Abstract
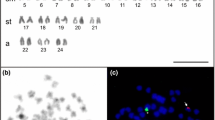
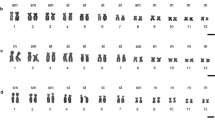
Chromosomal structural rearrangement in four scallops, Chlamys farreri (n = 19), Patinopecten yessoensis (n = 19), Chlamys nobilis (n = 16) and Argopecten irradians (n = 16), was studied by fluorescence in situ hybridization using histone H3 gene probes. The results show that histone H3 gene sites differ strikingly with regard to number, location, and intensity among, or even within these species. For example, two histone H3 gene loci were detected on the metaphase chromosomes of P. yessoensis, while one locus was found in the others. In P. yessoensis, differing intensities of hybridization signals were detected between homologues 5 and 11, and within homologue 11. These data suggest that the histone H3 gene is a qualified chromosome marker for the preliminary understanding of the historical chromosomal reconstructing of the Pectinidae family. The variable distribution patterns of the histone H3 gene suggest that gene duplication/diminution as well as chromosome rearrangements by inversion and translocation may have played important roles in the genomic evolution of Pectinidae. We also compiled our present results with former published data regarding the chromosome mapping of rDNAs in species of the Pectinidae family. Such comparative chromosomal mapping should improve our understanding of historical chromosomal reconstructions of modern-day scallops.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Albig W, Ebentheuer J, Klobeck G, Kunz J, Doenecke D (1996) A solitary human H3 histone gene on chromosome 1. Hum Genet 97:486–491
Barucca M, Olmo E, Schiaparelli S, Canapa A (2004) Molecular phylogeny of the family Pectinidae (Mollusca: Bivalvia) based on mitochondrial 16S and 12S rRNA genes. Mol Phylogenet Evol 31:89–95
Drabent B, Kardalinou E, Bode C, Doenecke D (1995) Association of histone H4 genes with the mammalian testis-specific H1t histone gene. DNA Cell Biol 14:591–597
Eirín-López JM, Fernanda Ruiz M, González-Tizón AM, Martínez A, Sánchez L, Méndez J (2004) Molecular evolutionary characterization of the mussel Mytilus histone multigene family: first record of a tandemly repeated unit of five histone genes containing an H1 subtype with“orphon” features. J Mol Evol 58:131–144
Eirín-López JM, González-Tizón AM, Martínez A, Méndez J (2002) Molecular and evolutionary analysis of mussel histone genes (Mytilus spp.): possible evidence of an “orphon origin” for H1 histone genes. J Mol Evol 55:272–283
Fernández-Tajes J, González-Tizón A, Martínez-Lage A, Méndez J (2003) Cytogenetics of the razor clam Solen marginatus (Mollusca: Bivalvia: Solenidae). Cytogenet Genome Res 101:43–46
Gajardo G, Parraguez M, Colihueque N (2002) Karyotype analysis and chromosome banding of the Chilean-Peruvian scallop Argopecten purpuratus (Lamarck, 1819). J Shellfish Res 21:585–590
Hirai H, Yamamoto MT, Taylor RW, Imai HT (1996) Genomic dispersion of 28S rDNA during karyotypic evolution in the ant genus Myrmecia (Formicidae). Chromosoma 105:190–196
Insua A, Freire R, Ríos J, Méndez J (2001) The 5S rDNA of mussels Mytilus galloprovincialis and M. edulis: sequence variation and chromosomal location. Chrom Res 9:495–505
Insua A, López-Piñón MJ, Méndez J (1998) Characterization of Aequipecten opercularis (Bivalvia: Pectinidae) chromosomes by different staining techniques and fluorescent in situ hybridization. Genes Genet Syst 73:193–200
Insua A, Méndez J (1998) Physical mapping and activity of ribosomal RNA genes in mussel Mytilus galloprovincialis. Hereditas 128:189–194
Komaru A, Wada KT (1985) Karyotypes of four species in the Pectinidae (Bivalvia: Pteriomorphia). Venus 44:249–259
Korobkov IA (1960) Family Pectinidae Lamarck, 1801. In: Orlov UA (ed) Osnovy Paleontologii, Mollusca-Loricata, Bivalvia and Scaphopoda. Academy Nauk USSR, Moscow, pp 82–85
López-Piñón MJ, Insua A, Méndez J (2005) Chromosome analysis and mapping of ribosomal genes by one- and two-color fluorescent in situ hybridization in Hinnites distortus (Bivalvia: Pectinidae). J Hered 96:52–58
Macgregor HC, Varley JM (1983) Working with animal chromosomes. John Wiley and Sons Ltd, Chichester and New York
Matsumoto M, Hayami I (2000) Phylogenetic analysis of the family Pectinidae (Bivalvia) based on mitochondrial cytochrome C oxidase subunit I. J Moll Stud 66:477–488
Miller DJ, Harrison PL, Mahony TJ, McMillan JP, Miles A, Odorico DM, ten Lohuis MR (1993) Nucleotide sequence of the histone gene cluster in the coral Acropora formosa (Cnidaria; Scleractinia): features of histone gene structure and organization are common to diploblastic and triploblastic metazoans. J Mol Evol 37:245–253
Pauls E, Affonso PR (2000) The karyotype of Nodipecten nodosus (Bivalvia: Pectinidae). Hydrobiologia 420:99–102
Pendás AM, Morán P, García-Vázquez E (1994) Organization and chromosomal location of the major histone cluster in brown trout, Atlantic salmon and rainbow trout. Chromosoma 103:147–152
Ran Y, Hammett RWK, Murray BG (2001) Phylogenetic analysis and karyotype evolution in the genus Clivia (Amaryllidaceae). Ann Bot 87:823–830
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Thiriot-Quiévreux C (2002) Review of the literature on bivalve cytogenetics in the last ten years. Cah Biol Mar 43:17–26
Tripputi P, Emanuel BS, Croce CM, Green LG, Stein GS, Stein JL (1986) Human histone genes map to multiple chromosomes. Proc Natl Acad Sci USA 83:3185–3188
Waller TR (1991) Evolutionary relationship among commercial scallops (Mollusca: Bivalvia: Pectinidae). In: Shumway SE (ed) Scallops: biology, ecology and aquaculture. Elsevier, Amsterdam, pp 1–73
Waller TR (1993) The evolution of ‘‘Chlamys’’ (Mollusca: Bivalvia: Pectinidae) in the tropical western Atlantic and eastern Pacific. Am Malacol Bull 10:195–249
Wang Y, Guo X (2004) Chromosomal rearrangement in Pectinidae revealed by rRNA loci and implications for bivalve evolution. Biol Bull 207:247–256
Wang Y, Xu Z, Guo X (2004) Differences in the rDNA-bearing chromosome divide the Asian-Pacific and Atlantic species of Crassostrea (Bivalvia, Mollusca). Biol Bull 206:46–54
Wang ZF, Tisovec R, Debry RW, Frey MR, Matera AG, Marzluff WF (1996) Characterization of a 55-kb mouse histone gene cluster on chromosome 3. Genome Res 6:702–714
Weiss H, Maluszynska J (2000) Chromosomal rearrangement in autotetraploid plants of Arabidopsis thaliana. Hereditas 133:255–261
Acknowledgements
The authors thank Dr. Ke Bi (University of Guelph, Canada) for critically reviewing the manuscript. This work is supported by grants from Hi-Tech Research and Development Program of China (2005AA603220) and National Natural Science Foundation of China (30300268).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, L., Bao, Z., Wang, S. et al. Chromosome rearrangements in Pectinidae (Bivalvia: Pteriomorphia) implied based on chromosomal localization of histone H3 gene in four scallops. Genetica 130, 193–198 (2007). https://doi.org/10.1007/s10709-006-9006-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-006-9006-8