Abstract
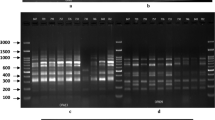
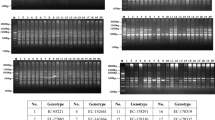
Randomly amplified polymorphic DNA (RAPD), inter-simple sequence repeat (ISSR) and a semi-random PCR system were used to analyze the genetic diversity of 16 Italian common bean landraces and their relationship to four commercial cultivars. Of the primers tested, 8 ISSR, 6 RAPD and 7 semi-random primers produced polymorphic and reproducible DNA fragments. A higher proportion of polymorphic bands were observed using ISSR (85%) and semi-random (90%) primers than RAPD (69%) method. The combination of any two semi-random markers allowed the identification of all 20 bean genotypes. In contrast ISSR (except for primer (CAC)3GC) and RAPD markers appeared to be less informative as more than two markers were necessary to achieve the same diagnostic level. Moreover, 7 ISSR, 2 RAPD and 8 semi-random exclusive bands were identified as putative population-specific markers. Semi-random and ISSR derived dendrograms showed similar tendencies in terms of genetic relatedness, whereas clustering of genotypes within groups was not similar when compared with the RAPD technique. Despite the different ability to resolve genetic variation among the investigated landraces, two major clusters with less than 60% (ISSR) and 40% (RAPD and semi-random) genetic similarity were formed with all three marker systems. The two groups were correlated with the phaseolin patterns and seed size of the landraces. The analysis showed that the cultivar ȁ8Lingua di Fuocoȁ9 and most of the landraces (13 out of 16) collected in Italy belong to the Andean gene pool, whereas only the three populations from Pratomagno belong to the Middle American gene pool.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
H. Akagi, Y. Yokozeki, A. Imagaki, A. Nkamura and T. Fujimura, Microsatellite DNA markers for rice chromosomes. Theor. Appl. Genet. 93 (1996) 866-873
B. Bornet, C. Muller, F. Paulus and M. Branchard, Highly informative nature of inter simple sequence repeat (ISSR) sequences amplified using tri- and tetra-nucleotide primers from DNA of cauliflower (Brassica oleracea var. botrytis L.). Genome 45 (2002) 890-896
L. Briand, A.E. Brown, J.M. Lenné and D.M. Teverson, Random amplified polymorphic DNA variation within and among bean landrace mixtures (Phaseolus vulgaris L.) from Tanzania. Euphytica 102 (1998) 371-377
CEE regulation n. 2081/92 Gazzetta ufficiale n. L 163 del 02/07/1996, pp. 0019–0021.
Y.M. Charters, A. Robertson, M.J. Wilkinson and G. Ramsay, PCR analysis of oilseed rape cultivars (Brassica napus L. ssp. oleifera) using 5′ anchored simple sequence repeat (SSR) primers. Theor. Appl. Genet. 92 (1996) 442-447
E.C. Chin, M.L. Senior, H. Shu and J.C. Smith, Maize simple repetitive DNA sequences: abundance and allele variation. Genome 39 (1996) 866-873
Trends in CIAT commodities. CaliColombia: CIAT (1993).
A.G. Connolly, I.D. Godwin, M. Cooper and I.H. DeLacy, Interpretation of randomly amplified polymorphic DNA data for fingerprinting sweet potato. Theor. Appl. Genet. 88 (1994) 332-336
T. Demeke, R.P. Adams and R. Chibbar, Potential taxonomic use of random amplified polymorphic DNA (RAPD): a case study in Brassica. Theor. Appl. Genet. 84 (1992) 990-994
J. Felsenstein, Confidence limits on phylogenesis: an approach using the bootstrap. Evolution 39 (1985) 783-791
M.E. Fernandez, A.M. Figueiras and C. Benito, The use of ISSR and RAPD marker for detecting DNA polymorphismgenotype identification and genetic diversity among barley cultivars with known origin. Theor. Appl. Genet. 104 (2002) 845-851
R. Freyre, R. Ríos, L. Guzmán, D.G. Debouck and P. Gepts, Ecogeographic distribution of Phaseolus spp. (Fabaceae) in Bolivia. Econ. Bot. 50 (1996) 195-215
S. Fukuoka, K. Hosaka and O. Kamijima, Use of random amplified polymorphic DNAs (RAPDs) for identification of rice accessions. Jpn. J. Genet. 67 (1992) 243-252
M.Z. Galvan, M.B. Aulicino, G. Medina and P.A. Baratti, Genetic diversity among Northwestern Argentinian cultivars of common bean (Phaseolus vulgaris L.) as revealed by RAPD markers. Genet. Resour. Crop Evol. 48 (2001) 251-260
M. Gawel, I. Wisniewska and A. Rafalski, Semi-specific PCR for the evaluation of diversity among cultivars of wheat and triticale. Cell Mol. Biol. Letters 7 (2002) 577-582
P. Gepts and F.A. Bliss, Phaseolin variability among wild and cultivated common beans (Phaseolus vulgaris) from Colombia. Econ. Bot. 40 (1986) 469-478
P. Gepts and F.A. Bliss, Dissemination pathways of common bean (Phaseolus vulgarisFabaceae) deduced from phaseolin electrophoretic variability. II. Europe and Africa. Econ. Bot. 42 (1988) 86-104
P. Gepts, A middle American and an Andean gene pool. In: P. Gepts (ed.) Genetic Resources of Phaseolus Beans. Dordrecht, the Netherlands: Kluwer (1988) pp. 375-390
P. Gepts, The use of molecular and biochemical markers in crop evolution studies. Evol. Biol. 27 (1993) 51-94
P. Gepts, T. Osborn, K. Rashka and F.A. Bliss, Phaseolin-protein variability in wild forms and landraces of the common bean (Phaseolus vulgaris): evidence for multiple centers of domestication. Econ. Bot. 40 (1986) 451-468
P. Gepts, K. Kmiecik, P. Pereira and F.A. Bliss, Dissemination pathways of common bean (Phaseolus vulgarisFabaceae) deduced from phaseolin electrophoretic variability. I. The Americas. Econ. Bot. 42 (1988) 73-85
J.E. Gilbert, R.V. Lewis, M.J. Wilkinson and P.D.S. Caligari, Developing an appropriate strategy to assess genetic variability in plant germplasm collections. Theor. Appl. Genet. 98 (1999) 1125-1131
L. Goulao, T. Valdiviesso, C. Santana and C.M. Oliveira, Comparison between phenetic characterisation using RAPD and ISSR markers and phenotypic data of cultivated chestnut (Castanea sativa Mill.). Genet. Resour. Crop Evol. 48 (2001) 329-338
J.C. Gower, Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53 (1966) 325-338
P. Jaccard, . Handbuch Biologische Arbeitsmethoden 11 (1928) 165-202
M.A. Johns, P.W. Skroch, J. Nienhuis, P. Hinrichsen, G. Bascur and C. Muñoz-Schick, Gene pool classification of common bean landraces from Chile based on RAPD and morphological data. Crop Sci. 37 (1997) 605-613
R.V. Kantety, X. Zeng, J.L. Bennetzen and B.E. Zehr, Detection of high levels of polymorphism among dent and popcorn inbreds using Inter-Simple Sequence Repeat (ISSR) amplification technique. Maize Genet. Coop. Newslett. 69 (1995) 132-134
T. Kojima and Y. Ogihara, High-resolution RFLP map of the long arm of chromosome 5A in wheats and its synteny among cereals. Genes Genet. Syst. 73 (1998) 51-58
G. Limongelli, G. Laghetti, P. Perrino and A.R. Piergiovanni, Variation of seed storage proteins in landraces of common bean (Phaseolus vulgaris L.) from BasilicataSouthern Italy. Euphytica 92 (1996) 393-399
L. Lioi, Geographical variation of phaseolin patterns in an Old World collection of Phaseolus vulgaris. Seed Sci. Technol. 17 (1989) 317-324
V. Llaca, A. Delgado Salinas and P. Gepts, Chloroplast DNA as an evolutionary marker in the Phaseolus vulgaris complex. Theor. Appl. Genet. 88 (1994) 646-652
Y. Loarce, R. Gallego and E. Ferrer, A comparative analysis of genetic relationships between rye cultivars using RFLP and RAPD markers. Euphytica 88 (1996) 107-115
F.L. Maciel, L.T.S. Gerald and S. Echeverrigaray, Random amplified polymorphic DNA (RAPD) markers variability among cultivars and landraces of common beans (Phaseolus vulgaris L.) of South-Brazil. Euphytica 120 (2001) 257-263
I. Metais, C. Aubry, B. Hamon, R. Jalouzot and D. Peltier, Description and analysis of genetic diversity between commercial bean lines (Phaseolus vulgaris L.). Theor. Appl. Genet. 101 (2000) 1207-1214
V. Miccolis, Aspetti generali ed innovativi del fagiolo fresco in Italia. Informatore agrario 23 (1995) 49-56
M. Morgante and A.M. Olivieri, PCR-amplified microsatellite markers in plant genetics. Plant J. 3 (1993) 175-182
J. Nowosielski, W. Podyma and D. Nowosielska, Molecular research on the genetic diversity of polish varieties and landraces of Phaseolus coccineus L. and Phaseolus vulgaris L. using the RAPD and AFLP methods. Cell Mol. Biol. Letters 7 (2002) 753-762
A.R. Piergiovanni, D. Cerbino and M. Brandi, The common bean populations from Basilicata (Southern Italy). An evaluation of their variation. Genet. Resour. Crop Evol. 47 (2000) 489-495
A. Prevost and M.J. Wilkinson, A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor. Appl. Genet. 98 (1999) 107-112
A. Rafalski, I. Wisniewska and M. Gawel, PCR-based system for evaluation of genetic relationship among inbred lines of maize and rye. J. Appl. Genet. 39A (1998) 90
A.P. Rodino, M. Santalla, I. Montero, P.A. Casquero and A.M. Ron De, Diversity of common bean (Phaseolus vulgaris L.) germplasm from Portugal. Genet. Resour. Crop Evol. 48 (2001) 409-417
F.J. Rohlf, NTSYS-PC: Numerical Taxonomy and Multivariate Analysis SystemVersion 2.02h. New York: Exeter Software (1993).
J. Rongwen, M.S. Akkaya, A.A. Bhagwat, U. Lavi and P. Cregan, The use of microsatellite DNA markers for soybean genotype identification. Theor. Appl. Genet. 90 (1995) 43-48
M.A. Saghai-Maroof, K.M. Soliman, R.A. Jorgensen and R.W. Allard, Ribosomal spacer length polymorphism in barley: Mendelian inheritance chromosomal location and population dynamics. Proc. Natl. Acad. Sci. USA 81 (1984) 8014-8019
S.S. Salimath, A.C. Oliveira de, I.D. Godwin and J.L. Bennetzen, Assessment of genome origins and genetic diversity in the genus Eleusine with DNA markers. Genome 38 (1995) 757-63
Y. Tao, J.M. Manners, M. Ludlow and R.J. Henzel, DNA hum (Sorghum bicolor L.) Moench). Theor. Appl. Genet. 86 (1993) 679-688
Toniolo L. 1989. Il fagiolo. In: Baldoni R. and Giardini L. (eds), Coltivazioni erbacee. Patron Editrice, pp. 302–314.
S.E. Wilkie, P.G. Isaac and P.G. Slater, Random amplified polymorphic DNA (RAPD) markers for genetic analysis in Allium. Theor. Appl. Genet. 86 (1993) 497-504
J.G.K. Williams, A.R. Kubelik, K.J. Livak, J.A. Rafalski and S.V. Tingey, DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 18 (1990) 6531-6535
W. Yang, A.C. Oliveira de, I. Godwin, K. Schertz and J.L. Bennetzen, Comparison of DNA marker technologies in characterizing plant genome diversity: variability in Chinese sorghums. Crop Sci. 36 (1996) 1669-1676
I. Yap and R.J. Nelson, WinBoot: a program for performing bootstrap analysis of binary data to determine the confidence limits of UPGMA-based dendrograms. Manila, Philippines: International Rice research Institute (1996).
K. Yu, S.J. Park and V. Poysa, Abundance and variation of microsatellite DNA sequences in beans (Phaseolus Vigna). Genome 42 (1999) 27-34
K.F. Yu and K.P. Pauls, Segregation of random amplified polymorphic DNA markers and strategies for molecular mapping in tetraploid alfalfa. Genome 36 (1993) 844-851
E. Zietkiewicz, A. Rafalski and D. Labuda, Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20 (1994) 176-183
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marotti, I., Bonetti, A., Minelli, M. et al. Characterization of Some Italian Common Bean (Phaseolus Vulgaris L.) Landraces by RAPD, Semi-random and ISSR Molecular Markers. Genet Resour Crop Evol 54, 175–188 (2007). https://doi.org/10.1007/s10722-005-3133-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-005-3133-4