Abstract
Microrganisms such as cyanobacteria have been often considered as exhibiting wide distribution mainly driven by environmental heterogeneity. Recently, however, new findings have evoked the role of previously neglected processes, such as dispersal limitation, determining the distribution of a wide range of microorganisms, including cyanobacteria. Here, we reviewed the biogeographic patterns of cyanobacteria with focus on molecular data and the evidences from the published literature for the processes driving these patterns. Also, considerations are made about concept of species, discordances in the taxonomic concepts, and level of taxonomic resolution, and how these affect the biogeographic study of cyanobacteria. From a overview, it can be observed that both environmental and historical factors are important to structure cyanobacteria diversity across time and space. Moreover, different species may exhibit significant differences in their distribution patterns, from possibly cosmopolitan species to other endemic species. However, distribution patterns are closely dependent on the concept of species, besides the taxonomic resolution, spatial and environmental scales, and the biases of the molecular methodologies applied in the studies. Thus, efforts to broaden sampling and sequencing of unknown and less-known species, as well as geographic regions and habitats poorly exploited, are crucial for a better understanding of cyanobacteria biogeography.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyanobacteria are a diverse group of microrganisms characterized as being oxygenic photosynthetic bacteria that possess chlorophyll a (Castenholz, 2001). Cyanobacteria possess a high ecological plasticity, thanks to which they occur in varied, often even extreme, habitats (Whitton & Potts, 2000). This group was responsible for critical events throughout the evolutionary history of life on Earth, such as the history of Earth-surface oxygenation (Lyons et al., 2014; Rasmussen et al., 2008; Sessions et al., 2009) and the evolution of eukaryotic cell through endosymbiosis (Margulis, 1970; Woese, 1987; McFadden, 2014). In addition, cyanobacteria play key roles in the ecosystems and biogeochemical cycles (Capone et al., 1997; Liu et al., 1997). Furthermore, in aquatic ecosystems, some taxa are capable of forming blooms, events characterized by the sudden increase in the abundance of one or some taxa in the environment. Also, many species from this group are able to release toxins (Carmichael, 2001; Paerl & Huisman, 2009), which can affect aquatic fauna as well as human populations. However, in spite of these negative impacts on ecosystems, cyanobacteria are economically important due to their potential use in pharmaceutical, food, and biotechnological industries (e.g., Singh et al., 2005; Gantar & Svircev, 2008; Gerwick et al., 2008; Pfeiffer et al., 2011; Zanchett & Oliveira-Filho, 2013).
Most research on cyanobacteria is guided by evolutionary, ecological, ecotoxicological, biochemical, and taxonomic concerns (for a review on common investigation on cyanobacteria see Sciuto & Moro, 2015). With the current development and improvement of molecular techniques and tools in microbial ecology, the microbial biogeography field has been deeply investigated (see Bell et al., 2005; Martiny et al., 2006, Ramette & Tiedje, 2007; Woodcock et al., 2007; Bell, 2010; Hanson et al., 2012; Naselli-Flores & Padisák, 2016; Padisák et al., 2016), including biogeography studies with focus on cyanobacteria (see Moreira et al., 2013). The methods for biogeographic studies in cyanobacteria and cyanotoxins are based on the identification and subsequent phylogenetic analyses of the isolates or environmental samples using predetermined genetic markers to establish their genetic diversity and geographic interaction (Neilan et al., 2003; Moreira et al., 2012). Thus, these studies are generally based on the discovery of new genera or species in new environments (e.g., Komárek, 1985; Couté et al., 2004; Couté & Bouvy 2004); endemisms (e.g., Sompong et al., 2005; Taton et al., 2006; Jungblut et al., 2010); potentials of dispersion and invasion (e.g., Padisák, 1997; Dyble et al., 2002; Wiedner et al., 2007; Vidal & Kruk, 2008; Sukenik et al., 2012; Wilk-Woźniak et al., 2016); and spatial distributions of species, populations and communities related to spatial and environmental variables (e.g., Papke et al., 2003; Drakare & Liess, 2010; Chamberlain et al., 2014).
The main goal of biogeography is to describe how species are distributed and the driving forces of these distribution patterns, and prokaryotic biogeography has been previously defined as the scientific field that documents the spatial distribution of prokaryotic taxa (Archaea and Eubacteria) in the environment at local, regional and continental scales (Ramette & Tiedje, 2007). To detect biogeographic patterns for microorganisms is crucial for understanding the distribution of biodiversity, since microorganisms are the most abundant and diverse group of Earth, being the life-support system of the planet (Pace, 1997; Hug et al., 2016; Locey & Lennon, 2016). Despite the classical idea of pure environmental selection on microbial distributions pervading the ecological literature for a long time (Baas-Becking, 1934), recent studies have evoked the existence of geographic isolation and dispersal barriers for some microbial groups from a variety of habitats, such as heterotrophic bacteria from soils (Cho & Tiedje, 2000), thermophilic cyanobacteria (Papke et al., 2003) and archaea (Whitaker et al., 2003) from hot springs. These recent hypothesis arguing against “cosmopolitan microbial rule” indicate a still very limited understanding of the mechanisms involved in microbial biogeography and how patterns of distribution vary among taxonomic groups and habitats, the main issues currently discussed in microbial ecology.
Despite the increased interest in exploring distribution patterns of cyanobacteria, however, a key issue in biogeography challenges the research in this field: the species concept. Since biogeography is concerned with the distribution of species, a well-defined species concept is required. Besides the existence of more than one concept of species (Dobzhansky, 1937; Wright, 1951; Slatkin, 1987), the concept of bacterial species in particular is considerably obscure, since its definition is typically defined coarsely (e.g., as > 97% similarity of 16S or 18S ribosomal RNA genes) (about bacterial species concept see Konstantinidis & Tiedje, 2005; Achtman & Wagner, 2008; Fraser et al., 2009; Caro-Quintero & Konstantinidis, 2011). Apart from these issues, cyanobacteria have a particular characteristic: the fact that they have traditionally been studied in botany and taxonomically classified in the same way as algae and plants. These issues, in addition to the scarcity of qualified professionals to identify and classify taxa, may potentially hamper studies of the group (Rejmánková et al., 2004), including biogeographic studies (as we will discuss throughout this review). This review explores the current status of the field of biogeography of cyanobacteria and highlights joint distribution/population genomics/biodiversity/workflows that aim to overcome some of the challenges commonly found in this field.
Cyanobacterial species concept
Although cyanobacteria have been widely studied in scientific literature, mainly with respect to aspects such as public health and bloom formation (Graham et al., 2009), little attention has been paid to their biodiversity (Nabout et al., 2013). These shortcomings in estimation of diversity and taxonomic classification are worsened by the confusion and overlap of the two taxonomic systems used to classify cyanobacteria: the Botanical Code and the Prokaryotic Code. As mentioned above, cyanobacteria were initially studied in botany, and for legacy reasons the International Code of Nomenclature for algae, fungi, and plants (Botanical Code) (McNeill et al., 2012) is still used for describing cyanobacteria in addition to the International Code of Nomenclature of Prokaryotes (Prokaryotic Code) (Parker et al., 2015). The history of cyanobacterial research in these two fields has also caused some discordance in taxonomic concepts and resulted in long discussions concerning the application of nomenclatural rules for cyanobacteria (Gaget et al., 2011; Oren, 2011; Oren & Garrity, 2014). Currently, the number of described cyanobacteria in the literature is controversial and the estimates range from 2000 (Sant’Anna et al., 2006) to 8000 species (Guiry, 2012) distributed among more than 300 genus. Nabout et al. (2013), in order to assume the underestimation of species, have estimated the number of unknown cyanobacteria species by using three models of discovery curves, and showed that the best model estimated a total of 6280 species (the number of species already described is 43% of this total).
With the huge advances that recent technologies have brought to microbiology, especially in the field of genomics, recent studies have focused their efforts on revising the taxonomy of cyanobacteria with a view to correcting mistakes caused by traditional taxonomics and clearing out the complicated evolutionary relationships of polyphyletic groups (e.g., Komárek, 2006; Komárek & Mareš, 2012; Komárek et al., 2014; Komárek, 2016). Today, the combination of microscopic data with molecular information is the most appropriate method for systematic and ecologic studies (Komárek et al., 2014; Komárek, 2016). More specifically, is considered as species within a single genus, those cyanobacterial strains or populations that belong to the “same genotype and morphotype with stable phenotypic features, and more or less stable and distinct ecological limits” (Komarek & Marěs, 2012; Komárek, 2016). Since the earliest-described species lack genetic information, previously described species are also undergoing thorough taxonomic revision (Castenholz, 2001; Komárek, 2006, 2016). Taxonomic revisions at the suprageneric level are also underway (e.g., Marěs, 2017), and currently it seems that the best defined way how to separate cyanobacterial species using molecular markers is the use of the 16S–23S internal transcribed spacer (ITS) region (Erwin & Thacker, 2008; Perkerson et al., 2011; Osorio-Santos et al., 2014).
It is also important to mention that, even if new efforts are made to access the genetic diversity of cyanobacteria, relating genetic diversity to a species concept is not easy. Cyanobacteria, in the same way as other prokaryotic organisms, have a set of essential genes (core genome) highly conserved and resistant to horizontal transfer (Shi & Falkowski, 2008; Larsson et al., 2011) and at same time a set of nonessential genes (accessory genome), that are more frequently subject to horizontal gene transfer and plays an important role in generating molecular diversity in cyanobacteria (Zhaxybayeva et al., 2006). Moreover, genome plasticity in cyanobacteria is evidenced by the broad distribution and hypervariability of mobile genetic elements (mainly insertion sequences), which can amount up to 10.95% of some genomes (Lin et al., 2010). Therefore, if populations of a same cyanobacterial species must have the same genotype, it is not trivial to think of which gene (or genes), and which cutoff value of nucleotide similarity, are more appropriate to consider. Phylogenetic reconstructions based on a single gene may be problematic, for instance, since they result in gene trees rather than species trees, ignoring the possibility of horizontal transfer which has been documented in prokaryotic rRNA operons (Ueda et al., 1999; Yap et al., 1999; Marěs, 2017).
Despite the genomics to be currently the most promising framework for revising the taxonomic classification of cyanobacteria, it is worth mentioning that these field has been moving forward at a pace that is relatively slower than currently observed for some of the other bacterial phyla (Alvarenga et al., 2017). Nowadays, the number of cyanobacterial genomes amounts for approximately 0.6% of all prokaryotic genomes available (Alvarenga et al., 2017). About 400 cyanobacterial genomes are available in public databases, in contrast to more than 30,000 complete genomes available for strains classified in 50 bacterial and 11 archaeal phyla (Land et al., 2015). Cyanobacteria, therefore, are still severely underrepresented in genomic databases when compared to other prokaryotes, and the currently available cyanobacterial genome databases are still lacking in taxonomic, environmental, and geographic diversity (Alvarenga et al., 2017). This scenario, in addition to providing an incomplete picture of this phylum, prevents the expansion of knowledge of the molecular biology of cyanobacteria, since sequences from neglected taxa may bring to light answers to meaningful questions (Richards, 2015).
From obtaining new genetic data of cyanobacteria in order to get closer to their real biodiversity, not only can their phylogeny be better resolved, but new approaches in ecological studies may be explored, such as in phylogenetic community ecology, phylogeography, and of course biogeography. As mentioned briefly above, biogeography is very sensitive to the species concept and taxonomic resolution, both closely related to access to biodiversity. This is because key concepts studied in biogeography are defined from a species definition: habitats/niches are defined as a particular combination of resources and conditions necessary for a particular species (Tiedje, 1993), and both dispersion and ecological drift (including here speciation and extinction) depends on the size of the populations (Slatkin, 1987), which in turn will depend on the definition of specie adopted. Considering the processes involved in the biodiversity distribution mentioned above, in the following sections, we present a synthesis of the biogeography studies on cyanobacteria, and the evidences presented in the literature of the driving factors shaping cyanobacteria distribution. At the same time, we made considerations regarding species concept, taxonomic resolution and other issues involved in the development of this research field. Finally, we discuss perspectives for this field and suggestions for future investigations that will contribute to a better understanding of the biogeography of cyanobacteria.
Environmental selection (current biotic and abiotic conditions)
One of the processes involved in the distribution of biodiversity widely studied in biogeography is the environmental selection (or environmental filtering), in which abiotic and biotic (biological interactions) conditions selects against species with different tolerances and habitat preferences. This process traditionally was thought to be a major mechanism structuring communities (Stein et al., 2014), and widely explored in ecological studies, mainly on animals and plants. In microbial ecology, the role of environmental heterogeneity also was thought to be the main factor shaping bacterial distribution, assumption known as Baas-Becking hypothesis (Baas-Becking, 1934). In this fundamental paradigm, everything is everywhere, but environment selects (Baas-Becking, 1934), the first proposition implies that microorganisms have dispersal abilities so high that the effects of past processes are suppressed; and the second one assumes that the current environmental characteristics determine the microbial distribution in the ecosystems. Although the idea of distribution purely selected by environmental conditions is currently under intense debate and review (as we will show in more detail in the following sections), a lot of evidence in the literature shows an important role of environmental heterogeneity in the biogeographic distribution of cyanobacteria.
At the same time, as cyanobacteria present a broad range of metabolic capacities that enable them to deal with a range of environmental conditions (Vasas et al., 2010)—this group exhibits high niche specialization (Whitton & Potts, 2000)—then, the occurrence of distributions restricted to certain habitat types can highlight the role of environmental selection. Several studies have shown that, in fact, the environmental heterogeneity influences the distribution of cyanobacteria in a variety of spatial scales and ecosystems (Tables 1 and 2). At local and regional spatial scales, patterns of cyanobacteria distribution in aquatic ecosystems are commonly related to the concentration of nutrients and variables related to solar radiation, such as water transparency and water body depth (e.g., Drakare & Liess, 2010; Tian et al., 2012; Chamberlain et al., 2014; Ren et al., 2014; Harris et al., 2016). Similarly, in terrestrial environments, the physical–chemical characteristics of soils and sediments appear to be important factors shaping the distribution of cyanobacteria (e.g., Garcia-Pichel et al., 2001; Thomasa & Dougil, 2006; Chamberlain et al., 2014).
Cyanobacterial blooms in aquatic ecosystems also show how changes in local and regional environmental characteristics can affect and alter the distribution patterns of these organisms. Events of cyanobacteria blooms show to what extent environmental variables (specifically the excess in nutrient concentrations and temperature increase) (Paerl et al., 2001; Paerl & Paul, 2012; Paerl & Otten, 2013) can rapidly affect and change the structure of cyanobacterial communities. Some genera in particular are widely studied because of their ability to form blooms (toxic or not) in freshwater environments, such as Microcystis, Anabaena, Cylindrospermopsis, and in marine ecosystems, such as Lyngbya, Synechococcus, Trichodesmium (Rastogi et al., 2015). On tropical blooms, for example, a meta-analysis showed that Microcystis blooms were more associated with higher total nitrogen concentrations, while Cylindrospermopsis blooms were more associated with higher maximum temperatures (Mowe et al., 2015). Also, environmental changes on a global scale (e.g., global warming) may potentially alter patterns of cyanobacterial blooms. More details on this issue could be found in the section: “Invasions and Global Climate Change”.
In extreme environments, cyanobacteria exhibit remarkable adaptability, notwithstanding the adverse environmental conditions of most inhospitable ecosystems in the Earth, including the frozen regions of the poles (e.g., Taton et al., 2006; Wood et al., 2008; Namsaraev et al., 2010), hypersaline environments (see Oren, 2015), and, on the other extreme, the high temperatures of hot springs (e.g., Papke et al., 2003; Miller et al., 2007; Ionescu et al., 2010). These commonly isolated ecosystems provide appropriate scenarios for assessing biogeographic patterns of distribution of highly specialized groups. In the polar environments, the distribution of cyanobacteria has been correlated with aeolian processes (Michaud et al., 2012), salinity (Jungblut et al., 2005) and soil chemical characteristics (Wood et al., 2008), whereas in thermal springs areas cyanobacteria distribution shows a relation with water temperature (Sompong et al., 2005).
Biotic variables (such as the abundance and richness of other organisms) may indicate that biological interactions are acting as important factors shaping cyanobacteria distribution. This relationship has already been studied and evidenced both in studies with experimental approach as well as through field data (e.g., Sullivan et al., 2003; Mühling et al., 2005; Agawin et al., 2007; Van Wichelen et al., 2010; Apple et al., 2011; Sønstebø & Rohrlack, 2011). A study specifically evaluating the diversity of Synechococcus in marine ecosystems (Mühling et al. 2005), for example, found evidence that viral infection may play an important role in determining the success of different Synechococcus strains. Positive and negative interactions with herbivores also influence cyanobacterial distribution patterns. For instance, copepods may facilitate cyanobacteria (Hong et al., 2013) and high abundances of generalist herbivores (e.g., Daphnia) can control cyanobacterial blooms when released from planktivorous fish predation (Sarnelle, 2007). Also, cyanobacterial blooms occurring in response to eutrophic conditions in water bodies may also be followed by associations with heterotrophic bacteria, several being capable of enhancing cyanobacterial growth (Berg et al., 2009).
In extreme environments such as poles and hot springs, biological interactions also play an important role in cyanobacteria distribution patterns. In the Antarctic continent, a study has observed an increase in the diversity of cyanobacteria from sub-Antarctic to continental Antarctica (Namsaraev et al., 2010), a pattern that could be explained by the disappearance of the vegetation cover since plants and mosses limit the amount of resources (nutrients available and light) for cyanobacteria. At the other extreme, in the alkaline hot springs of Yellowstone National Park, a strong negative association between the relative abundances of cyanobacteria and Chloroflexi (non-sulfur green bacteria) shows a likely competitive interaction between these two groups, possibly by habitat and/or limiting resources (Miller et al., 2009).
Thinking about continental and global distribution of cyanobacterial species, some taxa are studied because of their worldwide distributions. Before highlighting some of these studies, however, some terms used need to be clear. In biogeography, species considered as cosmopolitan are characterized by global distribution or distribution spanning several biogeographic provinces (Dijoux et al., 2014). It is important to mention here that this definition, which will be used henceforth, does not automatically imply in species that have highly efficient means of dispersal by being carried by wind or water (Fenchel & Finlay, 2004), but only concerns the distribution pattern seen today (dispersal abilities will be covered in the next section). Also, we will use the subcosmopolitan term, which was suggested by Padisák (2003) in order to distinguish these species from those cosmopolitan species, global occurrence of which is related to certain environments corresponding to species-specific adaptations.
Temperature is one of the majors studied factors shaping cyanobacteria distribution at global scales, and some subcosmopolitan species are restricted to the warmest or coldest climatic regions. Pantropical species are those that occur only roughly between the two Tropics and some cyanobacteria species found in this group are all Cylindrospermopsis species (except Cylindrospermopsis raciborskii (Woloszynska) Seenayya & Subba Raju; more about this species in the subsequent sections), Arthrospira fusiformis (Voronikhin) Komárek & J.W.G. Lund, Anabaena fuellebornii Schmidle, Anabaena iyengarii Bharadwaja, Anabaena leonardii Compère, Anabaena oblonga De Wildeman and Anabaenopsis tanganyikae (G.S. West) Woloszynska & V.V. Miller (Padisák, 2003). Well-known temperate species, on the other hand, appear to be restricted to temperate zones (Hoffmann, 1996; Vyverman, 1996), like Planktothrix rubescens (De Candolle ex Gomont) Anagnostidis & Komárek, Limnothrix redekei (Goor) Meffert, Dolichospermum solitarium (Klebahn) Wacklin, Hoffmann et Komárek, Dolichospermum flos-aquae Lyngbye Brébisson ex Bornet et Flahault) Wacklin, Hoffmann et Komárek, Dolichospermum lemmermannii (Richter in Lemmermann) Wacklin, Hoffmann et Komárek, Anabaenopsis arnoldii Aptekar, Anabaenopsis milleri Voronichin, Aphanizomenon flos-aquae Ralfs ex Bornet & Flahault and Cuspidothrix issatschenkoi (Usachev) P.Rajaniemi, Komárek, R.Willame, P. Hrouzek, K.Kastovská, L.Hoffmann & K.Sivonen.
Global distributions of cyanobacteria in the extreme cold environments, or cryoenvironments, have also received attention recently (e.g., Taton et al., 2006; Wood et al., 2008; Namsaraev et al., 2010). Based on clone-library and phylogenetic analysis (16S rRNA), Jungblut et al. (2010) evaluated the global distribution of cyanobacteria by comparing communities from the North America (High Arctic Canada) with those from analogous sites in Antarctica. This study have showed that several of the High Arctic ribotypes were found to be > 99% similar to Antarctic and alpine sequences. Moreover, more than 68% of all identified ribotypes at each site matched only cyanobacterial sequences from perennially cold terrestrial ecosystems, and were < 97.5% similar to sequences from warmer environments. These results show a subcosmopolitan distribution of ecotypes from different cyanobacterial taxa that, although they have a global distribution, they are highly specialized and adapted to the extreme cold conditions of the poles.
Some species of cyanobacteria that present subcosmopolitan distribution are of particular interest since specific strains are capable of forming harmful blooms. Among these, Microcystis aeruginosa (Kützing) Kützing is a bloom-forming freshwater-type cyanobacteria that abounds in eutrophic and hypertrophic freshwater bodies worldwide (Chorus & Bartam, 1999; de Figueiredo et al., 2006; Kardinaal et al., 2007; Vareli et al., 2009). This species has already been studied in biogeography and phylogeography and a huge of evidences of a subcosmopolitan distribution is found in the literature (Haande et al., 2007; Van Gremberghe et al., 2011). Studies analyzing populations from distinct geographic locations have revealed a high degree of intraspecific genetic similarity, indicating a robust species definition for this cyanobacterium, and this outlook was corroborated in studies using distinct molecular markers and taxonomic resolutions, such as the 16SrRNA marker (Neilan et al., 1997), cpcBA-IGS marker (Bittencourt-Oliveira et al., 2001), 16S-23S ITS marker (Haande et al., 2007; Van Gremberghe et al., 2011) and PC-IGS marker (Haande et al., 2007).
Van Gremberghe et al. (2011) studying several Microcystis ITS sequences from worldwide suggest a truly cosmopolitan distribution for this cyanobacterium, and interestingly, no genetic structuring according to climate conditions was found (several Microcystis ITS types were detected in a wide range of climates, indicating a broad tolerance or capacity for rapid local adaptation). Van Gremberghe et al. (2011) argue that is more plausible that the genetic structure of Microcystis populations is driven by a recent global expansion and founder effects that arise whenever new habitat patches are created (Boileau et al., 1992), which are then colonized by a random selection of strains from regional or possibly global sources. The rapid local adaptation in a wide array of environmental conditions may be a result of the genome plasticity of M. aeruginosa (Frangeul et al., 2008), allowing new genetic interactions and higher variance on which natural selection can act. Also, de novo mutations might be involved in rapid adaptation to novel environments given the large population sizes and short generation time of M. aeruginosa (van Gremberghe et al., 2011). Indeed, the distribution of nonneutral genes of M. aeruginosa has already been related to habitat specificity, for example to land use (Marmen et al., 2006, using the mcyD and mcyA genes), and an evidence of local expansion of M. aeruginosa as response to environmental adaptation was recently found (using a multilocus approach) (Tanabe & Watanabe, 2011).
Two groups of well-studied picocyanobacteria, Synechococcus and Prochlorococcus, also provide some evidences about processes that might structure populations on a global scale. Synechococcus and Prochlorococcus are marine cosmopolitan genera of cyanobacteria and the major primary producers in the world’s oceans (Li, 1994). The two genera are distinguishable by their possession of dissimilar light-harvesting apparatus (Ting et al., 2002) and even based on 16S rRNA gene (a very conservative genome region), several lineages within Synechococcus have been described (Fuller et al., 2003), while other potential lineages have been designated based on others genomic regions, such as ITS and ntcA sequences (Ahlgren & Rocap, 2006; Penno et al., 2006). It is widely known that different environmental pressures act not only between these two taxa, but also within each taxa, structuring well-known ecotypes (populations genetically different) in space according to different habitat preferences (e.g., Ernst et al., 2003; Zwirglmaier et al., 2008; Martiny et al., 2009; Ahlgren & Rocap, 2012; Huang et al., 2012; Mazard et al., 2012; Shibl et al., 2014) (Table 2). In a study on a global scale (Zwirglmaier et al., 2008), for instance, the distribution lineages (based on 16S rRNA) of these two cyanobacterial groups were related to environmental variables and, interestingly, Prochlorococcus appeared to be more influenced by the physical parameters (temperature and depth), while Synechococcus by chemical parameters (nutrients). Genetically distinct clades based on ITS sequences with very broad distributions have also been detected (Rocap et al., 2002; Chen et al., 2006), and for these two taxa, ITS diversity was comparable with Microcystis. However, Synechococcus and Prochlorococcus have presented a genetic structure clearly more pronounced when compared with Microcystis populations (Van Gremberghe et al., 2011).
Although the role of environmental selection on the distribution of several cyanobacterial taxa and habitats is well known, some factors must be taken into account. First, the spatial scale is an important factor that must be considered. Increasing environmental heterogeneity with the area along with the specificities of taxa in different habitats is the most common explanation for taxa–area patterns (Rosenzweig, 1995); thus, it is expected that environmental selection to be stronger at finer spatial scales than at broader ones. Moreover, this is certainly a taxon-dependent pattern. For example, in an environment where increasing the area leads to an increased environmental variation, the distribution of groups with greater environmental tolerance would tend to be less dependent on spatial scale and vice versa. In addition, this relationship can be found not only among species, but also among populations of the same taxon with different habitat preferences, or ecotypes—a possible approach to be explored in future studies with other cyanobacteria taxa, mainly species that were not yet, or very little, explored.
The second issue to be taken into account is the taxonomic resolution adopted in the study. As mentioned earlier, when microbial taxa are defined from molecular approaches and classified into operational taxonomic units (OTUs), they are defined grossly through similarity in the sequence of one or more regions of the genome. Thus, since habitats are defined as a particular combination of resources and conditions necessary for a particular taxon (Tiedje, 1993), turnover habitat estimates are sensitive to the taxonomic definition, and thus, some biogeographic patterns of microorganisms may be more detectable at fine resolutions (detecting more compositional variation) than at the coarsest resolutions (Hanson et al., 2012). For example, in a scenario where “all cyanobacteria” were defined as a single taxon, one would observe a cosmopolitan distribution group with an enormous niche amplitude, and the influence of environmental selection would not be detectable. Horner-Devine et al. (2004) described taxa–area relationships for nonphotosynthetic bacteria from salt marsh sediments using different OTU definitions (95, 97 and 99% 16S rDNA sequence similarity) at scales from centimeters to hundreds of meters. This study showed that the taxa varied in the space mainly due to the environmental heterogeneity, but the turnover rate of the taxa was dependent on the bacterial lineage and the taxonomic resolution adopted: the turnover was higher with the increase of the taxonomic resolution. That is to say that finer taxonomic resolutions (e.g., 99% of sequence similarity) actually tend to detect greater compositional variation and consequently have greater power of environmental selection detection.
Regarding Cyanobacteria, a study analyzing the spatial variation of Synechococcus marine populations (Mazard et al., 2012) showed that in all cutoff values (88, 91, 94, 97 and 99% of similarity of multilocus sequences) the environmental factors showed a correlation with the distribution of the populations. However, the OTU definition using the cutoff value of 94% provided the best separation of the OTUs in relation to the sampled sites and the environmental parameters and, therefore, the best resolution for detecting possible ecotypes. Similarly, Martiny et al. (2009) have showed that Prochlorococcus distribution was dependent on the degree of sequence identity used to define a taxon (using the 16S-23SrRNA ITS region): light correlates with broad-scale diversity (90% cutoff), whereas temperature with intermediate scale (95% cutoff). These approaches not only identifies ecological differences at the population level, but also allows the analysis of the biogeographic distribution of ecotypes as a function of environmental variation and evolutionary processes, an feasible approach to be adopted in future studies with other species of cyanobacteria from many other ecosystems.
Finally, it is important to mention that genetic diversity and morphological diversity do not automatically imply the existence of ecotypes (or even different species). One case that may be mentioned is the cyanobacterium M. aeruginosa. Although on the one hand a high level of genetic diversity in a high number of genotypes was detected in the genus (Neilan et al., 1995; Kondo et al., 2000; Bittencourt-Oliveira et al., 2001; Wilson et al., 2005; El Herry et al., 2008; Yoshida et al., 2008; Tanabe et al., 2009; Fathalli et al., 2011; Gaevsky et al., 2011), on the other hand a number of morphospecies previously described (based on colony morphology) are not supported by molecular data forming a clade of nearly identical 16S rDNA sequences (Lepère et al., 2000; Litvaitis, 2002). Based on this extremely low 16S sequence divergence, along with DNA-DNA hybridisation data, Otsuka et al. (2001) suggested, under the rules of the Bacteriological Code, merging all morphospecies into a single species. This example shows how the concatenated sequencing of several regions of the genome is important so that issues tangible to the ecological, evolution, and obviously taxonomy, are fully understood.
Historical processes (dispersion, past environmental selection, and drift)
In this section, we discuss the first statement of the Baas-Becking hypothesis, everything is everywhere, which implies that microorganisms have such a great dispersion capacity so as to quickly remove the effects of past processes. Historical processes that may influence the current distribution of organisms include dispersal, past environmental selection, and drift, which may lead to genetic divergence between populations and compositional variation between communities (Martiny et al., 2006). One of the main arguments behind the ‘everything is everywhere’ is that the small size, large population size, and consequently the high abundance of propagules increase the dispersion rate to levels where the dispersal limitation essentially does not exist. The high dispersal rate increases the similarity in the composition of communities and decreases rates of intra- and interpopulation differentiation through increased gene flow, in the case of cyanobacteria and other prokaryotes, through horizontal gene transfer (HGT) (Martiny et al., 2006; Hanson et al., 2012).
One commonly used approach to assess the role of the dispersal limitation in structuring biological communities is through distance–decay relationship: how communities become more dissimilar as the distance between them increases (neglecting the effect of environmental variation on space) (Nekola & White, 1999; Morlon et al., 2008). Thus, the effect of geographic distance should be relatively weak in habitats where the dispersal rate is high and vice versa. Hillebrand et al. (2001) were one of the first to record this relation for microbial taxa. This study showed that in all taxonomic groups analyzed (diatoms, ciliate, corals, and polychaetes), the species similarity declined significantly with distance, being one of the first evidence of dispersal limitation for microorganisms, which contradicts the classical hypothesis about the cosmopolitan dispersion of microorganisms.
Isolated and extreme environments (i.e., island-like habitats) can act as barriers to gene flow, lead to the isolation of microbial groups—and consequently to genetic divergence, and thus facilitate speciation processes. A study based on temporal phylogenetic approach calibrated using microfossil data from the extremophile cyanobacteria Chroococcidiopsis (Bahl et al., 2011) from cold and hot deserts showed that there was no relation between the genetic differentiation of the lineages (based on 16S-ITS-23S rRNA) of this taxon and the geographic distance, nor on a global scale or for each phylogenetically defined cluster. However, the common ancestry time of the lineages precedes the estimates for contemporary aridity in the desert regions where this taxon occurs, indicating that the distribution of Chroococcidiopsis was, at least in part, limited by barriers and/or invasive colonization. Another global-scale study with Synechococcus from hot springs (Papke et al., 2003) showed a negative correlation between geographic distance and genetic similarity (based on 16S-ITS-23S rRNA) among the lineages, suggesting that the nonrandom and noncosmopolitan distribution of Synechococcus populations is influenced, at least in part, by geographic isolation and dispersal limitation. Similarly, phylogeographic patterns of thermophilic cyanobacteria Mastigocladus laminosus Cohn ex Kirchner from thermal areas show a significant and positive correlation between genetic differentiation (based on 16S rRNA and nitrogen metabolism loci) and geographic distance, providing evidence for this species of a distribution pattern influenced by geographic isolation (Miller et al., 2007).
Although dispersal limitation seems to be an important factor in the distribution of some cyanobacterial groups, as shown above, other groups appear to have broad dispersal abilities and (sub)cosmopolitan distributions. A study in Jordanian hot springs (Ionescu et al., 2010), for instance, showed that some cyanobacterial isolates presented high similarity (> 99% 16S rRNA) with others from hot springs of other regions of the globe. Similarly, the aforementioned study by Jungblut et al. (2010) showed that many ribotypes from Arctic showed > 99% similarity (16S rRNA) with Antarctica and Alps sequences. Among these, further, an Arctic sequence showed 99.8% similarity to the sequence of Leptolyngbya antarctica (West & G.S.West) Anagnostidis & Komárek sequenced from Antarctica, indicating a high dispersion capacity for this species and thus a subcosmopolitan distribution pattern. However, as will be discussed below, these subcosmopolitan distributions records have used taxon definitions based on the 16S rRNA gene sequence similarity, a very conservative taxonomic definition.
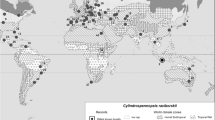
Among the examples of global dispersion, the best-known example is Cylindrospermopsis raciborskii. This species is one of the most notorious cylindrospermopsin (CYN) producers that can be found in freshwater habitats in the temperate, tropical, and subtropical regions of the world (Moreira et al., 2011). It is known that to make a definite determination of the native range of microbial species is extremely difficult: C. raciborskii was found for the first time in 1887 in the Nile, reported as C. kaufmannii (Schmidle) Huber-Pestalozzi, (Wołoszyńska, 1912); however, its Locus typicus is, indeed, the Rava Demangan pond in Java, Indonesia (Wołoszyńska, 1912). C. raciborskii expanded rapidly to Europe in the last century and is considered an invasive alien species of the temperate zone (Padisák, 1997).
Phylogeographic studies have showed that populations of C. raciborskii are geographically grouped. Neilan et al. (2003) based on 16S rRNA and cyanobacterium-specific short tandem repeat sequence (HIP1) clustered: (1) strains from the USA and Brazil; (2) European strains (Germany, Hungary, and Portugal); and (3) Australia strains. Gugger et al. (2005) based on 16S–23S internally transcribed spacer (ITS1) revealed the same continental cluster distribution and suggested that the current expansion of C. raciborskii in Europe—and in Central and North Americas—did not result from recent invasion and colonization by African or Australian strains (Padisák, 1997), but rather represent local strains that maintained “cryptic” populations over time and only recently proliferated due to climate change and variations in other environmental conditions (Padisák et al. 2016). Also, Moreira et al. (2012) based on multilocus sequences recently showed that C. raciborskii strains grouped into three well-supported distinct clusters: (1) European, (2) African/American, and (3) Asian/Australian, and also suggested the recent invasion of C. raciborskii in Portuguese and other European temperate environments.
C. raciborskii has been probably the only cyanobacterial species for which dispersal routes could be reconstructed and the speed of the dispersal could be estimated: it was estimated that less than a century was needed to colonize appropriate habitats all over the world (Padisák et al., 2016). The successful dispersion of C. raciborskii was largely attributed to its ability to tolerate travel along river courses (Padisák, 1997). Moreover, Piccini et al. (2011) proposed that phenotypic and genetic variability of C. raciborskii populations is linked to the existence of different ecotypes success of which is subject to the local environmental conditions. It is also speculated that unique physiological traits of C. raciborskii enable their proliferation in newly colonized ecosystems, currently exposed to greater environmental and temperature disturbances (Padisák et al., 2016). Further considerations about the invasive potential of C. raciborskii and the possible traits related to its invasiveness can be seen in the section: “Invasions and Global Climate Change”.
Phylogeographic studies with other cyanobacterial taxa also show how historical processes, such as dispersal limitation, adaptive radiation and allopatric speciation, can affect the global distributions of populations (Table 2). Another example that can be mentioned is the bloom-forming and nitrogen fixing filamentous cyanobacterium Dolichospermum (Anabaena) circinalis (Rabenhorst ex Bornet & Flahault) P.Wacklin, L.Hoffmann & J.Komárek. Worldwide blooms of D. circinalis are well known due to their production of neurotoxins as anatoxin-a and paralytic shellfish poisons (PSPs) (Padisák et al., 2016). Beltran & Neilan (2000) identified a geographic segregation of neurotoxin production in this cyanobacterium: American and European isolates of D. circinalis produce only anatoxin-a, while Australian isolates produce exclusively PSPs. Moreover, the phylogenetic structure of D. circinalis (based on 16S rRNA) suggested a monophyletic group with worldwide distribution, and the PSP- and non-PSP-producing D. circinalis formed two distinct 16S rRNA gene clusters (Beltran & Neilan, 2000). Although the phylogeographic structure of this cyanobacterium can not yet be fully understood, these results may suggest a truly (sub)cosmopolitan distribution pattern followed by local environmental adaptations.
Restricted and/or endemic distributions are interesting in biogeography because a combination of factors is required to emerge them, such as geographic isolation, temporal continuity for local adaptations to accumulate, and mechanisms to reduce the intensity of HGT (Souza et al., 2008). Overall, endemism can have two different kinds of origin: the most pure form of endemism is when a species evolves at a certain location and remains exclusive to that location; and the so-called relict endemism occurs as a result of habitat fragmentation or destruction and a subsequent extinction from all localities except one (Padisák et al., 2016). Regarding cyanobacteria, some examples can be mentioned: Aphanizomenon manguinii Bourrelly and Trichormus subtropicus (N.L. Gardner) Komárek & Anagnostidis, so far been recorded only on several islands in the Caribbean region (Komárek, 1985). Also many recently described species of Cylindrospermopsis which have so far been recorded only in Central America: C. acuminatocrispa Couté & M.Bouvy in a reservoir in NE Brazil (Couté & Bouvy, 2004), C. catemaco Komárková-Legnerová & R.Tavera in Lake Catemaco, Mexico (Komárková-Legnerová & Tavera, 1996) and C. taverae Komárek & Komárková-Legnerová in Central Mexico (Komárek & Komárková-Legnerová, 2002). And also a newly described species from Europe: C. sinuosa Couté, M.Leitão & H.Sarmento (Couté et al., 2004).
Polar regions are ideal locations for evaluating microbial endemism, since they contain parallel environments separated by large geographic distances and potential barriers to dispersal (Staley & Gosink, 1999), and indeed many possibly endemic cyanobacteria from Antarctica have been identified already (Taton et al., 2006; Comte et al., 2007; Jungblut et al., 2010; Michaud et al. 2012) (Table 1). In hot springs, another island-like habitat, distinct cyanobacterial populations were also characterized as possibly endemic (Papke et al., 2003; Sompong et al., 2005). Despite the importance of knowing endemic distributions, however, it is obvious that the detection of microbial endemism has some difficulties: the relation between the detection of endemism and the taxonomic resolution (as will be discussed below); the difficulty of confirming an endemic distribution because many cyanobacteria species (as well as other microorganisms) can inhabit an aquatic ecosystem unnoticed, since it may remain at a rather low biomass concentration and it does not form a conspicuous fraction of the community (Sukenik et al., 2012); and finally the difficulty of distinguishing whether a present taxon is in fact an active member of the community or a dormant transient (Curtis et al., 2002; Hanson et al., 2012). In addition, there several species that, for a long time, have been known only from the type locality (local endemics), which cannot be considered as real endemics as long as more surveys in the same habitat-types in other continents are missing (Padisák et al. 2016).
As for environmental selection, the influence of historical processes on the current distribution of microorganisms is dependent on the spatial scale and taxonomic resolution adopted in the study. Here, dispersion capacity is expected to decrease with increasing distance. One study with the archaeal Sulfolobus (Whitaker et al., 2003) have shown this relationship, in which in spatial scales ranging from meters to tens of thousands of kilometers, the magnitude of the dispersion rate is in fact associated with the spatial scale (migration is greater between closer sites). Again, it is clear that this relationship is taxon dependent, since different taxonomic groups have different rates of dispersion and/or establishment success, and more or less capacity to cover long distances.
Regarding taxonomic resolution, both dispersion and drift (including here speciation and extinction) depends on the size of the populations, which in turn will depend on the definition of species adopted. Again, using the definition of a taxon as “all cyanobacteria”, for example, would result in a large estimate of population size, high potential for dispersion, and a high probability of cosmopolitanism, and thus the influence of historical processes would be completely neglected. This relationship has already been evidenced in a study with marine populations of Prochlorococcus (Martiny et al., 2009), in which the distribution of the populations was correlated with the dispersal rate only in the finer taxonomic resolution examined (99.5% sequence similarity using the 16S–23S ITS region). This is a possible approach to be tested in future researches with other cyanobacterial taxonomic groups.
Finally, the relation between the detection of endemism and the taxonomic resolution is obvious. On the one hand, studies using a wide range of molecular techniques to evaluate spatial patterns of prokaryotes, such as 16S rDNA sequencing and DNA/DNA pair hybridization, indeed suggest that genera of prokaryotes are widely distributed (e.g., Hagstrom et al., 2000; Brandao et al., 2002; Zwart et al., 2002; Hedlund & Staley 2004). In addition, cosmopolitan distributions have already been recorded using a taxonomy definition based on the 16S rRNA gene (e.g., Garcia-Pichel et al., 1996; Ionescu et al., 2010; Jungblut et al., 2010). Since 1% divergence in the 16S rRNA gene being equal to isolation or reduced genetic exchange for < 10 million years (Jungblut et al., 2010), this gene is highly conserved and thought to underestimate the number of endemic species in a given habitat (Cho & Tiedje, 2000). On the other hand, when methods that offer a fine resolution are employed, bacteria appear to have more restricted and endemic distributions (e.g., Cho & Tiedje, 2000; Whitaker et al., 2003). This same pattern occurs in studies with cyanobacteria, where endemism and restricted distributions (suggesting low dispersion capacity and/or high niche specificity) are more detected at fine taxonomic resolutions than coarser (e.g., Papke et al., 2003; Miller et al., 2007; Mazard et al., 2012).
Invasions and global climate change
The entry, establishment, and spread of nonnative species to a new ecosystem, is frequently described as biological invasion (Vitousek et al., 1997; Ricciardi & Cohen, 2007). Biological invasion is considered as an extension of normal colonization processes such as succession (Elton, 1958), and it was proposed that the term invasive species will be restricted to describe nonnative species that expanded their geographic range, became abundant and have environmental and economic impact (Colautti & Macisaac, 2004). Invasions also may threaten global biodiversity by altering the structure and function of ecosystems and disrupting key biological interactions (Traveset & Richardson, 2006). Indeed, when invading new areas, phytoplankton species (including cyanobacteria) are able to cause irreversible environmental changes by outcompeting native species, changing food-web structures (Dufour et al., 2006), or reduce diversity (Borics et al., 2000, 2012). Moreover, biological invasions are important in biogeography since invasion involve dispersal ability, environmental adaptation, unique physiological traits (like dormant stages as dispersal units) and ecological plasticity.
The invasion of free-living microorganisms to new aquatic habitats is rather cryptic and difficult to detect therefore invasions of these “invisible invaders” have been rarely reported (Litchman, 2010). In that sense, algae and cyanobacteria are exceptions as they have visible characteristic of spectral signature and microscopic morphological features (Sukenik et al., 2012). This property partly contributed to the increased number of records on invasion of cyanobacteria taxa, such as C. raciborskii (Padisák, 1997; Dyble et al., 2002; Wiedner et al., 2007; Vidal & Kruk, 2008; Sukenik et al., 2012; Wilk-Woźniak et al., 2016), Raphidiopsis mediterranea Skuja (Wilk-Woźniak et al., 2016), M. aeruginosa (Moreira et al., 2014), and some species of the genus Aphanizomenon (Sukenik et al., 2012). Sedimentary time series and large-scale monitoring records show that both the expansion and abundance of cyanobacteria has increased significantly in the last 200 years, and more rapidly in the last 70 years (Taranu et al., 2015). Among the well-studied cyanobacteria due to their invasion potential, the most important freshwater invasive cyanobacteria are the species C. raciborskii (already mentioned in the section on “Historical processes”) and to a lesser extent Aphanizomenon ovalisporum Forti, which are native to tropical regions and in the last two decades have expanded to subtropical and temperate areas (Kokociński et al., 2009; Kaštovský et al., 2010; for a detailed review on the invasiveness of these two taxa see Sukenik et al., 2012).
As already mentioned, C. raciborskii was an originally Pantropical species and since the first comprehensive review on its distributional area and dispersal was published (Padisák, 1997), this cyanobacterium was documented in most parts of the globe (Padisák et al. 2016). According Wilk-Wozniak et al. (2016), the distribution of the records of this species suggests the following possible migration scenarios: (1) from the center of Africa toward the north of the continent; (2) from North Africa to Europe; (3) from Java to eastern Australia; (4) from Java through South Asia to Europe and (5) from Australia to Southern Africa and on to the eastern parts of North, Central and South America. Kokocinski et al. (2017) have showed that, in Europe, the recent invasion of C. raciborskii in the East-Central regions may be related to environmental factors, mainly temperature-related variables. Today, C. raciborskii is find in many areas from Europe, such as Germany (Mischke, 2001; Stuken et al., 2006), France (Briand et al., 2002; Druart & Briand, 2002; Cellamare et al., 2010), and Poland (Stefaniak & Kokocinski, 2005; Kokocinski & Soininen, 2012; Kobos et al., 2013); and others Mediterranean/subtropical regions including Portugal (Saker et al., 2004), Algeria (Bouaicha & Nasri, 2004), Italy (Mugnai et al., 2008; Barone et al., 2010), Tunisia (Fathalli et al., 2010), Egypt (Hamed, 2005; Mohamed, 2007), and Israel (Zohary & Shlichter, 2009; Alster et al., 2010). Recent expansion of the species in South America (Vidal & Kruk, 2008; Fabre et al., 2010) and Africa (van Vuuren & Kriel, 2008) toward higher latitudes is also documented (Padisák et al. 2016). Several studies published in the literature concerns its invasiveness potential (e.g., Padisák 1997; Sukenik et al., 2012; Wilk-Woźniak et al. 2016), and part of this effort is due to the fact C. raciborskii is a potentially toxic and bloom-forming cyanobacterium. Australian strains, for example, are known to produce cylindrospermopsin, and Brazilian strains have been reported to produce paralytic shellfish poisoning toxins (Neilan et al., 2003).
Since the invasion must be initiated with dispersion to new zones (by aeolian transport, migrating animals, and also facilitated by human activities) and the subsequent establishment to the new environment, the invader needs a set of traits that support its establishment and proliferation (Sukenik et al., 2012). Regarding C. raciborskii, and also other Nostocales cyanobacteria, can be mentioned: the ability to form dormant cells (akinetes) that may survive long and extreme dispersion routes and survive on unfavorable conditions; and the ability to fix atmospheric nitrogen in the absence of combined inorganic sources, thus extending the spectrum of ecosystems to which they can invade (Sukenik et al., 2012). This species may also utilize other limiting resources, such as phosphorus, more efficiently than other cyanobacteria due to high affinity and P storage capacity (Isvánovics et al., 2000; Wu et al., 2012). Moreover, an important feature of Cylindrospermopsis is its wide thermal tolerance (Briand et al., 2004), which is essential to maintain the populations during cold winters. Padisák (1997), for instance, have reported that akinetes of C. raciborskii germinate at temperatures < 24°C. Finally, allelopathy (ability to synthesize allelo-chemicals that inhibit other phytoplankton species or deter and reduce grazing) was suggested as a beneficial trait of C. raciborskii that contributes to its stable dominance and geographic expansion (Fastner et al., 2007; Figueredo et al., 2007; Paerl et al., 2011).
Besides C. raciborskii, some other potentially invasive species of cyanobacteria are mentioned in the literature. For instance, dispersal of halophilic species in temperate waters has been facilitated by winter de-icing of roads and improper treatment of industrial sewage (Kaštovský et al., 2010). In that sense, it can be mentioned the cyanobacterium Cuspidothrix issatschenkoi (Usachev) P.Rajaniemi, Komárek, R.Willame, P. Hrouzek, K.Kastovská, L.Hoffmann & K.Sivonen (Padisák et al., 2016), whose expansion in the twentieth century might be enhanced by gradual adaptation to typical freshwater environments and the ongoing climate warming (Kaštovský et al., 2010). Also, Planktothrix rubescens is another cyanobacterium whose invasive potential has been described in the literature (Kaštovský et al., 2010; Padisák et al., 2016). This species had its original distribution area covering southern central Norway and the western alpine area, especially large lakes in Switzerland, Austria Italy, Germany, Slovenia, and France (e.g., Barco et al., 2004; Jann-Para et al., 2004; Jacquet et al., 2005; Legnani et al., 2005; Ernst et al., 2009). In addition to appears to proliferate within its original area (Jacquet et al., 2005), increasing number of reports provide evidence for its dispersal both southward and eastward to Spain (Almodóvar et al., 2004; Barco et al., 2004), Portugal (Paulino et al., 2009), Central Italy (Messineo et al., 2006), Sicily (Naselli-Flores et al., 2007; Naselli-Flores, 2014), Greece (Vareli et al., 2009), Turkey (Albay et al., 2003; Akcaalan et al., 2007), Poland (Krupa & Czernas, 2003; Lenard, 2009), and Hungary (Vasas et al., 2014) (Padisák et al. 2016).
One can not fail to mention the role of global climate change in cyanobacterial invasion patterns. This is because global warming leads to worldwide proliferation of cyanobacterial species, thus increasing its invasiveness potential, and consequently increasing blooms events. In addition to local environmental changes that can potentially increase cyanobacteria proliferation (e.g., nutrient overenrichment of waters), climate change is a potent catalyst for the further expansion of cyanobacterial blooms (Paerl & Huisman, 2008, 2009; Häder & Gao, 2015). Rising temperatures favor cyanobacteria in several ways, thus increasing its potential for invasiveness: (1) cyanobacteria generally grow better at higher temperatures (often above 25°C), giving a competitive advantage at elevated temperatures (Elliott et al., 2006; Jöhnk et al., 2008); (2) global warming causes lakes to stratify earlier in spring and de-stratify later in autumn, which lengthens optimal growth periods; and finally, (3) global warming affects patterns of precipitation and drought, for example, more intense precipitation will increase surface and groundwater nutrient discharge into water bodies (Paerl & Huisman, 2008).
Harmful blooms of toxic cyanobacteria are of particular interest and concern because of their economic and health consequences (Rastogi et al., 2015). Harmful blooms due to excessive growth of certain cyanobacteria followed by the production of toxic compounds have been reported in many eutrophic to hypertrophic lakes, ponds, and rivers throughout the world (Rastogi et al., 2015) Many of these bloom events, for instance, are of toxic Microcystis species and strains that proliferate under current environmental alterations, including nutrient enrichment, global warming, and regional hydrologic changes (Paerl & Paul, 2012). Microcystin is among the most commonly occurring toxin produced by cyanobacteria in natural waters (Babica et al., 2006; Rastogi et al., 2015), and can cause liver complications and damage to the nervous system if ingested (Bláha et al., 2009). Moreover, it has been suggested that UV-B radiation may significantly influence strain composition of cyanobacterial blooms in favor of microcystin (MC) producers (Ding et al., 2013).
Another cyanobacterium potentially harmful-bloom forming is the already mentioned C. raciborskii. In a modeling effort, for instance, Mehnert et al. (2010) have demonstrated that under a scenario of climate change with an increase of 4°C in the water temperature, C. raciborskii would outcompete a native species (Aphanizomenon gracile Lemmermann). Also, temperature-dependent release of cylindrospermopsin and microcystin has already been reported for A. ovalisporum (Cirés et al., 2014), Planktothrix agardhii (Walls et al., 2018) and Microcystis (Dziallas & Grossart, 2011), implicating management complications associated with global change scenarios. However, it is worth to mention that not all harmful blooms are associated with an invasion process, since many cyanobacterial species are of broad geographic distribution and rapidly respond to current environmental changes. Furthermore, although temperature (and global warming) are recognized as important factors leading to the proliferation of cyanobacteria, the exact mechanisms and the role of environmental factors regulating harmful blooms are disputable and yet to be understood, requiring more efforts both through field data and through experiments and modeling approaches.
Concluding remarks and future research perspectives
The central and global importance of microorganisms in the natural ecosystems throughout the world is obvious, however, despite the increase of studies and data, the application of ecological and evolutionary theories in microbial systems is still very limited. From the body of evidence collected by that time, it is known that a variety of nonmutually exclusive factors influence the distribution of microbial diversity on the planet, such as environmental selection (e.g., Mazard et al., 2012), allopatric speciation (e.g., Dvorák et al., 2012), and dispersal limitation (e.g., Miller et al., 2007), which during a long time was assumed did not exist for the microorganisms (Baas-Becking, 1934; Finlay, 2002; Fenchel & Finlay, 2004). Regarding cyanobacteria, there is also an increase of interest in phylogeographic and biogeographic studies, from approaches that go beyond ecological and taxonomic traditional studies from a solely microscopic approach. However, the studies still comprise a very small variety of habitats and taxonomic groups explored (Tables 1 and 2). Due to the highly diverse character of these group, from unicellular to multicellular organisms with differentiated cells (heterocyst and akinetes), with free-living and associated/symbiotic life forms, and present in the most diverse habitats of the Earth, interesting perspectives for future biogeographic studies emerge.
As expected from the Baas-Becking hypothesis, environmental selection is indeed found to have some influence on cyanobacterial distribution patterns of various taxonomic groups and habitats (Tables 1 and 2). The studies reported here show that both in global scales (e.g., Zwirglmaier et al., 2008) and at local scales (e.g., Chamberlain et al., 2014), environmental characteristics influence, at least in some degree, the distribution of cyanobacteria. The key question here is how distinct taxa, and how distinct taxonomic resolutions adopted to define a “taxon”, respond to environmental variation. We think, therefore, that future biogeographic studies within this section will benefit from exploring issues such as: (i) what is the ecological relevance of different levels of taxonomic resolution adopted for a given taxon?; (ii) what level of taxonomic resolution is required for the detection of ecotypes of a given species and how important are the environmental variables that potentially structure the distribution of ecotypes?; (iii) how do different taxonomic and trait-based groups respond to environmental variation?; (iv) how the cyanobacterial traits are distributed and responding to environmental variation?
On the influence of historical processes, recent studies with contradictory evidences about microbial cosmopolitism have been debating this issue, including studies with cyanobacteria taxa (Table 2). Thanks to previous studies, it is clear that dispersal capacity is dependent on several factors, such as the traits and dispersion strategies of give taxon may influence your establishment success (Hanson et al., 2012 and cited references). Akinetes-forming cyanobacteria, for instance, may have a greater dispersion capacity than those that do not form these structures because it is more likely to remain viable until arrival in a favorable habitat (in addition to decreasing the likelihood of extinction in front to environmental fluctuations) (see discussion on C. raciborskii in the sections above). In addition, the type of habitat may have some influence on the microbial dispersion. For example, taxa from sediment may have a lower dispersion capacity relative to taxa from surface soil in the same ecosystem or geographic area.
In the context of historical processes, therefore, interesting questions still need to be explored in the future, such as: (i) how different taxonomic groups (defined at different levels of taxonomic resolution) and trait-based present patterns of dispersion and gene flow? (ii) in fluid/continuous habitats (e.g., lakes, oceans) the rate of dispersion is greater than in island-like/poorly flowing habitats (e.g., periphyton, soil crusts)?; (iii) what is the role of population isolation, HGT and genetic drift in cyanobacterial distribution patterns at large spatial scales and at fine spatial scales (using target molecular markers for each purpose)?; (iv) what are the dispersive routes of different taxa and how do they vary over time? (that is, to apply molecular clocks in order to put the spatial distribution of different cyanobacteria taxa into a temporal framework).
Finally, to answer all these questions, in addition to studies with field data, both manipulative experimental tests and temporal studies are needed. Experimental manipulations have the potential to be useful tools for understanding how environmental and spatial factors influence the distribution of microbial taxa because microorganisms provide much better controlled and more flexible experimental systems for testing ecological theory than larger organisms (Bell, 2010). For instance, one experimental study has identified a specific temporal window during which the effects of dispersal limitation were apparent, although a subtle environmental variation operating over time scales of a few days can quickly overcome this dispersal limitation (Bell, 2010). Also, another experimental study showed that the slope of the taxa–area relationship for natural bacterial communities inhabiting small aquatic islands is comparable to that found for larger organisms (Bell et al., 2005). These approaches can provide important insights into biogeography of cyanobacteria, providing the opportunity of analyzing dispersion, HGT, and environmental selection more precisely: for instance, testing taxa–area and distance–decay relationships, correlations among abundance, genetic diversity, and ecotypes diversification, and the comparison of these patterns among different species and clades of cyanobacteria on controlled environmental conditions.
Conclusions
Among prokaryotic organisms, cyanobacteria are one of the most morpho-physiologically distinct groups (from unicellular to multicellular life forms; with or without differentiated cells; with toxic, invasive, and bloom-forming capacities; etc.), and this feature provides interesting approaches for biogeographic studies that are still little explored. In an overview, biogeography of cyanobacteria is still in its infancy, and generalizations cannot yet be made. On the one hand, considered (sub)cosmopolitan species may actually represent ecotypes with restricted distributions; on the other hand, species with restricted or endemic distributions may actually be the result of an underestimation of occurrence due to insufficient sampling effort. Due to the ecological, sanitary, and economic importance, bloom-forming and invasive toxic species are still the most studied, and of which there are more data available, both ecological and genomic, in the literature. Thus, making efforts to know and map the real diversity of cyanobacteria in the most diverse ecosystems is the first step to leverage the study of biogeography of this group. The exploration of little-known geographic areas (e.g., subtropical environments) and habitats (e.g., terrestrial, microbial mats), as well as sampling and sequencing genomes from cyanobacteria of lesser-known taxa, is very much needed to improve our understanding on the biogeography of cyanobacteria.
So far, the fact that there are some cyanobacteria species with broad distributions and others with possible endemic distributions implies the nonuniformity of the responses of different taxa to ecological/evolutionary processes. To what extent these distribution patterns are dependent on species concept, level of taxonomic resolution, spatial and environmental scales, and the biases of the molecular methodologies applied in the studies, are key issues to be explored in future studies. We suggest that to answer these questions adequately, it is necessary to use multiple taxonomic definitions based on a variety of genetic markers (and biochemical and morphological characteristics, more easily accessible for cyanobacteria), and subsequent testing of hypothesis in the light of ecological and evolutionary theories. It should also be noted that taxonomic identification based on molecular methods, although has higher detectability of variation in biodiversity, actually provides data for the investigation of biogeographic patterns of microbial genes, not whole organisms. Therefore, another possible approach is to compare biogeographic patterns of cyanobacterial genes with whole genomes (already available for some cyanobacterial species) and microscopic-based definition, by analyzing how distribution patterns depend on the definitions of cyanobacterial species adopted.
References
Achtman, M. & M. Wagner, 2008. Microbial diversity and the genetic nature of microbial species. Nature Reviews Microbiology 6: 431–440.
Agawin, N. S. R., S. Rabouille, M. Veldhuis, L. Servatius, S. Hol, H. M. J. van Overzee & J. Huisman, 2007. Competition and facilitation between unicellular nitrogen-fixing cyanobacteria and non–nitrogen-fixing phytoplankton species. Limnology and Oceanography 52: 2233–2248.
Ahlgren, N. A. & G. Rocap, 2006. Culture isolation and culture-independent clone libraries reveal new marine Synechococcus ecotypes with distinctive light and N physiologies. Applied and Environmental Microbiology 72: 7193–7204.
Ahlgren, A. A. & G. Rocap, 2012. Diversity and distribution of marine Synechococcus: multiple gene phylogenies for consensus classification and development of qPCR assays for sensitive measurement of clades in the ocean. Frontiers in Microbiology 3: 1–24.
Akcaalan, R., M. Albay, C. Gürevin & F. C. Evik, 2007. The influence of environmental conditions on the morphological variability of phytoplankton in an oligo-mesotrophic Turkish lake. Annales de Limnologie- International Journal of Limnology 43: 21–28.
Albay, M., R. Akcaalan, H. Tufekci, J. S. Metcalf, K. A. Beattie & G. A. Codd, 2003. Depth profiles of cyanobacterial hepatotoxins (microcystins) in three Turkish freshwater lakes. Hydrobiologia 505: 89–95.
Almodóvar, A., G. G. Nicola & M. Nuevo, 2004. Effects of a bloom of Planktothrix rubescens on the fish community of a Spanish reservoir. Limnetica 23: 167–178.
Alster, A., R. N. Kaplan-Levy, A. Sukenik & T. Zohary, 2010. Morphology and phylogeny of a non-toxic invasive Cylindrospermopsis raciborskii from a Mediterranean Lake. Hydrobiologia 639: 115–128.
Alvarenga, D. O., M. F. Fiore & A. M. Varani, 2017. A metagenomic approach to cyanobacterial genomics. Frontiers in Microbiology 8: e809.
Apple, J. K., S. L. Strom, B. Palenik & B. Bianca, 2011. Variability in protist grazing and growth on different marine Synechococcus isolates. Applied and Environmental Microbiology 77: 3074–3084.
Baas-Becking, L. G. M., 1934. Geobiologie of Inleiding Tot de Milieukunde. W.P. Van Stockum & Zoon, The Hague.
Babica, P., J. Kohoutek, L. Bláha, O. Adamovský & B. Maršálek, 2006. Evaluation of extraction approaches linked to ELISA and HPLC for analyses of microcystin-LR, -RR and -YR in freshwater sediments with different organic material contents. Analytical and Bioanalytical Chemistry 385: 1545–1551.
Bahl, J., M. C. Y. Lau, G. J. D. Smith, et al., 2011. Ancient origins determine global biogeography of hot and cold desert cyanobacteria. Nature Communications 2: 163.
Barco, M., C. Flores, J. Rivera & J. Caixach, 2004. Determination of microcystin variants and related peptides present in a water bloom of Planktothrix (Oscillatoria) rubescens in a Spanish drinking water reservoir by LC/ESI-MS. Toxicon 44: 881–886.
Barone, R., G. Castelli & L. Naselli-Flores, 2010. Red sky at night cyanobacteria delight: the role of climate in structuring phytoplankton assemblage in a shallow, Mediterranean lake (Biviere di Gela, southeastern Sicily). Hydrobiologia 639: 43–53.
Bell, T., 2010. Experimental tests of the bacterial distance-decay relationship. The ISME Journal 4: 1357–1365.
Bell, T., D. Ager, J. Song, J. A. Newman, I. P. Thompson, A. K. Lilley & C. J. van der Gast, 2005. Larger islands house more bacterial taxa. Science 308: 1884.
Beltran, E. C. & B. A. Neilan, 2000. Geographical segregation of the neurotoxin-producing cyanobacterium Anabaena circinalis. Applied and Environmental Microbiology 66: 4468–4474.
Berg, K. A. C., K. Lyra, L. Sivonen, S. Paulin, P. Tuomi Suomalainen & J. Rapala, 2009. High diversity of cultivable heterotrophic bacteria in association with cyanobacterial water blooms. The ISME Journal 3: 314–325.
Bittencourt-Oliveira, M. C., M. C. Oliveira & C. J. S. Bolch, 2001. Genetic variability of Brazilian strains of the Microcystis aeruginosa complex (Cyanobacteria/Cyanophyceae) using the phycocyaninin tergenic spacer and flanking regions (cpcBA). Journal of Phycology 37: 810–818.
Bláha, L., P. Babica & B. Maršálek, 2009. Toxins produced in cyanobacterial water blooms—toxicity and risks. Interdisciplinary Toxicology 2: 36–41.
Boileau, M. G., P. D. N. Hebert & S. S. Schwartz, 1992. Nonequilibrium gene frequency divergence—persistent founder effects in natural-populations. Journal of Evolutionary Biology 5: 25–39.
Borics, G., I. Grigorszky, S. Szabó & J. Padisák, 2000. Phytoplankton associations in a small hypertrophic fishpond in East Hungary during a change from bottom-up to top-down control. Hydrobiologia 424: 79–90.
Borics, G., B. Tóthmérész, B. A. Lukács & G. Várbíró, 2012. Functional groups of phytoplankton shaping diversity of shallow lake ecosystems. Hydrobiologia 698: 251–262.
Bouaicha, N. & A. B. Nasri, 2004. First report of cyanobacterium Cylindrospermopsis raciborskii from Algerian freshwaters. Environmental Toxicology 19: 541–543.
Brandao, P. F. B., J. P. Clapp & A. T. Bull, 2002. Discrimination and taxonomy of geographically diverse strains of nitrile-metabolizing actinomycetes using chemometric and molecular sequencing techniques. Environmental Microbiology 4: 262–276.
Briand, J. F., C. Robillot, C. Quiblier-Lloberas, J. F. Humbert, A. Coute & C. Bernard, 2002. Environmental context of Cylindrospermopsis raciborskii (Cyanobacteria) blooms in a shallow pond in France. Water Research 36: 3183–3192.
Briand, J. F., C. Leboulanger, J. F. Humbert, C. Bernard & P. Dufour, 2004. Cylindrospermopsis raciborskii (Cyanobacteria) invasion at mid-latitudes: selection, wide physiological tolerance, or global warming? Journal of Phycology 40: 231–238.
Capone, D. G., J. P. Zehr, H. W. Paerl, B. Bergman & E. J. Carpenter, 1997. Trichodesmium, a globally significant marine cyanobacterium. Science 276: 1221–1229.
Carmichael, W. W., 2001. Health effects of toxin producing cyanobacteria: the “CyanoHABs”. Human and Ecological Risk Assessment 7: 1393–1407.
Caro-Quintero, A. & K. T. Konstantinidis, 2011. Bacterial species may exist, metagenomics reveal. Environmental Microbiology 14: 347–355.
Castenholz, R. W., 2001. Phylum BX. Cyanobacteria. Oxygenic Photosynthetic Bacteria. Springer-Verlag, New York: 473–599.
Cirés, S., L. Wörmer, A. Ballot, R. Agha, C. Wiedner, D. Velázquez, M. C. Casero & A. Quesada, 2014. Phylogeography of cylindrospermopsin and paralytic shellfish toxin-producing Nostocales cyanobacteria from Mediterranean Europe (Spain). Applied and Environmental Microbiology 80: 1359–1370.
Cellamare, M., M. Leitão, M. Coste, A. Dutartre & J. Haury, 2010. Tropical phytoplankton taxa in Aquitaine lakes (France). Hydrobiologia 639: 129–145.
Chamberlain, S. D., K. A. Kaplan, M. Modanu, K. M. Sirianni, S. Annandale & I. Hewson, 2014. Biogeography of planktonic and benthic cyanobacteria in coastal waters of the Big Island, Hawai’i. FEMS Microbiology Ecology 89: 80–88.
Chen, F., K. Wang, J. Kan, M. T. Suzuki & K. E. Wommack, 2006. Diverse and Unique Picocyanobacteria in Chesapeake Bay, Revealed by 16S-23S rRNA Internal Transcribed Spacer Sequences. Applied and Environmental Microbiology 72: 2239–2243.
Cho, J. C. & J. M. Tiedje, 2000. Biogeography and degree of endemicity of fluorescent Pseudomonas strains in soil. Applied and Environmental Microbiology 66: 5448–5456.
Chorus, I. & J. Bartam, 1999. Toxic cyanobacteria in water—a guide to their public health consequences. Monitoring and Management. E & FN Spon, London.
Colautti, R. I. & Hugh J. MacIsaac, 2004. A neutral terminology to define ‘invasive’ species. Diversity and Distributions 10: 135–141.
Comte, K., M. Sabacka, A. Carré-Mlouka, J. Elster & J. Komárek, 2007. Relationships between the Arctic and the Antarctic cyanobacteria; three Phormidium-like strains evaluated by a polyphasic approach. FEMS Microbiology Ecology 59: 366–376.
Couté, A. & M. Bouvy, 2004. A new species of the genus Cylindrospermopsis, C. acuminato-crispa spec.nova (Cyanophyceae, Nostocales) from Ingazeira reservoir, Northeast Brazil. Algological Studies 113: 57–72.
Couté, A., M. Leitao & H. Sarmento, 2004. Cylindrospermopsis sinuosa spec.nova (Cyanophyceae, Nostocales), une nuovelle espece du sud-ouest de la France. Algological Studies 111: 1–15.
Curtis, T. P., W. T. Sloan & J. W. Scannell, 2002. Estimating prokaryotic diversity and its limits. PNAS 99: 10494–10499.
de Figueiredo, D. R., A. Reboleira, S. C. Antunes, N. Abrantes, U. Azeiteiro, et al., 2006. The effect of environmental parameters and cyanobacterial blooms on phytoplankton dynamics of a Portuguese temperate lake. Hydrobiologia 568: 145–157.
Dijoux, L., F. Viard & C. Payri, 2014. The more we search, the more we find: discovery of a new lineage and a new species complex in the genus Asparagopsis. PLoS ONE 9: e103826.
Ding, Y., L. Song & B. Sedmak, 2013. UVB radiation as a potential selective factor favoring microcystin producing bloom forming cyanobacteria. PLoS ONE 8: e73919.
Dobzhansky, T. G., 1937. Genetics and the Origin of Species. Columbia University Press, Columbia.
Drakare, S. & A. Liess, 2010. Local factors control the community composition of cyanobacteria in lakes while heterotrophic bacteria follow a neutral model. Freshwater Biology 55: 2447–2457.
Druart, J. C. & J. F. Briand, 2002. First record of Cylindrospermopsis raciborskii (Woloszynska) Seenayya et Subba Raju (Cyanobacteria) in a lotic system in France. Annales de Limnologie. International Journal of Limnology 38: 339–342.
Dufour, P., G. Sarazin, C. Quiblier, S. Sane & C. Leboulanger, 2006. Cascading nutrient limitation of the cyanobacterium Cylindrospermopsis raciborskii in a Sahelian lake (North Senegal). Aquatic Microbial Ecology 44: 219–230.
Dvořák, P., P. Hašler & A. Poulíčková, 2012. Phylogeography of the Microcoleus vaginatus (Cyanobacteria) from three continents—a spatial and temporal characterization. Plos ONE 7: e40153.
Dyble, J., H. W. Paerl & B. A. Neilan, 2002. Genetic characterization of Cylindrospermopsis raciborskii (Cyanobacteria) isolates from diverse geographic origins based on nifH and cpcBA-IGS nucleotide sequence analysis. Applied and Environmental Microbiology 68: 2567–2571.
Dziallas, C. & H. Grossart, 2011. Increasing Oxygen Radicals and Water Temperature Select for Toxic Microcystis sp. PLoS ONE 6: e25569.
El Herry, S., H. Nasri & N. Bouaïcha, 2008. Morphological and phylogenetic analysis of colonies of Microcystis morphospecies isolated from the Lebna Dam, Tunisia. African Journal of Microbiology Research 2: 340–348.
Elliott, J. A., I. D. Jones & S. J. Thackeray, 2006. Testing the sensitivity of phytoplankton communities to changes in water temperature and nutrient load, in a temperate lake. Hydrobiologia 559: 401–411.
Elton, C. S., 1958. The ecology of invasions by animals and plants. London: Methuen. Progress in Physical Geography 31: 659–666.
Ernst, A., S. Becker, U. I. A. Wollenzien & C. Postius, 2003. Ecosystem-dependent adaptive radiations of picocyanobacteria inferred from 16S rRNA and ITS-1 sequence analysis. Microbiology 149: 217–228.
Ernst, B., S. J. Hoeger, E. O’Brien & D. R. Dietrich, 2009. Abundance and toxicity of Planktothrix rubescens in the pre-Alpine Lake Ammersee, Germany. Harmful Algae 8: 329–342.
Erwin, P. M. & R. W. Thacker, 2008. Cryptic diversity of the symbiotic cyanobacterium Synechococcus spongiarum among sponge hosts. Molecular Ecology 17: 2937–2947.
Fabre, A., C. Carballo, E. Hernandez, P. Piriz, L. Bergamio, L. Mwllo, S. Gonzalez, G. Pérez, J. G. León, S. Bonilla & C. Kruk, 2010. El nitrogeno y la relación zona eufótica de mezcia explican la presencia de cianobacterias en pequenos lagos subtropicales, artificiales de Uruguay. Pan- Ameriacan Journal of Aquatic Sciences 5: 112–125.
Fastner, J., J. Rücker, A. Stüken, K. Preussel, B. Nixdorf, I. Chorus, A. Köhler & C. Wiedner, 2007. Occurrence of the cyanobacterial toxin cylindrospermopsin in northeast Germany. Environmental Toxicology 22: 26–32.
Fathalli, A., A. B. Jenhani, S. Cires, M. Saker, M. Romdhane & V. Vasconcelos, 2010. First observation of the potentially toxic and invasive cyanobacterium Cylindrospermopsis raciborskii in Tunisan freshwaters: toxicity assessement and molecular characterization. Fresenius Environmental Bulletin 19: 1074–1083.
Fathalli, A., A. B. R. Jenhania, C. Moreira, M. Welker, M. Romdhane, A. Antunes & V. Vasconcelos, 2011. Molecular and phylogenetic characterization of potentially toxic cyanobacteria in Tunisian freshwaters. Systematic and Applied Microbiology 34: 303–310.
Fenchel, T. & B. J. Finlay, 2004. The ubiquity of small species: patterns of local and global diversity. BioScience 54: 777–784.
Figueredo, C. C., A. Giani & D. F. Bird, 2007. Does allelopathy contribute to Cylindrospermopsis raciborskii (cyanobacteria) bloom occurrence and geographic expansion? Journal of Phycology 43: 256–265.
Finlay, B. J., 2002. Global dispersal of free-living microbial eukaryote species. Science 296: 1061–1063.
Frangeul, L., P. Quillardet, A. M. Castets, J. F. Humbert, H. C. P. Matthijs, et al., 2008. Highly plastic genome of Microcystis aeruginosa PCC 7806, a ubiquitous toxic freshwater cyanobacterium. BMC Genomics 9: 274.
Fraser, C., E. J. Alm, M. F. Polz, B. G. Spratt & W. P. Hanage, 2009. The bacterial species challenge: making sense of genetic and ecological diversity. Science 323: 741–746.
Fuller, N. J., M. Dominique, P. Frédéric, D. Vaulot, A. F. Post & D. J. Scanlan, 2003. Clade-specific 16S ribosomal DNA oligonucleotides reveal the predominance of a single Marine Synechococcus clade throughout a stratified water column in the Red Sea. Applied and Environmental Microbiology 69: 2430–2443.
Gaevsky, N. A., V. I. Kolmakov, O. I. Belykh, I. V. Tikhonova, Y. Joung, T. S. Ahn, V. A. Nabatova & A. S. Gladkikh, 2011. Ecological development and genetic diversity of Microcystis aeruginosa from artificial reservoir in Russia. The Journal of Microbiology 49: 714–720.
Gaget, V., S. Gribaldo & N. T. de Marsac, 2011. An rpoB signature sequence provides unique resolution for the molecular typing of cyanobacteria. International Journal of Systematic and Evolutionary Microbiology 61: 170–183.
Gantar, M. & Z. Svircěv, 2008. Microalgae and cyanobacteria: food for thought. Journal of Phycology 44: 260–268.
Garcia-Pichel, F., L. Prufert-Bebout & G. Muyzer, 1996. Phenotypic and phylogenetic analyses show Microcoleus chthonoplastes to be a cosmopolitan cyanobacterium. Applied and Environmental Microbiology 62: 3284–3291.
Garcia-Pichel, F., A. López-Cortés & U. Nübel, 2001. Phylogenetic and morphological diversity of cyanobacteria in soil desert crusts from the Colorado Plateau. Applied and Environmental Microbiology 67: 1902–1910.
Gerwick, W. H., R. C. Coates, N. Engene, L. Gerwick, R. V. Grindberg, A. C. Jones & C. M. Sorrels, 2008. Giant marine cyanobacteria produce exciting potential pharmaceuticals. Microbe 3: 277–284.
Graham, L. E., J. M. Graham & L. W. Wilcox, 2009. Algae, 2nd ed. Pearson Benjamin Cummings, San Francisco.
Gugger, M., R. Molica, B. Le Berre, P. Dufour, C. Bernard & J.-F. Humbert, 2005. Genetic diversity of Cylindrospermopsis strains (Cyanobacteria) isolated from four continents. Applied and Environmental Microbiology 71: 1097–1100.
Guiry, M. D., 2012. How many species of algae are there? Journal of Phycology 48: 1057–1063.
Haande, S., A. Ballot, T. Rohrlack, J. Fastner, C. Wiedner & B. Edvardsen, 2007. Diversity of Microcystis aeruginosa strains (Chroococcales, Cyanobacteria) from East-African waterbodies. Archives of Microbiology 188: 15–25.
Häder, D. P. & K. Gao, 2015. Interactions of anthropogenic stress factors on marine phytoplankton. Frontiers in Environmental Science 3: e1.
Hagstrom, A., J. Pinhassi & U. L. Zweifel, 2000. Biogeographical diversity among marine bacterioplankton. Aquatic Microbial Ecology 21: 231–244.
Hamed, A. F., 2005. Survey of distribution and diversity of bluegreen algae (Cyanobacteria) in Egypt. Acta Botanica Hungarica 47: 117–136.
Hanson, C. A., J. A. Fuhrman, M. C. Horner-Devine & J. B. H. Martiny, 2012. Beyond biogeographic patterns: processes shaping the microbial landscape. Nature Reviews 10: 497–506.
Harris, J. M., P. Vinobaba, R. K. A. Kularatne & C. E. Kankanamge, 2016. Spatial and temporal distribution of cyanobacteria in Batticaloa Lagoon. Journal of Environmental Sciences 47: 211–218.
Hedlund, B. P. & J. T. Staley, 2004. Microbial endemism and biogeography. In Bull, A. T. (ed.), Microbial biodiversity and bioprospecting. ASM Press, Washington D.C.: 225–231.
Hillebrand, H., F. Watermann, R. Karez & U. G. Berninger, 2001. Differences in species richness patterns between unicellular and multicellular organisms. Oecologia 126: 114–124.
Hoffmann, L., 1996. Geographic distribution of freshwater bluegreen algae. Hydrobiologia 336: 33–40.
Hong, Y., M. A. Burford, P. J. Ralph, J. W. Udy & M. A. Doblin, 2013. The cyanobacterium Cylindrospermopsis raciborskii is facilitated by copepod selective grazing. Harmful Algae 29: 14–21.
Horner-Devine, M. C., M. Lage, J. B. Hughes & B. J. M. Bohannan, 2004. A taxa–area relationship for bacteria. Nature 432: 750–753.
Huang, S., S. W. Wilhelm, H. R. Harvey, K. Taylor, N. Jiao & C. Feng, 2012. Novel lineages of Prochlorococcus and Synechococcus in the global oceans. The ISME Journal 6: 285–297.
Hug, L. A., B. J. Baker, K. Anantharaman, et al., 2016. A new view of the tree of life. Nature Microbiology 1: 1–6.
Ionescu, D., M. Hindiyeh, H. Malkawi & A. Oren, 2010. Biogeography of thermophilic cyanobacteria: insights from the Zerka Ma’in hot springs (Jordan). FEMS Microbiology Ecology 72: 103–113.
Isvánovics, V., H. M. Shafik, M. Présing & S. Juhos, 2000. Growth and phosphate uptake kinetics of the cyanobacterium, Cylindrospermopsis raciborskii (Cyanophyceae) in throughfl ow cultures. Freshwater Biology 43: 257–275.
Jacquet, S., J.-F. Briand, C. Leboulanger, C. Avois-Jacquet, L. Oberhaus, B. Tassin, B. Vincon-Leite, G. Paolini, J.-C. Druart, O. Anneville & J.-F. Humberta, 2005. The proliferation of the toxic cyanobacterium Planktothrix rubescens following restoration of the largest natural French lake (Lac du Bourget). Harmful Algae 4: 651–672.
Jann-Para, G., I. Schwob & M. Feuillade, 2004. Occurrence of toxic Planktothrix rubescens blooms in lake Nantua, France. Toxicon 43: 279–285.
Jöhnk, K. D., J. Huisman, J. Sharples, B. Sommeijer, P. M. Visser & J. M. Stroom, 2008. Summer heatwaves promote blooms of harmfull cyanobacteria. Global Change Biology 14: 495–512.
Jungblut, A., I. Hawes, D. Mountfort, B. Hitzfeld, D. R. Dietrich, B. P. Burns & B. A. Neilan, 2005. Diversity within cyanobacterial mat communities in variable salinity meltwater ponds of McMurdo Ice Shelf, Antarctica. Environmental Microbiology 7: 519–529.
Jungblut, A. D., C. Lovejoy & W. F. Vincent, 2010. Global distribution of cyanobacterial ecotypes in the cold biosphere. The ISME Journal 4: 191–202.
Kardinaal, W. E. A., L. Tonk, I. Janse, S. Hol, P. Slot, J. Huisman & P. M. Visser, 2007. Competition for light between toxic and nontoxic strains of the harmful cyanobacterium Microcystis. Applied and Environmental Microbiology 73: 2939–2946.
Kaštovský, J., T. Hauer, J. Mareš, M. Krautová, T. Bešta, J. Komárek, J. Desortová, J. Heteša, A. Hindáková, V. Houk, E. Janeček, R. Kopp, P. Marvan, P. Pumann, O. Skácelová & E. Zapomelová, 2010. A review of the alien and expansive species of freshwater cyanobacteria and algae in the Czech Republic. Biological Invasions 12: 3599–3625.
Kobos, J., A. Błaszczyk, N. Hohlfeld, A. Toruńska-Sitarz, A. Krakowiak, A. Hebel, K. Sutryk, M. Grabowska, M. Toporowska, M. Kokocinski, B. Messyasz, A. Rybak, A. Napiorkowska Krzebietke, L. Nawrocka, A. Pelechata, A. Budzynska, P. Zagajewski & H. Mazur-Marzec, 2013. Cyanobacteria and cyanotoxins in Polish freshwater bodies. Oceanological and Hydrobiological Studies 42: 358–378.
Kokociński, M. & J. Soininen, 2012. Environmental factors related to the occurrence of Cylindrospermopsis raciborskii (Nostocales, Cyanophyta) at the north-eastern limit of its geographical range. European Journal of Phycology 47: 12–21.
Kokociński, M., D. Dziga, L. Spoof, K. Stefaniak, T. Jurczak, J. Mankiewicz-Boczek & J. Meriluoto, 2009. First report of the cyanobacterial toxin cylindrospermopsin in the shallow, eutrophic lakes of western Poland. Chemosphere 74: 669–675.
Kokociński, M., I. Gągała, I. Jasser, J. Karosienė, J. Kasperovičienė, J. Kobos, et al., 2017. Distribution of invasive Cylindrospermopsis raciborskii in the East-Central Europe is driven by climatic and local environmental variables. FEMS Microbiology Ecology 93: 1–8.
Komárek, J., 1985. Do all cyanophytes have a cosmopolitan distribution? Survey of freshwater cyanophyte flora of Cuba. Algological Studies 75: 11–29.
Komárek, J., 2006. Cyanobacterial taxonomy: current problems and prospects for the integration of traditional and molecular approaches. Algae 21: 349–375.
Komárek, J., 2016. Review of the cyanobacterial genera implying planktic species after recent taxonomic revisions according to polyphasic methods: state as of 2014. Hydrobiologia 764: 259–270.
Komárek, J. & J. Komárková-Legnerová, 2002. Contribution to the knowledge of planktic cyanoprokaryotes from central Mexico. Preslia 74: 207–233.
Komárek, J. & J. Mareš, 2012. An update to modern taxonomy (2011) of freshwater planktic heterocytous cyanobacteria. Hydrobiologia 698: 327–351.
Komárek, J., J. Kaštovský, J. Mareš & J. R. Johansen, 2014. Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) using a polyphasic approach. Preslia 86: 295–335.
Komárková-Legnerová, J. & R. Tavera, 1996. Cyanoprokaryota (Cyanobacteria) in the phytoplankton of Lake Catemaco (Veracruz, Mexico). Algological Studies 83: 403–422.
Kondo, R., G. Kagiya, S. Hiroishi & M. Watanabe, 2000. Genetic typing of a bloom-forming cyanobacterial genus Microcystis in Japan using 16S rRNA gene sequence analysis. Plankton Biology and Ecology 47: 1–6.
Konstantinidis, K. T. & J. M. Tiedje, 2005. Genomic insights that advance the species definition for prokaryotes. PNAS 102: 2567–2572.
Krupa, D. & K. Czernas, 2003. Mass appearance of cyanobacterium Planktothrix rubescens in Lake Piaseczno, Poland. Water Quality Research Journal of Canada 38: 141–152.
Kurmayer, R. & M. Gumpenberger, 2006. Diversity of microcystin genotypes among populations of the filamentous cyanobacteria Planktothrix rubescens and Planktothrix agardhii. Molecular Ecology 15: 3849–3861.
Land, M., L. Hauser, S. R. Jun, I. Nookaew, M. R. Leuze, T. H. Ahn, et al., 2015. Insights from 20 years of bacterial genome sequencing. Functional & Integrative Genomics 15: 141–161.
Larsson, J., J. A. A. Nylander & B. Bergman, 2011. Genome fluctuations in cyanobacteria reflect evolutionary developmental and adaptive traits. BMC Evolutionary Biology 11: 187.
Legnani, E., D. Copetti, A. Oggioni, G. Tartari, M.-T. Palumbo & G. Morabito, 2005. Planktothrix rubescens’ seasonal dynamics and vertical distributionin Lake Pusiano, North Italy. Journal of Limnology 64: 61–73.
Lenard, T., 2009. Metalimnetic bloom of Planktothrix rubescens in relation to environmental conditions. Oceanological and Hydrobiological Studies 38: 45–53.
Lepère, C., A. Wilmotte & B. Meyer, 2000. Molecular diversity of Microcystis strains (Cyanophyceae, Chroococcales) based on 16S rDNA sequences. Systematics and Geography of Plants 70: 275–283.
Li, W. K. W., 1994. Primary production of prochlorophytes, cyanobacteria, and eucaryotic ultraphytoplankton: measurements from flow cytometric sorting. Limnology and Oceanography 39: 169–175.
Lin, S., S. Hass, T. Zemojtek, P. Xiao, M. Vingron & R. Li, 2010. Genome wide comparison of cyanobacterial transposable elements, potential genetic diversity indicators. Gene 473: 139–149.
Litchman, E., 2010. Invisible invaders: non-pathogenic invasive microbes in aquatic and terrestrial ecosystems. Ecology Letters 13: 1560–1572.
Litvaitis, M. K., 2002. A molecular test of cyanobacterial phylogeny: inferences from constraint analyses. Hydrobiologia 468: 135–145.
Liu, H., H. A. Nolla & L. Campbell, 1997. Prochlorococcus growth rate and contribution to primary production in the equatorial and subtropical North Pacific Ocean. Aquatic Microbial Ecology 12: 39–47.
Locey, K. J. & J. T. Lennon, 2016. Scaling laws predict global microbial diversity. Proceedings of the National Academy of Sciences 113: 5970–5975.
Lu, X., C. Tian, H. Pei, W. Hu & J. Xie, 2013. Environmental factors influencing cyanobacteria community structure in Dongping Lake, China. Journal of Environmental Sciences 25: 2196–2206.
Lyons, T. W., C. T. Reinhard & N. J. Planavsky, 2014. The rise of oxygen in Earth’s early ocean and atmosphere. Nature 506: 307–315.
Marěs, J., 2017. Multilocus and SSU rRNA gene phylogenetic analyses of available cyanobacterial genomes, and their relation to the current taxonomic system. Hydrobiologia 811: 19–34.
Margulis, L., 1970. Origin of Eukaryotic Cells. Yale University Press, New Haven.
Marmen, S., D. Aharonovich, M. Grossowicz, L. Blank, Y. Z.Yacobi & D. J. Sher, 2006. Distribution and habitat specificity of potentially-toxic microcystis across climate, land, and water use gradients. Frontiers in Microbiology 7: 271.
Marmen, S., D. Aharonovich, M. Grossowicz, L. Blank, Y. Z. Yacobi & D. J. Sher, 2016. Distribution and Habitat Specificity of Potentially-Toxic Microcystis across Climate, Land, and Water Use Gradients. Frontiers in Microbiology 7: e271.
Martiny, J. B. H., B. J. M. Bohannan, J. H. Brown, et al., 2006. Microbial biogeography: putting microorganisms on the map. Nature Reviews 4: 102–112.
Martiny, A. C., A. P. K. Tai, D. Veneziano, F. Primeau & S. W. Chisholm, 2009. Taxonomic resolution, ecotypes and the biogeography of Prochlorococcus. Environmental Microbiology 11: 823–832.
Mazard, S., M. Ostrowski, F. Partensky & D. J. Scanlan, 2012. Multi-locus sequence analysis, taxonomic resolution and biogeography of marine Synechococcus. Environmental Microbiology 14: 372–386.
McFadden, G. I., 2014. Origin and evolution of plastids and photosynthesis in eukaryotes. Cold Spring Harbor Perspectives Biology 6: a016105.
McNeill, J., F. R. Barrie, W. R. Buck, V. Demoulin, W. Greuter, D. L. Hawksworth, et al., 2012. International Code of Nomenclature of algae, fungi, and plants (Melbourne Code). Regnum Vegetabile 154: 208.
Mehnert, G., F. Leunert, S. Cirés, K. D. Jöhnk, J. Rücker, B. Nixdorf & C. Wiedner, 2010. Competitiveness of invasive and native cyanobacteria from temperate freshwaters under various light and temperature conditions. Journal of Plankton Research 32: 1009–1021.
Messineo, V., D. Mattei, S. Melchiorre, G. Salvatore, S. Bogialli, R. Salzano, R. Mazza, G. Capelli & M. Bruno, 2006. Microcystin diversity in a Planktothrix rubescens population from Lake Albano (Central Italy). Toxicon 48: 160–174.
Michaud, A. B., M. Čabacká & J. C. Priscu, 2012. Cyanobacterial diversity across landscape units in a polar desert: Taylor Valley, Antarctica. FEMS Microbiology Ecology 82: 268–278.
Miller, S. R., R. W. Castenholz & D. Pedersen, 2007. Phylogeography of the thermophilic cyanobacterium Mastigocladus laminosus. Applied and Environmental Microbiology 73: 4751–4759.
Miller, S. R., A. L. Strong, K. L. Jones & M. C. Ungerer, 2009. Bar-coded pyrosequencing reveals shared bacterial community properties along the temperature gradients of two alkaline hot springs in Yellowstone National Park. Applied and Environmental Microbiology 75: 4565–4572.
Mischke, U., 2001. Der Neophyt Cylindrospermopsis raciborskii: eine Blaualge aus tropischen Regionen in Gewässern des Spree-Dahme Einzugsgebietes. In Krumbeck, H. & U. Mischke (eds), Gewässerreport (Nr. 6): Entwicklungen der Gewässer im Scharmützelseegebiet und angewandte Probleme des Gewässerschutzes. BTUCAR 6/2001, ISSN 1434-6834.
Mohamed, Z. A., 2007. First report of toxic Cylindrospermopsis raciborskii and Raphidiopsis mediterranea (Cyanoprokaryota) in Egyptian fresh waters. FEMS Microbial Ecology 59: 749–761.
Moreira, C., A. Fathalli, V. Vasconcelos & A. Antunes, 2011. Genetic diversity and structure of the invasive toxic cyanobacterium Cylindrospermopsis raciborskii. Current Microbiology 62: 1590–1595.
Moreira, C., V. Vasconcelos & A. Antunes, 2012. Phylogeny of microcystins: evidence of a biogeographical trend? Current Microbiology 66: 214–221.
Moreira, C., V. Vasconcelos & A. Antunes, 2013. Phylogeny and biogeography of Cyanobacteria and their produced toxins. Marine Drugs 11: 4350–4369.
Moreira, C., C. Spillane, A. Fathalli, V. Vasconcelos & A. Antunes, 2014. African Origin and Europe-Mediated Global Dispersal of The Cyanobacterium Microcystis aeruginosa. Current Microbiology 69: 328–633.
Morlon, H., R. Chuyong, R. Condit, S. Hubbell, D. Kenfack, D. Thomas, R. Thomas & J. L. Green, 2008. A general framework for the distance-decay of similarity in ecological communities. Ecology Letters 11: 904–917.
Mowe, M. A. D., S. M. Mitrovic, R. P. Lim, A. Furey & D. C. J. Yeo, 2015. Tropical cyanobacterial blooms: a review of prevalence, problem taxa, toxins and influencing environmental factors. Journal of Limnology 74: 205–224.
Mugnai, M. A., M. C. Margheri, C. Sili, S. Turicchia, E. Soldati, E. Maffettone, E. Funari, S. Scardala, M. Di Brizio & S. Ventura, 2008. The cyanobacterial community of Lake Trasimeno. Algological Studies 128: 37–64.
Mühling, M., N. J. Fuller, A. Millard, et al., 2005. Genetic diversity of marine Synechococcus and co-occurring cyanophage communities: evidence for viral control of phytoplankton. Environmental Microbiology 7: 499–508.
Nabout, J. C., B. S. Rocha, F. M. Carneiro & C. L. Sant’Anna, 2013. How many species of Cyanobacteria are there? Using a discovery curve to predict the species number. Biodiversity and Conservation 22: 2907–2918.
Namsaraev, Z., M. J. Mano, R. Fernandez & A. Wilmotte, 2010. Biogeography of terrestrial cyanobacteria from Antarctic ice-free areas. Annals of Glaciology 51: 3–7.
Naselli-Flores, L., 2014. Morphological analysis of phytoplankton as a tool to assess ecological state of aquatic ecosystems: the case of Lake Arancio, Sicily, Italy. Inland Waters 4: 15–26.
Naselli-Flores, L. & J. Padisák, 2016. Blowing in the wind: how many roads can a phytoplanktont walk down? A synthesis on phytoplankton biogeography and spatial processes. Hydrobiologia 764: 303–313.
Naselli-Flores, L., R. Barone, I. Chorus & R. Kurmayer, 2007. Toxic cyanobacterial blooms in reservoirs under a semiarid Mediterranean climate: the magnification of a problem. Environmental Toxicology 22: 399–404.
Neilan, B. A., D. Jacobs, A. E. Goodman, P. T. Cox & P. Hawkins, 1995. Genetic diversity and phylogeny of toxic cyanobacteria determined by DNA polymorphism within the phycocyanin locus. Applied and Environmental Microbiology 61: 3875–3883.
Neilan, B. A., D. Jacobs, T. del Dot, L. L. Blackall, P. R. Hawkins, P. T. Cox & A. E. Goodman, 1997. rRNA sequences and evolutionary relationships among toxic and nontoxic cyanobacteria of the genus Microcystis. International Journal of Systematic Bacteriology 47: 693–697.
Neilan, B. A., M. L. Saker, J. Fastner, A. Törökné & B. P. Burnes, 2003. Phylogeography of the invasive cyanobacterium Cylindrospermopsis raciborskii. Molecular Ecology 12: 113–140.
Nekola, J. C. & P. S. White, 1999. The distance-decay of similarity in biogeography and ecology. Journal of Biogeography 26: 867–878.
Oren, A., 2011. Cyanobacterial systematics and nomenclature as featured in the international bulletin of bacteriological nomenclature and taxonomy/international journal of systematic bacteriology/international journal of systematic and evolutionary microbiology. International Journal of Systematic and Evolutionary Microbiology 61: 10–15.
Oren, A., 2015. Cyanobacteria in hypersaline environments: biodiversity and physiological properties. Biodiversity and Conservation 24: 781–798.
Oren, A. & G. M. Garrity, 2014. Proposal to change general consideration 5 and principle 2 of the international code of nomenclature of prokaryotes. International Journal of Systematic and Evolutionary Microbiology 64: 309–310.
Osorio-Santos, K., N. Pietrasiak, M. Bohunická, L. H. Miscoe, L. Kováčik, M. P. Martin & J. R. Johansen, 2014. Seven new species of Oculatella (Pseudanabaenales, Cyanobacteria): taxonomically recognizing cryptic diversification. European Journal of Phycology 49: 450–470.
Otsuka, S., S. Suda, S. Shibata, H. Oyaizu, S. Matsumoto & M. M. Watanabe, 2001. A proposal for the unification of five species of the cyanobacterial genus Microcystis Kützing ex Lemmermann 1907 under the rules of the Bacteriological Code. International Journal of Systematic and Evolutionary Microbiology 51: 873–879.
Pace, N. R., 1997. A molecular view of microbial diversity and the biosphere. Science 276: 734–740.
Padisák, J., 1997. Cylindrospermopsis raciborskii (Woloszynska) Seenayya et Subba Raju, an expanding, highly adaptive cyanobacterium: worldwide distribution and review of its ecology. Archiv fǖr Hydrobiologie Supplementband Monographische Beitrage 107: 563–593.
Padisák, J., 2003. Phytoplankton. In O’Sullivan, P. E. & C. S. Reynolds (eds), The Lakes Handbook 1. Limnology and Limnetic Ecology. Blackwell Science Ltd., Oxford: 251–308.
Padisák, J., G. Vasas & G. Borics, 2016. Phycogeography of freshwater phytoplankton: traditional knowledge and new molecular tools. Hydrobiologia 764: 3–27.
Paerl, H. W. & J. Huisman, 2008. Blooms like it hot. Science 320: 57–58.
Paerl, H. W. & J. Huisman, 2009. Climate change: a catalyst for global expansion of harmful cyanobacterial blooms. Environmental Microbiology Reports 1: 27–37.
Paerl, H. W. & T. G. Otten, 2013. Harmful cyanobacterial blooms: causes, consequences, and controls. Microbial Ecology 65: 995–1010.
Paerl, H. W. & V. J. Paul, 2012. Climate change: Links to global expansion of harmful cyanobacteria. Water Research 46: 1349–1363.
Paerl, H. W., R. S. Fulton, P. H. Moisander & J. Dyble, 2001. Harmful freshwater algal blooms, with an emphasis on cyanobacteria. The Scientific World Journal 1: 76–113.
Paerl, H. W., N. S. Hall & E. S. Calandrino, 2011. Controlling harmful cyanobacterial blooms in a world experiencing anthropogenic and climatic induced change. Science of the Total Environment 409: 1739–1745.
Papke, R. T., N. Ramsing, M. M. Bateson & D. M. Ward, 2003. Geographical isolation in hot spring cyanobacteria. Environmental Microbiology 5: 650–659.
Parker, C. T., B. J. Tindall & G. M. Garrity, 2015. International code of nomenclature of prokaryotes. International Journal of Systematic and Evolutionary Microbiology. 59: 2107–2113.
Paulino, S., E. Valério, N. Faria, J. Fastner, M. Welker, R. Tenreiro & P. Pereira, 2009. Detection of Planktothrix rubescens (Cyanobacteria) associated with microcystin productionin a freshwater reservoir. Hydrobiologia 621: 207–211.
Penno, S., D. Lindell & A. F. Post, 2006. Diversity of Synechococcus and Prochlorococcus populations determined from DNA sequences of the N-regulatory gene ntcA. Environmental Microbiology 8: 1200–1211.
Perkerson, R. B., J. R. Johansen, L. Kováčik, J. Brand, J. Kaštovský & D. A. Casamatta, 2011. A unique pseudanabaenalean (cyanobacteria) genus Nodosilinea gen. nov. based on morphological and molecular data. Journal of Phycology 47: 1397–1412.
Pfeiffer, C., T. Bauer, B. Surek, E. Schomig & D. Grundemann, 2011. Cyanobacteria produce high levels of ergothioneine. Food Chemistry 129: 1766–1769.
Piccini, C., L. Aubriot, A. Fabre, et al., 2011. Genetic and eco-physiological differences of South American Cylindrospermopsis raciborskii isolates support the hypothesis of multiple ecotypes. Harmful Algae 10: 644–653.
Ramette, A. & J. M. Tiedje, 2007. Biogeography: an emerging cornerstone for understanding prokaryotic diversity, ecology, and evolution. Microbial Ecology 53: 197–207.
Rasmussen, B., I. R. Fletcher, J. J. Brocks & M. R. Kilburn, 2008. Reassessing the first appearance of eukaryotes and cyanobacteria. Nature 455: 1101–1104.
Rastogi, R. P., D. Madamwar & A. Incharoensakdi, 2015. Bloom dynamics of cyanobacteria and their toxins: environmental health impacts and mitigation strategies. Frontiers in Microbiology 6: 1254.
Rejmánková, E., J. Komárek & J. Komárková, 2004. Cyanobacteria—a neglected component of biodiversity: patterns of species diversity in inland marshes of northern Belize (Central America). Diversity & Distributions 10: 189–199.
Ren, Y., H. Pei, W. Hu, C. Tian, D. Hao, J. Wei & Y. Feng, 2014. Spatiotemporal distribution pattern of cyanobacteria community and its relationship with the environmental factors in Hongze Lake, China. Environmental Monitoring and Assessment 186: 6919–6933.
Ricciardi, A. & J. Cohen, 2007. The invasiveness of an introduced species does not predict its impact. Biological Invasions 9: 309–315.
Richards, S., 2015. It’s more than stamp collecting: how genome sequencing can unify biological research. Trends in Genetics 31: 411–421.
Rocap, G., D. L. Distel, J. B. Waterbury & S. W. Chisholm, 2002. Resolution of Prochlorococcus and Synechococcus Ecotypes by Using 16S-23S Ribosomal DNA Internal Transcribed Spacer Sequences. Applied and Environmental Microbiology 68: 1180–1191.
Rosenzweig, M. L., 1995. Species diversity in space and time. Cambridge University Press, Cambridge.
Saker, M. L., I. C. G. Nogueira & V. M. Vasconcelos, 2004. Distribution and toxicity of Cylindrospermopsis raciborskii (Cyanobacteria) in Portuguese freshwaters. Limnetica 22: 129–136.
Sant’Anna, C. L., M. T. P. Azevedo, L. F. Agujaro, M. C. Carvalho, L. R. Carvalho & R. C. R. Souza, 2006. Manual ilustrado para identificação e contagem de cianobactérias planctônicas de águas continentais brasileiras, 1st ed. Editora Interciência, Rio de Janeiro.
Sarnelle, O., 2007. Initial conditions mediate the interaction between Daphnia and bloom-forming cyanobacteria. Limnology and Oceanography 52: 2120–2127.
Sciuto, K. & I. Moro, 2015. Cyanobacteria: the bright and dark sides of a charming group. Biodiversity and Conservation 24: 711–738.
Sessions, A. L., D. M. Doughty, P. V. Welander, R. E. Summons & D. K. Newman, 2009. The continuing puzzle of the great oxidation event. Current Biology 19: 567–574.
Shi, T. & P. G. Falkowski, 2008. Genome evolution in cyanobacteria: the stable core and the variable shell. PNAS 105: 2510–2515.
Shibl, A. A., L. R. Thompson, D. K. Ngugi & U. Stingl, 2014. Distribution and diversity of Prochlorococcus ecotypes in the Red Sea. FEMS Microbiology Letters 356: 118–126.
Singh, S., B. N. Kate & U. C. Banerjee, 2005. Bioactive compounds from cyanobacteria and microalgae: an overview. Critical Reviews in Biotechnology 25: 73–95.
Slatkin, M., 1987. Gene flow and the geographic structure of natural-populations. Science 236: 787–792.
Sompong, U., P. R. Hawkins, C. Besley & Y. Peerapornpisal, 2005. The distribution of cyanobacteria across physical and chemical gradients in hot springs in northern Thailand. FEMS Microbiology Ecology 52: 365–376.
Sønstebø, J. H. & T. Rohrlack, 2011. Possible implications of chytrid parasitism for population subdivision in freshwater cyanobacteria of the genus Planktothrix. Applied and Environmental Microbiology 77: 1344–1351.
Souza, V., L. E. Eguiarte, J. Siefert & J. J. Elser, 2008. Microbial endemism: does phosphorus limitation enhance speciation? Nature Reviews 6: 559–564.
Staley, J. T. & J. J. Gosink, 1999. Poles apart: biodiversity and biogeography of sea ice bacteria. Annual Review of Microbiology 53: 189–215.
Stefaniak, K. & M. Kokocinski, 2005. Occurrence of invasive Cyanobacteria species in polimictic lakes of the Wielkopolska region (Western Poland). Oceanological and Hydrobiological Research 34(Supplement 3): 137–148.
Stein, A., K. Gerstner & H. Kreft, 2014. Environmental heterogeneity as a universal driver of species richness across taxa, biomes and spatial scales. Ecology Letters 17: 866–880.
Stuken, A., J. Rücker, T. Endrulat, K. Preussel, M. Hemm, B. Nixdorf, U. Karsten & C. Wiedner, 2006. Distribution of three alien cyanobacterial species (Nostocales) in northeast Germany: Cylindrospermopsis raciborskii, Anabaena bergii and Aphanizomenon aphanizomenoides. Phycologia 45: 696–703.
Sukenik, A., O. Hadas, A. Kaplan & A. Quesada, 2012. Invasion of Nostocales (cyanobacteria) to subtropical and temperate freshwater lakes—physiological, regional, and global driving forces. Frontiers in Microbiology 3: 1–9.
Sullivan, M. B., J. B. Waterbury & S. W. Chisholm, 2003. Cyanophages infecting the oceanic cyanobacterium Prochlorococcus. Nature 424: 1047–1051.
Tanabe, Y. & M. M. Watanabe, 2011. Local expansion of a panmictic lineage of water bloom-forming cyanobacterium Microcystis aeruginosa. Plos One 6: e17085.
Tanabe, Y., F. Kasai & M. M. Watanabe, 2009. Fine-scale spatial and temporal genetic differentiation of water bloom-forming cyanobacterium Microcystis aeruginosa: revealed by multilocus sequence typing. Environmental Microbiology Reports 1: 575–582.
Taranu, Z. E., I. Gregory-Eaves, P. R. Leavitt, et al., 2015. Acceleration of cyanobacterial dominance in north temperate subarctic lakes during the Anthropocene. Ecology Letters 18: 375–384.
Taton, A., S. Grubisic, P. Balthasart, D. A. Hodgson, J. Laybourn-Parry & A. Wilmotte, 2006. Biogeographical distribution and ecological ranges of benthic cyanobacteria in East Antarctic lakes. FEMS Microbiology Ecology 57: 272–289.
Thomasa, A. D. & A. J. Dougill, 2006. Distribution and characteristics of cyanobacterial soil crusts in the Molopo Basin, South Africa. Journal of Arid Environments 64: 270–283.
Tian, C., H. Pei, W. Hu & J. Xie, 2012. Variation of cyanobacteria with different environmental conditions in Nansi Lake, China. Journal of Environmental Sciences 24: 1394–1402.
Tiedje, J., 1993. Approaches to the comprehensive evaluation of prokaryotic diversity of a habitat. In Allksopp, D., R. R. Colwell & D. L. Hawksworth (eds), Microbial diverisity and ecosystem function. CAB International, Wallingford: 73–87.
Ting, C. S., G. Rocap, J. King & S. W. Chisholm, 2002. Cyanobacterial photosynthesis in the oceans: the origins and significance of divergent light-harvesting strategies. Trends in Microbiology 10: 134–142.
Traveset, A. & D. M. Richardson, 2006. Biological invasions as disruptors of plant reproductive mutualisms. Trends in Ecology and Evolution 21: 209–216.
Ueda, K., T. Seki, T. Kudo, T. Yoshida & M. Kataoka, 1999. Two distinct mechanisms cause heterogeneity of 16S rRNA. Journal of Bacteriology 181: 78–82.
van Gremberghe, I., F. Leliaert, J. Mergeay, et al., 2011. Lack of phylogeographic structure in the freshwater cyanobacterium Microcystis aeruginosa suggests global dispersal. Plos One 6: e19561.
Van Vuuren, S. J. & G. P. Kriel, 2008. Cylindrospermopsis raciborskii, a toxic invasive cyanobacterium in South African freshwaters. African Journal of Aquatic Science 33: 17–26.
van Wichelen, J., I. Van Gremberghe, P. Vanormelingen, et al., 2010. Strong effects of amoebae grazing on the biomass and genetic structure of a Microcystis bloom (Cyanobacteria). Environmental Microbiology 10: 2797–2813.
Vareli, K., E. Briasoulis, G. Pilidis & I. Sainis, 2009. Molecular confirmation of Planktothrix rubescens as the cause of intense, microcystin-synthesizing cyanobacterial bloom in Lake Ziros, Greece. Harmful Algae 8: 447–453.
Vasas, G., I. Bácsi, G. Surányi, M. M. Hamvas, C. Máthé, S. A. Nagy & G. Borbély, 2010. Isolation of viable cell mass from frozen Microcystis viridis bloom containing microcystin-RR. Hydrobiologia 63: 147–151.
Vasas, G., O. Farkas, G. Borics, T. Felföldi, G. Sramko, G. Batta, I. Bácsi & S. Gonda, 2014. Appearance of Planktothrix rubescens bloom with [D-Asp3, Mdha7] MC-RR in gravel pit pond of a shallow lake dominated area. Toxins 5: 2434–2455.
Vidal, L. & C. Kruk, 2008. Cylindrospermopsis raciborskii (Cyanobacteria) extends its distribution to Latitude 34°53′S: taxonomical and ecological features in Uruguayan eutrophic lakes. Pan-American Journal of Aquatic Sciences 3: 142–151.
Vitousek, P. M., H. A. Mooney, J. Lubchenco & J. M. Melillo, 1997. Human domination of Earth’s ecosystems. Science 277: 494–499.
Vyverman, W., 1996. The Indo-Malaysian North-Australian phycogeographical region revised. Hydrobiologia 336: 107–120.
Walls, J. T., K. H. Wyatt, J. C. Doll, E. M. Rubenstein & A. R. Rober, 2018. Hot and toxic: Temperature regulates microcystin release from cyanobacteria. Science of the Total Environment 611: 786–795.
Whitaker, R. J., D. W. Grogan & J. W. Taylor, 2003. Geographic barriers isolate endemic populations of hyperthermophilic archaea. Science 301: 976–978.
Whitton, B. A. & M. Potts, 2000. The Ecology of Cyanobacteria. Kluwer, Dordrecht, The Netherlands.
Wiedner, C., J. Rücker, R. Brüggemann & B. Nixdorf, 2007. Climate change affects timing and size of populations of an invasive cyanobacterium in temperate regions. Oecologia 152: 473–484.
Wilk-Woźniak, E., W. Solarz, K. Najberek & A. Pociecha, 2016. Alien cyanobacteria: an unsolved part of the “expansion and evolution” jigsaw puzzle? Hydrobiologia 764: 65–79.
Wilson, A. E., O. Sarnelle, B. A. Neilan, T. P. Salmon, M. M. Gehringer & M. E. Hay, 2005. Genetic variation of the bloom-forming cyanobacterium Microcystis aeruginosa within and among lakes: Implications for harmful algal blooms. Applied and Environmental Microbiology 71: 6126–6133.
Woese, C. R., 1987. Bacterial evolution. Microbiological Reviews 51: 221–271.
Wołoszyńska, J., 1912. O glonach planktonowych niektórych jezior jazuańskich, z uwzględnieniem glonów Sawy. Das phytoplankton einiger javanischer seen, mit berücksichtigung des Sawa-planktons 1912: 649–709.
Wood, S. A., A. Rueckert, D. A. Cowan & S. C. Cary, 2008. Sources of edaphic cyanobacterial diversity in the Dry Valleys of Eastern Antarctica. The ISME Journal 2: 308–320.
Woodcock, S., C. J. van der Gast, T. Bell, M. Lunn, T. P. Curtis, I. M. Head & W. T. Sloan, 2007. Neutral assembly of bacterial communities. FEMS Microbiology Ecology 62: 171–180.
Wright, S., 1951. The genetical structure of populations. Annals of Eugenics 15: 323–354.
Wu, Z., B. Zeng, R. Li & L. Song, 2012. Physiological regulation of Cylindrospermopsis raciborskii (Nostocales, Cyanobacteria) in response to inorganic phosphorus limitation. Harmful Algae 15: 53–58.
Yap, W. H., Z. S. Zhang & Y. Wang, 1999. Distinct types of rRNA operons exist in the genome of the actinomycetes Thermomonospora chromogena and evidence for horizontal transfer of an entire rRNA operon. Journal of Bacteriology 181: 5201–5209.
Yoshida, M., T. Yoshida, M. Satomi, Y. Takashima, N. Hosoda & S. Hiroishi, 2008. Intra-specific phenotypic and genotypic variation in toxic cyanobacterial Microcystis strains. Journal of Applied Microbiology 105: 407–415.
Zanchett, G. & E. C. Oliveira-Filho, 2013. Cyanobacteria and cyanotoxins: from impacts on aquatic ecosystems and human health to anticarcinogenic effects. Toxins 5: 1896–1917.
Zhaxybayeva, O., J. P. Gogarten, R. L. Charlebois, F. Doolittle & R. T. Papke, 2006. Phylogenetic analyses of cyanobacterial genomes: quantification of horizontal gene transfer events. Genome Research 16: 1099–1108.
Zohary, T. & M. Shlichter, 2009. Invasion pf Lake Kinneret by the N2-fixing cyanobacterium Cylindrospermopsis. Verhandlungen der Internationale Vereinigung fu¨r theoretische und angewandte Limnologie 30: 1251–1254.
Zwart, G., B. C. Crump, M. P. K. Agterveld, F. Hagen & S. Han, 2002. Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquatic Microbial Ecology 28: 141–155.
Zwirglmaier, K., L. Jardillier, M. Ostrowski, et al., 2008. Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environmental Microbiology 10: 147–161.
Acknowledgements
We thank the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), a foundation of the Ministério da Educação (MEC), for the scholarship granted to K.F.R. Research activities by L.D. has been supported by CNPq Productivity Fellowship (Grant 307886/2015-8). L.D.’s research project has been developed within the framework of the National Institutes for Science and Technology (INCT) in Ecology, Evolution and Biodiversity Conservation, supported by MCTIC/CNPq (Proc. 465610/2014-5) and FAPEG. The authors declare no conflicts of interest concerning the studies summarized here.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Judit Padisák
Rights and permissions
About this article
Cite this article
Ribeiro, K.F., Duarte, L. & Crossetti, L.O. Everything is not everywhere: a tale on the biogeography of cyanobacteria. Hydrobiologia 820, 23–48 (2018). https://doi.org/10.1007/s10750-018-3669-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-018-3669-x