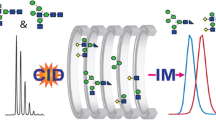
Abstract
Ion mobility mass spectrometry (IM-MS) is a promising analytical technique for glycomics that separates glycan ions based on their collision cross section (CCS) and provides glycan precursor and fragment masses. It has been shown that isomeric oligosaccharide species can be separated by IM and identified on basis of their CCS and fragmentation. These results indicate that adding CCSs information for glycans and glycan fragments to searchable databases and analysis pipelines will increase identification confidence and accuracy. We have developed a freely accessible database, GlycoMob (http://www.glycomob.org), containing over 900 CCSs values of glycans, oligosaccharide standards and their fragments that will be continually updated. We have measured the absolute CCSs of calibration standards, biologically derived and synthetic N-glycans ionized with various adducts in positive and negative mode or as protonated (positive ion) and deprotonated (negative ion) ions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein glycosylation is a prevalent post-translational modification, involving complex biosynthesis pathways, that modifies specific sites along the protein backbone. Glycosylation plays an important role in numerous biological processes, ranging from fertilization and immune response to cell−cell recognition and inflammation [1–3]. Advances in glycan characterisation technologies have increased our understanding of the molecular and structural roles of glycans in biological processes, which has led to the development of biotherapeutics including, monoclonal antibodies with specific glycoforms, that is driving drug design and vaccine programmes [4]. Despite the far-reaching functional and structural roles of glycans, supporting computational methods to store and handle the growing volumes of data still remain in their infancy, lagging behind other -omics bioinformatics activities. Furthermore, the absence of sophisticated tools directly impacts the quality of data reported in other databases, as exemplified by the carbohydrate stereochemistry and nomenclature misassignments in the Protein Data Bank [5].
Several initiatives to catalogue and organise glycan-related (structural and experimental) information have been released starting with CarbBank [6] and followed by the KEGG GLYCAN database [7], the Consortium for Functional Glycomics (CFG) [8], GLYCOSCIENCES.de [9], EUROCarbDB [10], GlycomeDB [11] and UniCarbKB [12]. A key impetus in glycomics is now perceptible in the move toward large-scale analysis of the structure and function of glycans. A diverse range of technologies and strategies is being applied to address the technically difficult problems of glycan structural analysis and subsequently the investigation of their functional roles [13].
Mass spectrometry (MS), either directly or in conjunction with chromatographic separation, is widely employed for the characterisation of oligosaccharides [14], yet studies remain hampered due to the inherent complexity of glycan structures. A technique that has the potential to improve glycan analysis is ion mobility-MS (IM-MS), in which ions are separated according to their mass, charge, size and shape. IM-MS separates ions based on the time required to traverse a region of neutral gas, generally helium or nitrogen, under the influence of a weak electric field and reports both an arrival time and mass-to-charge (m/z) value. The arrival time can be converted into a collision cross sections (CCS) which is an absolute value reflecting the rotationally averaged structure of the glycan ion. To date, only a limited number of reports apply IM-MS for glycomics [15–18], but we have demonstrated its potential including the separation of isomeric glycans, and by combining IM-MS with fragmentation to provide informative spectra of N-glycans released from sub-microgram amounts of human immunodeficiency virus gp120 [19–21].
Some commercially available IM-MS instruments use a non-uniform travelling wave (TW) field to transport ions through the IM cell and require calibration with known compounds to estimate CCSs. We have shown that when estimating CCSs using TW IM-MS instruments (TWCCS) it is vital to use known calibrant CCS values measured in the same IM gas and are of the same sample molecular class as the analyte (i.e., dextran CCSs measured in N2 for estimating N-glycans in N2) [20, 21]. Drift tube IM-MS instruments that utilize a uniform electric field on the other hand, do not require calibration and CCS (DTCCS) measurements are highly reproducible, precise and referred to as “absolute” in this report.
IM-MS derived CCSs have considerable potential as an additional dimension of structural information that is highly complementary to existing glycomics data storage and analysis pipelines. Because of the compatibility of ion mobility-mass spectrometry (IM-MS) with other separation techniques such as high-pressure liquid chromatography (U/HPLC), it is conceivable to implement CCS information in already existing databases. However, there are no tools available for supporting such data collections. Here, we address this deficiency by introducing the GlycoMob database that is tailored towards the storage of IM-MS data including absolute drift tube DTCCS and traveling wave TWCCS values, and how these data add new functionality to UniCarbKB.
Database implementation
GlycoMob (http://www.glycomob.org) is a modular extension of the freely available UniCarbKB framework that extends the MS resources provided by UniCarb-DB. The application is developed with the Play Framework using Java and Scala with a PostgreSQL database. The GlycoMob database schema (or collection of structures, experimental data and associated metadata) is embedded into the UniCarbKB database infrastructure. The integration of GlycoMob with UniCarb databases allows for improved data discovery and cross-referencing over the curated and experimentally verified data collections provided by the UniCarbKB initiative [22, 12].
Design and search features
GlycoMob aims to reinforce and implement data-sharing standards to provide a framework that supports new and current analytical technologies. As such, GlycoMob adopts the UniCarbKB user interface to retain familiarity across all UniCarb-related resources. The database is organized into three major sections including: (i) a complete listing of structures and compositions stored in the database; (ii) biological content-specific data collections, including glycans released from purified glycoproteins; and (iii) a description of analysed commercial glycans. Each section can be accessed from the GlycoMob homepage. In addition, GlycoMob can be searched in three ways: (i) by CSS value and underivatised mass; (ii) by glycoprotein; and (iii) by monosaccharide composition. Each search function can be accessed from the navigation bar and detailed information is provided. By default the search results will return matching structures or compositions.
Glycan structure CSS values
The first release of GlycoMob includes a broad range of masses and CCSs for native glycans as well as their fragments. In total, over 350 CCS values are attributed to 70 precursor glycan structures determined in both N2 and He drift gases. Furthermore, the database includes over 300 fragment ion measurements, predominately determined in the negative ion mode, which can be used to complement structure assignments using IM. As an example, the experimental CCS values obtained for native glycans and fragments from ribonuclease B are shown in Fig. 1 and the coverage of data accumulated from the standard glycoproteins is summarised in Fig. 2. Each entry is comprehensively annotated and includes: a description of the glycoprotein from which the glycans were released or, in the case of commercial standards, a reference to the supplier. Additional information on experimental conditions including gas pressures and voltages are also documented.
Screenshot showing the absolute CCSs of RNase B native glycans and fragments in helium and nitrogen. The CCS measurements are grouped together by the drift gas, selecting either of the tabs above the main table will display the relevant data sets. Users can automatically search the table content via the ‘Filter by CCS’ bar, additional search options include by composition and glycoprotein listed in the side panel or the main ‘Query’ page
Screenshot of the GlycoMob homepage as part of the UniCarbKB database. In the centre of the page is a concise summary of the data available including the glycoproteins analysed and associated number of CCS values for native structures and fragments. Embedded links in the table will redirect the user to the relevant data content pages
Experimental methods
GlycoMob has been populated initially with N-glycans released from four common well-characterised glycoproteins whose structures span all major classes, from high-mannose to complex. A detailed description of the experimental methods and procedures has been previously reported [20]. Briefly, N-linked glycans were released using hydrazine, from the glycoproteins porcine thyroglobulin, ribonuclease B, chicken ovalbumin, and bovine fetuin obtained from Sigma Chemical Co., Ltd. (Poole, Dorset, U.K.) and subsequently reacetylated prior to analysis. For the glycoprotein standards thyroglobulin and fetuin, sialic acids were removed by heating with 1 % acetic acid for 1 h at 70 °C. For electrospray analysis, samples were dissolved in water:methanol (1:1, v:v) at ∼1 mg/mL. Additionally, CSS measurements were generated for the widely used calibration standard dextran (Fluka and Sigma-Aldrich).
Measurements of absolute collision cross sections were performed using a Synapt G1 HDMS quadrupole/IMS/oa-ToF instrument (Waters Co., Manchester, U.K.) modified for drift tube operation [23]. The reported drift tube CCS represent averages of three (He) or two (N2) replicates acquired in independent measurements. In addition, high-resolution TW IM-MS measurements were performed on an unmodified Synapt HDMS G2-S 8000 m/z quadrupole/IMS/oa-ToF MS instrument (Waters Corporation, Manchester, U.K.). Based on these data, travelling wave CCS values were estimated as reported previously [20], and used to construct GlycoMob.
The coverage of GlycoMob also includes CSS (drift tube and travelling wave) measurements for synthetic pure high-mannose N-glycans (Dextra, Reading, UK) as [M+H]+, [M+Na]+, [M+K]+, [M-H]−, [M+Cl]− and [M+H2PO4]− ions in He and N2 drift gases [24] (Fig. 3).
High-mannose summary page. For each structure entry the CCSs measured in positive and negative mode using either nitrogen or helium drift gas are shown together with its adducts. Additionally, ion mobility data sets can be accessed from the ‘Glycoprotein Standards’ and ‘Dextran’ drop down lists in the right side panel. Users can select the embedded UniCarb-DB and UniCarbKB links to access relevant mass spectra and structural information, respectively. The structures are displayed using the hybrid Consortium for Functional Glycomics [25] and Oxford graphical notation [26, 27]
Bridging GlycoMob with UniCarb-related databases
A significant problem facing the glycobioinformatics community is the distribution of data. The principal objective of the UniCarbKB initiative is not only to connect related data collections, but also to provide descriptive metadata that is representative of the reported data. Efforts led by the Minimum Information for A Glycomics Experiment (MIRAGE) [28] project and the ontology work of GlycoRDF [29] aim to alleviate this situation by providing standardized data terms. In this context, fully characterized structures stored in GlycoMob (e.g., high-mannose standards) are linked to the tandem MS data repository UniCarb-DB. Similarly, users can access data stored in the curated glycan structure database UniCarbKB (and vice-versa), by following the links for individual glycoprotein entries and glycan compositions. Such connections allow researchers to easily navigate between databases and view experimental tandem MS spectra and related structural/glycoprotein information.
GlycoMob adopts GlycoRDF to provide a common standard for representing the stored information. GlycoRDF aims to enhance data interoperability by providing an ontology to standaridise glycomics data for building data-driven applications, which efficiently integrate heterogeneous datasets through the provision of a single SPARQL endpoint that can be used to query and return data from the semantic web. For example, such an endpoint enables users to simultaneously query GlycoMob, UniCarb and other RDF-compatible databases to answer complex research questions. The development of web applications to perform such queries is in its infancy, however, examples (using SPARQL) are provided on the UniCarbKB RDF site (http://rdf.unicarbkb.org).
Discussion
CCS measurements provide an additional parameter that can be used to improve the specificity of glycan identification. Here, we have described GlycoMob a novel database solution for storing and making available experimentally confirmed CSS values acquired from both TW and DT IM-MS instruments for the glycomics domain, which starts to address the lack of resources available for this emerging technology.
The database provides access to a broad dataset for CCS calibration of TW IM-MS instruments using native N-linked glycans released from naturally occurring and commercially available glycoproteins. Access to such data not only assists calibration, but also will provide an extra level of confidence in the characterization of N-glycan structures. Its availability will serve as a tool for the development of new analytical tools for structural data querying and spectra interpretation, in particular, when connected to liquid chromatography-MS-based workflows and the MS/MS database UniCarb-DB that contains masses, retention times and fragment information.
By providing a common interface to high-quality data, we believe that GlycoMob can become a vital resource for analyzing glycomic ion mobility data. As the technology improves and further data are generated, the database will be continually updated with new CCS values and associated data for glycans released by PNGaseF and hydrazinolysis, thereby providing a leading platform for IM-MS.
References
Spiro, R.G.: Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12(4), 43R–56R (2002)
Cummings, R.D., Pierce, J.M.: The challenge and promise of glycomics. Chem. Biol. 21(1), 1–15 (2014). doi:10.1016/j.chembiol.2013.12.010
Pang, P.C., Chiu, P.C., Lee, C.L., Chang, L.Y., Panico, M., Morris, H.R., Haslam, S.M., Khoo, K.H., Clark, G.F., Yeung, W.S., Dell, A.: Human sperm binding is mediated by the sialyl-Lewis(x) oligosaccharide on the zona pellucida. Science 333(6050), 1761–1764 (2011). doi:10.1126/science.1207438
Dalziel, M., Crispin, M., Scanlan, C.N., Zitzmann, N., Dwek, R.A.: Emerging principles for the therapeutic exploitation of glycosylation. Science 343(6166), 1235681 (2014). doi:10.1126/science.1235681
Crispin, M., Stuart, D.I., Jones, E.Y.: Building meaningful models of glycoproteins. Nat. Struct. Mol. Biol. 14(5), 354 (2007). doi:10.1038/nsmb0507-354a. discussion 354–355
Doubet, S., Albersheim, P.: CarbBank. Glycobiology 2(6), 505 (1992)
Hashimoto, K., Goto, S., Kawano, S., Aoki-Kinoshita, K., Ueda, N., Hamajima, M., Kawasaki, T., Kanehisa, M.: KEGG as a glycome informatics resource. Glycobiology 16(5), 63R–70R (2006)
Raman, R., Venkataraman, M., Ramakrishnan, S., Lang, W., Raguram, S., Sasisekharan, R.: Advancing glycomics: implementation strategies at the consortium for functional glycomics. Glycobiology 16(5), 82R–90R (2006). doi:10.1093/glycob/cwj080
Lutteke, T., Bohne-Lang, A., Loss, A., Goetz, T., Frank, M., von der Lieth, C.: GLYCOSCIENCES.de: an internet portal to support glycomics and glycobiology research. Glycobiology 16(5), 71R–81R (2006)
von der Lieth, C., Freire, A., Blank, D., Campbell, M., Ceroni, A., Damerell, D., Dell, A., Dwek, R., Ernst, B., Fogh, R., Frank, M., Geyer, H., Geyer, R., Harrison, M., Henrick, K., Herget, S., Hull, W., Ionides, J., Joshi, H., Kamerling, J., Leeflang, B., Lutteke, T., Lundborg, M., Maass, K., Merry, A., Ranzinger, R., Rosen, J., Royle, L., Rudd, P., Schloissnig, S.: EUROCarbDB: an open-access platform for glycoinformatics. Glycobiology 21(4), 493–502 (2011)
Ranzinger, R., Frank, M., von der Lieth, C., Herget, S.: Glycome-DB.org: a portal for querying across the digital world of carbohydrate sequences. Glycobiology 19(11), 1563–1567 (2009)
Campbell, M.P., Peterson, R., Mariethoz, J., Gasteiger, E., Akune, Y., Aoki-Kinoshita, K.F., Lisacek, F., Packer, N.H.: UniCarbKB: building a knowledge platform for glycoproteomics. Nucleic Acids Res. 42(1), D215–D221 (2014). doi:10.1093/nar/gkt1128
Kailemia, M.J., Ruhaak, L.R., Lebrilla, C.B., Amster, I.J.: Oligosaccharide analysis by mass spectrometry: a review of recent developments. Anal. Chem. 86(1), 196–212 (2014). doi:10.1021/ac403969n
Novotny, M.V., Alley Jr., W.R., Mann, B.F.: Analytical glycobiology at high sensitivity: current approaches and directions. Glycoconj. J. 30(2), 89–117 (2013). doi:10.1007/s10719-012-9444-8
Both, P., Green, A.P., Gray, C.J., Sardzik, R., Voglmeir, J., Fontana, C., Austeri, M., Rejzek, M., Richardson, D., Field, R.A., Widmalm, G., Flitsch, S.L., Eyers, C.E.: Discrimination of epimeric glycans and glycopeptides using IM-MS and its potential for carbohydrate sequencing. Nat. Chem. 6(1), 65–74 (2014). doi:10.1038/nchem.1817
Li, H., Bendiak, B., Siems, W.F., Gang, D.R., Hill Jr., H.H.: Carbohydrate structure characterization by tandem ion mobility mass spectrometry (IMMS)2. Anal. Chem. 85(5), 2760–2769 (2013). doi:10.1021/ac303273z
Plasencia, M.D., Isailovic, D., Merenbloom, S.I., Mechref, Y., Novotny, M.V., Clemmer, D.E.: Resolving and assigning N-linked glycan structural isomers from ovalbumin by IMS-MS. J. Am. Soc. Mass Spectrom. 19(11), 1706–1715 (2008). doi:10.1016/j.jasms.2008.07.020
Fenn, L.S., McLean, J.A.: Structural resolution of carbohydrate positional and structural isomers based on gas-phase ion mobility-mass spectrometry. Phys. Chem. Chem. Phys. 13(6), 2196–2205 (2011). doi:10.1039/c0cp01414a
Harvey, D.J., Scarff, C.A., Crispin, M., Scanlan, C.N., Bonomelli, C., Scrivens, J.H.: MALDI-MS/MS with traveling wave ion mobility for the structural analysis of N-linked glycans. J. Am. Soc. Mass Spectrom. 23(11), 1955–1966 (2012). doi:10.1007/s13361-012-0425-8
Hofmann, J., Struwe, W.B., Scarff, C.A., Scrivens, J.H., Harvey, D.J., Pagel, K.: Estimating collision cross sections of negatively charged N-glycans using traveling wave ion mobility-mass spectrometry. Anal. Chem. 86(21), 10789–10795 (2014). doi:10.1021/ac5028353
Pagel, K., Harvey, D.J.: Ion mobility-mass spectrometry of complex carbohydrates: collision cross sections of sodiated N-linked glycans. Anal. Chem. 85(10), 5138–5145 (2013). doi:10.1021/ac400403d
Campbell, M., Hayes, C., Struwe, W., Wilkins, M., Aoki-Kinoshita, K., Harvey, D., Rudd, P., Kolarich, D., Lisacek, F., Karlsson, N., Packer, N.: UniCarbKB: putting the pieces together for glycomics research. Proteomics 11(21), 4117–4121 (2011)
Bush, M.F., Hall, Z., Giles, K., Hoyes, J., Robinson, C.V., Ruotolo, B.T.: Collision cross sections of proteins and their complexes: a calibration framework and database for gas-phase structural biology. Anal. Chem. 82(22), 9557–9565 (2010). doi:10.1021/ac1022953
Struwe, W.B., Benesch J.L., Harvey D.J., Pagel K.: Collision cross sections of high-mannose N-glycans in commonly observed adduct states – identification of gas-phase conformers unique to [M−H]− ions. Analyst (2015). doi:10.1039/C5AN01092F
Varki, A.: Essentials of Glycobiology, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor (2009)
Harvey, D.J., Merry, A.H., Royle, L., Campbell, M.P., Dwek, R.A., Rudd, P.M.: Proposal for a standard system for drawing structural diagrams of N- and O-linked carbohydrates and related compounds. Proteomics 9(15), 3796–3801 (2009). doi:10.1002/pmic.200900096
Harvey, D.J., Merry, A.H., Royle, L., Campbell, M.P., Rudd, P.M.: Symbol nomenclature for representing glycan structures: extension to cover different carbohydrate types. Proteomics 11(22), 4291–4295 (2011). doi:10.1002/pmic.201100300
York, W.S., Agravat, S., Aoki-Kinoshita, K.F., McBride, R., Campbell, M.P., Costello, C.E., Dell, A., Feizi, T., Haslam, S.M., Karlsson, N., Khoo, K.H., Kolarich, D., Liu, Y., Novotny, M., Packer, N.H., Paulson, J.C., Rapp, E., Ranzinger, R., Rudd, P.M., Smith, D.F., Struwe, W.B., Tiemeyer, M., Wells, L., Zaia, J., Kettner, C.: MIRAGE: the minimum information required for a glycomics experiment. Glycobiology 24(5), 402–406 (2014). doi:10.1093/glycob/cwu018
Ranzinger, R., Aoki-Kinoshita, K.F., Campbell, M.P., Kawano, S., Lutteke, T., Okuda, S., Shinmachi, D., Shikanai, T., Sawaki, H., Toukach, P., Matsubara, M., Yamada, I., Narimatsu, H.: GlycoRDF: an ontology to standardize glycomics data in RDF. Bioinformatics 31(6), 919–925 (2015). doi:10.1093/bioinformatics/btu732
Acknowledgments
This work was supported by the Australian National eResearch Collaboration Tools and Resources project [NeCTAR RT016 to M.P.C] and Biotechnology and Biological Sciences Research Council (BBSRC) [BB/L017733/1 to W.S, D.J.H, J.L.P.B], the Free University Berlin and the Max Planck Society.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Struwe, W.B., Pagel, K., Benesch, J.L.P. et al. GlycoMob: an ion mobility-mass spectrometry collision cross section database for glycomics. Glycoconj J 33, 399–404 (2016). https://doi.org/10.1007/s10719-015-9613-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-015-9613-7