Abstract
Stripe rust, caused by Puccinia striiformis f. sp. tritici, is one of the most devastating foliar diseases of common wheat (Triticum aestivum L.) worldwide. Growing resistant cultivars is the most effective approach to control the disease. To determine inheritance of stripe rust resistance and map the resistance gene in a common wheat line D31, developed from Triticum sphaerococcum Perc. (accession number AS348), F1, F2, and BC1 progenies derived from the Taichung 29 × D31 cross were firstly inoculated with Chinese PST race CYR32 during whole growth stages under the field conditions. Genetic analysis indicated that the resistance to CYR32 in the line D31 was conferred by one recessive gene, temporarily designated as YrSph. A total of 400 simple sequence repeat (SSR), 315 pairs of sequence-related amplified polymorphism and 42 pairs of target region amplified polymorphism markers were screened, and four SSR markers and three TRAP markers were found to be polymorphic between the resistant and susceptible DNA bulks as well as their parents. Genetic linkage was tested on segregating F2 population and indicated that all of the ten markers were linked to the resistance gene, two of which flanked the locus at 8.5 and 6.9 cM, respectively. The SSR markers mapped the resistance gene on chromosome arm 2AS. The results of chromosome location and pedigree analysis indicate that YrSph was probably a novel stripe rust resistance gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe (yellow) rust, caused by Puccinia striiformis f. sp. tritici Eriks., is one of the most damaging diseases of common wheat (Triticum aestivum L.) worldwide, especially in many cool and moist environments (Stubbs 1985; Wellings and McIntosh 1990; Wellings et al. 2003; McIntosh and Brown 1997; Chen 2005, 2007). So far, breeding and release of resistant cultivars has been the most effective and economical approach to control the disease. Currently, more than 49 officially named Yr genes at 49 loci (Yr1–Yr49) and many temporarily designated genes have been reported (McIntosh et al. 2003, 2007; Herrera-Foessel et al. 2011; Lagudah 2011). The varietal resistances, however, are usually short-lived because of rapid virulence changes in pathogen populations. In China, especially southwest China, stripe rust is one of the most destructive diseases of wheat (Li and Liu 1957; Li et al. 1984; Li and Zeng 2000; Wan et al. 2004) and has been considered the most important disease of wheat since the first major epidemic in 1950. China also has the largest epidemic region in the world (Stubbs 1988). More unfortunately, the recent appearance and spread of new stripe rust races CYR31, CYR32, and CYR33 have caused breakdown of numerous formerly resistant sources of stripe rust resistance or Yr genes, studies have shown that Yr5, Yr10, Yr11, Yr12, Yr13, Yr14, Yr15, Yr24, Yr26, YrZH84, and some other genes are still effective while Yr1, Yr2, Yr3, Yr4, Yr6, Yr7, Yr8, Yr9, and certain other genes have lost their effectiveness (Wan et al. 2007). Therefore, exploring new and effective stripe rust resistance genes and transferring to the existing cultivars has become an important task for the breeding wheat with stripe rust resistance (Yang et al. 1994; McIntosh and Brown 1997).
Molecular marker, including random amplified polymorphic DNA (RAPD), restriction fragment length polymorphism (RFLP), amplified restriction fragment polymorphism (AFLP), simple sequence repeat (SSR), and resistance gene analog polymorphism (RGAP), have been widely used for tagging resistance genes to stripe rust in wheat during the past years (Sun et al. 1997; Chague et al. 1999; Peng et al. 1999; Robert et al. 1999; Ma et al. 2001; Shi et al. 2001; Sun et al. 2002; Wang et al. 2002; Suenaga et al. 2003; Yan et al. 2003; Eriksen et al. 2004; Luo et al. 2005; Bariana et al. 2006; Chicaiza et al. 2006; Lin and Chen 2007; McIntosh et al. 2010). Sequence-related amplified polymorphism (SRAP) markers, which are PCR-based markers that amplify open reading frames and produce a number of co-dominant markers per amplification, were recently developed as a useful molecular marker system (Li and Quiros 2001). SRAP markers are more consistent and repeatable than RAPDs, and are less labor-intensive and time-consuming to produce than AFLP techniques (Welsh and McClelland 1990; Li and Quiros 2001; Ferriol et al. 2003; Budak et al. 2004; Gulsen et al. 2005). It is easily adapted to efficiently perform high throughput data collection from thousands or even millions of individuals, which is critical to any large-scale plant breeding program. It has been successfully applied in several species for different purposes (Budak et al. 2004; Ferriol et al. 2003; Sun et al. 2007; Rahman et al. 2007; Li et al. 2010). The other useful PCR-based markers technique namely, target region amplification polymorphism (TRAP) which amplify intragenic polymorphism has been reported by Hu and Vick (2003). TRAPs are amplified by a forward or fixed primer using gene/EST sequence information in the database and a reverse or arbitrary primer that is similar to that of a SRAP primer except for AT- or GC-rich cores that anneal with introns and exons, respectively. TRAPs were effectively used in assessing genetic diversity among wild sunflower (Helianthus annuus L.) and spinach (Spinacia oleracea L.) (Hu et al. 2003; 2007), genetic fingerprinting of lettuce (Lactuca sativa L.) (Hu et al. 2005), tagging gene in sunflower (Rojas-Barros et al. 2005), and constructing genetic linkage maps in wheat, common bean (Phaseolus vulgaris L.), cultivated sugarcane (Saccharum officinarum L.) (Liu et al. 2005; Miklas et al. 2006; Alwala et al. 2008).
Triticum sphaerococcum Perc. (2n = 6x = 42, AABBDD), including 17 varieties, is a member of the primary gene pool and mainly distributed into the northwestern of India with dry and hot climate (Dong 1982). Lan et al. (2003) demonstrated that T. sphaerococcum (accession number AS348) were highly resistant in both seedling and adult-plant stages to stripe rust races prevalent (including CYR32) in the field of China. Wheat new line D31, derived from the cross between AS348 and a susceptible line 94-3854, also is an excellent resistance to stripe rust race CYR32 (which is currently prevalent in China) in the field. The objective of the present study was to identify the stripe rust resistance gene in D31 and map the gene using molecular markers.
Materials and methods
Wheat materials
An F2 population with 144 plants and 43 BC1 plants, derived from the cross between a resistant wheat line D31 and a susceptible variety Taichung 29, were used for the mapping of stripe rust resistance gene. D31, a new wheat line and highly resistant to a mixture of predominant Chinese PST races (including CYR31, CYR32, and CYR33) at both the seedling and adult stages under field conditions, was developed from the cross AS348/2/94-3854 by Triticeae Research Institute, Sichuan Agricultural University. Furthermore, T. sphaerococcum accession number AS348 and wheat line 94-3854, the parents of wheat line D31, were used for comparing the responses to confer the sources pedigree of the resistance gene.
Evaluations for stripe rust resistance
The parents and genetic populations were inoculated in the field with PST isolate CYR32, which provided by the Plant Protection Institute of Gansu Academy of Agricultural Sciences, Gansu, China. The spreader rows (SY95-71, a susceptible line, was used as check in the field) were artificially inoculated in the seedling of the two-leaf stage. To ensure precision of stripe rust resistance, we follow-up investigated rust responses of per plant from the seedling to the adult-plant stages. At last, when rust was fully developed on the susceptible check, SY95-71, infection types (IT) were scored based on a scale of 0, 0;, 1, 2, 3, and 4, where 0 = immunity, 0; = necrotic flecks, and 1–4 = highly resistant, resistant, susceptible, and highly susceptible, respectively (Liu 1988).
Molecular markers analyzes
Genomic DNA of individual plant was extracted from the third or fourth healthy leaf using cetyltrimethylammonium bromide (CTAB) method as essentially described by Saghai et al. (1984). Resistant and susceptible bulks comprising equal amounts of DNA from ten resistant and ten susceptible F2 plants, respectively, were used for bulked segregant analysis (Michelmore et al. 1991). The primers sequences of 400 SSR markers were obtained from grain genes (website) and published by Röder et al. (1998) and Pestsova et al. (2000). A total of 315 primer combinations (15 forward primers in combination with 21 reverse primers) of SRAP reported by Li et al. (2010), and 42 primer combinations (six fixed primers in combination with seven arbitrary primers) of TRAP reported by Hu and Vick (2003), were synthesized. All of the SSR, SRAP and TRAP primers were screened on the two parents and the resistant and susceptible bulks. The candidate markers subsequently identified in DNA of each of the F2 population.
Statistical analyzes and genetic mapping
Chi-squared (χ2) analyzes were performed to check goodness of fit of observed segregations for stripe rust response with the expected ratios. Linkage between DNA markers and the resistance gene was established with MAPMARKER/EXP 3.0b (Lander et al. 1987). Markers were placed with a LOD threshold of 3.0 and a maximum distance of 30 cM. The Kosambi function was applied to convert recombination fractions into map distances (Kosambi 1944). The genetic map was drawn with the software Mapdraw V2.1 (Liu and Meng 2003).
Results
Inheritance of stripe rust resistance in D31
At both seedling and adult-plant stages, D31 and AS348 showed highly resistant reaction (IT 0;) against CYR32, whereas 94-3854 and Taichung 29 were susceptible (IT 3–4) by follow-up investigating rust responses of per plant. When the F1, F2, and BC1 progenies of Taichung 29/D31 were tested with CYR32, all 15 F1 plants were susceptible (IT 3–4). The 144 F2 plants segregated into 38 resistant (IT 0–2) and 106 susceptible (IT 3–4), which conformed to 1R:3S segregation ratio (χ 2 = 0.148, 1 df, P ≥ 0.05). Moreover, the BC1 population (backcross with D31) segregated into 20 resistant (IT 0–0;) and 23 susceptible (IT 3–4), which is shown to 1R:1S segregation ratio (χ 2 = 0.093, 1 df, P ≥ 0.05). Comprehensive analyzes above data suggested that the stripe rust resistant in D31 derived from T. sphaerococcum Perc. (accession number AS348), and was conferred by a single recessive gene, tentatively designated YrSph (Table 1).
Linkage analysis and genetic map
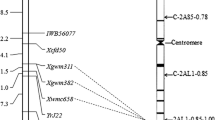
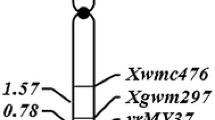
Bulked segregant analysis was used to identify SSR markers linking to the gene YrSph. Among screened 400 SSR markers, four SSR markers Xwmc149, Xwmc246, Xwmc198, and Xgwm372 were polymorphic between the resistant and susceptible DNA bulks. Subsequent linkage analysis based on the phenotype and genotype data of the 144 F2 population plants with the four polymorphic SSR markers, indicated that the resistance gene YrSph was linked to the four SSR loci with genetic distances ranging from 8.5 to 38.5 cM, in which the closest flanking SSR loci was Xwm246-2AS (Figs. 1, 3). According to the SSR genetic linkage map of common wheat (Somers et al. 2004), the four SSR markers were all located on the short arm of the chromosome 2A, suggesting that YrSph is on the short arm of 2A. In addition, three SSR markers, Xwmc246, Xwmc198, and Xgwm372 exhibited co-dominant inheritance and a 1:2:1 segregation ratio, whereas another SSR allele Xwmc149 showed dominant and a 1:3 segregation ratio in the F2 population (Table 2).
Agarose gel electrophoresis detection result of microsatellite DNA products amplified by primer Xwmc246 among individuals of F2. M marker, P R resistance parent D31, P S susceptible parent Taichung 29, R resistance individual, S susceptible individual, asterisk notes exchange between marker and agronomic trait, pound notes heterozygote, arrow notes polymorphic bands
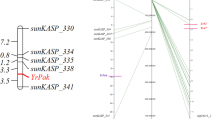
Two molecular marker techniques, SRAP and TRAP have been further used to detect molecular markers for YrSph to obtain more nearly direct markers. Of a total of 315 primer pairs from combinations of 15 forward and 21 reverse SRAP primers reported by Li et al. (2010), and 42 combinations of six arbitrary and seven fixed TRAP primers reported by Hu and Vick (2003), respectively, screened between the resistant and susceptible DNA bulks as well as their parents. Six primer combinations that generated strong and repeatable polymorphic bands were selected to test F2 population plants (Fig. 2). As an example, Fig. 2 showed the banding pattern of F2 population plants screened with SRAP primers F-me15 and R-me6 for marker F-me15/R-me6. In addition, all six markers, including three SRAP markers, F-me15/R-me6, F-me14/R-me10, and F-me4/R-me1, and three TRAP markers, FIX1/Sa12-700, FIX5/Sa4-700, and FIX5/Ga5-800 exhibited dominant inheritance and a 1:3 segregation ratio in the F2 population (Table 2). Mapmaker/EXP version 3.0b at minimum LOD of 3.0 was used for the linkage analysis. All of these six markers were linked to the resistance gene YrSph, in which the closest flanking marker was F-me15/R-me6 with a genetic distance of 6.9 cM (Fig. 3).
Polyacrylamide gel electrophoresis detection result of SRAP DNA products amplified by primer combinations F-me15/R-me6 among individuals of F2. M marker, P R resistance parent D31, P S susceptible parent Taichung 29, Rp resistant pool, Sp susceptible pool, R resistance individual, S susceptible individual, arrow notes polymorphic bands
Discussion
To date, more than 40 permanently and many temporarily designated Yr genes have been reported (Chen 2005; McIntosh et al. 2007; Herrera-Foessel et al. 2011; Lagudah 2011). Due to the new races emerging and lacking effective resistance genes, exploring new and more effective stripe rust resistance genes has become an important task for wheat breeders and pathologists. Here we identified a stripe rust resistance gene YrSph in wheat line D31 and mapped it on the short arm of chromosome 2A. Classical genetic analysis showed that D31 was controlled by a single recessive gene and conferred effective all-stage resistance against Chinese PST race CYR32 and other current pathogen populations, including CYR31 and CYR33. Therefore, the YrSph gene should be useful for developing cultivars with stripe rust resistance. Besides YrSph, three other stripe rust resistance genes Yr1, Yr17, and Yr32 have been previously located on wheat chromosome 2A as well. The Yr1 gene was first found and named in Chinese 166, and genetic analysis showed that it was a single dominant gene (Lupton et al. 1962). The Yr17 gene was report in VPM1 (Maia et al. 1967), which was supposed to have been introgressed from Aegilops ventricosa (2n = 4x = 28, DVDVMVMV) (Bariana et al. 1994) and located on the short arm of 2A using RFLP analysis (Robert et al. 1999) and displayed yellow rust resistance at the seedling stage (Bariana et al. 2006). The Yr32 gene, originally derived from the common wheat cultivars Senat, was located on the long arm of 2A (Eriksen et al. 2004). In this study, although YrSph is also located on the short arm of 2A, we conclude that it is different from Yr1, Yr17, and Yr32 from pedigree analysis and chromosome locations and maybe a novel stripe rust resistance gene.
As wheat new line D31, derived from T. sphaerococcum (accession number AS348), has the same genome with common wheat, it is an excellent bridge parent to transferring its resistance gene into different wheat cultivars. Highly resistance at both seedling and adult-plant stages to stripe rust races prevalent (including CYR32) after nearly 5 years of field test evaluation in the southwestern of China, and common wheat background makes D31 a desirable resistant donor to wheat breeding programs. Based on epidemiological considerations in China, it has been suggested that different resistance genes for controlling wheat stripe rust should be deployed in the defined over-summering, over-wintering, and eastern spring epidemic regions (Li and Zeng 2002; Wan et al. 2007). This would be best achieved by the use of resistant cultivars with multiple or different resistance genes, or effective multilines capable of reducing the build-up of inoculum. So far, growing multiple-resistance-genes cultivars by pyramiding of YrSph and other efficient resistance genes (including Yr5, Yr10, Yr11, Yr12, Yr13, Yr14, Yr15, Yr24, Yr26, YrZH84) in China, would promote utilization of the resistance gene. In fact, we have used D31 in breeding programs in recent years in Triticeae Research Institute, Sichuan Agricultural University by crossed or backcrossed with the main wheat cultivars (such as Chuannong 16 and Shumai 482) in Sichuan province, and obtained some resistance lines with good agronomic traits. More fortunately, we have demonstrated YrSph to a new gene and identified its flanking SSR and SRAP markers in this study. These results should accelerate its application in our breeding programs.
References
Alwala S, Kimbeng CA, Veremis JC, Gravois KA (2008) Linkage mapping and genome analysis in a Saccharum interspecific cross using AFLP, SRAP, and TRAP markers. Euphytica 164:37–51
Bariana HS, McIntosh RA (1994) Characterization and origin of rust and powdery mildew resistance in VPM1 wheat. Euphytica 77:53–61
Bariana HS, Parry N, Barclay IR, Loughman R, McLean RJ, Shankar M, Wilson RE, Willey NJ, Francki M (2006) Identification and characterization of stripe rust resistance gene Yr34 in common wheat. Theor Appl Genet 112:1143–1148
Budak H, Shearman RC, Parmaksizand I, Dweikat I (2004) Comparative analysis of seeded and vegetative biotype buffalo grasses based on phylogenetic relationship using ISSRs, SSRs, RAPDs, and SRAPs. Theor Appl Genet 109:280–288
Chague V, Fahima T, Sun GL, Dahan A, Sun GL, Korol AB, Ronin YI, Grama A, Roder MS, Nevo E (1999) Isolation of microsatellite and RAPD markers flanking the Yr15 gene of wheat using NILs and bulked segregant analysis. Genome 42:1050–1056
Chen XM (2005) Epidemiology and control of stripe rust (Puccinia striiformis f. sp. tritici) on wheat. Can J Plant Pathol 27:314–337
Chen XM (2007) Challenges and solutions for stripe rust control in the United States. Aust J Agric Res 58:648–655
Chicaiza O, Khan IA, Zhang X, Brevis JC, Chen X, Dubcovsky J (2006) Registration of five wheat isogenic lines for leaf rust and stripe rust resistance genes. Crop Sci 46:485–487
Dong YS (1982) World wheat (in Chinese). China Agriculture Press, Beijing
Eriksen L, Afshari F, Christiansen MJ, McIntosh RA, Jahoorand A, Wellings CR (2004) Yr32 for resistance to stripe rust present in the wheat cultivar Carstens V. Theor Appl Genet 108:567–575
Ferriol M, Pico B, Nuez F (2003) Genetic diversity of a germplasm collection of Cucurbita pepo using SRAP and AFLP markers. Theor Appl Genet 107:271–282
Gulsen O, Shearman RC, Vogel KP, Lee DJ, Baenziger PS, Heng-Moss TM, Budak H (2005) Nuclear genome diversity and relationships among naturally occurring buffalo grass genotypes determined by sequence-related amplified polymorphism. HortScience 40:537–541
Herrera-Foessel SA, Lagudah ES, Huerta-Espino J, Hayden M, Bariana HS, Singh D, Singh RP (2011) New slow rusting leaf rust and stripe rust resistance genes Lr67 and Yr46 in wheat are pleiotropic or closely linked. Theor Appl Genet 122:239–249
Hu J, Vick BA (2003) Target region amplification polymorphism (TRAP), a novel marker technique for plant genotyping. Plant Mol Biol Rep 21:289–294
Hu J, Ochoa OE, Truco MJ, Vick BA (2005) Application of the TRAP technique to lettuce (L.) genotyping. Euphytica 144:225–235
Hu JG, Mou BQ, Vick BA (2007) Genetic diversity of 38 spinach (Spinacia oleracea L.) germplasm accessions and 10 commercial hybrids assessed by TRAP markers. Genet Resour Crop Evol 54:1667–1674
Kosambi DD (1944) The estimation of map distance from recombination values. Ann Eugen 12:172–175
Lagudah ES (2011) Molecular genetics of race non-specific rust resistance in wheat. Euphytica 179:81–91
Lan XJ, Liu DC, Zheng YL (2003) Studies on the inheritance of stripe rust resistance in Triticum sphaerococcum. Acta Phytopathol Sin 1:91
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li ZQ, Liu HW (1957) Discussions about decreases in resistance to wheat stripe rust of BimA1. Acta Bot Boreal-Occident Sin 2:93–102
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Li ZQ, Zeng SM (2000) Wheat rusts in China. China Agricultural Press, Beijing
Li ZQ, Zeng SM (2002) Wheat rusts in China. China Agriculture Press, Beijing
Li ZQ, Shang HS, Yin XL, Qiang ZF, Zhao YQ, Lu HP, Hong XW, Song WZ, Liu SJ (1984) Studies on the breakdown of lovrin cultivars of wheat to stripe rust (Puccinia striiformis West). Sci Agric Sin 1:68–74
Li AX, Liu QC, Wang QM, Zhang LM, Zhai H, Liu SZ (2010) Construction of molecular linkage maps using SRAP markers in sweet potato. Acta Agron Sin 8:1286–1295
Lin F, Chen XM (2007) Genetics and molecular mapping of genes for race-specific all-stage resistance and non-race-specific high-temperature adult-plant resistance to stripe rust in spring wheat cultivar Alpowa. Theor Appl Genet 114:1277–1287
Liu XK (1988) Study on the yellow rust resistance to common wheat (T. aetivum). Plant Prot 15:33–39
Liu RH, Meng JL (2003) MapDraw: a microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas 3:317–321
Liu ZH, Anderson JA, Hu J, Friesen TL, Rasmussen TL, Faris JD (2005) A wheat intervarietal genetic linkage map based on microsatellite and target region amplified polymorphism markers and its utility for detecting quantitative trait loci. Theor Appl Genet 111:782–794
Luo PG, Ren ZL, Zhang HQ (2005) Identification, chromosome location, and diagnostic markers for a new gene (YrCN19) for resistance to wheat stripe rust. Phytopathology 95:1266–1270
Lupton FCH, Mace RCF (1962) Inheritance of resistance to yellow rust (Puccinia glumarum Erikss and Henn.) in seven varieties of wheat. Trans Br Mycol Soc 45:21–45
Ma JX, Zhou RG, Dong YS, Wang LF, Wang XM, Jia JZ (2001) Molecular mapping and detection of the yellow rust resistance gene Yr26 in wheat transferred from Triticum turgidum L using microsatellite markers. Euphytica 120:219–226
Maia N (1967) Obtention des blés tendres resistants au pietin-verse par croisements interspecifiques blés X Aegilops. Comptes Rendus des Séances de l’Académie d’Agriculture de France 53:149–154
McIntosh RA, Brown GN (1997) Anticipatory breeding for resistance to rust diseases in wheat. Annu Rev Phytopathol 35:311–326
McIntosh RA, Yamazaki Y, Devos KM, Dubcovsky J, Rogers WJ, Appels R (2003) Catalogue of gene symbols for wheat. In: Proceedings of the 10th international wheat genetics symposium, vol 4, Paestum, Italy, 1–6 Sept 2003
McIntosh RA, Devos KM, Dubcovsky J, Rogers WJ, Morris CF, Appels R, Somers DJ, Anderson OA (2007) Catalogue of gene symbols for wheat: 2007 suppl
McIntosh RA, Dubcovsk J, Rogers WJ, Morris C, Appels R, Xia XC (2010) Catalogue of gene symbols for wheat: 2010 suppl. Annu Wheat Newsl
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions using segregation populations. Proc Natl Acad Sci USA 88:9828–9832
Miklas PN, Hu J, Grünwald NJ, Larsen KM (2006) Potential application of targeted region amplified polymorphism (TRAP) markers for mapping and tagging disease resistance traits in common bean. Crop Sci 46:910–916
Peng JH, Fahima T, Röder MS, Li YC, Dahan A, Grama A, Ronin YI, Korol AB, Nevo E (1999) Microsatellite tagging of the stripe rust resistance gene YrH52 derived from wild emmer wheat, Triticum dicoccoides, and suggestive negative crossover interference on chromosome 1B. Theor Appl Genet 98:862–872
Pestsova E, Ganal MW, Röder MS (2000) Isolation and mapping of microsatellite markers specific for the D genome of bread wheat. Genome 43:689–697
Rahman M, Peter BEM, Li GY (2007) Development of SRAP, SNP, and multiplexed SCAR molecular markers for the major seed coat color gene in Brassica rapa L. Theor Appl Genet 115:1101–1107
Robert O, Abelard C, Dedryver F (1999) Identification of molecular markers for the detection of the yellow rust resistance gene Yr17 in wheat. Mol Breed 5:167–175
Röder MS, Korzun V, Wendehake K (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rojas-Barros P, Jan CC, Hu J (2005) Mapping of a recessive branching gene in RHA 271 using molecular markers. u. In: Proceedings of the 27th Sunflower Research Workshop, Fargo, ND, 12–13 Jan 2005
Saghai MA, Biyashev RM, Yang GP, Zhang Q, Allard RW (1994) Extraordinarily polymorphic microsatellite DNA in barley: species diversity, chromosomal locations, and population dynamics. Proc Natl Acad Sci USA 91:5466–5470
Shi ZX, Chen XM, Line RF, Leung H, Wellings CR (2001) Development of resistance gene analog polymorphism markers for the Yr9 gene resistance to wheat stripe rust. Genome 44:509–516
Stubbs RW (1985) Stripe rust. In: Roelfs AP, Bushnell WR (eds) The cereal rusts, vol II. Academic, New York, pp 61–101
Stubbs RW (1988) Pathogenicity analysis of yellow (stripe) rust of wheat and its significance in a global context. In: Simmonds NW, Rajaram S (eds) Breeding strategies for resistance to the rusts of wheat. CIMMYT, Mexico, pp 23–38
Suenaga K, Singh RP, Huerta-Espino J, William HM (2003) Microsatellite markers for genes Lr34/Yr18 and other quantitative trait loci for leaf rust and stripe rust resistance in bread wheat. Phytopathology 93:881–890
Sun GL, Fahima T, Korol AB, Turpeinen T, Grama A, Ronin YI, Nevo E (1997) Identification of molecular markers linked to the Yr15 stripe rust resistance gene of wheat originated in wild emmer wheat, Triticum dicoccoides. Theor Appl Genet 95:622–628
Sun Q, Wei Y, Ni C, Xie C, Yang T (2002) Microsatellite marker for yellow rust resistance gene Yr5 introgressed from spelt wheat. Plant Breed 121:539–541
Sun ZD, Wang ZN, Tu JX, Zhang JF, Yu FQ, Peter BEV, Li GY (2007) An ultradense genetic recombination map for Brassica napus, consisting of 13551 SRAP markers. Theor Appl Genet 114:1305–1317
Wan AM, Zhao ZH, Chen XM, He ZH, Jin SL, Jia QZ, Yao G, Yang JX, Wang BT, Li GB, Bi YQ, Yuan ZY (2004) Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici in China in 2002. Plant Dis 88:896–904
Wan AM, Chen XM, He ZH (2007) Wheat stripe rust in China. Aust J Agric Res 58:605–619
Wang LF, Ma JX, Zhou RH, Wang XM, Jia JZ (2002) Molecular tagging of the yellow rust resistance gene Yr10 in common wheat, P.I. 178383 (Triticum aestivum L.). Euphytica 124:71–73
Wellings CR, McIntosh RA (1990) Puccinia striiformis f. sp. tritici in Australasia: pathogenic changes during the first 10 years. Plant Pathol 39:316–325
Wellings CR, Wright DG, Keiper F, Loughman R (2003) First detection of wheat stripe rust in western Australia: evidence for a foreign incursion. Aust Plant Pathol 32:321–322
Welsh J, McClelland M (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res 18:7213–7218
Yan GP, Chen XM, Line RF, Wellings CR (2003) Resistance gene-analog polymorphism markers co-segregating with the YR5 gene for resistance to wheat stripe rust. Theor Appl Genet 106:636–643
Yang ZM, Tang BR, Shen KQ, Xia XC (1994) A strategic program in wheat resistance breeding-building and utilization of sources of second-line resistance against rusts and mildew in China. Acta Agron Sin 20:385–394
Acknowledgments
This work was supported by the National Basic Research Program of China (973 Program and 2011CB100100) and the National Basic Research Special Program of China (Grant No. 2010CB134402). The authors are grateful to Prof. Q. Z. Jia, Plant Protection Research Institute, Gansu Academy of Agricultural Sciences, Lanzhou, People’s Republic of China, for providing the stripe rust isolates.
Author information
Authors and Affiliations
Corresponding author
Additional information
Shi-Sheng Chen and Guo-Yue Chen contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Chen, SS., Chen, GY., Chen, H. et al. Mapping stripe rust resistance gene YrSph derived from Tritium sphaerococcum Perc. with SSR, SRAP, and TRAP markers. Euphytica 185, 19–26 (2012). https://doi.org/10.1007/s10681-011-0593-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0593-9