overall set. Comparison of our performance in batch 0 and total set by error metrics (Table 3) shows that QMPFF3 and ARROW performance batch 0 is statistically better than on total set. All log D values for our QMPFF3 and ARROW submissions with and without pKa corrections and with empirical hydrophobic corrections with their statistical errors are presented in Table S1.
Finally, our torsional restraint technique for GAFF performed really well. Torsional restraints calculations improved LogD predictions significantly: computed MAE value for GAFF-TR model is 2.32 vs 2.75 for unrestrained calculations using initial starting geometries and 2.52 for unrestrained calculations using optimal starting geometries. The full results of all GAFF logD calculations are presented in Table 2. Detailed comparison with all GAFF related submissions are presented on Fig. S1 and Table S2. Some GAFF calculation have special peculiarities such as United Atom model for cyclohexane or ELBA water model. The closest analogue of our GAFF-TR calculation is column 10 in Table S2 which is neutral GAFF, presumably, without pKa corrections. While it has a slightly better RMSD 2.61 vs 2.71, GAFF-TR does better on overall correlation coefficient R 0.75 vs 0.65 and Kendall’s τ 0.54 vs 0.49.
Comparison of our predictions for water-cyclohexane distribution coefficient based on the QMPFF3-pKa, ARROW-pKa, and GAFF-TR to experiment for all 53 SAMPL5 molecules. Also shown for comparison are the predictions for COSMO-RS. The correlation coefficient R for QMPFF3-pKa, ARROW-pKa, GAFF-TR and COSMO-RS are 0.58, 0.60, 0.75, and 0.84 respectively
Conclusion
One of the great things about having a firm deadline is that it illuminates exactly where your team and your methods are. The FF parameters, the tools to obtain them, and the MD code we used are all relatively new and we were writing/finalizing some of these during the challenge. After the SAMPL5 challenge, we uncovered errors in our dH/dL calculation that were responsible for a part of our systematic hydrophobic shift, but not for all of it. The ARROW functional form development was accelerated specifically for the challenge and was implemented during the competition. Additionally, we were running and checking QM benchmarks for several new atom types that the SAMPL5 molecules required. The workflow lessons for us are that we became really short of time; that our parametrization procedures need to be much more automatic than they are now; and that while major code additions benefit from deadlines, they also suffer from them.
Scientifically, we drew several conclusions from the SAMPL5 challenge. First, in absolute terms, we see that our methods are not yet where we wish them to be. Some of the areas of improvement were clearly shown by the challenge: we need a better description of alkanes, better sampling, the latter both with brute force (longer simulation times) and with clever techniques (meta-dynamics and restraints), and more automated parametrization workflow. There are certainly other directions which we have not digested and formulated yet.
Our second goal was to see whether our polarizable FF’s show a systematic improvement over a non-polarizable FF (GAFF). On this goal the evidence is inconclusive. In batch 0 the performance was in the desired order: ARROW-pKa > QMPFF3-pKa > GAFF-TR. However, on the full set GAFF-TR, outperformed both polarizable FF’s. We would like to say that this was due to the extra attention we devoted to Batch 0 compounds, but we cannot be certain. Some of the remaining 40 molecules may simply be more challenging. It is also possible that some of them may need better sampling than the 500 ps we used for QMPFF3-pKa and ARROW-pKa, while for GAFF-TR we used torsional space corrections to logD that improved the agreement with experiment.
Our third aim was to compare our techniques to those of other groups’. Based on the Kendall’s τ metric, in batch 0, our most complex FF, ARROW, placed first among all MD FF-based methods. Furthermore, all three of our FF methods and some of their variants were at top of the rankings for batch 0. This is very satisfying. On the full set our performance was significantly worse, with exception of GAFF-TR which placed a respectable second amongst MD-FF methods. Again, the authors of the COSMO-RS technique [56] deserve much praise for their clearly superior entry.
Fourth, we are very pleased by the utility of torsional restraints. As mentioned above, the computed MAE value for GAFF-TR model is 2.32 vs 2.75 for unrestrained calculations using initial starting geometries and 2.52 for unrestrained calculations using optimal starting geometries, a very significant improvement. Of note are also the relatively short simulation times permitted by this technique. Essentially GAFF-TR used the least computational time of all submitted MD methods yet performed better than advanced force field calculations. This approach will be useful to other groups attempting similar calculations.
Participating in the SAMPL5 distribution coefficient challenge was incredibly useful for our group. We are happy to see that our force-fields perform relatively well; but we also see clearly that we have much room for improvements in both the models and in the workflow.
References
Anisimov VM, Lamoureux G, Vorobyov IV, Huang N, Roux B, MacKerell AD (2005) Determination of electrostatic parameters for a polarizable force field based on the classical Drude oscillator. J Chem Theory Comput 1(1):153–168. doi:10.1021/ct049930p
Anisimov VM, Vorobyov IV, Roux B, MacKerell AD Jr (2007) Polarizable empirical force field for the primary and secondary alcohol series based on the classical Drude model. J Chem Theory Comput 3:1927–1946
Applequist J (1977) An atom dipole interaction model for molecular optical properties. Acc Chem Res 10(3):79–85. doi:10.1021/ar50111a002
Applequist J (1993) Atom charge transfer in molecular polarizabilities: application of the Olson–Sundberg model to aliphatic and aromatic hydrocarbons. J Phys Chem 97(22):6016–6023. doi:10.1021/j100124a039
Cisneros GA (2012) Application of Gaussian electrostatic model (GEM) distributed multipoles in the AMOEBA force field. J Chem Theory Comput 8(12):5072–5080. doi:10.1021/ct300630u
Donchev AG, Galkin NG, Illarionov AA, Khoruzhii OV, Olevanov MA, Ozrin VD, Subbotin MV, Tarasov VI (2006) Water properties from first principles: simulations by a general-purpose quantum mechanical polarizable force field. Proc Natl Acad Sci USA 103(23):8613–8617. doi:10.1073/pnas.0602982103
Donchev AG, Galkin NG, Pereyaslavets LB, Tarasov VI (2006) Quantum mechanical polarizable force field (QMPFF3): refinement and validation of the dispersion interaction for aromatic carbon. J Chem Phys 125(24):244107. doi:10.1063/1.2403855
Donchev AG, Galkin NG, Illarionov AA, Khoruzhii OV, Olevanov MA, Ozrin VD, Pereyaslavets LB, Tarasov VI (2008) Assessment of performance of the general purpose polarizable force field QMPFF3 in condensed phase. J Comput Chem 29(8):1242–1249. doi:10.1002/jcc.20884
Patel S, Brooks CL 3rd (2004) CHARMM fluctuating charge force field for proteins: I parameterization and application to bulk organic liquid simulations. J Comput Chem 25(1):1–15. doi:10.1002/jcc.10355
Patel S, Mackerell AD Jr, Brooks CL 3rd (2004) CHARMM fluctuating charge force field for proteins: II protein/solvent properties from molecular dynamics simulations using a nonadditive electrostatic model. J Comput Chem 25(12):1504–1514. doi:10.1002/jcc.20077
Piquemal J-P, Chelli R, Procacci P, Gresh N (2007) Key role of the polarization anisotropy of water in modeling classical polarizable force fields. J Phys Chem A 111(33):8170–8176. doi:10.1021/jp072687g
Piquemal JP, Cisneros GA, Reinhardt P, Gresh N, Darden TA (2006) Towards a force field based on density fitting. J Chem Phys 124(10):104101. doi:10.1063/1.2173256
Piquemal J-P, Gresh N, Giessner-Prettre C (2003) Improved formulas for the calculation of the electrostatic contribution to the intermolecular interaction energy from multipolar expansion of the electronic distribution. J Phys Chem A 107(48):10353–10359. doi:10.1021/jp035748t
Ponder JW, Wu C, Ren P, Pande VS, Chodera JD, Schnieders MJ, Haque I, Mobley DL, Lambrecht DS, DiStasio RA, Head-Gordon M, Clark GNI, Johnson ME, Head-Gordon T (2010) Current status of the AMOEBA polarizable force field. J Phys Chem B 114(8):2549–2564. doi:10.1021/jp910674d
Ren P, Ponder JW (2003) Polarizable atomic multipole water model for molecular mechanics simulation. J Phys Chem B 107(24):5933–5947. doi:10.1021/jp027815+
Rick SW, Stuart SJ, Berne BJ (1994) Dynamical fluctuating charge force fields: application to liquid water. J Chem Phys 101:6141–6156
Shi Y, Xia Z, Zhang J, Best R, Wu C, Ponder JW, Ren P (2013) Polarizable atomic multipole-based AMOEBA force field for proteins. J Chem Theory Comput 9(9):4046–4063. doi:10.1021/ct4003702
Cole DJ, Vilseck JZ, Tirado-Rives J, Payne MC, Jorgensen WL (2016) Biomolecular force field parameterization via atoms-in-molecule electron density partitioning. J Chem Theory Comput 12(5):2312–2323. doi:10.1021/acs.jctc.6b00027
Pereyaslavets LB, Finkelstein AV (2012) Development and testing of PFFSol1.1, a new polarizable atomic force field for calculation of molecular interactions in implicit water environment. J Phys Chem B 116(15):4646–4654. doi:10.1021/jp212474p
Halgren TA, Damm W (2001) Polarizable force fields. Curr Opin Struct Biol 11(2):236–242
Warshel A, Kato M, Pisliakov AV (2007) Polarizable force fields: history, test cases, and prospects. J Chem Theory Comput 3(6):2034–2045. doi:10.1021/ct700127w
Khoruzhii O, Butin O, Illarionov A, Leontyev I, Olevanov M, Ozrin V, Pereyaslavets L, Fain B (2014) Polarizable force fields for proteins. In: Protein modelling. Springer, Berlin, pp 91–134
Hagler AT (2015) Quantum derivative fitting and biomolecular force fields: functional form, coupling terms, charge flux, nonbond anharmonicity, and individual dihedral potentials. J Chem Theory Comput 11(12):5555–5572. doi:10.1021/acs.jctc.5b00666
Mackerell AD, Bashford D, Bellott M, Dunbrack R, Evanseck J, Field M, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE III, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102:3586–3616
Weiner SJ, Kollman PA, Nguyen DT, Case DA (1986) An all atom force field for simulations of proteins and nucleic acids. J Comput Chem 7(2):230–252. doi:10.1002/jcc.540070216
Cornell W, Cieplak P, Bayly C, Gould I, Merz K, Ferguson D, Spellmeyer D, Fox T, Caldwell J, Kollman P (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Jorgensen WL, Tirado-Rives J (1988) The OPLS potential functions for proteins—energy minimizations for crystals of cyclic peptides and crambin. J Am Chem Soc 110:1657–1666
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) Development and Testing of the OPLS All-Atom Force Field on Conformational Energetics and Properties of Organic Liquids. J Am Chem Soc 118:11225–11236
Levitt M, Lifson S (1969) Refinement of protein conformations using a macromolecular energy minimization procedure. J Mol Biol 46(2):269–279
Allinger NL, Yuh YH, Lii JH (1989) Molecular mechanics. The MM3 force field for hydrocarbons. J Am Chem Soc 111(23):8551–8566. doi:10.1021/ja00205a001
Martin MG, Siepmann JI (1998) Transferable potentials for phase equilibria. 1. United-atom description of n-alkanes. J Phys Chem B 102(14):2569–2577. doi:10.1021/jp972543+
Mobley DL, Wymer KL, Lim NM, Guthrie JP (2014) Blind prediction of solvation free energies from the SAMPL4 challenge. J Comput Aided Mol Des 28(3):135–150. doi:10.1007/s10822-014-9718-2
Nicholls A, Mobley DL, Guthrie JP, Chodera JD, Bayly CI, Cooper MD, Pande VS (2008) Predicting small-molecule solvation free energies: an informal blind test for computational chemistry. J Med Chem 51(4):769–779. doi:10.1021/jm070549+
Geballe MT, Skillman AG, Nicholls A, Guthrie JP, Taylor PJ (2010) The SAMPL2 blind prediction challenge: introduction and overview. J Comput Aided Mol Des 24(4):259–279. doi:10.1007/s10822-010-9350-8
Geballe MT, Guthrie JP (2012) The SAMPL3 blind prediction challenge: transfer energy overview. J Comput Aided Mol Des 26(5):489–496. doi:10.1007/s10822-012-9568-8
Bannan CC, Burley KH, Mobley DL (2016) Blind prediction of cyclohexane–water distribution coefficients from the SAMPL5 challenge
Rustenburg AS, Dancer J, Lin B, Ortwine DF, Mobley DL, Chodera JD (2016) Measuring experimental cyclohexane/water distribution coefficients for the SAMPL5 challenge
Donchev AG, Ozrin VD, Subbotin MV, Tarasov OV, Tarasov VI (2005) A quantum mechanical polarizable force field for biomolecular interactions. Proc Natl Acad Sci USA 102(22):7829–7834. doi:10.1073/pnas.0502962102
Halgren TA (1996) Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J Comput Chem 17(5–6):490–519. doi:10.1002/(sici)1096-987x(199604)17:5/6<490:aid-jcc1>3.0.co;2-p
Misquitta AJ (2013) Charge transfer from regularized symmetry-adapted perturbation theory. J Chem Theory Comput 9(12):5313–5326
Williams HL, Chabalowski CF (2001) Using Kohn − Sham orbitals in symmetry-adapted perturbation theory to investigate intermolecular interactions. J Phys Chem A 105(3):646–659. doi:10.1021/jp003883p
Misquitta AJ, Szalewicz K (2002) Intermolecular forces from asymptotically corrected density functional description of monomers. Chem Phys Lett 357(3–4):301–306. doi:10.1016/S0009-2614(02)00533-X
Misquitta AJ, Jeziorski B, Szalewicz K (2003) Dispersion energy from density-functional theory description of monomers. Phys Rev Lett 91(3):033201
Werner HJ, Knowles PJ, Knizia G, Manby FR, Schütz M (2012) Molpro: a general-purpose quantum chemistry program package. Wiley Interdisciplinary Reviews: Computational Molecular Science 2(2):242–253
Řezáč J, Hobza P (2011) Extrapolation and scaling of the DFT-SAPT interaction energies toward the basis set limit. J Chem Theory Comput 7(3):685–689. doi:10.1021/ct200005p
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81(1):511–519. doi:10.1063/1.447334
Hoover WG (1985) Canonical dynamics: equilibrium phase-space distributions. Phys Rev A 31(3):1695–1697
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690. doi:10.1063/1.448118
Khoruzhii O, Donchev AG, Galkin N, Illarionov A, Olevanov M, Ozrin V, Queen C, Tarasov V (2008) Application of a polarizable force field to calculations of relative protein-ligand binding affinities. Proc Natl Acad Sci USA 105(30):10378–10383. doi:10.1073/pnas.0803847105
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005) GROMACS: fast, flexible, and free. J Comput Chem 26(16):1701–1718. doi:10.1002/jcc.20291
Kumar S, Rosenberg JM, Bouzida D, Swendsen RH, Kollman PA (1992) THE weighted histogram analysis method for free-energy calculations on biomolecules. I. The method. J Comput Chem 13(8):1011–1021. doi:10.1002/jcc.540130812
Kästner J (2011) Umbrella sampling. Wiley Interdis Rev Comput Mol Sci 1(6):932–942. doi:10.1002/wcms.66
Villa A, Mark AE (2002) Calculation of the free energy of solvation for neutral analogs of amino acid side chains. J Comput Chem 23(5):548–553. doi:10.1002/jcc.10052
MacCallum JL, Tieleman DP (2003) Calculation of the water–cyclohexane transfer free energies of neutral amino acid side-chain analogs using the OPLS all-atom force field. J Comput Chem 24(15):1930–1935. doi:10.1002/jcc.10328
Radzicka A, Wolfenden R (1988) Comparing the polarities of the amino acids: side-chain distribution coefficients between the vapor phase, cyclohexane, 1-octanol, and neutral aqueous solution. Biochemistry 27(5):1664–1670. doi:10.1021/bi00405a042
Klamt A, Eckert F (2000) COSMO-RS: a novel and efficient method for the a priori prediction of thermophysical data of liquids. Fluid Phase Equilib 172(1):43–72. doi:10.1016/S0378-3812(00)00357-5
Misquitta A, Stone A (2007) CamCASP: a program for studying intermolecular interactions and for the calculation of molecular properties in distributed form. University of Cambridge, Cambridge
Acknowledgments
The authors are grateful to the organizers and participants of SAMPL5 for arranging this competition on distribution coefficients, which enabled us to test our in-house software and assess our force-fields. We also would like to thank Anthony Stone for a beautiful and succinct way of visualizing potential maps [57] that inspired us to do the same. We are also thankful to everyone else in our company (in particular Meredith Robert) for supporting our participation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Ganesh Kamath, Igor Kurnikov, Boris Fain and Leonid Pereyaslavets have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kamath, G., Kurnikov, I., Fain, B. et al. Prediction of cyclohexane-water distribution coefficient for SAMPL5 drug-like compounds with the QMPFF3 and ARROW polarizable force fields. J Comput Aided Mol Des 30, 977–988 (2016). https://doi.org/10.1007/s10822-016-9958-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9958-4