Abstract
Detection of genetic relationships between 19 chickpea cultivars and five accessions of its wild progenitor Cicer reticulatum Ladizinsky were investigated by using RAPD and ISSR markers. On an average, six bands per primer were observed in RAPD analysis and 11 bands per primer in ISSR analysis. In RAPD, the wild accessions shared 77.8% polymorphic bands with chickpea cultivars, whereas they shared 79.6% polymorphic bands in ISSR analysis. In RAPD analysis 51.7% and 50.5% polymorphic bands were observed among wild accessions and chickpea cultivars, respectively. Similarly, 65.63% and 56.25% polymorphic bands were found in ISSR analysis. The dendrogram developed by pooling the data of RAPD and ISSR analysis revealed that the wild accessions and the ICCV lines showed similar pattern with the dendrogram of RAPD analysis. The ISSR analysis clearly indicated that even with six polymorphic primers, reliable estimation of genetic diversity could be obtained, while nearly 30 primers are required for RAPD. Moreover, RAPD can cause genotyping errors due to competition in the amplification of all RAPD fragments. The markers generated by ISSR and RAPD assays can provide practical information for the management of genetic resources. For the selection of good parental material in breeding programs the genetic data produced through ISSR can be used to correlate with the relationship measures based on pedigree data and morphological traits to minimize the individual inaccuracies in chickpea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chickpea (Cicer arietinum L.) is the third most important pulse crop in the world and ranks first in the Indian subcontinent and Mediterranean basin. India is the largest producer of chickpea, accounting for 66% of the world production (FAO 2004). The average annual yield world-wide (0.78 ton/ha) is considered to be somewhat lower than its potential yield (Singh et al. 1994; Sudupak et al. 2002). However, chickpea productivity is not enough to fulfill the requirement of an increasing population. One major reason for the low productivity of cultivated chickpea, Cicer arietinum, is its narrow genetic base and its sexual incompatibility with other Cicer wild types in natural inter-specific crosses.
Therefore, many chickpea breeding programmes are focused on improving the genetic potential both to increase yield and to provide protection against abiotic and biotic stresses. In order to enhance genetic potential, there must be a comprehensive understanding of the amount and pattern of genetic variation that exists within and between the available cultivated and wild accessions. World germplasm collections of cultivated chickpea are lacking in diversity that many include traits needed for effective improvement of the crop (Robertson et al. 1997; Collard et al. 2003a). However, this may be overcome by looking to the wild relatives to widen the genetic bases of breeding programmes through interspecific hybridization (Singh and Ocampo 1997).
Several different molecular methods are available for the identification of cultivars and genetic diversity. The Polymerase chain reaction (PCR) methods using arbitrary primers have been widely utilized in the last 10 years. Random amplified polymorphic DNA’s (RAPD’s) is one of the most popular techniques, which has been used for measuring genetic diversity in several plant species, including chickpea (Moussa et al. 1996; Sant et al. 1999; Collard et al. 2003a) and lens (Duran et al. 2004). According to Ratnaparkhe et al. (1998) ISSR technique is more reliable than the RAPD technique and generates larger numbers of polymorphisms per primer. RAPD and ISSR markers are highly polymorphic and are useful in studies on genetic diversity in chickpea (Ratnaparkhe et al. 1998); phylogeny in chickpea (Iruela et al. 2002); gene tagging in chickpea (Rajesh et al. 2002) and evolutionary biology (Reddy et al. 2002). ISSR markers have already been used in chickpea (Collard et al. 2003a and Rajesh et al. 2003). Amplified fragment length polymorphism (AFLP) analysis was used to evaluate the genetic variation among cultivated chickpea and wild Cicer relatives (Nguyen et al. 2004) and in pigeonpea (Panguluri et al. 2006). Both RAPD and ISSR remain attractive options despite availability of sophisticated techniques because they are easy, quick, simple and economical. Neither sequence information nor any prior genetic studies are required for this analysis.
To date there have been very few reports investigating the level of genetic variation between the accessions of Cicer and wild Cicer for the generation of intraspecific or interspecific populations of chickpea. Previous research in genetic mapping of chickpea populations has utilized populations derived from interspecific crosses between chickpea and Cicer reticulatum, in order to detect polymorphisms between parents and marker segregation (Simon and Muehlbauer 1997; Santra et al. 2000; Winter et al. 2000; Shan et al. 2005). Chickpea-breeding aims at developing high yielding cultivars that combine long lasting resistance against Ascochyta blight and Fusarium wilt with tolerance to abiotic stresses such as drought, cold and salt. Use of RAPD and ISSR for genetic diversity analysis of Indian accessions has not been demonstrated. The present study was conducted to determine the genetic variability between and within Cicer accessions using RAPD and ISSR markers and to provide a better understanding of the genetic diversity of the Cicer accessions.
Materials and methods
Plant material
The plant material comprised of 24 accessions of chickpea including 19 cultivars of Cicer arietinum L. var. macrospermum Jaub. et Spach and five accessions of its wild progenitor Cicer reticulatum Ladizinsky (Table 1). All the above material was obtained from Pulse Research Laboratory, Indian Agricultural Research Institute, New Delhi, India.
Plant DNA extraction and purification
Total plant genomic DNA was isolated following the protocol given by Doyle and Doyle (1987) as modified by Panguluri et al. (2006).
RAPD reaction
PCR amplification was performed as described by Williams et al. (1990) with some modifications. Sixty random decamer primers, twenty each of series OP (AS), OP (F) and OP (I) were obtained from Operon Technologies Ltd., (Alameda, California). 29 primers were used for analysis out of 60 RAPD primers. The reaction mixture (25 μl) contained 10 × -assay buffer, 2.5 mM MgCl2, 400 μM dNTP’s (Bangalore geni), 5.0ρ moles of primer, 50 ng template DNA and 1 U of Taq DNA Polymerase (Bangalore genei). Amplification was carried out in a thermo-cycler (Biometra) for 40 cycles, each consisting of a denaturation step at 94°C for 1 min, annealing at 55°C for 1 min and an extension step at 72°C for 2 min. An initial denaturation step at 94°C for 5 min, and a final synthesis step of 8 min at 72°C were also included.
ISSR reaction
PCR amplification was performed as described by Williams et al. (1990) with some modifications. Ten primers (UBC primers) were obtained. Six primers {UBC-112 (GACA)4 (ISSR-1)}, UBC-864 {(ACTG)4 (ISSR-2)}, UBC-841 {(GACAC)4 (ISSR-5)}, UBC-810 {(TGGA)4 (ISSR-6)}, UBC-827 {(GACT)4 (ISSR-7)} and UBC-820 {(CAC)5 (ISSR-10)} were used for analysis. Amplification was carried for 40 cycles, each consisting of a denaturation step at 94°C for 1 min, annealing at 45°C & 54°C for 1 min. and an extension step at 72°C for 2 min. An initial denaturation step at 94°C for 5 min, and a final synthesis step of 8 min at 72°C were also included.
Agarose gel electrophoresis for RAPD and ISSR
Reaction products were mixed with 4 μl of 6 × loading dye (0.25% Bromophenol Blue, 0.25% Xylene Cynol and 40% Sucrose, w/v), electrophoresed on 1.4% agarose gel, stained with ethidium bromide and visualized under UV light.
Data analysis for RAPD and ISSR
DNA fragment profiles representing a consensus of two independent replicates were scored in a binary fission with ‘0’ indicating the absence and ‘1’ indicating presence of band. Using the binary data, a similarity matrix was constructed using the Jaccard coefficient, which was further subjected to UPGMA clustering analysis and a dendrogram was generated. A cophenetic matrix was constructed using the matrix that was used to generate the clusters. A correlation between the cophenetic matrix and the similarity matrix was determined by using MXCOMP module. All the above analysis was done using the software package NTSYS-PC (version 2.02e).
Results and discussion
Differences between genotypes with regard to agronomic characters, morphological characters, biochemical characters (e.g. storage proteins, isozymes), and molecular characteristics are either indirect or direct representations of differences at the DNA level and are therefore expected to provide information about genetic relationships. The assessment of genetic diversity is important not only for crop improvement but also for efficient management and conservation of germplasm resources. For this purpose 19 cultivars of chickpea and five different accessions of its wild relative Cicer reticulatum were analyzed by using 29 reproducible RAPD markers and 6 ISSR markers.
RAPD analysis
RAPD analysis revealed a good polymorphism among chickpea cultivars. Sixty random (each twenty of OP- (AS), OP-(I) and OP-(F)) primers were taken and only 29 primers were found to be polymorphic. On an average 6 bands per primer and 74.4% polymorphic bands were observed in a total of 176 bands. From RAPD data 49.4% of common bands and 50.5% (Fig. 1) of polymorphic bands were observed among chickpea cultivars and 48.3% of common bands and 51.7% of polymorphic bands were observed among wild accessions. The primer OP (F)-3 gave rise to maximum bands (11) and OP (F)-12 showed the least number of bands (2).
Agarose gel showing the amplified product using OP (AS) – 12 primer. M; I kb ladder, Lane 1–8; Pusa-256, Pusa-362, Pusa-372, Pusa-391, Pusa-1003, Pusa-1101, Pusa-1103 and Pusa-1105, Lane 9–13; ICC 17121, ICC 17123, ICC 177124, ICC 17160 and ICC 17163, Lane 14–24; BGD-72, BGD-112, DG-36, DG-46, DG-51, DG-63, DG-71, DG-72, ICCV-2, ICCV-10 and Annegiri
RAPD dendrogram
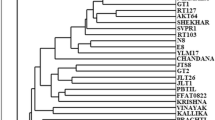
From the UPGMA dendrogram (Fig. 2), it is discernible that the material taken for the analysis can be divided in to two major clusters. Wild accessions are grouped into one cluster and all chickpea cultivars in another cluster, which is sub divided into groups and sub groups. ICCV cultivars and Annegiri clustered into one group and all Pusa accessions were clustered together in another group. The similarity coefficient varied from 0.84 to 0.96 in cultivars, while it varied from 0.71 to 0.81 in wild accessions.
Although the Cicer species are predominantly self-pollinating, more variation was observed among them. The reason for this genetic variation could be that the specific accessions were heterozygous at some marker loci. Similar observations were reported in pea and lentil (Simon and Muehlbauer 1997), in chickpea (Moussa et al. 1996; Sant et al. 1999). Considerable variation was observed between wild accessions and cultivated chickpea (77.8%) in RAPD analysis. Iruela et al. (2002) showed that RAPD markers successfully identified genetic variation in Cicer. The variation identified was greater than that revealed by the isozymes or seed storage proteins used in previous studies of genetic relationships among annual Cicer species (Ahmad et al. 1992; Labdi et al. 1996; Tayyer and Wainess 1996). As large amount of genetic variation exists between chickpea cultivars and its wild accessions, this can be used efficiently for gene tagging and genome mapping of wild and cultivar crosses to introgress disease and insect resistance into the cultivated genotypes.
ISSR analysis
Ten primers were taken for ISSR analysis and six primers (ISSR 1, ISSR 2, ISSR 5, ISSR 6, ISSR 7 and ISSR 10) were found to be polymorphic. On an average 11 bands per primer and 80% polymorphic bands were observed in a total of 64 bands. The wild species shared 20.3% common bands and 79.6% polymorphic bands with ISSR markers. 43.75% common bands and 56.25% polymorphic bands were found among cultivated chickpea varieties and 34.37% common bands and 65.63% polymorphic bands were found among wild accessions. Out of six polymorphic ISSR primers, ISSR-5 (Fig. 3) given the maximum bands (16) and ISSR-7 showed least number of bands (7).
Agarose gel showing the amplified product using ISSR-5 primer. M; I kb ladder, Lane 1–8; Pusa-256, Pusa-362, Pusa-372, Pusa-391, Pusa1003, Pusa-1101, Pusa-1103 and Pusa-1105, Lane 9–13; ICC 17121, ICC 17123, ICC 177124, ICC 17160 and ICC 17163, Lane 14–24; BGD-72, BGD-112, DG-36, DG-46, DG-51, DG-63, DG-71, DG-72, ICCV-2, ICCV-10 and Annegiri
ISSR dendrogram
A close look at UPGMA dendrogram (Fig. 4) revealed that the material taken for analysis is divided into two major clusters. All wild accessions except ICC17163 are grouped together in the first cluster. Chickpea cultivars formed a major group in the second cluster. This group has been divided into two sub groups in which Pusa-1105 and Annegiri are together, whereas the other cultivars constitute a separate group. The similarity coefficient values for the cultivated chickpea lines in this UPGMA dendrogram range between 0.76 and 1.00, whereas the values range between 0.58 and 0.76 in case of wild accessions.
It was found that all the chickpea cultivars present in a cluster have similar pedigree or share a common parent. When compared to the RAPD dendrogram, the ISSR dendrogram showed more correlation with the pedigree data, which shows that the ISSR markers are the most efficient marker system, because of their capacity to reveal several informative bands from single amplification (a mean of 11 informative bands per primer). Similar observations were reported by Bornet and Branchard (2001) and Fernandez et al. (2002) in barley and Qian et al. (2001) in rice.
Since ISSR markers are dominant, the similarity at the sequence level of monomorphic bands can be questioned. But numerous studies have new verified that most co-migrating fragments are identical by descent, at least at the intraspecific level (Wu et al. 2000; Sales et al. 2001). Rajesh et al. (2003) reported that genetic relationship analysis based on ISSRs supports the morphological and crossability data, ISSRs prove tobe an efficient marker system.
The diversity thus observed with microsatellites in the chickpea germplasm is probably due to the use of landraces throughout most of the Indian subcontinent (Malhotra et al. 1987 and Sant et al. 1999), and even today these landraces are being used for the development of elite cultivars. However, the genetic diversity between the various landraces still remains tobe studied and molecular markers will be greatly useful in quantifying this diversity.
RAPD and ISSR dendrogram
To decrease the inaccuracies of the independent techniques, a dendrogram was developed by pooling the data of both RAPD and ISSR. Two major clusters were observed in this UPGMA dendrogram (Fig. 5). All the wild accessions grouped together into one major cluster, whereas all the chickpea cultivars formed a second cluster. The second major cluster is again divided into many groups, sub groups and sub-sub groups. The similarity coefficient for the chickpea cultivars in the dendrogram varies from 0.8 to 0.94, whereas for wild accessions ranges from 0.71 to 0.80. When the dendrogram was correlated with the pedigree data it was found that very few cultivars with common parents clustered together. The dendrogram showed similar clustering pattern for the wild accessions with that of RAPD, but in the case of cultivars, it showed much variation with the dendrogram of both RAPD and ISSR.
The chickpea cultivars DG-63 and DG-71 clustered together as they have similar pedigree. Similarly, Pusa-1101 and Pusa-1103 are in the same cluster as they have similar pedigree data. The degree of variation within Cicer arietinum and Cicer reticulatum accessions depended on the accessions. This observation was consistent with the study by Simon and Muehlbauer (1997), who detected variation within single C. reticulatum accession (PI 489777), used to generate an interspecific mapping population. Our results are in accordance with Iruela et al. (2002) reported that low level of genetic diversity within C. arietinum compared to the wild species. Shan et al. (2005) showed that a natural hybrid could be useful for bridging crosses to introduce genes to chickpea from incompatible species given that C. reticulatum was the wild progenitor of chickpea.
The observed genetic variation within specific accessions could be due to the mixture of homozygous plants or the specific accessions were heterozygous at some marker loci. Although Cicer species are predominantly self-pollinating, a low level of out crossing has been previously reported (Gowda 1981). The amount of genetic variation detected within C. arietinum was less than that detected within the wild C. reticulatum. This indicated that the wild C. reticulatum might offer potential novel sources of genetic variation, which may be introduced into cultivars to broaden the genetic base of chickpea through interspecific hybridization. Successful crosses have been made between the two most closely related wild species (C. reticulatum and C. echinospermum) and the cultivated species (Ladizinsky and Adler 1976; Singh and Ocampo 1997; Collard et al. 2003b; Nguyen et al. 2004).
ISSR analysis is more economical and reliable than that of RAPD. Earlier studies also reported that ISSR technique generates large number of polymorphisms in chickpea (Collard et al. 2003a). The phylogenetic relationship between Cicer species from this study was overall consistent with most previous studies (review by Croser et al. 2003; Nguyen et al. 2004; Sudupak 2004; Sudupak et al. 2004).
Conclusion
The present investigation demonstrates the potential of RAPD and ISSR fingerprinting in detecting polymorphism among chickpea cultivars and wild accessions. The ISSR analysis clearly indicated that even with six polymorphic primers reliable estimation of genetic diversity could be obtained, while nearly 30 primers are required for RAPD. Moreover, RAPD can cause genotyping errors due to competition in the amplification of all RAPD fragments. C. arietinum cultivars had the narrowest genetic variation while its wild C. reticulatum accessions had much greater genetic variation, which could be used in chickpea improvement. However, the genetic diversity was unbalanced and varied considerably between species in the world collections of wild annual Cicer germplasm. The number of accessions and genetic diversity in the wild annual Cicer germplasm was very limited overall compared to other major collections, such as wheat, barley and rice (Virk et al. 1995; Abbo et al. 2003). The markers generated by ISSR and RAPD assays can provide practical information for the management of genetic resources. For the selection of good parental material in breeding programs the genetic data produced through ISSR can be used to correlate with the relationship measures based on pedigree data and morphological traits to minimize the individual inaccuracies in chickpea. Further, large amount of genetic variation which exists between chickpea cultivars and its wild accessions can be used efficiently for gene tagging and genome mapping of wild and cultivar crosses to introgress the disease and insect resistance into the cultivated genotypes.
Abbreviations
- ISSR:
-
Inter Simple Sequence Repeats
- RAPD:
-
Random Amplified Polymorphic DNA
References
Abbo S, Berger J, Turner NC (2003) Evaluation of cultivated chickpea: four bottlenecks limit diversity and constrain adaptation. Func Plant Biol 30:1081–1087
Ahmad F, Slinkard AE (1992) Genetic relationships in the genus Cicer L. as revealed by polyacrylamide gel electrophoresis of seed storage proteins. Theor Appl Genet 84:688–692
Bornet B, Branchard M (2001) Non anchored inter simple sequence repeats (ISSR) markers reproducible and specific tools for genome finger printing. Plant Mol Biol Rep 19:209–215
Collard BCY, Pang ECK, Taylor PWJ (2003a) Selection of wild Cicer accessions for the generation of mapping populations segregating for resistance to ascochyta blight. Euphytica 130:1–9
Collard BCY, Pang ECK, Ades PK, Taylor PWJ (2003b) Preliminary investigation of QTLs associated with seedling resistance to ascochyta blight from Cicer echinospermum, a wild relative of chickpea. Ther Appl Genet 107:719–729
Croser JS, Ahmad F, Clarke HJ, Siddique KHM (2003) Utilization of wild Cicer in chickpea improvement-progress, constraints, and prospects. Aust J Agri Res 54:429–444
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Duran Y, Fratini R, Garcia P, Perez de la Vega M (2004) An intersubspecific genetic map of Lens. Theor Appl Genet 108:1265-1270.
FAO (2004) http://faostat.fao.org
Fernandez ME, Fygueiras AM, Benito C (2002) The use of ISSR and RADP markers for detecting DNA polymorphism genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet 104:845-851.
Gowda CLL (1981) Natural outcrossing in chickpea. Int Chickpea Newsl 5:6
Iruela M, Rubio J, Cubero JI, Gil J, Milan T (2002) Phylogenetic analysis in the genus Cicer and cultivated chickpea using RAPD and ISSR markers. Theor Appl Genet 104:643–651
Labdi M, Robertson LD, Singh KB, Charrier A (1996) Genetic diversity and phylogentic relationships among the annual Cicer species as revealed by isozyme polymorphism. Euphytica 88:181–188
Ladizinsky G, Adler A (1976) Genetic relationships among the annual species of Cicer L. Theor Appl Genet 48:197–203
Malhotra RS, Pundir RPS, Slinkard AE (1987) Genetic resources of chickpea. In: Saxena MC, Singh KB (eds) The chickpea. CAB International, Wallingford, UK, pp 67–81
Moussa EH, Millan T, Gil J, Cubero JI (1996) Variability and genome length estimation in chickpea (Cicer arietinum L.) revealed by RAPD analysis. J Genet Breed 51:83–85
Nguyen TT, Taylor PWJ, Redden RJ, Ford R (2004) Genetic diversity estimates in Cicer using AFLP analysis. Plant Breed 123:173–179
Panguluri SK, Janaiah K, Govil JN, Kumar PA, Sharma PC (2006) AFLP fingerprinting in Pigeonpea (Cajanus cajan (L.) Millsp.) and its wild relatives. Genet Resour Crop Evol 53:523–531
Qian W, Ge S and Hang DY (2001) Genetic variation with in and among populations of a wild rice Oryza granulata from china detected by RAPD and ISSR markers. Theor Appl Genet 102:440–449
Rajesh PN, Tullu A, Gil J, Gupta VS, Ranjekar PK, Muehlbaur FJ (2002) Identification of an STMS marker for the double-podding gene in chickpea. Theor Appl Genet 105:604–607
Rajesh PN, Sant VJ, Gupta VS, Muehlbaur FJ and Ranjekar PK (2003) Genetic relationships among annual and perennial wil species of Cicer using inter simple sequence repeat (ISSR) polymorphism. Euphytica 129:15–23
Ratnaparkhe MB, Santra DK, Tullu A, Muehlbauer FJ (1998) Inheritance of inter simple sequence repeat polymorphism and linkage with fusarium wilt resistance gene in chickpea. Theor Appl Genet 96:348–353
Reddy MP, Sarla N, Siddiq EA (2002) Inter simple sequence repeats (ISSR) polymorphism and its application in plant breeding. Euphytica 128:9–17
Robertson LD, Ocampo B, Singh KB (1997) Morphological variation in wild annual Cicer species in comparision to the cultigen. Euphytica 95:309–319
Sales E, Nebauer SG, Mus M, Segura J (2001) Population genetic study in the Balearic endemic plant species Digitalis minor (Scrophuiariaceae) using RAPD markers. Am J Bot 88:1750–1759
Sant VJ, Patankar AG, Sarode ND, Mhase LB, Sainani MN, Deshmukh RB, Ranjekar PK, Gupta VS (1999) Potential of DNA markers in detecting divergence and in analyzing heterosis in Indian elite chickpea cultivars. Theor Appl Genet 98:1217–1225
Santra DK, Tekeoglu M, Ratnaparkhe MB, Kaiser WJ, Muehlbauer FJ (2000) Identification and mapping of QTLs conferring resistance to ascochyta blight in chickpea. Crop Sci 40:1606–1612
Shan F, Clarke HC, Plummer JA, Yan G, Siddique KHM (2005) Geographical patterns of genetic variation in the world collections of wild annual Cicer characterized by amplified fragment length polymorphism. Theor Appl Genet 110:381–391
Simon CJ, Muehlbauer FJ (1997) Construction of chickpea linkage map and its comparison with the maps of pea and lentil. J Heridity 88:115–119
Singh KB, Ocampo B (1997) Exploration of wild Cicer species for yield improvement in chickpea. Theor Appl Genet 95:418–423
Singh KB, Malhotra RS, Halila MH, Knights EJ, Verma MM (1994) Current status and future strategy in breeding chickpea for resistance to biotic and abiotic stresses. Euphytica 73:137–149
Sudupak MA, Akkaya MS, Kence A (2002) Analysis of genetic relationships among perennial and annual Cicer species growing in Turkey using RAPD markers. Theor Appl Genet 105:1220–1228
Sudupak MA (2004) Inter- and intra-species inter simple sequence repeat (ISSR) variation in the genus Cicer. Euphytica 135:229–238
Sudupak MA, Akkaya MS, Kence A (2004) Genetic relationships among perennial and annual Cicer species growing in Turkey assessed by AFLP fingerprinting. Theor Appl Genet 108:937–944
Tayyer RI, Waines JG (1996) Genetic relationships among annual species of Cicer (Fabaceae) using isozyme variation. Theor Appl Genet 92:245–254
Virk PS, Newbury HJ, Jacckson MT, Ford-Lioyd BV (1995) The identification of duplicate accessions within a rice germplasm collection using RAPD analysis. Theor Appl Genet 90:1049–1055
Willams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphism amplified by arbitraray primers are useful as genetic markers. Nuceic Acids Res 18:6531–6535
Winter P, Benko-Iseppon AM, Huttel B, Ratnaparkhe MB, Tullu A, Sonnante G, Pfaff T, Tekeoglu M, Santra D, Sant VJ, Rajesh PN, Kahl G, Muehlbauer FJ (2000) A linkage map of the chickpea (Cicer arietinum L.) genome based on recombinant inbred lines from a C. arietinum × C. reticulatum cross: localization of resistance genes for fusarium races 4 and 5. Theor Appl Genet 101:1155–1163
Wu J, Krutovski KV, Strauss SH (2000) Nuclear DNA diversity, population differentiation and phylogetic relationships in the California closed-cone pines based on RAPD and allozyme markers. Genome 42:893–908
Acknowledgements
Authors are grateful to Dr. S S Yadav, Principal Scientist and Dr. Jitender Kumar, Senior Scientist, Pulse Research Laboratory, IARI, New Delhi for providing seed material.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rao, L.S., Usha Rani, P., Deshmukh, P.S. et al. RAPD and ISSR fingerprinting in cultivated chickpea (Cicer arietinum L.) and its wild progenitor Cicer reticulatum Ladizinsky. Genet Resour Crop Evol 54, 1235–1244 (2007). https://doi.org/10.1007/s10722-006-9104-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-006-9104-6