Abstract
EST-SSR from Medicago truncatula Gaertn. and Glycine max (L.) Merr. were tested for transferability in various species of Onobrychis (O. pyrenaica Sennen, O. argentea Boiss. and O. viciifolia Scop.). Repeatable amplification was obtained for 81% of the microsatellites and 52% were polymorphic. Six selected SSRs from M. truncatula were used to fingerprint and estimate the genetic similarity of a set of 23 accessions of O. viciifolia. PCA analysis discriminated among the different Onobrychis species and the sainfoin accessions were clustered in a single major group. This grouping is discussed in terms of the history of cultivation of sainfoin in Spain. The selected SSRs will allow fingerprinting and genetic studies in Onobrychis species, solving the lack of available SSR markers in this species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sainfoin (Onobrychis viciifolia Scop.) is a traditional European fodder legume that thrives on well-drained alkaline soils in many parts of Central and Southern Europe. Cultivated sainfoin is an allotetraploid species (2n = 4x = 28, Hayot-Carbonero et al. 2010) obtained from wild botanical forms (Badoux 1965). Sainfoin domestication occurred at the end of the XVI century in the French part of the Rhine Valley. Soon, the common and giant types were selected: the first is characterized by its rusticity and persistence, the second by its higher production level and vigour (Gasparin 1846).
Despite their wide spread in Europe, a decline in the area of sainfoin, along with other temperate legumes, was mainly due to the expansion of grass production based on cheap inorganic nitrogen fertilizers in the 1970s. This intensive ruminant production system has contributed to high air and water pollution within the EU (Gill et al. 2009; Rochon et al. 2004). More recently, the move to more extensive and sustainable production systems has increased the interest for forage legumes since they offer a potential to mitigate these effects while providing local protein sources that are becoming of increasing importance in animal feeds. In this new scenario, sainfoin is appearing as a clear home-grown alternative for fodder production. Sainfoin has some anthelmintic (Hoste et al. 2006) and nutritional advantages in comparison with other forage legumes, most notably due to its condensed tannins content (Mueller-Harvey 2006), which may lead to a better dietary protein utilisation (Min et al. 2003), lower methane emissions and risk of bloat (Puchala et al. 2005; Ramirez-Restrepo and Barry 2005) in ruminant livestock. Agronomically, it’s positive characteristics include a deep taproot, that allows the plant to be very resistant to drought and of course, being a legume, there is a high level of residual fertility after sainfoin ley has been ploughed (Koivisto and Lane 2001).
While advances in plant breeding have led to improved lucerne varieties, sainfoin production has remained relying on old cultivars. Characterisation of existing germplasm seems to be necessary in order to preserve these genetic resources and provide alternative approaches for developing further breeding programmes. While morphological characterization is a necessary step in plant breeding, several molecular tools are being used to complement plant variety characterization and identification. Among these, microsatellites or simple sequence repeats (SSRs) have become one of the most useful molecular marker systems in plant breeding. They are widely used in cultivar fingerprinting, genetic diversity assessment, molecular mapping, and marker assisted breeding due to their high polymorphism, repeatability and because they are codominant.
In legumes, the availability of SSRs has allowed to make progress in the characterization and assessment of the genetic diversity of species like Medicago (Falahati-Anbaran et al. 2004), SSR cross-species amplification has been assessed (Peakall et al. 1998) and cross-amplified SSRs have been used for genetic mapping in Trifolium repens (L.) (Zhang et al. 2007). While there are not microsatellites available for O. viciifolia, the screening of SSRs identified in Medicago and Glycine spp. offers the possibility to find similar regions on the Onobrychis genome that would allow a quick and reliable characterization of the sainfoin material.
Microsatellites isolation can be time and money consuming in the absence of abundant DNA sequences in a species, restricting sometimes their use to a few important agricultural crops. Still, the amount of SSRs identified is increasing at a steady pace, especially in legumes model species, and the development of SSR markers through data mining has also become an efficient and low cost option for many plant species. An alternative approach consists in the utilization of SSR loci from related species (Smulders et al. 1997). The transferability of SSR loci often depends on genetic relatedness, and while high rates of transferability across species within the same genus (>50%) has been reported in tree species (Liewlaksaneeyanawin et al. 2004; Wünsch and Hormaza 2002) and among legumes (Eujayl et al. 2004; Gaitán-Solís et al. 2002; Peakall et al. 1998), the transferability across genera and beyond seems to be lower (Peakall et al. 1998; White and Powell 1997). On the other side, SSRs identified in ESTs (EST-SSR) markers show higher transferability rates and are expected to be more conserved than genomic SSR markers (Scott et al. 2000).
In the absence of microsatellite loci available in the genus Onobrychis, in this work the transferability of microsatellite markers from Medicago and Glycine to Onobrychis species was evaluated. A group of microsatellites were selected and these were used to provide an initial characterization of a collection of sainfoin accessions and to carry out a preliminary evaluation of the genetic diversity among them.
Materials and methods
Plant material and genomic DNA extraction
Twenty-three accessions of sainfoin were used in this study (Table 1), among them 12 European accessions represent the commercial material available, nine are accessions collected from seed production growers in the North-eastern part of Spain and the remaining two are selections obtained aiming respectively for persistency and for production from the CITA (Zaragoza, Spain). Two related wild species from the Pyrenees range, O. pyrenaica Sennen and O. argentea Boiss., were included to check their relatedness to O. viciifolia and between themselves, and Trifolium repens was used as an outgroup (Table 1). Additionally, Medicago truncatula Gaertn. and Trifolium repens L. accessions were used as positive controls when testing SSR transferability.
Ten plants per population were randomly selected and young healthy leaves from each plant were collected for DNA extraction. Genomic DNA was extracted using the modified (Doyle and Doyle 1987) DNA extraction protocol described by Hormaza (1999). Extracted DNA was quantified spectrophotometrically and diluted to 10 ng/μl before PCR amplification. For testing SSR transferability, individual DNAs were used, for variety characterization an equivalent amount of diluted DNA of the ten plants was pooled and used for SSR amplification.
SSR analysis
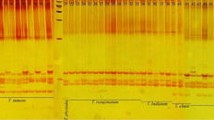
Twenty seven selected microsatellites from Medicago truncatula and Glycine max (L.) Merr. (Gutierrez et al. 2005; Peakall et al. 1998; Zhang et al. 2007) previously cross-amplified in other legume species (Gutierrez et al. 2005; Peakall et al. 1998; Zhang et al. 2007) were tested for transferability in O. viciifolia (Table 2). PCR reactions were performed in 20 μl volumes containing 20 mM Tris–HCl, pH 8.4, 50 mM KCl, 4 mM MgCl2, 0.1 mM each dNTP, 0.2 mM each primer, 40 ng genomic DNA and 0.45 units Taq polymerase (Life Technologies, USA). Reactions were carried out on a iCycler thermocycler (BioRad, USA) using the following temperature profile: an initial step of 2 min at 94°C, 35 subsequent cycles of 45 s at 94°C, 45 s at Ta and 1 min at 72°C, and a final step of 5 min at 72°C. Ta being the annealing temperature, optimised for each SSR locus. PCR products were separated by electrophoresis using 3% ‘Metaphor’ (FMC Bioproducts) agarose gels in 1× TBE buffer at 5 V/cm, stained with ethidium bromide and visualised under UV light. M. truncatula and T. repens DNA was used as positive controls in the PCR reactions. Band scoring was carried out using DNA size standards (10 bp or 1 kb, Life Technologies) depending on the SSR amplification range. Those SSRs that were amplified in Onobrychis and that allowed a clear scoring of SSR alleles were selected for the characterization of all the accessions. Thus 6 SSRs (Table 3) were selected and evaluated in the 25 Onobrychis accessions and T. repens. All PCR amplifications were done at least twice in order to ensure the results reproducibility.
Data analysis
The polymorphism information content (PIC) of each of the analysed SSRs was calculated using the formula: \( PIC = 1 - \sum\nolimits_{j = 1}^{n} {Pij^{2} } \) (Botstein et al. 1980) where Pij is the frequency of the ‘ith’ allele for marker ‘i’ in the ‘jth’ population and summation extends over n alleles. For the estimation of genetic similarity SSR alleles were scored as presence or absence in all the accessions and a similarity matrix was obtained from the proportion of shared fragments (Nei and Li 1979), these calculations were performed using the NTSYS-pc 2.02 package (Rohlf 2002). To visualize differences between populations, the similarity matrix was used to run a Principal Component Analysis (PCA) in Genalex 6.1 (Peakall and Smouse 2006). Scatter plot representation was used to represent the first two Eigen values.
Results
The 27 SSRs analysed were correctly amplified in Medicago truncatula, their original species, or in Trifolium repens, a species in which they had already been tested for cross-amplification (Table 2). In Onobrychis species, consistent amplification was obtained for 22 (81%) of the 27 microsatellites tested (Table 2) once an appropriate annealing temperature was optimised for each microsatelite. However, amplification sizes of the different SSRs were usually larger in Onobrychis viciifolia (79–865 bp) than that originally described in Medicago (79 to 240 bp).
Fourteen of these 22 SSRs were polymorphic in all three Onobrychis species (Table 2) and six of them were selected for characterization of the sainfoin genotypes as they allowed the detection of clear bands and easy fragment recognition (Table 2). A total of 35 alleles were detected using the six microsatellites in the 25 Onobrychis populations. Each SSR amplified 5–7 alleles in Onobrychis and the average number of alleles per SSR was 5.83 (Table 3). The PIC values were calculated for each SSR on the 25 Onobrychis accessions (Table 3). This value ranged from 0.45 to 0.85 with the average being 0.72. Microsatellite BI74 was the SSR that revealed the higher polymorphism in the accessions tested.
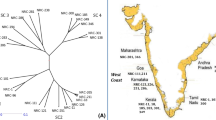
While O. viciifolia accessions showed high genetic similarity, it was lower between O. viciifolia and Onobrychis wild species, O. argentea and O. pyrenaica (Fig. 1). These two species seemed to be closely related to each other as their similarity was almost 0.95. However, T. repens, was remarkably distinguished, with only a 0.23 of similarity with Onobrychis accessions.
The analysis of 6 SSRs in 23 populations of O. viciifolia and in O. pyrenaica, O. argentea and T. repens, produced unique fingerprints for each population, allowing the differentiation of all the 26 accessions. The first two Eigen vector of the PCA analysis explained 60.1% of the observed variation (Fig. 1). From this analysis a clear differentiation was observed between T. repens and all the Onobrychis species. Another group brought together the two wild species O. argentea and O. pyrenaica, clearly separated from O. viciifolia accessions and the T. repens genotype.
All 23 O. viciifolia accessions were clustered in a genetically diverse group (Fig. 2). Within these accessions ‘Cotswold common’, ‘Somborne’, ‘Esparcette’, ‘Ambra’, ‘Sepial’, ‘Mezquita de Jarque’ and ‘Villahermosa del Rio’ scattered in a large clump, that appeared to be distant from the more tightly related ‘Ukrania’, ‘Incoronata’, ‘Ambra’, ‘Selection 1’, ‘Lagueruela’, ‘Loarre’, ‘Reznos’, ‘Visnovsky’, ‘Polonia’, ‘Korunga’, ‘Yubilena’, ‘Torrecilla de Cameros’, ‘Villahoz’, ‘Selection 2’, ‘Graus’ and ‘Tartareu’. The British accessions, ‘Cotswold Common’ and ‘Somborne’, were the most genetically distant from the East European accessions like ‘Visnovsky’, ‘Polonia’ and ‘Korunga’. The French and Italian ‘Ambra’, ‘Sepial’, ‘Incoronata’ and ‘Fakir’ spread the British and East European accessions. Spanish accessions like ‘Mezquita de Jarque’, ‘Selection 1’, ‘Lagueruela’, ‘Loarre’, ‘Reznos’, ‘Torrecilla de Cameros’, ‘Villahoz’, ‘Selection 2’, ‘Graus’ and ‘Tartareu’. were distributed among French, Italian and East European accessions.
Discussion
In order to characterize and evaluate the genetic diversity of cultivated sainfoin, microsatellite loci derived from related legume species were evaluated for transferability in O. viciifolia and the wild species O. argentea and O. pyrenaica. Transferable SSR loci were selected on the basis of polymorphism and reproducibility and these were used for the characterization and genetic similarity evaluation of 23 sainfoin accessions.
Cross-genus transferability
The proportion of microsatellites transferable between Medicago and Onobrychis (81%) was higher than data previously reported for cross-genus amplification. Intra-genus amplification was usually around 50% (Eujayl et al. 2004; Peakall et al. 1998) and declined quickly when reaching out the genus. Zhang et al. (2007) found 18–22% of transferability from Medicago to Trifolium, but Peakall et al. (1998) reported only 1–3% of transferability of Glycine’s SSR among other legumes genus. In this work, EST-SSR were used and these are expected to be more conserved than genomic SSR (Peakall et al. 1998; Scott et al. 2000). Additionally, the SSR loci assayed for transferability had already been shown to be conserved among different genera. Thus the high level of transferability detected here indicates that selecting cross-conserved SSR for transferability across new species is a good strategy for the identification of transferable SSR in a new species. In spite of the conserved nature of the EST-SSRs, which may limit their polymorphism, a high proportion of microsatellites were found to be polymorphic in Onobrychis (63% of transferable SSRs). Six of these were used for the analysis and showed high PIC values (average 0.72) indicating their usefulness for molecular studies in O. viciifolia.
The number of allele per locus observed in Onobrychis ranged from 5 to 7, which is slightly lower than previously reported in lucerne (4–14) (Falahati-Anbaran et al. 2004). Since Onobrychis is a tetraploid species and accessions were analysed in bulks of 10 individuals the number of alleles detected seem to be low. This could be caused for various reasons. First the detection method used, agarose electrophoresis, is not sensitive enough to differentiate alleles with small size differences and it may not be scoring alleles that are being amplified. Nevertheless, working with a tetraploid species, an initial evaluation of the accessions by horizontal electrophoresis allows the visualization of the amplified loci and facilitates a preliminary evaluation of the markers. Otherwise more sensitive detection methods like capillary electrophoresis would provide fragment patterns even more difficult to score due to the stutter of the markers. On the other hand, the bulking of DNA samples and their co-amplification by PCR cause less frequent alleles to be lost and only most common alleles to be detected. This may cause that a number of alleles present in the population could not be detected, leading therefore to an under estimation of the number of alleles detected per loci in each population.
In various SSR (like MtBC47B06F1 and MtBB44F02R1) various loci seemed to be amplified. The generation of multiple products during cross-species amplification may occur by mutation, rearrangements and duplications in the flanking region and/or changes in the number of repeats (Peakall et al. 1998), as reported by Gutierrez et al. (2005) using EST-SSR in legumes.
Genetic similarity
The level of polymorphism shown by the six microsatellites selected made it possible to produce a unique fingerprint of each population. The PCA analysis based on the coefficient of similarity differentiated all O. viciifolia accessions from T. repens and from O. argentea and O. pyrenaica wild species, that were highly similar amongst themselves. This is surprising because morphologically O. argentea is more similar to O. viciifolia than to O. pyrenaica.
All sainfoin varieties shared a similarity coefficient above 0.80, matching results from Sardaro et al. (2003) that reported by ALFP a similarity level of 0.73 among sainfoin accession using AFLP analysis. The high similarity found among O. viciifolia accession suggested that they were more closely related between themselves than varieties of Trigonella foenum-graecum L. (0.66, Dangi et al. 2004) or Phaseolus coccineus L. or vulgaris L. (0.6 and 0.75, Sicard et al. 2005) which are more ancient domesticated legume crops.
The distribution of the O. viciifolia genotypes in the PCA analysis was in relation with the geographic distribution of these accessions, with the British accessions more distant from the rest, East Europeans clustered together and distant from the British ones, and French and Italian accessions in between these two groups. The British cultivars ‘Cotswold Common’ and ‘Somborne’, were clearly differentiated from the other material and this was probably caused by the isolation of British germplasm and low seed exchanges with the rest of the continent from early 1980 s (Aldrich 1984). Only East European accession ‘Esparcette’ from Poland, appeared closer to the British accessions, and was provided by the same seed producer, Cotswold Seeds, and this similarity may be due to common ancestry or hybridisation.
On the other side, Spanish accessions showed a high genetic similarity with both East European and French accessions. Sainfoin originally sown in Spain was the “common” type and giant sainfoin was introduced experimentally by the Botanical Gardens of Madrid in 1791 (Muller 1893). Since then there have been successive introductions, the most important being promoted by the Ministry of Agriculture at the end of the 1960 s. Thus, foreign giant sainfoins (mainly introduced from France) could have mixed with native forms (Pujol 1974) given that sainfoin is thought to be an allogamous species. Nowadays, part of the seed demand is met with imports from Eastern Europe countries. This situation together with seed transfer between farmers and commercial enterprises is contributing further to the contamination of local ecotypes (Delgado et al. 2008). This is particularly true for “Selection 1” and ‘Selection 2’ which were obtained from French and East European material. ‘Villahermosa del Rio’ was found to be the most distinct accession of all and therefore represent an interesting source of variability for breeding purposes.
The results obtained indicate that conservation of SSR allows successful cross-genus amplification of Medicago and Glycine microsatellite markers in Onobrychis and Trifolium, which belong to the same family (Leguminosae) but different genera. Transferable selected microsatellites were useful to fingerprint Onobrychis accessions and to estimate their genetic diversity. The availability of these markers will be useful to accelerate the implementation of appropriate germplasm conservation strategies, to assist in the selection process for development of synthetic varieties with high level of heterosis or to assess seed purity in sainfoin collections.
References
Aldrich DTA (1984) Lucerne, red clover and sainfoin—herbage production. In: Thomson DJ (ed) Forage legumes, occasional symposium 16, BGS, Hurley, pp 126–131
Badoux S (1965) Étude des caractères morphologiques physiologiques et agronomiques de populations d’esparcette (Onobrychis spp.). Rech Agron Suisse 4:111–190
Botstein D, White RL, Skolnic M, Davis RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Dangi RS, Lagu MD, Choudhary L, Ranjekar PK, Gupta VS (2004) Assessment of genetic diversity in Trigonella foenum-graecum and Trigonella caerulea using ISSR and RAPD markers. BMC Plant Biol 4:13
Delgado I, Salvia J, Buil I, Andrés C (2008) The agronomic variability of a collection of sainfoin accessions. Span J Agric Res 6:401–407
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Eujayl I, Sledge MK, Wang L, Chekhoskiy K, Zwonitzer JC, Mian MAR (2004) Medicago truncatula EST-SSRs reveal cross-species genetic markers for Medicago spp. Theor Appl Genet 108:414–422
Falahati-Anbaran M, Habashi AA, Esfahany M, Mohammadi SA, Ghareyazie B (2004) Population genetic structure based on SSR markers in alfalfa (Medicago sativa L.) from various regions contiguous to the centres of origin of the species. J Genet 86:59–63
Gaitán-Solís E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-Species amplification in Phaseolus ssp. Crop Sci 42:2128–2136
Gasparin A (1846) Cours d’agriculture. Librairie Agricole de la Maison Rustique IV, Paris, p 780
Gill M, Smith P, Wilkinson JM (2009) Mitigating climate change: the role of domestic livestock. Animal 4:323–333
Gutierrez MV, Patto MCV, Huguet T, Cubero JI, Moreno MT, Torres AM (2005) Cross-species amplification of Medicago truncatula microsatellites across three major pulse crops. Theor Appl Genet 110:1210–1217
Hayot-Carbonero C, Mueller-Harvey I, Brown TA, Smith L (2010) Sainfoin (Onobrychis viciifolia): a beneficial forage legume. Plant Genet Res (In press)
Hormaza JI (1999) Early selection in cherry combining RAPDs with embryo culture. Sci Hortic 79:121–126
Hoste H, Jackson F, Athanasiadou S, Thamsborg S, Hoskin SO (2006) The effects of tannin-rich plants on parasitic nematodes in ruminants. Trends Parasitol 22:253–261
Koivisto JM, Lane GPF (2001) Sainfoin, worth another look. FAO, http://www.fao.org/ag/AGP/doc/Gbase
Liewlaksaneeyanawin C, Ritland CE, El-Kassaby YA, Ritland K (2004) Single-copy, species-transferable microsatellite markers developed from loblolly pine ESTs. Theor Appl Genet 109:361–369
Min BR, Barry TN, Attwood GT, McNabb WC (2003) The effect of condensed tannins on the nutrition and health of ruminants fed fresh temperate forages: a review. Anim Feed Sci Technol 106:3–19
Mueller-Harvey I (2006) Unravelling the conundrum of tannins in animal nutrition and health. J Sci Food Agric 86:2010–2037
Muller JT (1893) Diccionario universal de agricultura. Elias & Co, Barcelona, p 786
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Peakall R, Smouse P (2006) Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Gilmore S, Keys W, Morgante M, Rafalski A (1998) Cross-species amplification of soybean (Glycine max) Simple sequence repeats (SSRs) within the genus and other legume genera: implications for the transferability of SSRs in plants. Mol Biol Evol 15:1275–1287
Puchala R, Min BR, Goetsch AL, Sahlu T (2005) The effect of a condensed tannin-containing forage on methane emission by goats. J Anim Sci 83:182–186
Pujol M (ed) (1974) El fomento de la producción forrajeropratense en la provincia de Huesca. Ministerio de Agricultura, Pesca y Alimentación, Madrid, Spain, p 182
Ramirez-Restrepo CA, Barry TN (2005) Alternative temperate forages containing secondary compounds for improving sustainable productivity in grazing ruminants. Anim Feed Sci Technol 120:179–201
Rochon JJ, Doyle CJ, Greef JM, Hopkins A, Molle G, Sitzia M, Scholefield D, Smith CJ (2004) Grazing legumes in Europe: a review of their status, management, benefits, research needs and future prospects. Grass Forage Sci 59:197–214
Rohlf FJ (2002) NTSYS-pc, numerical taxonomy and multivariate analysis system, version 2.2. Exeter Software, New York
Sardaro S, Molinari L, Albertini E, Rosellini D, Negri V, Falcinelli M (2003) Molecular distinctiveness of wild populations of Poa pratensis, Lolium perenne and Onobrychis viciifolia. Sementi elette 49:47–49
Scott KD, Eggler P, Seaton G, Rosetto M, Ablett EM, Lee LS, Henry RJ (2000) Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100:723–726
Sicard D, Nanni L, Porfiri O, Bulfon D, Papa R (2005) Genetic diversity of Phaseolus vulgaris L. & Phaseolus coccineus L. landraces in central Italy. Plant Breed 124:464–472
Smulders MJM, Bredemeijer G, Rus-Kortekaas W, Arens P, Vosman B (1997) Use of short microsatellites to generate polymorphisms among Lycopersicon esculentum cultivars and accessions of other Lycopersicon species. Theor Appl Genet 94:264–272
White G, Powell W (1997) Cross-species amplification of SSR loci in the Meliaceae family. Mol Ecol 6:1195–1197
Wünsch A, Hormaza JI (2002) Molecular characterization of sweet cherry (Prunus avium L.) genotypes using peach (Prunus persica (L.) Batsch) SSR sequences. Heredity 89:56–63
Zhang Y, Sledge MK, Bouton JH (2007) Genome mapping of white clover (Trifolium repens L.) and comparative analysis within the Trifolieae using cross-species SSR markers. Theor Appl Genet 114:1367–1378
Acknowledgments
We gratefully acknowledge T. Bespín, A. M. Cachi, S. Sancho and M. E. Guerra for technical assistance; Dr F. Fillat for expert advices in collecting wild species. This work has been partly supported by the Marie Curie Actions Project no MRTN-CT-2006-035805 and by the DGA Grupo de Excelencia A-11.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Demdoum, S., Muñoz, F., Delgado, I. et al. EST-SSR cross-amplification and genetic similarity in Onobrychis genus. Genet Resour Crop Evol 59, 253–260 (2012). https://doi.org/10.1007/s10722-011-9681-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-011-9681-x