Abstract
Kin recognition has been demonstrated by plant biomass allocation and morphology traits as well as by nitrogen (N) uptake, but has not been examined from a nutrient-niche view yet. In this study, four species with distinct lifestyles, including Glycine max (L.) Merr. (herbaceous legume), Belamcanda chinensis (L.) DC. (herbaceous non-legume), Caesalpinia pulcherrima (L.) Sw. (woody legume), and Populus tomentosa (L.) Carr. (woody non-legume) were used to demonstrate kin recognition by estimating their biomass and allocation, as well as nutrient niches based on their uptake efficiency for N, phosphorus (P), sulfur (S), potassium (K), calcium (Ca), magnesium (Mg), and iron (Fe). For G. max, kin recognition was achieved by increased biomass, and by reduced nutrient-uptake efficiency of N, P, S, K, Ca, Mg, and Fe (decreased nutrient niches) to decrease nutrient competition among kin plants compared to the strangers. Although B. chinensis and C. pulcherrima had no biomass response, kin plants of B. chinensis increased, whereas C. pulcherrima decreased their S-uptake efficiency compare to strangers. Therefore, kin competition occurred in B. chinensis through increased nutrient niche whereas kin recognition occurred in C. pulcherrima through decreased nutrient niche. By comparison, P. tomentosa showed the co-occurrence of kin recognition and competition by increased root allocation and decreased P-uptake efficiency. These findings suggest that the biomass allocation and plant nutrient niches based on their nutrient-uptake efficiency can be used as potential parameters to identify kin recognition.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
According to the resource-partitioning theory, stronger competition might occur among intraspecific plants because they exhibit more niche overlap and similarity in nutrient use as compared to interspecific plants (Cheplick and Kane 2004; Silvertown 2004; Mo et al. 2016). However, the kin selection theory predicts that altruism towards relatives could be possible through the evolution of kin recognition (Hamilton 1964). Recent studies proposed kin recognition by showing the kin responses of the siblings living together in numerous plant species (Murphy and Dudley 2009; File et al. 2012; Simonsen et al. 2014). Studying kin recognition in plants can contribute to our understanding of plant interactions (Chu et al. 2008; Milla et al. 2009), community structure and diversity in natural ecosystems (Callaway and Mahall 2007; Brooker and Kikvidze 2008). Plants respond to kin recognition by increasing their fitness and displaying their plasticity in competitive traits in the presence of their relatives (Cheplick 1992; File et al. 2012). For example, siblings of Plantago lanceolata and Ipomoea hederacea give more reproductive output (Tonsor 1989; Biernaskie 2011) than strangers. When the fitness is not available, competitive traits, e.g., root distribution (Ninkovic 2003; Dudley and File 2007; Caffaro et al. 2013; Fang et al. 2013; Semchenko et al. 2014) and leaf distribution (Ninkovic 2003; Murphy and Dudley 2009; Lepik et al. 2012; Milla et al. 2012), which involved in soil nutrient acquisition and light and CO2 capture, should be invited to identify kin interactions (File et al. 2012).
More recently, several studies attempted to explore kin recognition by estimating nitrogen (N) uptake of plants using 15N isotope (Zhang et al. 2016). Besides N, various soil elements are essential for maintaining cell structure and physiological activities of plants (Buchanan et al. 2000; Taiz and Zeiger 2006; Xu et al. 2016), e.g., phosphorus (P) is the primary component of phospholipids, coenzymes, and high-energy phosphates, which drive the metabolic cycles of photosynthesis and respiration. Sulfur (S) is a component of many proteins, amino acids (cysteine, cystine, and methionine), and vitamins B1 and biotin. Potassium (K) is required to activate enzymes related to various metabolic reactions. Magnesium (Mg) is a constituent of chlorophyll molecules, and calcium (Ca) is indispensable to the growth of meristematic cells (Maathuis 2009). To realize species coexistence, plants can develop their nutrient niches reflected by their uptake efficiency for these essential elements. As a result, nutrient niches could be a potential signal of kin recognition. However, so far it remains unexplored (Cheplick 1992; Zhang et al. 2016).
Plant kin recognition responses are species specific (Murphy and Dudley 2009; Lepik et al. 2012). Diverse species have distinct growth and resource-use strategies for light and soil nutrients (Tilman 1988). Thus, the kin recognition responses might vary on the types of nutritional elements among plant species, e.g., legumes can fix atmospheric N via their associated rhizobium, leading to differentiation in N utilization as compared to non-legume plants (Franco and de Faria 1997). Nevertheless, most of previous studies focused on the annual herbaceous plants with the same lifestyle (Dudley and File 2007; Murphy and Dudley 2009; Biedrzycki et al. 2010; Biernaskie 2011; Lepik et al. 2012); however, the mechanism of kin recognition in perennial plants, particularly in shrub or woody plants (Bais et al. 2006), as well as, kin responses on the uptake of essential elements remains unclear. Accordingly, we assumed that kin recognition could be achieved by decreasing the nutrient-uptake efficiency (i.e., reduced nutrient-niches) of kin plants. To test this hypothesis, the seeds of Glycine max (herbaceous legume), Belamcanda chinensis (herbaceous non-legume), Caesalpinia pulcherrima (woody legume), and the branches of Populus tomentosa (woody non-legume) were collected. We mainly aimed to demonstrate which nutritional element(s) could be regarded as the potential parameter(s) to identify kin recognition among plant species with different lifestyles.
Materials and methods
Material and experimental design
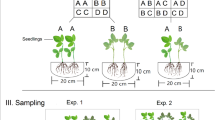
Seeds of G. max (cultivars of Shengdou No.5), B. chinensis, and C. pulcherrima were collected from Xishuangbanna Botanic Garden, Chinese Academy of Science in Yunnan Province (N 21°41′42.00″; E101°25′45.68″, 559 m above sea level). Cutting branches of the staminiferous (the pistilliferous plants were restrained in this district) P. tomentosa plants were collected from a plantation in Henan Province (N35°18′13.71″; E113°55′15.05″, 310 m above sea level). The offspring seeds/branches of each species were collected from four mother plants of the same cultivar growing in the same plot within a distance of approximately 5 m. For each species, the offspring plants from the same mother plant were defined as siblings, whereas those from different mother plants were referred to as strangers. We used the one-factor experimental design of relatedness. For each treatment, two seedlings, either siblings (kin groups) or strangers (strangers groups), from four mother plants were planted as pairs per cylinder pot without any barriers for root contact. Twenty-four replicates were used for each treatment of each species.
Growth condition
All seeds collected from the mother plants of G. max, B. chinensis, and C. pulcherrima were allowed to germinate in the Petri dishes containing only distilled water. After 6 days, the healthy and similar-sized seedlings were selected to eliminate the size effect. The cutting branches of P. tomentosa from different stock plants were cut into similar shape and prepared for cottage plantation until rooted. All plant seedlings were then transplanted into soil cylinder pots on October 2, 2015 in a greenhouse at the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China. The cylinder pots were 10 cm in height and 20 cm in diameter, and the distance between the two seedlings was 10 cm. The soil contained 50% roseite and 50% humus. The temperature of the glasshouse was maintained at 28 °C in the day and at 18 °C in the night, with a light:dark photoperiod of 16:8 h, the photosynthetic photon fluency rate of 180 μM m−2 s−1, and the relative humidity of approximately 60%. The pots were watered every two days to maintain soil moisture at 70% field water holding capacity. The plants were not fertilized during the experiment.
Biomass and allocation measurements
After the plants grew for 100 days, they were harvested individually. Subsequently, roots were washed in distilled water to remove any elements bound to their surface. G. max, C. pulcherrima, and P. tomentosa plants were separated into leaf, stem, and root parts, whereas B. chinensis plants were separated into shoots and roots, because they have no obvious stems. All plant materials were oven-dried at 65 °C for 48 h, and weighed for each plant part and whole dry biomass for each species.
Element concentration and uptake efficiency measurements
The individual plant leaves, stems, and roots of each species were ground with a ball mill (MM2, Retsch, Haan, Germany). Subsequently, 60 mg of each pulverized sample was packaged in silver paper to determine the concentration of N and S using the vario MACRO cube (Elementar Analysensysteme, GmbH, Germany). Samples (150 mg) were prepared to analyze the contents of P, K, Ca, Mg, and Fe. According to the method of Zarcinas et al. (1987), the samples were placed in a 100-ml microwave jar and digested with 5 ml of nitric acid for 1 h, followed by the addition of 2 ml hydrogen peroxide (H2O2). Subsequently, they were allowed to cool to about 25 °C and then transferred into 15-ml volumetric flasks. The sample volumes were made constant by adding ultrapure water. The total elemental concentration in the digests was determined by inductively coupled plasma-optical emission spectrometry (ICP-OES) according to the method of Fassel et al. (2008). All values were expressed in mg kg−1 DW plant.
Calculation and statistics
The root and leaf allocations were calculated by the ratio of root biomass to total plant biomass and the ratio of leaf biomass to total biomass, respectively. The nutrient-uptake efficiency was calculated by (element concentrations in root × root biomass + element concentrations in shoot × shoot biomass)/root biomass/100 days (Moreau et al. 2015), and the unit was expressed as mg g−1 day−1. A two-way analysis of variance (ANOVA) was used to test the effects of species, kinship, and their interaction on biomass and nutrient-uptake efficiency with SPSS 16 (SPSS Inc., Chicago, IL, USA). The differences in the biomass and nutrient-uptake efficiency of siblings and strangers were considered significant at P < 0.05.
Results
Biomass and allocation
Plant biomass was significantly dependent on the species, kinship, and their interaction (Table 1). The kin plants of G. max showed higher shoot and root biomass than the stranger plants (Fig. 1a). The allocation to root and leaf tissues was not significantly different between the kin and strangers of G. max (the root allocation ratio was 0.164 for kin and 0.169 for strangers while leaf allocation ratio was 0.389 for kin and 0.366 for strangers). The species B. chinensis and C. pulcherrima showed no significant difference in the shoot and root biomass, as well as in the root and leaf allocation between kin and strangers (Fig. 1; P > 0.05). For P. tomentosa, the kin plants showed higher root biomass and root allocation (the root allocation ratio was 0.172 for kin and 0.132 for strangers) than strangers, but shoot biomass and leaf allocation (kin ratio was 0.724 and strangers was 0.720) was not different between the kin and stranger groups (Fig. 1d).
Plant shoot (leaf and stem) and root biomass when a G. max, b B. chinensis, c C. pulcherrima, and d P. tomentosa growing with kin or strangers. The values were mean ± SE of 12 replicates. The asterisks (*) on bars indicate significant differences between kin and stranger of each subarea biomass including leaf, stem and root in each species at P < 0.05 level
Nutrient-uptake efficiency
Significant interactive effects between species and kinship were observed on the uptake efficiency for N, S, K, Ca, and Mg (Table 1). Compared to strangers, kin plants of G. max showed lower uptake efficiency for N (P = 0.039), P (P = 0.024), S (P = 0.043), K (P = 0.001), Ca (P = 0.02), and Mg (P = 0.044; Fig. 2a). However, kin plants of B. chinensis showed higher (P = 0.01; Fig. 2b) whereas kin of C. pulcherrima (P = 0.027; Fig. 2c) showed lower S-uptake efficiency than strangers. P. tomentosa showed decreased P-uptake efficiency (P = 0.031) compared to the strangers (Fig. 2d). No significant difference was observed in the uptake efficiency for other elements between kin and strangers in each species.
The nutrient niches consist of nutrient uptake efficiency of N, P, S, K, Ca, Mg and Fe in plant species of a G. max, b B. chinensis, c C. pulcherrima, and d P. tomentosa growing with kin or strangers. The values were logarithmic mean ± SE (Log (mg g−1 day−1) of 12 replicates. The value of the midpoint in each radar map was 1, the asterisks (*) on side of element indicate significant differences of corresponding element(s) between kin and strangers at P < 0.05 level
Discussion
Numerous studies have suggested that kin recognition could occur mainly based on the fitness and morphological traits of plant species, but it is seldom evaluated according to their nutrient-uptake efficiency or nutrient niches. Here, we showed that the nutrient-uptake efficiency is species-specific for various essential elements, and the nutrient niches together with biomass allocation could be a proper indicator of kin recognition.
Kin recognition is generally identified through the better performance of plants living with relatives (File et al. 2012). This is confirmed by the higher biomass of kin plants than the strangers of G. max, but is not reflected in the biomass response of B. chinensis and C. pulcherrima. A possible explanation is that the biomass responses are not universal among the self-incompatible plants (Masclaux et al. 2010). Moreover, we observed no kin recognition responses in root or shoot allocation for both species. This indicates that a decreased root allocation might be not necessary for kin recognition (Dudley and File 2008; Zhang et al. 2016). By comparison, the P. tomentosa kin plants showed a higher competitive ability than its strangers through higher root allocation (Dudley and File 2007). Thus, various biomass responses among these species again confirmed that plant kin recognition was species-specific (de Kroon 2007; Lepik et al. 2012).
Although plant biomass and competitive traits have been suggested to be the powerful indicators of kin recognition, they are often affected by environmental factors (Cheplick 1992; File et al. 2012). Consequently, there could be other more proper kin recognition indicators, based on the fact that plants recognize their kin perhaps through reducing the competition and mutual accommodation of their resources to cooperate with their relatives (Lepik et al. 2012). Moreover, the field experiments demonstrate that three or four resources are limiting in any plant community (Tilman 1982). Thus, the limited elements that play a key role in plant growth might respond to kin recognition based on Liebig’s law of the minimum. In this study, the nutrient-uptake efficiency for N, P, S, K, Ca, and Mg showed kinship and species interactive effects and can be considered as the potential signals or complementary parameters to identify kin interactions when the fitness is unavailable.
A variation in the nutrient-uptake efficiency between kin and stranger plants suggests a complementary kin interaction as compared to the biomass response. Given the increased nutrient-uptake efficiency exhibits a higher competition for soil nutrients (Kuzyakov and Xu 2013), the nutrient-uptake efficiency of many essential elements can reflect the plant nutrient niches to a certain extent. The decreased uptake efficiency of N, P, S, K, Ca, Mg, and Fe elements indicate reduced nutrient niches in the kin plants of G. max compared to the strangers (Fig. 1a), leading to mutual accommodation of kin. It is similar to a previous study that siblings showed significantly decreased uptake of total N (Zhang et al. 2016). Moreover, the decreased nutrient uptake in siblings yet led to more biomass production, indicating relatively higher nutrient-use efficiency. Similarly, kin recognition was observed in C. pulcherrima based on its lower S-uptake efficiency despite without any change in its biomass. On the contrary, the higher S-uptake efficiency suggested higher competition in the kin plants of B. chinensis compared to its strangers. Although the kin plants of P. tomentosa showed significantly increased root allocation, but did not enhance their uptake efficiency of nutritional elements compared to the strangers (Zhang et al. 2016). Instead, they achieved kin recognition through their reduced P-uptake efficiency (Fig. 2d). As greater root allocation commonly refers to the enhanced belowground competitive ability (Dudley and File 2007), kin cooperation and competition co-occurred in P. tomentosa plants. Such increased root allocation response could be mainly ascribed to the competition for space or water in the given pot size. Based on above information, we demonstrated that the nutrient-uptake efficiency of plants, together with biomass responses, can identify kin recognition more precisely.
We observed kin recognition response in terms of biomass allocation and nutrient-uptake efficiency was species-specific for various essential elements in plants of distinct lifestyles, e.g., annual versus perennial and legume versus non-legume plants. These results indicate that kin recognition could be related to plant lifestyles and strategies for nutrient absorption and transmission (Graves et al. 2006) and nutrient niches (McKane et al. 2002). The reason is that nutrients might be used mainly for transformation and utilization in annual plants, but also for storage in perennial plants, thus affecting their biomass allocation (Zhu et al. 2011). Here, we adopted the same type of soil and nutrient supply level for different plant species. Compared to the three perennial plants, the kin response of annual G. max on the compromise of all measured elements could be attributed to its fast growth and short life cycle with strong nutrient requirements (Aerts and Chapin 2000). This is reflected in our result that G. max showed higher nutrient-uptake efficiency than the other species (Fig. 2).
For perennial plants, the kin response of nutrient uptake exhibits element selectivity based on the identity of plant species. Because legumes usually contain abundant proteins, especially sulfur-containing amino acids (Jukanti and Chibbar 2012), lead to S-uptake as a sensitive response to kin recognition. Thus, kin recognition is achieved by decreasing the S-uptake efficiency of G. max and C. pulcherrima in kin plants compared to strangers. However, the kin response on the S-uptake of the non-legume plants of B. chinensis was opposite to that of legumes. This could be ascribed to different S requirements of these plants. Further studies should be performed to explain why these plants are sensitive to uptake of S. The kin recognition response in P. tomentosa on the uptake of P element could depend on its limited supply and high requirement for fast growth in the long-term lifestyle. However, our study was confined to only a limited number (four) of plant species because of the difficulty in seed collection. Therefore, more rigorous investigation of kin recognition in vast number of plant species should be performed for the better understanding of plant interaction among plants with distinct lifestyles.
In summary, our study shows significant species-specific kin responses in terms of both biomass allocation and nutrient niches based on the nutrient uptake efficiency of soil essential elements. Therefore, nutrient niches together with biomass allocation can be regarded as the potential parameters to identify plant kin recognition. Considering the distinct roles of various essential elements in plant growth and development, more studies should be focused on specific micronutrients or some specific functional elements in response to plant kin recognition in future.
References
Aerts R, Chapin FS (2000) The mineral nutrition of wild plants revisited: A re-evaluation of processes and patterns. Adv Ecol Res 30:1–67
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266
Biedrzycki ML, Jilany TA, Dudley SA, Bais HP (2010) Root exudates mediate kin recognition in plants. Commun Integr Biol 3:1–8
Biernaskie ML (2011) Evidence for competition and cooperation among climbing plants. Proc R Soc B 278:1989–1996
Brooker RW, Kikvidze Z (2008) Importance: an overlooked concept in plant interaction research. J Ecol 96:703–708
Buchanan BB, Gruissem W, Jones RL (2000) Biochemistry and molecular biology of plants. American Society of Plant Physiologists, Rockville
Caffaro MM, Vivanco JM, Botto J, Rubio G (2013) Root architecture of Arabidopsis is affected by competition with neighbouring plants. Plant Growth Regul 70:141–147
Callaway RM, Mahall BE (2007) Family roots. Nature 448:145–146
Cheplick GP (1992) Sibling competition in plants. J Ecol 80:567–575
Cheplick GP, Kane KH (2004) Genetic relatedness and competition in Triplasis purpurea (Poaceae): resource partitioning or kin selection? Int J Plant Sci 165:623–630
Chu CJ, Maestre FT, Xiao S, Weiner J, Wang YS, Duan ZH, Wang G (2008) Balance between facilitation and resource competition determines biomass-density relationships in plant populations. Ecol Lett 11:1189–1197
de Kroon H (2007) Ecology: how do roots interact? Science 318:1562–1563
Dudley SA, File AL (2007) Kin recognition in an annual plant. Biol Lett 3:435–438
Dudley SA, File AL (2008) Yes, kin recognition in plants! Biol Lett 4:69–70
Fang SQ, Clark RT, Zheng Y, Iyer-Pascuzzi AS, Weitz JS, Kochian LV, Edelsbrunner H, Liao H, Benfey PN (2013) Genotypic recognition and spatial responses by rice roots. Proc Natl Acad Sci USA 110:2670–2675
Fassel VA, Kniseley RN, Chem A (2008) Inductively coupled plasma: optical emission spectroscopy. Anal Chem 46(13):1110A–1120A
File AL, Murphy GP, Dudley SA (2012) Fitness consequences of plants growing with siblings, reconciling kin selection, niche partitioning and competitive ability. Proc R Soc B 279:209–218
Franco AA, de Faria SM (1997) The contribution of N2-fixing legumes to land reclamation and sustainability in the tropics. Soil Biol Biochem 29:897–903
Graves JH, Peet RK, White PS (2006) The influence of carbon & mdash; nutrient balance on herb and woody plant abundance in temperate forest understories. J Veg Sci 17(2):217–226
Hamilton WD (1964) The genetical evolution of social behavior II. J Theor Biol 7:1–52
Jukanti AK, Chibbar RN (2012) Gaur1 PM, Gowda1 CLL. nutritional quality and health benefts of chickpea (Cicer arietinum L): a review. Br J Nutr 108:S11–S26
Kuzyakov Y, Xu X (2013) Competition between roots and microorganisms for nitrogen: mechanisms and ecological relevance. New Phytol 198(3):656–669
Lepik A, Abakumova M, Zobel K, Semchenko M (2012) Kin recognition is density-dependent and uncommon among temperate grassland plants. Funct Ecol 26:1214–1220
Maathuis FJM (2009) Physiological functions of mineral macronutrients. Curr Opin Plant Biol 12(3):250–258
Masclaux F, Hammond RL, Meunier J, Gouhier-Darimont C, Keller L, Reymond P (2010) Competitive ability not kinship affects growth of Arabidopsis thaliana accessions. New Phytol 185(1):322–331
McKane RB, Johnson LC, Shaver GR, Knute J, Nadelhoffer KJ, Rastetter EB, Fry B, Giblin AE, Kiellandk K, Kwiatkowski BL, Laundre JA, Murray G (2002) Resource-based niches provide a basis for plant species diversity and dominance in arctic tundra. Nature 415(3):68–70
Milla R, Forero DM, Escudero A, Iriondo JM (2009) Growing with siblings, a common ground for cooperation or for fiercer competition among plants? Proc R Soc B 276:2531–2540
Milla R, Velez del BA, Escudero A, Iriondo JM (2012) Kinship rivalry does not trigger specific allocation strategies in Lupinus angustifolius. Ann Bot 110:165–175
Mo Y, Yang R, Liu L, Gu X, Yang X, Wang Y, Zhang X, Li H (2016) Growth, photosynthesis and adaptive responses of wild and domesticated watermelon genotypes to drought stress and subsequent re-watering. Plant Growth Regul 79(2):229–241
Moreau D, Pivato B, Bru D, Busset H, Deau F, Faivre C, Matejicek A, Strbik F, Philippot L, Mougel C (2015) Plant traits related to nitrogen uptake influence plant-microbe competition. Ecology 96:2300–2310
Murphy GP, Dudley SA (2009) Kin recognition: competition and cooperation in Impatiens (Balsaminaceae). Am J Bot 96:1990–1996
Ninkovic V (2003) Volatile communication between barley plants affects biomass allocation. J Exp Bot 54:1931–1939
Semchenko M, Saar S, Lepik A (2014) Plant root exudates mediate neighbour recognition and trigger complex behavioural changes. New Phyto 204:631–637
Silvertown J (2004) Plant coexistence and the niche. Trends Ecol Evol1 9:605–611
Simonsen AK, Chow T, Stinchcombe JR (2014) Reduced plant competition among kin can be explained by Jensen’s inequality. Ecol Evol 4(23)::4454–4466
Taiz L, Zeiger E (2006) Sunderland: sinauer associates. Plant Physiol pp:100–119
Tilman D (1982) Resource competition and community structure. Princeton University Press, Princeton
Tilman D (1988) Plant strategies and the dynamics and structure of plant communities. Monogr Popul Biol 26:1–360
Tonsor SJ (1989) Relatedness and intraspecific competition in Plantago lanceolate. Am Nat 134:897–906
Xu H, Liu C, Lu R, Guo G, Chen Z, He T, Gao R, Li Y, Huang J (2016) The difference in responses to nitrogen deprivation and re-supply at seedling stage between two barley genotypes differing nitrogen use efficiency. Plant Growth Regul 79(1):119–126
Zarcinas BA, Cartwright B, Spouncer LR (1987) Nitric acid digestion and multi-element analysis of plant material by inductively coupled plasma spectrometry. Commun Soil Sci Plan 18(1):131–146
Zhang L, Liu Q, Tian Y, Xu X, Ouyang H (2016) Kin selection or resource partitioning for growing with siblings, implications from measurements of nitrogen uptake. Plant Soil 398(1–2):79–86
Zhu L, Li Z, Ketola T (2011) Biomass concentrations and nutrient uptake of plants cultivated on artificial floating beds in China’s rural area. Ecol Eng 37(10):1460–1466
Acknowledgements
We thanked Luyang Dong from China Internet Network Information Center for help looking after the plant seedlings. This study is supported by National Natural Science Foundation of China (31470560).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Li, J., Xu, XL. & Liu, YR. Kin recognition in plants with distinct lifestyles: implications of biomass and nutrient niches. Plant Growth Regul 84, 333–339 (2018). https://doi.org/10.1007/s10725-017-0343-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0343-7