Abstract
Orychophragmus violaceus, a ground covering plant that is widely distributed in China. It has both high economical value in food, forage, health care and ornamental value in gardening. In this study, the genetic diversity of 245 individuals from nine populations in China were investigated using the inter-simple sequence repeat markers. Of the 100 primers screened, eight were highly polymorphic. Using these primers, 162 discernible DNA fragments were generated with 150 (92.59%) being polymorphic, indicating a pronounced genetic variation at the species level. Also, there were high levels of polymorphism at the population level with the percentage of polymorphic bands ranging from 85.74 to 90.06%. Analysis of molecular variance showed that the genetic variation within populations was 80.80% and the variance among populations was 16.43%. The Nei’s GST (0.1643) and gene flow among populations (Nm = 2.5760) revealed large gene exchanges among populations. O. violaceus belongs to out-crossing plants. It is capable of reproducing by self-sowing, thus can influence population genetic structure. The pronounced genetic variation within populations tells us that O. violaceus is a proper plant for genetic research and that there is great potential of breeding this species for gardening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Information on genetic diversity patterns can provide insight into the evaluating and utilizing of the germplasm resources. Knowledge of how genetic variation is partitioned among populations may have important implications not only to evolutionary biology but also to conservation biology. Hence, reliable estimates of population differentiation are crucial to understanding the connectivity among populations and represent important tools in the development of conservation strategies (Balloux and Lugon-Moulin 2002; Neel and Ellstrand 2003). In addition, environmental barriers and life history may all form the genetic structure of populations (McCue et al. 1996; Donnelly and Twonson 2000).
Orychophragmus violaceus (Linn.) O.E. Schulz, belongs to the Cruciferae family, and is a biennial herb. This plant is widely distributed in the northeast, northwest, north, east and middle of China. It is found in a variety of environments, including plains, mountains, roadsides, adjacent to buildings and so on (Zhou et al. 1987). O. violaceus is commonly used for forage, health care and gardening (Luo et al. 1998; Xiao and Luo 1994; Ren et al. 1998; Weng and Huang 2001), and is highly regarded for its great economic and ornamental usages. Various topics have been studied on O. violaceus, including distant hybridization, karyotypes, tissue culture, isozymes and genetic transformation (Li et al. 1994; Zhao et al. 1995; Liang and Tao 1997; Li 1998; Luo et al. 2000; Wu et al. 2002; Mei et al. 2003). In recent years, many cities in China (Beijing, Nanjing, Shanghai, Hangzhou et al.) have used O. violaceus as a ground cover plant in gardens, streets and understory in a large scale (Fig. 1; Zhang and Dai 2005). The research on ornamental characters, morphological variations were observed a few years ago, when many types of variations in this plant was found (Zhang et al. 2005; Fig. 2). Up to now, genetic diversity of it has not been reported, this made it difficult to perform effective breeding program and to protect the resources.
The inter-simple sequence repeats (ISSR) is a newly developed modification of SSR (simple sequence repeats)-based marker systems (Ziekiewica et al. 1994). It has been recognized as useful molecular markers in the analysis of genetic diversity and population genetic analysis (Lu et al. 2006; Wang et al. 2005; Kar et al. 2005; Esayas et al. 2005). The ISSR has advantages, e.g., low quantities of template DNA requirement, no need of sequence data for primer construction, random distribution throughout the genome, generation of many informative bands per reaction, and reliable and reproducible production (Ziekiewica et al. 1994; Nagaoka and Ogihara 1997; Ge 2001). Consequently, the ISSR has been widely used in marker assisted selection, genetic diversity analysis, DNA fingerprinting, and evolution and molecular ecology (Zhao et al. 2007a, b; Vijayan et al. 2006; Carvalho et al. 2005; Bolibok et al. 2005; Nybom 2004; Wang 2002).
In the present study, we applied the ISSR markers to examine the genetic diversity of O. violaceus in three areas, Beijing, Nanjing and Shanghai, where large populations were found. Morphological studies indicate lots of variations within each population (Zhang and Dai 2005). We were particularly interested in the level of differentiation among populations and the levels of diversity within populations. This information help us understand the genetic background and will provide a reference for the utilization of genetic resources, species protection and including future breeding program.
Materials and methods
Plant materials
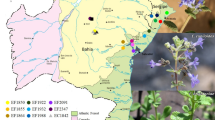
Specimens were collected from nine populations of O. violaceus in or around Beijing, Nanjing and Shanghai. Figure 3 shows the distribution of O. violaceus in some literature and the three areas that were sampled. Table 1 shows the locations and the climate factors of these populations. The size of the individual populations was 30 × 30 m, and the distance between populations within the three groups was farther than 3 km. Individual plants were systematically and randomly sampled, in which a total of less than 10% of the whole population was randomly sampled. 25–35 individual plants were collected with an interplant distance of at least 5 m to increase the possibility of detecting potential individual variation (Jin and Lu 2003).
Young leaves were collected from each sample plant and dried in silica gel for subsequent DNA extraction.
DNA extraction
Total DNA was extracted from young leaves following the CTAB (cetyltrimethyl ammonium bromide) method described by Doyle and Doyle (1987) with one modification. The quality of DNA was determined in 1.0% agarose gels. The purified total DNA was quantified by a BioradSmartspec3000 UV–Vis spectrophotometer. DNA samples were adjusted to concentration of 20 ng/μL with ddH2O and subjected to PCR amplification.ISSR PCR amplification.
ISSR PCR amplification
PCR amplification was done in a Techne TC-512 thermocycler using 10 μL reaction mixture containing 20 ng DNA, 1.0 μL of 10× buffer (200 mmol/L Tris–HCl; 200 mmol/L KCl; 100 mmol/L (NH4)2SO4; 15 mmol/L MgCl2), 0.2 mM dNTP, 0.4 μM primer and 0.5 U of Taq DNA polymerase. The PCR schedule was 94°C for 5 min followed by 45 cycles of 94°C for 45 s, 54–60°C for 1 min, 72°C for 1.5 min and a final extension of 7 min at 72°C. ISSR-PCR products were separated by electrophoresis on 6% urea polyacrylamide gels and silver stained to visualize the bands. One hundred primers (UBC primer set no. 9, Biotechnology Laboratory, University of British Columbia, Canada) were screened initially. Two DNA templates from each population were chosen randomly for PCR amplification. Fifty ISSR primers were screened on these selected individuals by comparing the effects of magnesium concentrations and annealing temperature during amplification. Eight primers that produced clear and reproducible fragments were selected for the full survey of all 245 individuals (Table 2).
Data analysis
Only distinct, reproducible, well-resolved fragments were scored as present (1) or absent (0) for each of the ISSR markers. The Polymorphic bands (PPB), allele number (Ao), effective allele number (Ae), Nei’s (1973) expected heterozygosity (He), Shannon index of diversity (I), gene diversity among subpopulations (Hs) and of the total population (Ht), gene differentiation (Gst), gene flow (Nm) and gene distance (D) were analyzed using POPGENE software, version 1.31 (Yeh et al. 1999). The Analysis of Molecular Variance (AMOVA program version 1.55, Excoffier et al. 1992) was performed to describe the genetic structure and variation among groups, among populations within groups and among individuals. A dendrogram was constructed by an unweighted paired group method of cluster analysis using arithmetic averages (UPGMA) on NTSYS-pc program (version 1.21, Rohlf 2000).
Results
Genetic diversity within populations
The ten primers generated 162 bands ranging in size from 200 to 1500 bp (Fig. 4), corresponding to an average of 20.2 bands per primer. Of these bands, 92.59% (150 in total) were PPB at the species level. Assuming Hardy–Weinberg equilibrium, the Nei’s gene diversity (He) ranged from 0.3330 (SG) to 0.3659 (BY) for O. violaceus, with an average of 0.3464 at the population level. Shannon index (I) ranged from 0.4938 (SG) to 0.5369 (BY), with an average of 0.5150 at the population level. Among the nine populations, BY from Beijing region exhibited the greatest variability (PPB 90.06%, He 0.3659, I 0.5369). In contrast, genetic diversity in population SG was lowest with PPB 86.98%, He 0.3330, I 0.4938 (Table 3). Considering the groups of populations, those from Beijing group had, on average, higher levels of diversity than those from Shanghai group.
Genetic structure of populations
To assess the overall distribution of genetic diversity, the AMOVA program was used to analyze the distance matrix. The genetic differentiation among populations in AMOVA was highly significant (P < 0.001; Table 4). A large proportion of genetic variation (80.8%) existed among individuals within the populations, whereas 16.43% resided among populations. Nei’s (1973) estimator of population substructure (G ST) also indicated a low level of population differentiation (G ST = 0.1643). These G ST translated into correspondingly high levels of gene flow (Nm), with 2.5760 migrants exchanged between populations (on average) each generation. There have been concerns about the direct use of G ST to estimate gene flow (Whitlock and McCauley 1999). However, Tn some extent, G ST remains a useful measure of the average effects of gene flow (Neigel 2002).
The genetic distance and UPGMA analysis
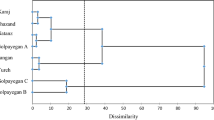
The UPGMA analysis indicated that there are two clusters (Fig. 5). The first cluster contains population NB from Nanjing, and populations SB and SF from Shanghai. The second cluster contains of population NL from Nanjing, population SG from Shanghai, and populations BS, BT, BY and BH from Beijing. The four populations from Beijing regions were clustered together, whereas, Nanjing and Shanghai populations had relatively closer relation.
Discussion
Genetic diversity of Orychophragmus violaceus
ISSRs can provide more polymorphism than isozymes and RAPD markers (Wolfe and Liston 1998; Ge and Sun 1999; Camacho and Liston 2001). By using ISSR primers, we found a high level genetic diversity for the species of O. violaceus with 92.59% of bands being polymorphic in all nine populations. The percentage of PPB in each population ranged from 85.74 to 90.06%. The genetic variation detected was a little higher than other species (Brassica campestris L., Raphanus sativus L. and Brassica juncea L.) in this family (He et al. 2002; Wu et al. 1997; Shen et al. 2004; An et al. 1999; Rabanni et al. 1998; Burton et al. 2004). In comparison, O. violaceus presents high genetic diversity both within and among populations.
The level and pattern of genetic diversity detected by ISSRs in the present study highly agreed with the analysis of morphological characters, karyotypes and isozymes of O. violaceus. Firstly, the analysis of variation of morphological characters of O. violaceus revealed that there were many kinds of variations in petal color, petal shape, flower size, leaf color, leaf shape, plant shape and other features. Many peculiar variations were found, e.g., speckled petals, petal stain, toothed petal margin and speckled leaves. These characters were significantly different among populations and within populations (Zhang et al. 2005). The karyotypes of O. violaceus also had relatively high diversity among populations, individuals and cells (Zhang et al. 2006), and high percentage of B chromosomes and aneuploids were observed in the nucleus. Li et al. (1994) pointed that the variants of O. violaceus have evolved with considerable hereditary variation during the phylogenesis of the species, which can be seen in the significantly different of the locations of secondary constrictions, the chromosome arm ratio and karyotypes. In addition, analysis of POD in different environments of O. violaceus indicated that there were great differences among plants in isoenzyme patterns and relative enzymatic activity (Li 1998).
The high genetic variability among population in the plant may be a consequence of sexual reproduction, mutations of somatic cells, selection, gene flow, genetic drift and changing environment (Gao and Yang 2006). The cross-pollination mechanism, sexual reproduction, high seed ratio and self-sown ability to produce offspring of O. violaceus could result in the accumulation of abundant genetic variation during the long evolution history of the species. The variations of O. violaceus was also explained in cytology, autotetraploid or allotetraploid chromosomes lead to the diversity of alleles, and the change of the number of somatic chromosomes was due to cytomixis and chromosome fractionation during the process of mitosis and meiosis. Differences of gene dosage, modification and regulation will occur in different cells of different plants (Wu et al. 1996).
Population genetic structure
ISSR markers revealed that O. violaceus populations had more genetic variation within populations than among populations. Little differentiation was detected among populations. There are many factors that determine the genetic structure of plant populations, including reproductive biology, gene flow, seed dissemination and nature selection. Reproductive biology is one of the most important factors (Zhao et al. 2007a, b). According to Hamrick and Godt (1989), outcrossing plant species tend to exhibit between 10 and 20% genetic variation among populations while selfing species exhibit, on average, 50% variation between populations. Studies on the biology of flowering and pollination indicate O. violaceus is an outcrosser (Zhang et al. 2007). It is also supported by the genetic differentiation (G ST = 0.1643) among populations of O. violaceus, which was very close to the average genetic differentiation (G ST = 0.144) in outbred populations (Bussell 1999).
The variation among individuals within populations was the main source of variation of O. violaceus. The reasons for this population genetic structure are as follows: (1) O. violaceus is widely distributed in the northeast, northwest, north, east and middle of China and is found in various kinds of environments. The strong adaptability and wide distribution results in little differentiation between populations. (2) O. violaceus exchanged its genes mainly by seeds and pollen. The spread of seeds and pollen is the main way of gene flow in natural populations of plants (Li and Chen 2004). Seeds and pollen are both small and dispersed by wind. Due to the high frequency of strong winds, the effects of long distance dispersal of seeds and pollen by wind are similar. (3) Gene flow is the most important factor in making population genetic structure homogeneous. The greater the amount of gene flow among populations, the less gene differentiation occurs (Slatkin 1985). The large gene flow (Nm = 2.5760 > 1) of O. violaceus could counteract most of the gene differentiation which is caused by genetic drift within populations.
We also found little genetic differentiation among three groups, which suggests that the populations from different sides are poorly differentiated. Only 2.77% of the total variation was found among three geographic regions (Beijing, Nanjing, and Shanghai). This small vitiation might be due to the continuous distribution in China. Though there is no direct transport of seeds and pollen among three regions, gene flow could influence the evolution process of different regions. Another possible reason is that the populations we selected were cultured populations not natural population. Seeds were collected around the cities and it is difficult to know their real source.
Implications for development
The information about population genetic diversity represents population adaption to environments which conditions on the level of its adaptive evolution. It is also of critical importance to the conservation and management of plants, including the assessment of the conservation value and status of special populations (Hogbin and Peakall 1999; Bawa and Ashton 1991; Hamrick and Godt 1996). The high genetic diversity maintained within populations of O. violaceus is encouraging. In addition, the plant possesses great ornamental value and developmental potential. It is necessary to protect existing natural populations and its habitat in order to preserve as much genetic variety as possible. Further research on the reproductive mechanism and law of inheritance and variation of some characters (petal color and shape, leaf shape) of O. violaceus will help us to understand its natural hybrids and promote its better utilization in the future.
References
An XH, Chen BY, Fu TD, Liu HL (1999) Genetic diversity of Chinese landraces in Brassica napus was analysed by RAPD markers. J Huazhong Agric Univ 6:524–527
Balloux F, Lugon-Moulin B (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Bawa KS, Ashton PS (1991) Conservation of rare trees in tropical rain forests: a genetic perspective. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, pp 62–67
Bolibok H, Pakoczy-Trojanowska M, Hronada A, Pietrzykowski R (2005) Efficiency of different PCR-based marker systems in assessing genetic diversity among winter rye (Secale cereale L.) inbred lines. Euphytica 146:109–116
Burton WA, Ripley VL, Potts DA, Salisbury PA (2004) Assessment of genetic diversity in selected breeding lines and cultivars of canola quality Brassica juncea and their implications for canola breeding. Euphytica 136:181–192
Bussell JD (1999) The distribution of random amplified polymorphic DNA (RAPD) diversity amongst populations of Isotoma petraea (Lobeliaceae). Mol Ecol 8:775–789
Camacho JC, Liston A (2001) Population structure and genetic diversity of Botrychium pumicola (Ophioglossaceae) based on inter-simple sequence repeats (ISSR). Am J Bot 81:965–972
Carvalho A, Matos M, Lima-Brito J, Guedes-Pinto H, Benito C (2005) DNA fingerprint of F1 interspecific hybrids from the triticeae tribe using ISSRs. Euphytica 143:93–99
Donnelly MJ, Twonson H (2000) Evidence for extensive genetic differentiation among populations of the malaria vector Anopheles arabiensis eastern Africa. Insect Mol Biol 9:357–367
Doyle JJ, Doyle JK (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bulletin 19:11–15
Esayas A, Endashaw B, Tomas B (2005) Inter-simple sequence repeat (ISSR) variation in forest coffee trees (Coffea arabica L.) populations from Ethiopia. Genetica 124:213–221
Excoffier L, Smousse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Gao L, Yang B (2006) Genetic diversity of wild Cymbidium goeringii (Orchidaceae) populations from Hubei based on ISSR analysis. Biodivers Sci 3:250–257
Ge S (2001) Application of DNA molecular marker in conservation biology. In: Zou YP, Ge S, Wang XD (eds) Molecular marker in systematic and evolutionary botany. Science Press, Beijing, pp 140–149
Ge XJ, Sun M (1999) Reproductive biology and genetic diversity of a cryptoviviparous mangrove Aegiceras corniculatum (Myrtinaceae) using allozyme and inter-simple sequence repeat (ISSR) analysis. Mol Ecol 8:2061–2069
Hamrick JL, Godt MJW (1989) Allozyme diversity in plant species. In Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and germplasm resources. Sinauer Associaties, Sunderland, pp 43-63
Hamrick JL, Godt MJW (1996) Conservation genetics of endemic plant species. In: Avise JC, Hamrick JL (eds) Conservation genetics: case histories from nature. Chapman and Hall, New York, pp 281–304
He YT, Tu JX, Fu TD, Li DR, Chen BY (2002) Genetic diversity of germplasm resources of Brassica campestris L. in China by RAPD markers. Acta Agronom Ica Sinica 5:697–703
Hogbin PM, Peakall R (1999) Evaluation of the contribution of genetic research to the management of the endangered plant Zieria prostrate. Conserv Biol 13:514–522
Jin Y, Lu BR (2003) Sampling strategy of genetic diversity. Biodivers Sci 11:155–161
Kar PK, Vijayan K, Mohandas TP, Nair CV, Saratchandra B, Thangavelu K (2005) Genetic variability and genetica structure of wild and semi-domestic populations of Tasar silkworm (Antheraea mylitta) ecorace daba as revealed through ISSR markers. Genetica 125:173–183
Li J (1998) Analysis of POD in different Orychophragmus violaceus. J Southwest Agric Univ 20:223–225
Li HS, Chen GZ (2004) Genetic diversity of mangrove plant Sonneratia caseolaris in Hainan island based on ISSR analysis. Acta Ecologica Sinica 24:1656–1662
Li ZX, Cao XD, Liu DX, Liu J, Zhang ZS (1994) A study on the karyotype of some Chinese variants of Zhuge Cai, Orychophragmus violaceus. Acta Agronomica Sinica 20:595–600
Liang MS, Tao Z (1997) Separation of Orychophragmus violaceus gene promoters. Southwest China J Agric Sciences 3:1–5
Lu Z, Wang Y, Peng Y, Korpelainen H, Li C (2006) Genetic diversity of populus cathayana Rehd populations in southwestern China revealed by ISSR markers. Plant Sci 170:407–412
Luo P, Huang BQ, Yin JM, Chen ZL, Chen YH, Lan ZQ (1998) A new forage genetic resource Orychophragmus violaceus (L.) O. E. Schulz. Genet Resour Crop Evol 45:491–494
Luo P, Ye Q, Lan ZQ (2000) A study on floral biology of seedings in vitro in Orychophragmus violaceus: Induction of flowers in seedings of O. violaceus cultured in vitro. Plant Cell Tiss Organ Cult 63:73–75
McCue KA, Buckler ES, Holtsford TP (1996) A hierarrchical view of gentic structure in the rare annual plant Clarkia springvillensis. Conserv Biol 10:1425–1434
Mei DS, Li YC, Hu Q (2003) Investigation of male sterile lines intergeneric somatic hybrids of Brassica napus (+) Orychophragmus violaceus and B. napus (+) Sinapis arvensis. Chinese J Oil Crop Sci 1:72–75
Nagaoka T, Ogihara Y (1997) Applicability of inter-simple sequence repeat polymorphism in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor Appl Genet 93:133–139
Neel MC, Ellstrand NC (2003) Conservation of genetic diversity in the endangered plant Eriogonum ovalifolium var. vineum (Polygonaceae). Conserv Genet 4:337–352
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Neigel JE (2002) Is F ST obsolete? Conserv Genet 3:167–173
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 5:1143–1150
Ren QJ, Yu JP, Zhang GL (1998) The integrative utilization of Orychophragmus violaceus Resources. Chinese Wild Plant Resour 2:24–25
Rohlf FJ (2000) NTSYS-pc. Numerical taxonomy and multivariate analysis system, Version 2.1. Exeter Software, Setautket, New York
Shen JX, Fu TD, Yang GS (2004) Relationship between hybrid performance and genetic diversity based on SSR and ISSR in Brassica napus L. Scientia Agricultura Sinica 4:477–483
Slatkin M (1985) Gene flow in natural populations. Annu Rev Ecol Syst 16:393–430
Vijayan K, Srivatsava PP, Nair CV, Awasthi AK, Tikader A, Sreenivasa B, Urs SR (2006) Molecular characterization and markers associated with yield traits in mulberry using ISSR markers. Plant Breed 125:298–301
Wang JB (2002) ISSR markers and their applications in plant genetics. Hereditas 24:613–616
Wang S, Miao X, Zhao W, Huang B, Fan M, Li Z, Huang Y (2005) Genetic diversity and population structure among strains of the entomopathogenic fungus, Bieauveria bassiana, as revealed by inter-simple sequence repeats (ISSR). Mycol Res 109:1364–1372
Weng DB, Huang XF (2001) Evaluation on the protein quality of Orychophragmus violaceus. Acta Boanica Boreali-Occidentalia Sinica 21:673–677
Whitlock MC, Mccauley DE (1999) Indirect measures of gene flow and migration: FST ≠ 1/(4Nm + 1). Heredity 82:117–125
Wolfe AD, Liston A (1998) Contributions of PCR-based methods to plant systematics and evolutionary biology. In: Soltis DE, Soltis PS, Doyle JJ (eds) Molecular systematics of plants II: DNA sequencing. Kluwer, New York, pp 43–86
Wu YY, Jiang JY, Shuai SW, Yao LZ (1996) A study on the cytogenetics of Orychophragmus violaceus. Southwest China J Agric Sci 3:38–41
Wu NF, Li RG, Wu XM, Zhu L, Fan YL, Qian XZ (1997) RAPD molecular markers and genetic diversity among 40 cultivars of Brassica napus in China. Chinese Biodiver 5(246):246–250
Wu YY, Wang B, Taylor PWJ (2002) Comparaison study on several molecular markers between rapeseed and Orychophragmus violaceus. Chinese J Oil Crop Sci 2:22–25
Xiao L, Luo P (1994) The research presence and development prospect of Orychophragmus violaceus. Acta Botanica Boreali-Occidentalia sinica 14:237–241
Yeh FC, Yang RC, Boyle T (1999) POPGENE Version 1.31, Microsoft window-based freeware for population genetic analysis. University of Alberta and Centre for International Forestry Research, Edmonton
Zhang LJ, Dai SL (2005) The value of development of Orychophragmus violaceus and its landscape utilization. Beijing Landscape 4:43–45
Zhang LJ, Qin HM, Wang M, Dai SL (2005) Variation of morphological characteristics of Orychophragmus violaceus. Biodivers Sci 13:535–545
Zhang LJ, Duan C, Dai SL (2006) A studies on karyotypes of four populations in Orychophragmus violaceus from Beijing, The 2006 doctoral forum of China, pp 2504–2512
Zhang LJ, Han Y, Dai SL (2007) The pollination biology of Orychophragmus violaceus. Northern Horticul 12:119–121
Zhao Y, Wang ML, Wang TJ (1995) Low temperature pollen preservation of Orychophragmus violaceus. Southwest China J Agric Sci 3:65–69
Zhao WG, Zhou ZH, Miao XX, Zhang Y, Wang SB, Huang JH, Xiang H, Pan YL, Huang YP (2007a) A comparison of genetic variation among wild and cultivated Morus species (Moraceae: Morus) as revealed by ISSR and SSR markers. Biodivers Conserv 16:275–290
Zhao Y, Chen XY, Wang XR, Pian RQ (2007b) ISSR analysis of genetic diversity among Lespedeza bicolor populations. J Plant Genetic Resour 2:195–199
Zhou TY, Guan KJ, Guo RL (1987) Flora Reipublicae Popularis Sinicae. Tomus 33. Science Press, Beijing, pp 40–43
Zhou JM, Wei ZM, Xu ZH, Liu SG, Luo P (1997) Agrobacterium-mediated transformation of Orychophragmus violaceus and regeneration of transgenic plants. Acta Phytophysiologica Sinica 23:21–28
Ziekiewica E, Ratalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, LJ., Dai, SL. Genetic variation within and among populations of Orychophragmus violaceus (Cruciferae) in China as detected by ISSR analysis. Genet Resour Crop Evol 57, 55–64 (2010). https://doi.org/10.1007/s10722-009-9450-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-009-9450-2