Abstract
As a counterweight to genetic erosion, landraces could become important germplasm resources for breeding. In the present study the genetic diversity of 62 Greek and Eastern European oat accessions (seven cultivars and 55 common and red oat landraces collected from 1904 to 1960) was studied using simple sequence repeats and 31 morphological descriptors. High levels of polymorphism were detected and 209 distinctive alleles were identified; on average 14.65 alleles per primer. Both, the principal component analysis derived from the morphological data, and the unweighted pair group method with arithmetic mean from the molecular analysis clustered the oat genotypes according to their type (common and red oats). In addition, the analysis of molecular variation showed that there were noteworthy differences within designated geographic regions. Moreover, a more discrete subpopulation structure was identified based on allele frequency using a Bayesian clustering approach. A clear distinction was evident among the red and common oats suggesting that they should be considered as separate species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oat is a western Mediterranean cereal (Loskutov 2008) with a fairly recent agricultural history, since its cultivation began a few millennia later than the one of wheat and barley, and archaeological records documented oats—along with rye—as weed contaminants (Newton et al. 2010). In more recent years, this grain was mainly used as hay or silage; but its’ main utilization has been as a livestock feed. Nowadays, oat is considered as one of the most important cereal crops in the world (being widely used for human consumption as a source of valuable nutrients; Boczkowska and Tarczyk 2013) and has received considerable attention with respect to collection and conservation.
More than 200,000 accessions of wild and cultivated Avena species are stored in seed gene banks worldwide (Diederichsen 2008). Landraces in particular, made a significant contribution to the current oat germplasm, since several modern varieties trace their origin back to a restricted number of them. Different definitions for landraces have been given through time, the most comprehensive being “landrace is a highly diverse population or mixture of genotypes naturally developed in a certain region under the influence of the regionally prevailing conditions of climate, soil and management, without or with only little mass selection” (Banga 1944). In contemporary breeding approaches, it is of great importance to preserve, characterize and utilize plant material that can provide useful genes in the genetic pool of the cultivated oats. A large number of oat cultivars was derived from individual selections from landraces, or crosses involving these selections. For instance, more than 100 cultivars were developed worldwide from ‘Red Rustproof’ oat landrace (Coffman 1977).
However, as a result to monoculture and pure lines plant breeding, landraces were widely replaced by modern cultivars in farming (Hammer et al. 2003). Furthermore, each modern cultivar presents a fraction of the genetic diversity comprised in the source population or the species (Wesenberg et al. 1992); hence the oat germplasm is suffering an ongoing genetic errosion (Loskutov 1998; Fu et al. 2003). Since the loss of genetic diversity has unpredictable consequences, a continuing effort is required to conserve, characterize and maintain wild populations, old landraces, and Avena populations in general.
The goal of the current study was to uniquely fingerprint and examine the genetic diversity existing among mostly Greek oat landraces using simple sequence repeats (SSR) molecular markers and morphological agronomic traits. The genetic structure of this germplasm collection was studied using a model-based Bayesian clustering method to assign genotypes to distinct gene pools. This study also offers vital baseline information for the association of Greece’s locally preserved germplasm with landraces of broader Eastern European origin, for the first time in literature.
Materials and methods
Plant material
A total of 55 Avena landraces originally collected from 1904 to 1960 (acquired from the USDA, Vavilov Institute of Plant Genetic Resources-VIR and the Agricultural Research Institute of Cyprus-ARI), plus seven commercial cultivars were used in the current study (Table 1).
Morphology analysis
The entries were characterised at the experimental fields of NAGREF (Cereals Institute, Thermi-Thessaloniki, Greece) in unreplicated field plots. Seeding for the evaluation was performed in autumn (mid-November 2013) and harvesting in summer (mid-June 2014). Climate conditions are reported in Online Resource 1. Thirty-one morphological primary descriptors—according to the ECPGR Avena descriptors (IBPGR 1985)—were recorded from at least 30 plants within each entry (Online Resource 2). SPSS Statistics v 20 (IBM) was used to construct a principal component analysis (PCA) plot from the combined morphological descriptors.
DNA extraction and PCR amplification
Ten leaves from 15-day-old individual seedlings were harvested and DNA was extracted using the NucleoSpin® kit (Macherey–Nagel GmbH & Co, Düren, Germany) following the manufacturer’s instructions. DNA concentration was determined spectrometrically and its quality was established by agarose gel electrophoresis. After an initial screening, ten carboxyfluorescein (FAM) labeled SSR primer pairs derived from an oat genomic library (Li et al. 2000) were chosen because of their consistency in amplification and polymorphism in the oat panel (Table 2).
Amplification reactions were set up in a 10 μL volume of a mixture containing 25 ng of genomic DNA, 1x Type-it® Multiplex PCR Master Mix (Qiagen, Venlo, Limburg, Netherlands) and 0.1 μM of each primer. PCR amplification was performed in a Bio-Rad PTC-200 thermocycler (Applied Biosystems, Foster City, CA, USA) under the following temperature profile: 5 min at 95 °C followed by 35 cycles, each one included 30 s at 95 °C, 90 s at 57 °C, 30 s at 72 °C and a final extension for 30 min at 60 °C. SSR markers were analyzed on an ABI 3130 genetic analyzer. Size standard GeneScan 500 LIZ™ (Applied Biosystems, Foster City, CA, USA) was included with each sample to define allele sizes. Data were analyzed using GeneMapper ® (Applied Biosystems, Foster City, CA, USA).
Data analysis
Allele fragments were converted into a binary data matrix table. FreeTree (Pavlicek et al. 1999) was used to calculate the genetic similarities among taxa and for the construction of the bootstrapped dendrogram, which was depicted with the implementation of TreeView (http://taxonomy.zoology.gla.ac.uk/rod/treeview.html). The level of primer informativeness was estimated by means of the discriminating power (Dj), as described by Tessier et al. (1999). Genotypic variations were assessed across various populations by means of analysis of molecular variance (AMOVA) using GenAlEx 6.4 (Peakall and Smouse 2006). The significance of the resulting variance components and the inter-population genetic distances were tested using 999 random permutations. Mantel test was conducted using the XLSTAT statistical software (Addinsoft), in order to test the goodness of fit between morphological and molecular markers.
Bayesian model-based clustering approach to identify the genetic structure in the oat germplasm was performed using Structure 2.3.4 (Pritchard et al. 2000). The structure algorithm was run using the admixture model, with 10 independent replicate runs per K value (number of clusters) ranging from 1 to 10. Each run involved a burning period of 100,000 iterations and a post burning simulation length of 100,000. Validation of the most likely number of clusters K was performed with the Structure Harvester (http://taylor0.biology.ucla.edu/structureHarvester).
Results
Diversity and morphological characters
Differences were recorded between oat accessions for most of the descriptors (Online Resource 2). Among the agronomic descriptors, plant height was a variable trait that ranged from 105 (PI 264854) to 160 cm (RCAT013285), with an average of 131.33 cm. Variations were also recorded for leaf length (34.33 cm for PI 258580 vs 53.97 cm for RCAT012905, with an average of 44.12 cm) and leaf width (1.5 cm for PI 258580 vs 3.17 cm for NLD037 3694, with an average of 2.19 cm). Days to heading ranged from 161 days (PI 258580 and PI 264838) to 179 days (RUS00110528). Finally, the size of grain (weight of 1000 seeds) was also a variable character, since it fluctuated from 13 (RUS00111655) to 37 g (PI 264845).
Growth habit was rather a uniform trait since the majority of landraces (82 %) had a semi prostrate growth habit; seven Greek landraces had a prostrate growth habit and only two Greek landraces (CZE047 03C070360 and CZE047 03C070417) had an erect growth habit. Stem thickness was rather constantly thin and intermediate thin, since only five eastern European landraces (NLD037 3694, CZE047 03C070203, CZE047 03C070206, CZE047 03C070359 and RUS00110528) had a thick stem. Strikingly, only one genotype (RUS00111655) was found to have a unilateral panicle, naked grain and four fertile spikelets per floret. Finally, all landraces had low shedding at maturity (Online Resource 2).
Thirty-one scored attributes were combined in order to produce a PCA plot. The plot analysis classified the oat genotypes into two groups that mainly reflected their geographical origin (Greek and Eastern European) and their type (common—red oats; Fig. 1). Furthermore, it was detected that certain Greek landraces were more discrete from the core of the groups, having a greater heterogeneity.
Plot of PCA analysis of the combined 31 morphological data. Black encircled, are the Greek landraces. Numbers correspond to accessions shown in Table 1
Genetic diversity of oat landraces and cultivars
The SSR primers used to assess the genetic information in the oat collection, revealed a high level of diversity in landraces displaying a total of 209 alleles, with an average of 14.65 alleles per primer. The majority of alleles were polymorphic, since only five of them were common among all genotypes. The total number of alleles per marker varied from three (primer AM38) to 39 (primer AM3; Table 2), while 91 different electrophoretic patterns were detected. Furthermore, 49 alleles were unique, while 105 alleles had a frequency lower than 0.05. The discrimination power (Dj) varied from 0.29 (primer AM38) to 0.99 (primer AM04) with a mean of 0.83, illustrating that this primer set was very informative.
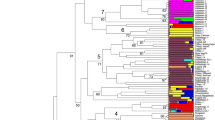
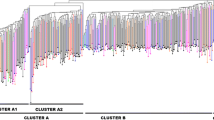
Genetic similarity calculated among the oat collection varied from 0.36 (landraces PI 258580 and NLD037 3695) to a maximum of 0.91 (landraces NLD037 3693 and NLD037 3695, as well as, landraces ARI 3-7 and ARI 6-2) with a mean similarity of 0.56. The UPGMA dendrogram classified the 62 Avena genotypes into two main clusters (100 % bootstrap value) according to their origin (Fig. 2). The first group which was the largest, contained all the oat landraces of Eastern European origin, a few Greek landraces and the majority of the commercial cultivars. Moderate to high bootstrapped values were detected among the oat landraces, while the highest genetic affinity was detected among two genotypes from Turkey (NLD037 3693 and NLD037 3695). Interestingly, the Greek landraces and variety ‘Flega’ that were clustered in this group, had low bootstrap supporting values. The second group included the majority of Greek, all the Cypriot landraces and the commercial cultivar ‘Kassandra’. The UGPMA clustering carried out for the SSR data showed a noteworthy within group difference among geographic regions of Greece and Cyprus, well supported by significant bootstrap values (Fig. 2). The highest affinity within the group was detected among the Greek landraces PI 264840 and PI 264843. Correlation coefficient among the morphological assessments and the molecular data was significant (r = 0.51, p < 0.001).
Genetic structure of oat landraces and cultivars
The pattern of structure and the allocation of genetic diversity were additionally analyzed with a Bayesian-based approach implemented in the Structure 2.3.4 software (Pritchard et al. 2000). We investigated the range from K = 1 to K = 10 and calculated the posterior probability for each value of K using the estimated log likelihood of K. The obvious optimum for the ad hoc quantity based on the second order rate of change of the likelihood function with respect to ΔK was observed for K = 2 (Fig. 3). As a result, the entire oat accessions (four regional populations) were successfully assigned to two groups (K = 2, ΔK = 4.15); inferred clusters were calculated with more than 90 % probability intervals. In addition, a minor peak at K = 4 (ΔK = 1.09) was identified (data not shown). Structure analysis divided the oat genotypes into two groups.
Bayesian cluster analysis of the optimum K cluster. 1: Greek landraces, 2: Cypriot landraces, 3: Eastern European landraces and 4: commercial cultivars. Percentages of genotype membership to clusters as inferred for K = 2 and K = 4.The color in each bar plot represents the probability of each individual belonging to a given group
The first population (Greek genotypes) was assigned to group I with a 0.764 proportion of membership. The 32 accessions of Greek origin contained several fairly admixed genotypes. Eventhough the majority of these entries had a group I (red) membership value higher than 0.9, there were some landraces that had admixed genotypes and were assigned to group II (green; landraces PI 264849, PI 264850, RUS00110218, RUS00110220, RUS00111516, RUS00111655 and ‘Flega’). Cypriot accessions (five landraces) were also clustered in the first group, having a 0.968 proportion of membership. Furthermore, this population contained five rather genetically homogenous genotypes, since the only entry having a membership value lower than 0.9 (ARI 1–5). All Eastern European oats were assigned to group II with a 0.978 % of membership. Finally, the fourth population contained the remaining five commercial varieties (‘Firth’, ‘Wistar’, ‘Lutz’, ‘Fuchs’ and ‘Freddy’) that were assigned to group II having a 0.879 proportion of membership. Bayesian clustering and admixture analysis assisted the quantification of associations among oat landraces, according to their origin and revealed the presence of some sort of hierarchical structure in our entries. Furthermore, structure analysis for K = 4 permitted us a better understanding of a more detailed genetic association, since it became evident that the commercial varieties were affiliated to specific Greek and Eastern European landraces (Fig. 3).
Analysis of molecular variance revealed that a high proportion of the total genetic diversity (78 %) was allocated within the four populations. The highest variability was recorded for the Greek genotypes (SS = 439.125), followed by Eastern European landraces (SS = 266.750) and commercial cultivars (SS = 52.800). Furthermore, it was also revealed that the genetic distance between clusters was significant (Fst = 0.219; p = 0.010). Fst values suggested the presence of divergence between populations. The closest genetic proximity was found among the Eastern European and commercial cultivars (Fst = 0.138; p = 0.001) and the more diverged populations were the Cypriot and the commercial cultivars (Fst = 0.415; p = 0.005).
Discussion
Agronomic traits and morphological analysis has been repeatedly used when characterizing large data sets of Avena taxa in genebanks (Diederichsen 2008, 2009), or smaller sets of oat species, cultivars and landraces (Souza and Sorrells 1991; Sheikhehpour et al. 2014; Boczkowska et al. 2014). As a result, morphological description has become a valuable source of information for breeding and agronomic research programs (Boczkowska et al. 2014). In the current study and in order to study the genetic diversity in our oat collection, 31 morphological traits were scored. It was detected that the resulting grouping reflected the principal area of origin; since Greek landraces were typically clustered separately from Eastern European oat landraces.
Eventhough morphological traits are generally employed in order to estimate genetic variation since their measurements are not laborious, still, diversity estimation based on morphology alone has limitations. Unfortunately, traits are heavily influenced by the environment, are limited in number and possibly unintentional selection for traits with agronomical value (i.e. selection for lightly colored seeds etc.) has further reduced them. As a result, oats germplasm has also been studied by a number of molecular markers types; for example AFLP (Fu et al. 2004, 2005), ISSR (Boczkowska and Tarczyk 2013) RAPD (Baohong et al. 2003), SSR (Li et al. 2000, 2007; Fu et al. 2003; Nersting et al. 2006; Montilla-Bascón et al. 2013), or with an array of combined molecular techniques (He and Bjørnstad 2012; Boczkowska et al. 2014). In the current study, microsatellites were employed to estimate the genetic diversity in the oat collection, since they can be used to examine the allelic diversity changes and could be linked in the future to genomic regions that control diverse traits; thus enabling efficient and precise transfer of useful alleles from landraces to modern cultivars (Montilla-Bascón et al. 2013).
The molecular marker genetic analysis of the oat collection revealed a high level of variability among accessions. Moreover, this diversity was additionally apparent within the common oat (A. sativa L.) landrace germplasm that exhibited a total of 35 unique alleles, followed by the red oats (A. byzantina C. Koch), and the commercial cultivars, with eight and six unique alleles, respectively. Twenty three common oat landraces had unique alleles (ten of them were Greek), while three out of five Cypriot genotypes also contained unique alleles; something that underlines the distinctiveness of this germplasm. Finally, five Greek A. byzantina landraces (out of 15) had unique alleles. Montilla-Bascón et al. (2013) inquired the genetic diversity and the population structure among oat cultivars and reported that genetic variability was more evident in the common oat germplasm. Nonetheless, Diederichsen (2008) reported that eventhough red oat (byzantina type) accessions account for the 11.6 % of the PGRC oat collection still they covered the 34.7 % of all morphological groups. This means, that red oats are much richer in variety, since they comprise a great proportion of morphological groups. This is an indication that a significant conservation of genetic variation exists in landraces and thus offering opportunities for oat breeding.
Unfortunately, to date the vast genetic diversity of oats has not been fully utilized. Even though 564 A. byzantina landraces are catalogued in the EURISCO database (http://eurisco.ipk-gatersleben.de/), only 203 red oat cultivars have been developed. On the contrary, 1756 A. sativa landraces occur and 4464 common oat cultivars have been produced respectively. This has a major impact on the genetic erosion of the species. Fu et al. (2003) proved that modern plant breeding reduces genetic diversity by studying oat cultivars released from 1886 to 2001. They concluded that a significant decrease of alleles was evident in cultivars bred after 1970, with severe consequences to pest and disease resistance. In that term, the genetic material characterized in the current study has a great significance; since several cultivars were bred from its genetic pool. For instance, the Greek landrace CIav 357 collected from Trace (Greece) in 1904 (http://www.ars-grin.gov/), was used for the selection of ‘Markton’, a globaly milestone variety, that further produced numerous smut and rust resistant oats varieties.
In the present study it was detected from both morphological (Fig. 1) and molecular data (Fig. 2) that there is a clear disparity among common and red oat landraces, as well as, a clear geographical clustering, which is in agreement with previous studies (Odonoughue et al. 1994; Fu et al. 2005; Newell et al. 2011; Montilla-Bascón et al. 2013). Avena sativa cultivars differed substantially from the A. sativa landraces, and, for K = 4, they emerged as a separate group (Fig. 3). However, the red oat cultivar (‘Kassandra’) was clustered with the red oat landraces, signifying either lesser improvement of the red oat (A. byzantina) cultivars in contrast to common oats (A. sativa), or due to the more ‘active’ participation of red oat landraces in its lineage (Montilla-Bascón et al. 2013). Furthermore, it was established that the primary region of landraces, correlates to the degree of genetic affinity. This has also been reported from Diederichsen (2008) that concluded a strong positive correlation between the number of accessions originating from a given country and the number of morphological groups found.
Data support the assumption that common and red oats were domesticated independently of each other (Zohary and Hopf 2000). Loskutov (2008) proposed that A. byzantina had a different evolution history than A. sativa. It was suggested that the big-seeded A. sterilis L. endured modifications in the mode of floret dispersal and led to the establishment of the cultivated form of A. byzantina, while A. sativa was evolved from the smaller-seeded forms of A. ludoviciana Durieu (Loskutov and Rines 2011). Furthermore, the red oats are correlated to the Mediterranean climate and are winter annuals, opposed to common oats that are grown as spring annuals at temperate-climate geographical zones (Coffman 1961). However, the biological concept of oats contradicts to the taxonomy of the Avena species. Rodionova et al. (1994) identified 32 A. sativa and 17 A. byzantina botanical varieties and additionally distinct 19 botanical forms; summing up to 62 morphologically distinguishable taxonomical types within the cultivated hexaploid oat (Diederichsen 2008). Danert (1972) categorized 19 botanical varieties of A. sativa and considered A. byzantina as a distinct species. On the contrary, Baum (1977) suggested that A. byzantina is not as a separate Avena species and it should be regarded as A. sativa. This has caused great nomenclature confusion, especially among different genebanks and thus it is realistic to accept that effective communication regarding oat genetic resources is far from accomplished when it comes to formal taxonomical names (Diederichsen 2004).
Furthermore, to make matters even more complicated, the mutual presence of both Avena types in the same regions over the centuries, has partially homogenized their genetic background. Especially in the northern region of Greece, where both the common hexaploid and red oat species overlap, such an admixture is expected. As a consequence, some Greek landraces were clustered among accessions of the Balkan region, while certain Southern Greek and Cypriot A. sativa accessions were grouped with the red oat landraces. This was also reported by Montilla-Bascón et al. (2013), that noted that some landraces described as common oats were grouped with red oat accessions and shared a specific A. byzantina allele. As a result, it was concluded that geographic proximity of these common oat accessions with their most related red oat landraces resulted from a potential cross among the landraces.
Interestingly, all Greek common oat landraces that were donated from VIR (RUS00110218, RUS00110220, RUS00111516 and RUS00111655) were clustered with the Eastern European A. sativa landraces. This may be attributed to the substantial germplasm exchange among different regions of origin that escalated during the last 150 years and has made it doubtlessly challenging to discriminate accessions between geographical origin and certain character states (Diederichsen 2008). Thus, the current diversity and genetic markup of Avena landraces maintained, may differ from the endemic landraces collected from Vavilov in the beginning of the twentieth century (Diederichsen 2008).
Conclusion
Overall, both morphological and molecular data support noteworthy divergence among designated geographical areas. Furthermore, a more distinct subpopulation organization was identified grounded on allele frequency using a Bayesian assembling method. A strong discrepancy was apparent among the red and common oats suggesting that they ought to be considered as distinct species.
References
Banga O (1944) Veredeling van tuinbouwgewassen. Zwolle, pp 211
Baohong G, Zhou X, Murphy JP (2003) Genetic variation within Chinese and western cultivated oat accessions. Cereal Res Commun 31:339–346
Baum BR (1977) Oats: wild and cultivated, a monograph of the genus Avena L. (Poaceae). Canada Department of Agriculture, Ottawa
Boczkowska M, Tarczyk E (2013) Genetic diversity among Polish landraces of common oat (Avena sativa L.). Genet Resour Crop Evol 60:2157–2169
Boczkowska M, Nowosielski J, Nowosielska D, Podyma W (2014) Assessing genetic diversity in 23 early Polish oat cultivars based on molecular and morphological studies. Genet Resour Crop Evol 61:927–941
Coffman FA (1961) Origin and history. In: Coffman FA (ed) Oats and oat improvement. American Society of Agronomy, Madison, pp 15–40
Coffman FA (1977) Oat history, identification and classification. USDA-ARS Tech. Bull. No. 1516. U.S. Gov. Print. Office, Washington, DC
Danert S (1972) Zur Systematik von Avena sativa L. und zur Beurteilung von Avena nuda Hoj. [Taxonomy of Avena sativa L. and interpretation of Avena nuda Hojer]. Kulturpflanze 19:45–51 (in German)
Diederichsen A (2004) Case studies for the use of infraspecific classifications in managing germplasm collections of cultivated plants. Acta Hortic 634:127–139
Diederichsen A (2008) Assessments of genetic diversity within a world collection of cultivated hexaploid oat (Avena sativa L.) based on qualitative morphological characters. Genet Resour Crop Evol 55:419–440
Diederichsen A (2009) Duplication assessments in Nordic Avena sativa accessions at the Canadian national genebank. Genet Resour Crop Evol 56:587–597
Fu YB, Peterson GW, Scoles G, Rossnagel B, Schoen DJ, Richards KW (2003) Allelic diversity changes in 96 Canadian oat varieties released from 1886 to 2001. Crop Sci 43:1989–1995
Fu YB, Kibite S, Richards KW (2004) Amplified fragment length polymorphism analysis of 96 Canadian oat varieties released between 1886 and 2001. Can J Plant Sci 84:23–30
Fu YB, Peterson GW, Williams D, Richards KW, Fetch MJ (2005) Patterns of AFLP variation in a core subset of cultivated hexaploid oat germplasm. Theor Appl Genet 111:530–539
Hammer K, Gladis T, Diederichsen A (2003) In situ and on-farm management of plant genetic resources. Eur J Agron 19:509–517
He X, Bjørnstad A (2012) Diversity of North European oat analyzed by SSR, AFLP and DArT markers. Theor Appl Genet 125:57–70
IBPGR (1985) Oat doscriptors. International Board for Plant Genetic Resources, Rome
Li CD, Rossnagel BG, Scoles GJ (2000) The development of oat microsatellite markers and their use in identifying relationships among Avena species and oat cultivars. Theor Appl Genet 101:1259–1268
Li RZ, Wang SW, Duan LS, Li ZH, Christoffers MJ, Mengistu LW (2007) Genetic diversity of wild oat (Avena fatua) populations from China and the United States. Weed Sci 55(2):95–101
Loskutov IG (1998) The collection of wild oat species of C.I.S. as a source of diversity in agricultural traits. Genet Resour Crop Evol 45:291–295
Loskutov IG (2008) On evolutionary pathways of Avena species. Genet Resour Crop Evol 55:211–220
Loskutov IG, Rines HW (2011) Avena. In: Kole C (ed) Wild crop relatives: genomic and breeding resources. Springer, Berlin, pp 109–183
Montilla-Bascón G, Sánchez-Martín J, Rispail N, Rubiales D, Mur L, Langdon T, Griffiths I, Howarth C, Prats E (2013) Genetic diversity and population structure among oat cultivars and landraces. Plant Mol Biol Rep 31:1305–1314
Nersting L, Andersen S, von Bothmer R, Gullord M, Jørgensen R (2006) Morphological and molecular diversity of Nordic oat through one hundred years of breeding. Euphytica 150:327–337
Newell MA, Cook D, Tinker NA, Jannink JL (2011) Population structure and linkage disequilibrium in oat (Avena sativa L.): implications for genome-wide association studies. Theor Appl Genet 122:623–632
Newton AC, Akar T, Baresel JP, Bebeli PJ, Bettencourt E, Bladenopoulos KV, Czembor JH et al (2010) Cereal landraces for sustainable agriculture. A review. Agron Sustain Dev 30(2):237–269
Odonoughue LS, Souza E, Tanksley SD, Sorrells ME (1994) Relationships among North-American oat cultivars based on restriction-fragment-length-polymorphisms. Crop Sci 34:1251–1258
Pavlicek A, Hrda S, Flegr J (1999) FreeTree-freeware program for construction of phylogenetic trees on the basis of distance data and bootstrapping/jack-knife analysis of the tree robustness. Application in the RAPD analysis of the genus Frenkelia. Folia Biol (Praha) 45:97–99
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure from multilocus genotype data. Genetics 155:945–959
Rodionova NA, Soldatov VN, Merezhko VE, Jarosh NP, Kobyljanskij VL (1994) Flora of cultivated plants, vol 2, part 3, Oat. Kolos, Moscow
Sheikhehpour S, Bahraminejad S, Cheghamirza K (2014) Morphological and molecular genetic variations of oat genotypes grown in Kermanshah, Iran. Mol Biol Rep 41:4023–4030
Souza E, Sorrells ME (1991) Relationships among 70 North American oat germplasm: I. Cluster analysis using quantitative characters. Crop Sci 31:599–605
Tessier R, David J, This P, Boursiquot JM, Charrier A (1999) Optimization of the choice of molecular markers for varietal identification in Vitis vinifera L. Theor Appl Genet 98:171–177
Wesenberg DM, Briggle LW, Smith DH (1992) Germplasm collection, preservation and utilization. In: Marshall HG, Sorrells ME (eds) Oat science and technology. American Society of Agronomy, Madison, pp 793–820
Zohary D, Hopf M (2000) Domestication of plants in the Old World, 3rd edn. Oxford University Press, Oxford
Acknowledgments
This work was partially supported by the start-up funding of AK from CUT.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nikoloudakis, N., Bladenopoulos, K. & Katsiotis, A. Structural patterns and genetic diversity among oat (Avena) landraces assessed by microsatellite markers and morphological analysis. Genet Resour Crop Evol 63, 801–811 (2016). https://doi.org/10.1007/s10722-015-0284-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-015-0284-9