Abstract
The main goal of this study is to gain insight into the relationship between the genetic profile of cultivated rice (Oryza spp.) accessions and their resistance to rice blast. Therefore, the genetic and phenotypic variability of a set of 350 cultivated rice accessions originating from Africa (Benin, Mali and Nigeria, Ivory Coast etc.) was examined. Seventy-seven fluorescent amplified fragment polymorphism (AFLP) markers were used to gain insight into the genetic variation and to classify the germplasm collection. In addition, the rice germplasm was assessed for its resistance to blast disease caused by Pyricularia oryzae in upland field conditions. Huge differences in responses of rice accessions to P. oryzae were observed, ranging from highly susceptible to highly resistant. Twelve percent of all accessions were highly resistant to P. oryzae. Based on their AFLP marker profile these highly resistant accessions could be separated from the other accessions. Stepwise regression revealed that the best prediction of the blast resistance level was achieved with a maximum number of 13 AFLP markers. Marker CTA22 was the most important for accurate prediction of blast resistance, this marker was present in all highly resistant accessions. It can be concluded that AFLP markers are a valuable tool to screen rice accessions for their susceptibility towards blast disease and that, based on a subset of markers, it is possible to predict the resistance to rice blast.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Blast disease, caused by the filamentous ascomycete fungus Pyricularia oryzae (Couch and Kohn 2002) is one of the most important biotic constraints to rice production in Benin (Vodouhe et al. 1981; Afouda et al. 2007). It is the most widespread disease in African rice production causing yield losses of up to 100% (Séré et al. 2013). In Benin, blast disease hotspots have recently been mapped by the Africa Rice Center (AfricaRice) and partners, revealing the presence of isolates belonging to varying pathotypes in the rice growing areas. Extensive and uncontrolled use of fungicides is injurious to human health and poses environmental safety concerns. The deployment of host resistance in rice cultivation has been recognized to be the most economical and effective way of controlling the disease and reducing the need for chemical applications. In this regard, more than 100 single resistance genes (R genes) and around 100 quantitative trait loci (QTL) have been identified and some were mapped on rice chromosomes (Ballini et al. 2008; Sharma et al. 2012). Unfortunately, country-specific field evaluation of a set of differential lines, carrying most of identified genes showed, the capacity of new aggressive races of P. oryzae to overcome a number of dominant genes (R genes) in African (Pi1, Pi7, Pi5, Pikp, Pia, Pita2, Piks, Pi3, Pik, Pita, Piz, Pikh and Pikm) (Baboy et al. 1995; WARDA 1999; Odjo et al. 2011). Currently, blast disease poses a significant threat to rice food security in Benin (Odjo et al. 2011). In fact, P. oryzae is highly variable, and loss of resistance in varieties is quite common (Zeigler et al. 1994; Jia et al. 2003). This often appears when resistance is based on a single resistance (R) gene (Mackill and Bonman 1992; Dai et al. 2010). Scientists argued that each of the R genes known so far only acts against a subset of P. oryzae isolates. Over time, varieties may thus become susceptible when new more virulent isolates will occur. However, a continued search for new R genes and their pyramiding into genotypes remains necessary for efficient disease management (Wang et al. 2010; RoyChowdhury et al. 2012). The approach by genes pyramiding will confer overlapping resistance spectra to multiple isolates/races of P. oryzae (Hittalmani et al. 2000). It was also shown that quantitative resistance that provides broad spectrum resistance and is controlled either by several minor or major genes, durably protects the host plant against P. oryzae (Wisser et al. 2005; RoyChowdhury et al. 2012; Wang et al. 2013).
There are only two types of cultivated rice species in the world: Oryza sativa L. and O. glaberrima Steud. Both are commonly grown in Africa. O. glaberrima or African rice was domesticated in West Africa approximately 3500 years ago (Jones et al. 1997). This indigenous rice possesses useful traits such as weed competitiveness, tolerance to various abiotic stressors (drought, iron, acidity, salinity) as well as pest and disease resistance (Pham 1992; Jones et al. 1997; Futakuchi and Sié 2009; Sié et al. 2010; Thiémélé et al. 2010). However, O. glaberrima is becoming rare in African cropping systems where O. glaberrima is mostly replaced by O. sativa cultivars due to the former rice’s negative traits such as poor yield, grain shattering, poor resistance to lodging (Linares 2002; National Research Council 1996). O. sativa or Asian rice comprises two major groups: (1) the Japonica Group with sticky, short-grained genotypes that are mainly cultivated upland; and (2) the Indica Group with non-sticky, long-grained genotypes that are mostly grown in lowland conditions. Asian rice was introduced some 500 years ago to Africa by Portuguese sailors. Today, most varieties grown in Africa belong to O. sativa. Successful artificial interspecific hybridizations of both O. glaberrima and O. sativa have led to a series of popular hybrids known as NERICA (“New Rice for Africa”), which combines the resilience of the African species with the productivity of the Asian ones (Pham 1992; Jones et al. 1997; Futakuchi and Sié 2009; Sié et al. 2010; Thiémélé et al. 2010).
To date, only limited information about agricultural characteristics including pest and disease resistance of African rice genepool is available. Germplasm without consistent data on diseases and pest tolerance are of little value for breeding programs and sustainable paddy production. The present work studied genetic variation a set of cultivated rice accessions (O. sativa and O. glaberrima) of diverse African origins (Benin, Mali, Nigeria, Ivory Coast etc.) and examined their reactions to blast disease. Linkages between the genetic structure/markers and disease resistance was be investigated. Finally, a sub-set germplasm was proposed for a thorough evaluated of agronomic traits.
Materials and methods
Source of the germplasm collection
The 350 rice (5 O. sativa and 345 O. glaberrima) accessions used in this study (Suppl. Table 1), were obtained from the germplasm collection of AfricaRice, a Consultative Group for International Agricultural Research (CGIAR) centre located in Cotonou, Benin. Three reference accessions (WAB0006871 and WAB0007880 described as O. glaberrima rice and WAB0006881 as O. sativa species) were also included in the study as taxonomic and blast resistance/susceptibility references.
DNA extraction and AFLP analysis of rice accessions
Genomic DNA extraction from each accession was done according to the CetylTrimethyl Ammonium Bromide (CTAB) method described by Saghai-Maroof et al. (1984). Five seeds from each of the 350 rice samples were grown in a growth chamber at 27 °C. Approximately 200 mg of leaf samples were harvested on 21 days-old seedlings and ground in liquid nitrogen in 2 mL of CTAB using a pestle and mortar. Then, 500 μL of each of the obtained suspensions were transferred to centrifuge tubes and incubated in a water bath for 30 min at 60 °C. An equal volume of chloroform:isoamyl alcohol (24:1, v/v) was added to each tube. The suspension was gently mixed by inverting the tubes. Tubes were centrifuged at 12,000 rpm for 10 min and the supernatant recovered and mixed with an equal volume of ice-cold isopropanol. DNA was recovered as a pellet by centrifugation at 12,000 rpm for 5 min, washed with 100 μL of 70% ethanol, dried under vacuum and dissolved in 30 μL of TE buffer. Two μL of RNase were added to the DNA and incubated for 1 h. DNA concentrations were measured using the Quantus™ Fluorometer (Promega) and adjusted to approximatively 250 ng/μL.
AFLP polymorphic markers were used to analyse the genomic DNA using the modified protocol described by Vos et al. (1995). To this end, DNA (250 ng/μL) was completely digested with 5 U of each MseI and EcoRI restriction enzymes and ligated to 5 pmol EcoRI and 25 pmol MseI adapters. Pre-amplification of ligation products was done with specific EcoRI and MseI primers without additive nucleotides and over 20 cycles of 94 °C during 30 s, 56 °C during 60 s, and 72 °C during 60 s were run in a thermal cycler. Concentration of the amplified DNA was checked on a 1.5% agarose gel. Next, DNA was diluted 25 times in TE buffer. For selective amplification, three primer combinations with three selective bases at the end (EcoRI + ACC/MseI + CTC, EcoRI + ACC/MseI + CTA, EcoRI + ACC/MseI + CAT) were used to reveal a maximum number of polymorphic markers in the rice samples. EcoRI primer was end-labeled with NED fluorescent dye. Amplified products were mixed with GeneScan-500 ROX size standard (Applied Biosystems Inc., USA) and deionized formamide. Finally, they were loaded on an Applied Biosystems electrophoresis instrument 3130 for electrophoretic size separation with performance optimized polymer (POP 4).
Field screening for resistance to blast in Benin
Benin republic has two rainy seasons during which blast disease can naturally occur. The AfricaRice Cotonou experimental site was identified as a disease hotspot (Odjo et al. 2011). Two field experiments were successively conducted under upland rainfed ecology in 2015 during the two rainy seasons, i.e. from April to July and from August to October, respectively. The first experiment was implemented using an augmented randomized complete block design with 17 control varieties replicated in the design whereas an Alpha Lattice design with four replicates was used during the second experiment. For each of the 350 accessions, 18 seeds were sown in a plot of three 50 cm long rows with a spacing of 20 cm between rows and 10 cm between plants. Fourteen days prior to sowing of the latter accession seeds, three infesting rows, consisting of three blast-susceptible reference varieties (Maratelli, CO39 and IRBLTA2-PI) were sown perpendicularly to the test plots in order to spread the disease. Thinning of the accessions was done 14 days after sowing to maintain only one seedling grown in each hill. A pre-drilling base application of 300 kg/ha of NPK fertilizer (15-15-15) was applied at sowing of the test accessions. Next, 300 kg/ha of urea was applied in two equal splits at 21 and 42 days after sowing. Hand-weeding and watering were done as per need. No treatment against diseases and insects was done. Disease symptoms were scored on the inner plants of each row of each plot leaving aside the first and last plant in each row. Scoring was done weekly starting from the appearance of the first symptoms until harvest. The IRRI 1–9 scale (2013) was used and four to five scoring events were performed during the two experiments. Mean Disease Index (DI) was obtained by calculating the average disease scores for each accession. Four disease resistance classes were considered: resistant (R): DI < 3, moderately resistant (MR): 3 < DI < 4, moderately susceptible (MS): 4 < DI < 5 and susceptible (S): DI > 5. A representative sub-set of the total germplasm approximate genetic diversity was then selected according to their blast response and variability for AFLP markers for efficient use in crop improvement programmes. Actions were taken to avoid duplicate the most similarly similar accessions.
Statistical data analysis
AFLP electropherograms were analyzed by GeneMapper Software Version 4.0 using the marker GeneScan-500 ROX in the range size of 30–500. Genotype marker results were automatically compiled in a standard binary Excel format (1; present, 0: absent). Polymorphism Information Content (PIC) values were calculated according to Botstein et al. (1980). Genetic dissimilarity coefficients using the Jaccard distance were calculated to determine the genetic relationship between accessions and the hierarchical clustering tree was constructed using Unweighted Neighbor Joining (UWNJ). Analysis of molecular variance (AMOVA) was carried out to estimate molecular diversity at each hierarchical level among and within the groups identified and genetic diversity parameters calculated using GenAlEx 6.502. DI values were subjected to a Linear Discriminant Analysis (LDA) with identified polymorphic AFLP markers based on continuous and categorical approaches using the R software (R Core Team 2017) to define the linear combination of AFLP markers which best separate accessions according to blast resistance. In addition, stepwise regression models were constructed for selecting an appropriate subset of markers to predict the rice blast resistance.
Results
Genetic diversity detected by AFLP markers among assembled rice germplasm
The three primer combinations generated a total of 10,384 bands out of which 72% were polymorphic at 77 different loci (putative loci/genome landmarks). Average Polymorphic Information Content (PIC) detected was 24 ± 4%. The primer combination EcorI-ACC:Mse-CAT appeared to be most informative with a PIC of 29% (Table 1). Jaccard’s genetic dissimilarity coefficients ranged from 0.03 to 0.98 across all the germplasm.
The UWNJ tree based on Jaccard’s dissimilarity coefficients derived from the 77 polymorphic AFLP markers evidenced three main genetic clusters: one with 184 accessions including the O. glaberrima control WAB0006871; a second with 83 accessions including the only O. sativa control (WAB0006881, Moroberekan), and a third with 86 accessions including the O. glaberrima control WAB0007880 (Fig. 1). Dissimilarity coefficients varied from 0.05 to 0.84 in cluster 1 and from 0.03 to 0.59 in cluster 3, whereas a larger range was observed in cluster 2 (0.03–0.98). The highest Shannon’s diversity index (I) was found within cluster 2 with I = 0.4 followed by cluster 1 with 0.32, whereas a much smaller I-value was observed in cluster 3 (0.26). Expected heterozygosity averaged 0.26, 0.21 and 0.17 in cluster 2, cluster 1 and cluster 3, respectively (Table 2). A high proportion of genetic diversity was most apparent in cluster 2, which comprised all O. sativa accessions, including control variety WAB0006881. Cluster 1 and cluster 3 contained exclusively O. glaberrima accessions. AMOVA used to determine the percentage of genetic variation within and among the three identified clusters (Table 3), revealed that the within-cluster diversity explained most of the genetic variation (77%), whereas only 23% of total variance was due to differences between the three clusters (P < 0.001).
UWNJ tree cluster analysis showing accessions reaction to blast disease. Colours indicate the disease status: resistant accessions (Severity < 3) are written in red; moderate resistant (3 < Severity < 4) in blue; moderate susceptible (4 < Severity < 5) in green and susceptible (Severity > 5) are indicated in black. (Color figure online)
Frequency of blast disease among 350 cultivated rice accessions
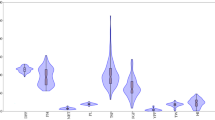
DI ranged from 0 to 9 in the first experiment, while for the second experiment, DI ranged from 0.33 to 7 (Fig. 2). A significant positive association (R2 = 0.64, P < 0.001) was found for blast disease index between the two successively conducted field experiments from April to October 2015. In other words, blast disease score distributions were similar in the two experiments. However, we observed that disease pressure was lower during the second experiment. Nevertheless, resistance/susceptibility status of accessions did not change. Average disease scores recorded on the susceptible control varieties WAB0007142 (CO 39) and GSOR301253 (Maratelli) were 8 and 7.6, respectively. Out of the total 350 tested accessions, 12% were highly resistant, 24% moderately resistant, 2 9% moderately susceptible and 35% susceptible. The highest number of resistant accessions was found in cluster 2 (20), whereas cluster 1 contained the highest number of moderately resistant rice accessions (50).
Relationship between AFLP markers and blast disease resistance/susceptibility
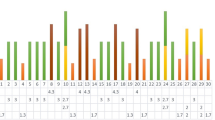
Among the major genetic clusters identified based on 77 polymorphic AFLP markers, we observed a sub-cluster A within cluster 2 that contained the highest number of resistant accessions compared to cluster 1 and 3. This affinity in dispersal of accessions reaction spectrum, gives an indication that AFLP markers may help to detect genetic differences in the resistance levels of rice accessions by distances correlation (Fig. 1). LDA using the total of 77 AFLP markers was performed (Fig. 3) revealing a significant association of AFLPs to accessions’ reactions in the field (R2 = 0.37). A series of models were performed to better examine the relation of genetic diversity with blast disease. Stepwise regression analysis revealed that only 13 out of 77 polymorphic markers were necessary to explain accurately accessions reactions pattern (Table 4). Based on 13 most significant AFLP markers, we attempted to predict three disease classes (Resistant: DI < 3, moderate resistant: 3 < DI < 4 and susceptible with DI > 4) by a General Linear Model (GLM). Thi s GLM showed significant coefficients (P < 0.05) for a total of seven markers with disease status (Table 5). The probability of accessions assignment to the right disease class was 68%. Better results were obtained when we considered only two disease classes (DI < 3 and DI > 3). In case a binary classification was used, the algorithm was able to explain 90% of the responses and the coefficients of a number of eight markers were found to be significant (Suppl. Table 2).
Genetic differentiation for response of 350 rice accessions to blast disease by Linear Discriminant Analysis using AFLP markers colours indicate the disease status: resistant accessions (Severity < 3) are written in red; moderate resistant (3 < Severity < 4) in blue; moderate susceptible (4 < Severity < 5) in green and susceptible (Severity > 5) are indicated in black. (Color figure online)
Among the significant markers, CTA22-366 with a size of 366 base pairs was particularly present in highly resistant accessions. Markers utility was such that at molecular level, when CTA22-366 was present, the observed status of the accessions was always resistant. But when this marker was absent, we found that accessions were either resistant or susceptible. A representative sub-set of 42 rice accessions (12% of the total germplasm) of the total 350 rice accessions studied was proposed (based on their molecular representativeness and diversified blast resistance responsiveness) for breeding purposes. This selected rice sub-set consisted of 26 resistant accessions, 9 moderately resistant and 7 susceptible accessions.
Discussion
The PIC value obtained for the studied germplasm collection (mean = 24%) was lower than those reported by Semon et al. (2005) and Dramé et al. (2011) who assessed genetic diversity of 198 and 82 O. glaberrima accessions, respectively using Simple Sequence Repeats (SSR) markers. They reported PIC values of 34 and 52%, respectively. According to Vieira et al. (2016), there are some advantages of SSR markers in polymorphism detection due to multi-allelic and codominant nature. Furthermore, a higher accessions number (350) was examined in the present study and might contain more genetically similar samples.
The partitioning of molecular variance in the total germplasm indicated significant genetic variation both among clusters (23%) and within clusters (77%). Genetic diversity was found to be more within than between clusters. High genetic variance estimates within-groups ranging from 71 to 95% have been reported in earlier studies of African cultivated rice (Ming et al. 2010; Salem and Sallam 2016; Ndjiondjop et al. 2017). Our results provide a global insight into how the germplasm stratification manifests, which revealed an asymmetric division of the collection. Cluster 2, including all O. sativa accessions from Benin as well as the control WAB0006881 showed the highest level of genetic diversity. This result is in close agreement with many other studies, suggesting a significant reduction in genetic diversity in O. glaberrima (Li et al. 2011; Wang et al. 2014). Within cluster 2, the sub-cluster A grouped 20 O. glaberrima accessions together with the six O. sativa accessions. As it sometimes happens that farmers grow both species in the same field, it is possible there was a gene flow between O. sativa and O. glaberrima. Spontaneous interspecific hybridizations is not uncommon in rice fields (Second 1982; Barry et al. 2007; Nuijten et al. 2009). Thereby, the pollen flow is usually from O. sativa to O. glaberrima (Sano 1989). According to Jusu (1999) and Semon et al. (2005), interspecific hybridizations result in new varieties.
Among all screened rice accessions, 12% were highly resistant and 24% were moderately resistant to blast disease. The other accessions were moderately susceptible or susceptible (29 and 35%, respectively). Results suggest a large genetic variability in blast resistance studied within the germplasm. Accessions showing high blast disease resistance might possess a single resistance gene. Quantitative variability in blast resistance resulting from multiple gene control could be detected within the category of moderately resistant accessions (Vasudevan et al. 2014). Out of 353 tested rice accessions (including the three controls), 50 were moderately resistant in cluster 1, whereas only 15 and 23 moderately resistant were found in cluster 2 and cluster 3, respectively. This suggests a higher level of quantitative resistance within cluster 1 than within the other clusters. It would be interesting to compare field resistance levels with artificial inoculation results using specific Beninese blast isolates. There is also a need to evaluate the yield of the identified resistant material for breeding purposes. Genebanks and breeders will more likely conserve O. glaberrima germplasm when it is high-yielding (Wambugu et al. 2013). The remaining, lower-yielding resistant germplasm can be used by breeders as resistance gene donors.
Diversity in cultivated rice accessions was found to correlate with varying phenotypic blast disease reactions. Based on their AFLP pattern, susceptible and resistant accessions could be distinguished. A number of AFLP markers were significantly associated with blast resistance. These associations were strongest when using CTA22-366 marker. Marker utility for blast resistance screening will be examined through future genetic studies by developing hybrid individuals and backcross populations. This study suggested a total of 13 AFLP markers which have the potential to identify a highly variable germplasm core set that is easier to evaluate and would facilitate the conservation and use in future breeding programs. Particularly, this will help formulate future strategies for improving rice blast resistance. Clustering identified accession groups that are genetically distinct from and that included different levels of blast resistance. Plant genetic diversity information is important pre-breeding information for selecting parents that can be used in intra and inter-crossing to broaden the genetic base of modern rice cultivars (Thakur et al. 2015). The choice of donor parents could be based on the knowledge on the accession’s genetic group. AFLPs was confirmed to be a very useful molecular tool for exploration of relationship of genetic diversity with blast resistance. This new resistant germplasm is an additive value for broadening horizon towards the identification of new blast R genes.
References
Afouda L, Gnikpo S, Sere Y, Gumedzoe Y (2007) Prevalence des principales maladies sur des varietes de riz cultivees dans le departement de l’alibori au Nord-Benin. J de la Recherche Scientifique de l’Universite de Lome 9:1027–1988
Baboy L, Mateso R, Sabiti K (1995) Tests de résistance au champ vis-à-vis de la pyriculariose du riz au Zaire. Tropicultura 13:93–98
Ballini E, Morel JB, Droc G, Price A, Courtois B, Notteghem JL, Tharreau D (2008) A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microbe Interact 21:859–868
Barry MB, Pham JL, Noyer JL, Billo C, Courtois B, Ahmadi N (2007) Genetic diversity of the two cultivated rice species (O. sativa & O. glaberrima) in Maritime Guinea. Evidences for inter-specific recombination. Euphytica 154:127–137
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Couch BC, Kohn LM (2002) A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae, from M. grisea. Mycologia 94:683–693
Dai Y, Jia Y, Correll J, Wang X, Wang Y (2010) Diversification and evolution of the avirulence gene AVR-Pita1 in field isolates of Magnaporthe oryzae. Fung Genet Biol 47:973–980
Dramé KN, Sanchez I, Gregorio G, Ndjiondjop MN (2011) Suitability of a selected set of simple sequence repeats (SSR) markers for multiplexing and rapid molecular characterization of African rice (Oryza glaberrima Steud.). Afr J Biotechnol 10:6675–6685
Futakuchi K, Sié M (2009) Better exploitation of African rice (Oryza glaberrima Steud.) in varietal development for resource-poor farmers in West and Central Africa. Agric J 4:96–102
Hittalmani S, Parco A, Mew TV, Zeigler RS, Huang N (2000) Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. Theor Appl Genet 100:1121–1128
IRRI (International Rice Research Institute) (2013) Standard evaluation system for rice, 5th edn. International Rice Research Institute, Manila
Jia Y, Bryan GT, Farrall L, Valent B (2003) Natural variation at the Pi-ta rice blast resistance locus. Phytopathology 93:1452–1459
Jones MP, Dingkuhn M, Aluko GK, Semon M (1997) Interspecific Orza sativa × O.glaberrima Steud. Progenies in upland rice improvement. Euphytica 92:237–246
Jusu MS (1999) Management of genetic variability in rice (Oryza sativa L. and O. glaberrima Steud.) by breeders and farmers in Sierra Leone. PhD-thesis Wageningen University, The Netherlands
Li ZM, Zheng XM, Ge S (2011) Genetic diversity and domestication history of African rice (Oryza glaberrima) as inferred from multiple gene sequences. Theor Appl Genet 123:21–31
Linares OF (2002) African rice (Oryza glaberrima): history and future potential. Proc Natl Acad Sci USA 99:16360–16365
Mackill DJ, Bonman JM (1992) Inheritance of blast resistance in near-isogenic lines of rice. Phytopathology 82:746–749
Ming H, Fang-min X, Li-yun C, Xiang-qian Z, Jojee L, Madonna D (2010) Comparative analysis of genetic diversity and structure in rice using ILP and SSR markers. Rice Sci 17:257–268
National Research Council (1996) Lost crops of Africa. Volume 1: grains. National Academy Press, Washington, p 380
Ndjiondjop M-N, Semagn K, Gouda AC, Kpeki SB, Dro Tia D, Sow M, Goungoulou A, Sie M, Perrier X, Ghesquiere A, Warburton ML (2017) Genetic variation and population structure of Oryza glaberrima and development of a mini-core collection using DArTseq. Front Plant Sci 8:1748. https://doi.org/10.3389/fpls.2017.01748
Nuijten E, van Treuren R, Struik PC, Mokuwa A, Okry F, Teeken B, Richards P (2009) Evidence for the emergence of new rice types of interspecific hybrid origin in West African farmers’ fields. PLoS ONE 4(10):e7335. https://doi.org/10.1371/journal.pone.0007335
Odjo T, Ahohuendo BC, Onasanya A, Akator K, Séré Y (2011) Analysis of Magnaporthe oryzae population structure in Benin. Afr J Agric Res 6:6183–6188
Pham JL (1992) Evaluation des ressources génétiques des riz cultivés en Afrique par hybridation intra et interspécifique. Thèse Docteur et sciences, Université de Paris XI ORSAY (France), p 236
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
RoyChowdhury M, Jia Y, Cartwright RD (2012) Structure, function and coevolution of rice blast resistance genes. Acta Agric Sin 38:381–393
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci 81(24):8014–8018
Salem KFM, Sallam A (2016) Analysis of population structure and genetic diversity of Egyptian and exotic rice (Oryza sativa L.) genotypes. C R Biol 339:1–9
Sano Y (1989) The direction of pollen flow between two co-occurring rice species, Oryza sativa and O. glaberrima. Heredity 63:353–357
Second G (1982) Origin of the genic diversity of cultivated rice (Oryza spp.): study of the polymorphism scored at 40 isozyme loci. Jpn J Genet 57:25–57
Semon M, Nielsen R, Jones MP, McChouch SR (2005) The population structure of African cultivated rice O. glaberrima (Steud.): evidence for elevated levels of linkage disequilibrium caused by admixture with O. sativa and ecological adaptation. Genetics 169:1639–1647
Séré Y, Fargette D, Abo ME, Wydra K, Bimerew M, Onasanya A, Akator SK (2013) Managing the major diseases of rice in Africa. In: Wopereis MCS, Johnson DE, Ahmadi N, Tollens E, Jalloh A (eds) Realizing Africa’s rice promise. CABI (H ISBN 9781845938123), pp 2013–228
Sharma T, Rai A, Gupta S, Vijayan J, Devanna B, Ray S (2012) Rice blast management through host-plant resistance: retrospect and prospects. Agric Res 1:37–52
Sié M, Ogunbayo SA, Dakouo D, Sanou I, Dembélé Y, N’dri B, Dramé KN, Sanni KA, Toulou B, Glele RK (2010) Evaluation of intra and interspecific rice varieties adapted to valley bottom conditions in Burkina Faso. Afr J Plant Sci 4:308–318
Thakur S, Singh PK, Das A, Rathour R, Variar M, Prashanthi SK, Singh AK, Singh UD, Chand D, Singh NK, Sharma TR (2015) Extensive sequence variation in rice blast resistance gene Pi54 makes it broad spectrum in nature. Front Plant Sci 6:345. https://doi.org/10.3389/fpls.2015.00345
Thiémélé D, Boisnard A, Ndjiondjop M, Chéron S, Séré Y, Aké S, Ghesquière A, Albar L (2010) Identification of a second major resistance gene to Rice yellow mottle virus, RYMV2, in the African cultivated rice species, O. glaberrima. Theor Appl Genet 121:169–179
Vasudevan K, Vera Cruz CM, Gruissem W, Bhullar NK (2014) Large scale germplasm screening for identification of novel rice blast resistance sources. Plant Sci 5:1–9. https://doi.org/10.3389/fpls.2014.00505
Vieira MLC, Santini L, Diniz AL, Munhoz CF (2016) Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol 39:312–328
Vodouhe SR, Ojegui M, Amadji F (1981) Impact of blast on rice cultivation in People’s Republic of Benin. In: Proceedings of the symposium on rice resistance to blast, Montpellier, France, 18–21 March. Service de Pathologie Végétale, Montpellier, France, pp 27–33
Vos P, Hogers R, Bleeker M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23:4407–4414
Wambugu PW, Furtado A, Waters DL, Nyamongo DO, Henry R (2013) Conservation and utilization of African Oryza genetic resources. Rice 6:29
Wang JC, Wen JW, Liu WP, Yan SR, Wang JQ, Ren JP (2010) Interaction studies between rice and Pyricularia grisea in Jilin Province, P. R. China. In: Jia, Y (ed) Proceedings of the 5th international rice blast conference, USA. USDA-DBNRRC, p 90
Wang JC, Jia Y, Wen JW, Liu WP, Liu XM, Li L, Jiang ZY, Zhang JH, Guo XL, Ren JP (2013) Identification of rice blast resistance genes using international monogenic differentials. Crop Prot 45:109–116
Wang M, Yu Y, Haberer G, Marri PR, Fan C, Goicoechea JL, Zuccolo A, Song X, Kudrna D, Ammiraju JS, Cossu RM (2014) The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat Genet 46:982–988
WARDA (1999) Program report 1996–1997. West Africa Rice Development Association, Bouaké, Côte d’Ivoire
Wisser RJ, Sun Q, Hulbert SH, Kresovich S, Nelson RJ (2005) Identification and characterization of regions of the rice genome associated with broad-spectrum, quantitative disease resistance. Genetics 169:2277–2293
Zeigler RS, Tohme J, Nelson R, Levy M, Correa Victoria FJ (1994) Lineage exclusion: a proposal for linking blast population analysis to resistance breeding. In: Zeigler RS, Leong SA, Teng PS (eds) Rice blast disease. CAB International, Wallingford, pp 267–292
Acknowledgements
Funding of this Research work by the Monsanto’s Beachell-Borlaug International Scholars Program (MBBIS) and the Flemish Fund for Scientific Research (BOF) with support of AfricaRice. Special thanks to the AfricaRice genebank for providing seed and the related information.
Authors contribution
OINY carried out the field works, the genotyping, data analysis and drafted the manuscript. KA participated in project design, genotyping, data analysis and revised the manuscript. SL participated in statistical analysis of data and revised the manuscript. DS, AD, WV, PVD, and GH participated in project design, data analysis and revised the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yelome, O.I., Audenaert, K., Landschoot, S. et al. Exploring genetic diversity and disease response of cultivated rice accessions (Oryza spp.) against Pyricularia oryzae under rainfed upland conditions in Benin. Genet Resour Crop Evol 65, 1615–1624 (2018). https://doi.org/10.1007/s10722-018-0638-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-018-0638-1